Abstract
Autophagy is a highly conserved mechanism to overcome various stresses and recycle cytoplasmic components and organelles. Ubiquitin-like (UBL) protein Atg12 is a key protein involved in autophagosome formation through stimulation of Atg8 conjugation to phosphatidylethanolamine. Here, we describe the identification of the autophagy-related gene Acatg12, encoding an Atg12 homologous protein in the cephalosporin C producing fungus Acremonium chrysogenum. Disruption of Acatg12 impaired the delivery and degradation of eGFP-Atg8, indicating that the autophagic process was blocked. Meanwhile, conidiation was dramatically reduced in the Acatg12 disruption mutant (∆Acatg12). In contrast, cephalosporin C production was increased twofold in ∆Acatg12, but fungal growth was reduced after 6 days fermentation. Consistent with these results, the transcriptional level of the cephalosporin biosynthetic genes was increased in ∆Acatg12. The results extend our understanding of autophagy in filamentous fungi.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Autophagy is primarily a cytoplasmic degradation process via vacuole or lysosome for coping with various stresses and maintaining cellular homeostasis. This process is highly conserved in eukaryotes [1]. Macroautophagy, microautophagy and chaperone-mediated autophagy are three major autophagic pathways [2]. In macroautophagy (generally called autophagy), a double-membrane vesicle termed autophagosome engulfs cellular constituents (cargoes) such as redundant and damaged organelles and proteins, and fuses with the vacuole membrane, resulting in degradation of cargoes by the lysosomal hydrolases of the vacuole. In microautophagy, the substrates are directly engulfed through invagination of the vacuole membrane. So far, more than 40 autophagy-related genes (Atg) have been found in Saccharomyces cerevisiae [3]. Some of their homologous genes were also identified in filamentous fungi, but their function and mechanism are poorly understood.
Two ubiquitin-like (UBL) protein conjugation systems, consisting of Atg8 and Atg12, are essential for autophagy [2]. The conjugation production of Atg8 and phosphatdylethanolamine (PE) has a pivotal role in phagophore expansion [4]. After the C-terminal arginine is removed by the cysteine protease Atg4, Atg8 is bonded with an E1-like enzyme Atg7, subsequently transferred to an E2-like enzyme Atg3, and eventually conjugated to PE. Another UBL protein, Atg12, is also dependent on Atg7, but it uses Atg10 as E2-like enzyme to form the Atg12-Atg5 conjugate [5, 6]. The Atg12-Atg5 conjugate with an E3-like activity facilitates Atg8-PE conjugation by accelerating the transfer of Atg8 from Atg3 to PE [7]. The Atg12-Atg5 conjugate forms a complex with Atg16 [8]. Both UBL protein conjugation systems are located in the pre-autophagosomal structure (PAS) [9].
Acremonium chrysogenum is well known for producing the β-lactam antibiotic cephalosporin C (CPC) and its derivatives [10]. The CPC biosynthetic enzymes are located in different cellular compartments [11]. These enzymes are encoded by six biosynthetic genes (pcbAB, pcbC, cefD1, cefD2, cefEF and cefG), three transporter genes (cefM, cefP and cefT) and a regulatory gene (cefR) [11, 12]. CPC production is also regulated by various factors such as carbon source, nitrogen source, phosphorus source, pH and oxygen [13]. Recently, autophagy was found to have an effect on antibiotic production. In Penicillium chrysogenum, penicillin production was enhanced by deletion of atg1 [14]. Similarly, disruption of Acatg1 of A. chrysogenum resulted in a remarkable increase in CPC production but a significant reduction of fungal growth at the late stage of fermentation [15]. However, the Acatg11 disruption mutant showed a reduced CPC production [16]. These results indicate that different autophagy-related genes have distinct effects on secondary metabolite production, possible through different pathways. In this study, an autophagy-related gene, Acatg12, was identified in A. chrysogenum and its function was investigated. The results extend our understanding of autophagy in filamentous fungi.
Materials and Methods
Strains, Plasmids, Culture Media and Growth Conditions
Strains and plasmids used in this study are listed in Table S1. The cephalosporin C producing strain A. chrysogenum CGMCC 3.3795, which was purchased from the China General Microbiological Culture Collection Center, was used as the wild-type strain (WT). TSA medium, LPE medium and the modified MDFA medium were used for the growth, sporulation and fermentation of A. chrysogenum and its derivatives as described previously [17]. For Agrobacterium tumefaciens-mediated transformation (ATMT), minimal medium, induction medium and co-cultivation medium were used [18]. Escherichia coli was grown at 37 °C in Luria–Bertani (LB) medium supplemented with antibiotics when required.
Cloning of the Acatg12 gene
The DNA and cDNA of Acatg12 were amplified with primers Acatg12-F/Acatg12-R and cloned into pEASY-Blunt (TransGen, China), respectively. DNA sequencing was performed by GENEWIZ (Beijing, China).
Constructions of the Acatg12 Disruption Mutant and Its Complemented Strain
To obtain the Acatg12 disruption mutant (∆Acatg12), a 1192 bp upstream and a 1117 bp downstream DNA fragments of Acatg12 were amplified from WT with primers Acatg12UF/Acatg12UR and Acatg12DF/Acatg12DR, respectively. For gene disruption, the plasmid pAgAcatg12 was constructed (Fig. S1). ATMT was used for fungal transformation as described previously [16, 18]. The bleomycin-resistant and hygromycin B-sensitive (bleRhphS) transformants were selected and confirmed by PCR using the gene outside primers Acatg12Out-F/Acatg12Out-R and inside primers Acatg12In-F/Acatg12In-R. Southern hybridization was performed to further confirm ∆Acatg12 as previously described [16, 18]. A 534 bp DNA fragment amplified from WT with primers Probe-F/Probe-R was used as a probe. For Southern hybridization, the fungal genomic DNA was digested with ScaI.
To complement ∆Acatg12, the 1535 bp and 1194 bp DNA fragments containing the putative promoter and terminator regions of Acatg12 were amplified with primers Acatg12CF1/Acatg12CR1 and Acatg12CF2/Acatg12CR2, respectively. Plasmid pAgAtg12C was constructed for complementation of ∆Acatg12 (Fig. S2). Finally, pAgAtg12C was introduced into ∆Acatg12 via ATMT. The hygromycin B resistant (hphR) transformants were selected and confirmed as the complemented strain (Acatg12C) by PCR with primers Acatg12In-F/Acatg12In-R.
Fluorescence Microscopy
Fluorescence microscopy was performed on an Axio-observer A1 microscope (Carl Zeiss) as described previously [16, 18]. Images of cells were captured by Zeiss AxioCam MR camera with the AxioVision software. For detection of eGFP-Acatg8, pAgHBPEAT containing the expressing cassette of eGFP-Acatg8 was transformed into ∆Acatg12. Transformants were selected on the TSA medium containing 50 µg/ml hygromycin B and 500 µg/ml cefotaxime and confirmed by PCR.
Determination of Conidiation and CPC Production
Fungal conidiation and CPC production were determined as described previously [17].
RNA Isolation and Real-Time Reverse Transcription PCR (RT-PCR)
RNA was isolated as described previously [17]. The RNA samples were treated with RQ1 RNase free DNase (Promega, USA). The PrimeScript™ RT reagent Kit (TaKaRa, Japan) was used for reverse transcription. Synthesis of cDNA and real-time RT-PCR were performed as described previously [17].
Western Blot Analysis
To examine the autophagy of ∆Acatg12 during fermentation, Western blot analysis of AcAtg8 was performed as previously described [16, 18]. Total cellular extracts from WT and ∆Acatg12 were collected after fermentation for 48, 72, 96 and 120 h. Proteins were separated by SDS-PAGE followed by immunoblotting with anti-AcAtg8 serum. GAPDH was used as control [16, 18].
Statistical Analysis
All data were expressed as mean ± SD. Differences were determined by 2-tailed Student’s t test between two groups in Excel (Microsoft). Statistical significance is indicated as * for P < 0.05 and ** for P < 0.01.
Results
Identification and Characterization of an Autophagy-Related Gene Acatg12 in A. chrysogenum
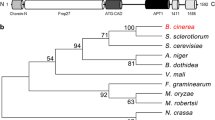
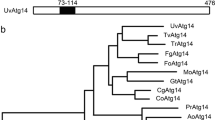
The whole genomic sequence of A. chrysogenum CGMCC 3.3795 (unpublished) was searched in the non-redundant protein database of National Center for Biotechnology Information (NCBI) by using the BLASTx program. A query sequence encoding a protein with 40% identity to Saccharomyces cerevisiae Atg12 and 63% identity to Sordaria macrospora Atg12 was identified and designated Acatg12 (GenBank accession No. MG987012). Comparing its genomic and cDNA sequences, one intron was identified (extending from 394 to 484 with respect to the putative translation start point). Acatg12 encodes a protein of 151 amino acids with a calculated molecular mass of 15.6 kDa. A conserved glycine residue involved in covalent interaction between Atg12 and Atg5 was found at the C-terminus of AcAtg12. Most residues involved in interactions with Atg3, Atg5 and Atg8 are conserved in AcAtg12 (Fig. 1).
Multiple sequence alignment of AcAtg12 and its orthologs. Alignment was performed with the following sequences: AcAtg12 from A. chrysogenum, ScAtg12 from S. cerevisiae (P38316.1), SmAtg12 (XP_003349162.1, excluding C-terminal amino acids) from Sordaria macrospora, AnAtg12 (Q5BCH0.2) from Aspergillus nidulans, CeAtg12(CCD61524.1) from Caenorhabditis elegans, HsAtg12 (O94817.1) from Homo sapiens, MoAtg12 (Q51P78.1) from Magnaporthe oryzae, NcAtg12 (Q7S083.1) from Neurospora crassa, PcAtg12 (ABO31083.1) from P. chrysogenum. Pound sign indicates the non-canonical Atg8 interaction motif in yeast and human. Asterisk stands for the Atg12 residues essential for binding with Atg3 in human. Black spots mark the residues involved in non-covalent interaction with Atg5 in yeast. The C-terminal conserved glycine residue and the phenylalanine residue essential for the function of Atg12-Atg5 conjugate are labeled by red color. (Color figure online)
Acatg12 is Essential for Autophagy in A. chrysogenum
To identify the function of Acatg12, an Acatg12 disruption mutant (∆Acatg12) was constructed via homologous recombination (Fig. S3). Finally, ∆Acatg12 was isolated and confirmed by PCR and Southern hybridization (Fig. S3). RT-PCR analysis showed that transcription of Acatg12 was completely abolished in ∆Acatg12 and restored in the complemented strain Acatg12C (Fig. S3).
To analyze the effect of Acatg12 on autophagy, the autophagy marker Acatg8 gene fused to eGFP was introduced into ∆Acatg12. 3 × 107 spores of each fungal strain were inoculated in MDFA medium and cultured for 20 h. For autophagy induction, the fungal mycelia were switched into water and incubated for 4 h. To prevent the degradation of autophagic bodies, 2 mM PMSF (phenylmethanesulfonyl fluoride, a cysteine protease inhibitor) was added. In ∆Acatg12, the eGFP-AcAtg8 signal was diffused in the cytoplasm (Fig. 2a). In contrast, eGFP-AcAtg8 was almost fully accumulated in the vacuole of WT. These results indicated that engulfment of AcAtg8 was inhibited in ∆Acatg12. In consistent with this, eGFP-AcAtg8 proteolysis assay showed that degradation of eGFP-Acatg8 was also inhibited in ∆Acatg12 (Fig. 2b), indicating that Acatg12 is essential for the autophagy of A. chrysogenum.
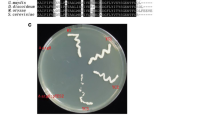
Detection of autophagy in WT and ∆Acatg12. a Fluorescence microscopy of eGFP-Acatg8 in WT and ∆Acatg8. DIC, differential interference contrast. The autophagy was induced when fungal mycelia were switched into water containing 2 mM PMSF after cultured in MDFA for 20 h. b Western blot analysis of eGFP-Acatg8 degradation in WT::eGFP-Acatg8 and ∆Acatg12::eGFP-Acatg8 after 3 days cultured in MDFA. Protein was separated on a 12% SDS-PAGE gel. GAPDH was used as control. WT::eGFP-Acatg8, expression of eGFP-Acatg8 in WT; ∆Acatg12:eGFP-Acatg8, expression of eGFP-Acatg8 in ∆Acatg12
Disruption of Acatg12 Reduces Conidiation of A. chrysogenum
To further study the function of Acatg12, conidiations of WT, ∆Acatg12 and Acatg12C were analyzed on LPE medium. 1 × 106 spores of each strain were spread on LPE medium. After incubation for 7 days, the number of conidia was counted and the mycelium dry weight was measured (Fig. 3a, b). The conidia number of ∆Acatg12 was dramatically decreased to one-sixth of the wild type. As expected, the conidiation was restored in strain Acatg12C. These results demonstrated that Acatg12 is important for the conidiation of A. chrysogenum. When additional carbon sources were supplemented in the medium, the conidiation of ∆Acatg12 was partially restored (Fig. 3c, d).
Conidiation of WT, ∆Acatg12 and Acatg12C. a Conidia formation in WT, ∆Acatg12 and Acatg12C. b Mycelial dry weight in WT, ∆Acatg12 and Acatg12C was measured after growth in LPE medium for 7 days. c Conidia formation of WT, ∆Acatg12 and Acatg12C in the LPE medium supplied with different nitrogen sources. Ammonium chloride (NH4Cl), ammonium nitrate (NH4NO3), sodium nitrate (NaNO3), ammonium sulfate ((NH4)2SO4), methionine (Met) or proline (Pro) was added in the LPE medium at the final concentration of 1.2%. CK, the medium without supplement of additional carbon or nitrogen sources. d Conidia formation of WT, ∆Acatg12 and Acatg12C in the LPE medium supplied with different carbon sources. Glucose, glycerol, sucrose, mannitol or inositol was added in LPE medium at the final concentration of 1.2%. Error bars represent standard deviations from three independent experiments. The ** and *indicate P < 0.01 and 0.05 relative to the control strain WT, respectively
Disruption of Acatg12 Increases CPC Production and Expression of CPC Biosynthetic Genes
To investigate the effect of Acatg12 disruption on CPC production, fermentation of WT, ∆Acatg12 and Acatg12C was performed. CPC production of ∆Acatg12 increased twofold with respect to WT (Fig. 4a). In Acatg12C, CPC production was restored to the wild-type level. WT, ∆Acatg12 and Acatg12C showed comparable growth rates at logarithmic phase (Fig. 4b). The biomass of ∆Acatg12 was only slightly reduced at the stationary phase compared to WT and Acatg12C. These results indicated that the increased CPC production was mainly due to the disruption of Acatg12.
Cephalosporin C production and biomass of WT, ∆Acatg12 and Acatg12C. a CPC production of WT, ∆Acatg12 and Acatg12C during fermentation in MDFA medium. b Biomass of WT, ∆Acatg12 and Acatg12C during fermentation in MDFA medium. Error bars represent standard deviations from three independent experiments
To further address the reasons of increased CPC in ∆Acatg12, transcriptional level of the cephalosporin biosynthetic genes pcbAB, pcbC, cefD1, cefD2, cefEF and cefG was determined via real-time RT-PCR. The transcriptional levels of most of these genes, except cefEF, were significantly increased in ∆Acatg12 on the 4th or 5th day, just before CPC production reached the peak (Fig. 5), while cefD2 reached the highest level in the 3rd day of fermentation. Then, pcbAb, pcbC and cefG maintained higher levels of expression in ∆Acatg12 until day 7. The transcriptional level of cefEF in ∆Acatg12 increased significantly only after six days of fermentation. These results indicate that one reason for the increase in CPC production was the transcriptional increase in cephalosporin biosynthetic genes caused by Acatg12 disruption.
Discussion
Fluorescence microscopy demonstrated that transport and degradation of eGFP-Acatg8 were inhibited in ∆Acatg12, indicating that Acatg12 is essential for Acatg8-mediated autophagy. In addition to the autophagy-related function, disruption of Acatg12 also led to a decrease in conidia formation. Similarly, disruption of atg12 in maize impaired the recycling of nitrogen and decreased seed yield [19]. Autophagy is required for nutrient supply in anthers and lipid metabolism of rice [20]. In S. cerevisiae and S. pombe, atg12 was essential for ascosporogenesis during nitrogen starvation [21]. Similar to ∆Acatg1 [15], supplementation of additional carbon sources partially restored the conidiation of ∆Acatg12, suggesting that carbon source homeostasis dependent on autophagy could be the reason of conidiation reduction.
Furthermore, we found that disruption of Acatg12 increased CPC production. In A. oryzae, disruption of MoATG8 resulted in the accumulation of methionine synthase [22]. It is known that methionine stimulates CPC production by increasing the transcription of biosynthesis genes, so disruption of Acatg12 could accumulate methionine synthase and in turn increase the concentration of endogenesis methionine. Recently, our group found that enhancement of endogenesis SAM concentration through reconstructing methionine cycle dramatically increase CPC production [22]. As the late steps of the cephalosporin biosynthesis pathway take place in peroxisomes [11], we speculate that the accumulated peroxisomes due to the deficiency of autophagy also increase CPC production. Since autophagy is primarily a degradation process, it is possible that some positive regulators that stimulate cephalosporin biosynthesis are not recycled at a normal rate due to the impairment of autophagy, thus resulting in an increase in CPC production.
References
Yorimitsu T, Klionsky DJ (2005) Autophagy: molecular machinery for self-eating. Cell Death Differ 2(12 Suppl):1542–1552. https://doi.org/10.1038/sj.cdd.4401765
Feng Y, He D, Yao Z, Klionsky DJ (2014) The machinery of macroautophagy. Cell Res 24(1):24–41. https://doi.org/10.1038/cr.2013.168
Shpilka T, Welter E, Borovsky N, Amar N, Shimron F, Peleg Y, Elazar Z (2015) Fatty acid synthase is preferentially degraded by autophagy upon nitrogen starvation in yeast. Proc Natl Acad Sci USA 112(5):1434–1439. https://doi.org/10.1073/pnas.1409476112
Nakatogawa H (2013) Two ubiquitin-like conjugation systems that mediate membrane formation during autophagy. Essays Biochem 55:39–50. https://doi.org/10.1042/bse0550039
Mizushima N, Noda T, Yoshimori T, Tanaka Y, Ishii T, George MD, Klionsky DJ, Ohsumi M, Ohsumi Y (1998) A protein conjugation system essential for autophagy. Nature 395(6700):395–398. https://doi.org/10.1038/26506
Tanida I, Mizushima N, Kiyooka M, Ohsumi M, Ueno T, Ohsumi Y, Kominami E (1999) Apg7p/Cvt2p: a novel protein-activating enzyme essential for autophagy. Mol Biol Cell 10(5):1367–1379
Hanada T, Noda NN, Satomi Y, Ichimura Y, Fujioka Y, Takao T, Inagaki F, Ohsumi Y (2007) The Atg12-Atg5 conjugate has a novel E3-like activity for protein lipidation in autophagy. J Biol Chem 282(52):37298–37302. https://doi.org/10.1074/jbc.C700195200
Kuma A, Mizushima N, Ishihara N, Ohsumi Y (2002) Formation of the approximately 350-kDa Apg12-Apg5.Apg16 multimeric complex, mediated by Apg16 oligomerization, is essential for autophagy in yeast. J Biol Chem 277(21):18619–18625. https://doi.org/10.1074/jbc.M111889200
Suzuki K, Kirisako T, Kamada Y, Mizushima N, Noda T, Ohsumi Y (2001) The pre-autophagosomal structure organized by concerted functions of APG genes is essential for autophagosome formation. EMBO J 20(21):5971–5981. https://doi.org/10.1093/emboj/20.21.5971
Ozcengiz G, Demain AL (2013) Recent advances in the biosynthesis of penicillins, cephalosporins and clavams and its regulation. Biotechnol Adv 31(2):287–311. https://doi.org/10.1016/j.biotechadv.2012.12.001
Martin JF, Ullan RV, Garcia-Estrada C (2010) Regulation and compartmentalization of beta-lactam biosynthesis. Microb Biotechnol 3(3):285–299. https://doi.org/10.1111/j.1751-7915.2009.00123.x
Evers ME, Trip H, van den Berg MA, Bovenberg RAL, Driessen AJM (2004) Compartmentalization and transport in beta-lactam antibiotics biosynthesis. Adv Biochem Eng Biotechnol 88:111–135. https://doi.org/10.1007/b99259
Schmitt EK, Hoff B, Kück U (2004) Regulation of cephalosporin biosynthesis. Adv Biochem Eng Biotechnol 88:1–43. https://doi.org/10.1007/b9925614
Bartoszewska M, Kiel JA, Bovenberg RA, Veenhuis M, van der Klei IJ (2011) Autophagy deficiency promotes beta-lactam production in Penicillium chrysogenum. Appl Environ Microbiol 77(4):1413–1422. https://doi.org/10.1128/AEM.01531-10
Wang H, Pan Y, Hu P, Zhu Y, Li J, Jiang X, Liu G (2014) The autophagy-related gene Acatg1 is involved in conidiation and cephalosporin production in Acremonium chrysogenum. Fungal Genet Biol 69:65–74. https://doi.org/10.1016/j.fgb.2014.06.004
Liu J, Hao T, Hu P, Pan Y, Jiang X, Liu G (2017) Functional analysis of the selective autophagy related gene Acatg11 in Acremonium chrysogenum. Fungal Genet Biol 107:67–76. https://doi.org/10.1016/j.fgb.2017.08.006
Long LK, Yang J, An Y, Liu G (2012) Disruption of a glutathione reductase encoding gene in Acremonium chrysogenum leads to reduction of its growth, cephalosporin production and antioxidative ability which is recovered by exogenous methionine. Fungal Genet Biol 49(2):114–122. https://doi.org/10.1016/j.fgb.2011.12.004
Khang CH, Park S-Y, Rho H-S, Lee Y-H, Kang S (2006) Filamentous fungi (Magnaporthe grisea and Fusarium oxysporum). Methods Mol Biol 344:403–420. https://doi.org/10.1385/1-59745-131-2:403
Li F, Chung T, Pennington JG, Federico ML, Kaeppler HF, Kaeppler SM, Otegui MS, Vierstra RD (2015) Autophagic recycling plays a central role in maize nitrogen remobilization. Plant Cell 27(5):1389–1408. https://doi.org/10.1105/tpc.15.00158
Kurusu T, Koyano T, Hanamata S, Kubo T, Noguchi Y, Yagi C, Nagata N, Yamamoto T, Ohnishi T, Okazaki Y, Kitahata N, Ando D, Ishikawa M, Wada S, Miyao A, Hirochika H, Shimada H, Makino A, Saito K, Ishida H, Kinoshita T, Kurata N, Kuchitsu K (2014) OsATG7 is required for autophagy-dependent lipid metabolism in rice postmeiotic anther development. Autophagy 10(5):878–888. https://doi.org/10.4161/auto.28279
Mukaiyama H, Kajiwara S, Hosomi A, Giga-Hama Y, Tanaka N, Nakamura T, Takegawa K (2009) Autophagy-deficient Schizosaccharomyces pombe mutants undergo partial sporulation during nitrogen starvation. Microbiology 155:3816–3826. https://doi.org/10.1099/mic.0.034389-0
Liu J, Gao W, Pan Y, Liu G (2018) Metabolic engineering of Acremonium chrysogenum for improving cephalosporin C production independent of methionine stimulation. Microb Cell Fact 17(1):87. https://doi.org/10.1186/s12934-018-0936-5
Acknowledgements
We are grateful to Prof. Juan F. Martín (Universidad de León, Spain) for providing the plasmid pJL43-RNAi. We thank Prof. Seogchan Kang (Penn State University, USA) and Prof. Xingzhong Liu (Institute of Microbiology, CAS) for providing plasmid pAg1-H3. This work was supported by grants from the National Natural Science Foundation of China (31670091) and Construction of the Registry and Database of Bioparts for Synthetic Biology of the Chinese Academy of Sciences (ZSYS-016).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chen, C., He, J., Gao, W. et al. Identification and Characterization of an Autophagy-Related Gene Acatg12 in Acremonium chrysogenum. Curr Microbiol 76, 545–551 (2019). https://doi.org/10.1007/s00284-019-01650-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-019-01650-7