Abstract
The microbial diversity and the monthly fluctuations in Medicago sativa field soil in response to hydrogen gas were investigated. Illumina high-throughput sequencing was used to analyze the bacteria in raw and hydrogen-treated rhizosphere soil. Among the 18 soil samples, the abundance change of the predominant phyla Proteobacteria and Actinobacteria showed opposite trends. The diversity index analysis of the nine leguminous soil samples showed the highest diversity of the microbial community in July. In the other nine soil samples treated with hydrogen, the microbial diversity decreased and the diversity of soil microorganisms in September was higher than that in July, but not significantly so. The heat map analysis revealed that the microbial community composition of the soil samples was different before and after the hydrogen treatment. After the soil samples were treated with hydrogen, the dominant genera were Nocardioide, Pseudomonas, Janibacter, Microbacterium, Microvirga, Streptomyces, and Phenylobacterium in May; Bradyrhizobium, Haliangium, Sphingomonas, Blastocatella, Lysobacter, and Sphingopyxis in July; and Aeromicrobium, Pseudonocardia, Lentzea, and Skermanella in September. This study indicates that time and hydrogen gas have significant effects on the diversity of microbes in M. sativa rhizospheric soil.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Agriculture is of considerable economic significance for China. Inorganic fertilizers are used to obtain high yields, which cause serious damage to soil and health. It is generally thought that soil microorganisms play an important role in soil ecosystems, biogeochemical cycles, and ecological processes such as decomposition and mineralization of organic matters, absorption of mineral nutrients, and nitrogen cycling [11, 17, 27]. To assess the magnitude of changes in the bacterial community, it is necessary to understand the bacterial diversity and monthly changes in agricultural soil [3, 10].
Leguminous plants produce hydrogen gas as a by-product during the process of nitrogen fixation [23]. It has been estimated that 240,000 L of hydrogen gas were produced per hectare of legumes over the growing season [7]. For some legumes, hydrogen gas is assimilated because of the uptake hydrogenase (Hup); as a result, no hydrogen gas is released to the rhizosphere. However, for the other legumes, the uptake hydrogenase is absent; nevertheless, no hydrogen gas can be detected [34]. The previous work on this phenomenon has indicated that hydrogen gas is absorbed by soil microbes called hydrogen-oxidizing bacteria [15]. Dong proposed the “theory of hydrogen fertilizer”, that is, the hydrogen released from the process of nitrogen fixation can promote the growth of plants by changing the specific soil microbial populations in the rhizosphere [7]. Many studies have also highlighted that the soil microbial diversity after the hydrogen gas treatment is important to the protection of the ecosystem and the promotion of plant growth [7, 14, 34].
Soil microbes are featured by multiple varieties and large quantities, but a large number of non-culture microbes become the biggest obstacle for the traditional microbial ecological research. The new approaches to the study of microbial communities are advanced molecular technologies [16]. He [14] reported that the populations of Proteobacteria, particularly β- and γ-Proteobacteria, were stimulated in soil samples by a hydrogen treatment system using three methods. The result was similar to those of a study in which the populations of β- and δ-Proteobacteria were induced [34]. Bloem and his coworkers [2] found that microbial biomass in winter wheat fields showed peak levels in spring and autumn, which could indicate an increase in bacterial growth and activity. Smit et al. [29] demonstrated that the bacterial community in July differs from that in the other months using both cultivation-based and molecule-based methods. Considerable progress has been made in this field with respect to soil microbial diversity and monthly fluctuations; however, the experiments on the soil microbial diversity of leguminous plants in different months are rare.
In this study, Illumina high-throughput sequencing was used to investigate the microbial communities of the raw and hydrogen-treated rhizosphere soil of Medicago sativa collected separately in May, July, and September 2016 on the basis of the PCR of the bacterial 16S rDNA gene.
Materials and Methods
Soil Samples and DNA
The experiment sites were located in Tongchuan (Shaanxi, NW China). M. sativa rhizospheric soil samples were collected separately in May, July, and September 2016. The S-shaped sampling method was used to collect the rhizosphere soil samples. All of the rhizosphere soil samples were pooled together, mixed well, and then divided into three sets; the sets were kept in plastic bags at − 20 °C for future analysis. Half of each set of the three samples were treated with the hydrogen treatment equipment according to the method described by Dong et al. [6]. The classification of samples is presented in Table 1.
Further, 0.25 g of soil was filtered through a 2-mm-pore sieve and then used for DNA extraction with the PowerSoil® DNA Isolation Kit (Mo Bio Laboratories Inc., Carlsbad, CA, USA). The concentration and purity were measured on 1.8% agarose gels (Biowest agarose, Spain).
High-Throughput Sequencing
The bacterial 16S rDNA gene was amplified with a set of primers targeting the V3 + V4 region. The primers were 338F (5′-ACTCCTACGGGAGGCAGCA-3′) and 806R (5′-GGACTACHVGGGTWTCTAAT-3′) [33]. All the PCR reactions were carried out in a reaction system (50 µL) containing 0.2 µL of Q5 high-fidelity DNA polymerase (New England BioLabs), 10 µL of the buffer, and 10 µL of a high GC Enhancer. PCR amplification was conducted according to the following protocols: initial denaturation at 95 °C for 5 min, followed by 15 cycles. Each cycle involved denaturation at 95 °C for 1 min, annealing at 50 °C for 1 min, and extension at 72 °C for 1 min, with a final elongation step at 72 °C for 7 min. The purified PCR products were quantified with Nanodrop 2000 Illumina high-throughput sequencing conducted at Biomarker Technologies (Beijing, China).
Bioinformatic Analyses
The sequencing analysis was performed using Trimmomatic for sequence filtering. The Uchime algorithm [9] in the Mothur [26] software was used to remove the chimeric sequence to obtain a high-quality Tags sequence. The OTU was clustered by 97% similarity using the UCLUST algorithm [32] in the QIIME software and annotated according to the taxonomy databases of Silva (http://www.arb-silva.de/) (bacteria) and UNITE (http://unite.ut.ee/) (Fungi). On the basis of the abundance of each sample species in the OTU list, four common biodiversity indices were calculated using the Summary single command in the application software Mothur. The Bate diversity analysis was performed using the QIIME software to compare the differences in the diversity of species for various species.
Results
Microbial Diversity Index
Shannon and Simpson indices were used to estimate the diversity of microorganisms in the samples. The higher the Shannon index was, the higher was the diversity of the community; in contrast, the larger the Simpson index was, the lower was the diversity of the community [13]. The coverage of each sample was greater than 99%, which indicated that the information of microbial species was fully reflected (Table 2).
For the rhizosphere soil samples, the Shannon index of the “b” group was the largest, and the Simpson index was the smallest. The Shannon index of the “a” group was lower, and the Simpson index was higher. The results indicated that the diversity was the highest in July and the lowest in May.
Changes in the Shannon and Simpson indices proved that the microbial diversity of soil samples treated with hydrogen decreased. The diversity of soil microorganisms in September was slightly higher than that in July.
Analysis of Community Structure
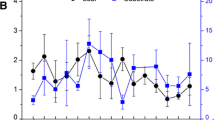
Ten phyla and undeclared groups were obtained from the 18 soil samples: Proteobacteria, Actinobacteria, Gemmatimonadetes, Acidobacteria, Chloroflexi, Bacteroidetes, Verrucomicrobia, Nitrospirae, Candidate-division TM7, and Cyanobacteria. The dominant phyla in the 18 soil samples were Actinobacteria and Proteobacteria. The abundance of Actinobacteria in the soil samples of leguminous plants decreased; the change trend of Proteobacteria was opposite to that of Actinobacteria; the abundance of Nitrospirae was the highest in July. The abundance trends of Actinobacteria and Proteobacteria in the soil samples treated with hydrogen gas were similar to those of the soil samples of legumes, and the abundance of Nitrospirae was the highest in July.
Compared with that of the soil samples of leguminous plants, the abundance of Actinobacteria increased by 29.5% in May and 20.3% in September and decreased by 14.6% in July; the abundance of Proteobacteria increased by 14.1% in May, 42.0% in July, and 9.8% in September, which implied that hydrogen gas had an enrichment effect on these microorganisms (Fig. 1).
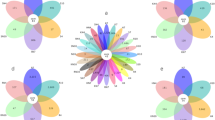
The changes in the bacterial diversity and the abundance of different soil samples are shown in the heat map (Fig. 2). The color gradient from blue to red indicates the relative abundance from low to high. Repeated groups of this experiment were clustered. From the degree of the color distribution, we inferred that the dominant genera had changed significantly after the hydrogen treatment. The genera that appeared in May were Nocardioide, Pseudomonas, Janibacter, Microbacterium, Microvirga, Streptomyces, and Phenylobacterium; in July, the genera that emerged were Bradyrhizobium, Haliangium, Sphingomonas, Blastocatella, Lysobacter, and Sphingopyxis; with Aeromicrobium, Pseudonocardia, Lentzea, and Skermanella presented in September.
Principal Component Analysis
In Fig. 3, the horizontal axis represents the first principal component (PC1), the vertical axis represents the second principal component (PC2), and the percentages represent the contribution of the principal component to the sample difference. The more similar the composition of the samples was, the closer was the distance between the samples in the PCA plots (Fig. 3). The figure shows that the contribution rates of PC1 and PC2 were 53.55% and 21.06%, respectively, which indicated that these two principal components were the major factors for explaining the structural differences in the microbial community structure. In the direction of PC1, the soil samples in May and the soil samples in July and September were significantly distributed in two regions, which indicated that time had a significant effect on the changes in the structure of the samples at the level of PC1. In the case of PC2, the raw soil samples could be clearly distinguished from the samples treated with hydrogen, which showed that the hydrogen gas influenced the microbial community structure changes at the PC2 level, particularly in May. This might be attributed to the fact that the hydrogen concentration was low (0.553 ppmv) in May and the soil microorganisms underwent significant changes after the hydrogen treatment.
Discussion
Soil microbes are of considerable significance in the process of energy flow and material cycling in ecosystems [31]. At present, one of the most extensive methods of studying microbial diversity is to amplify and sequence part of the 16S rDNA gene. V3–V4 is the most accurate variable region to recognize the genera. In this study, the temporal dynamics of the soil microbial diversity in legumes was analyzed by Illumina high-throughput sequencing. The time of soil samples collection in this survey was limited, but the sample collections were still representative. Therefore, the results of this study reflect to a certain extent the response of M. sativa soil samples to hydrogen.
Among the 18 soil samples, the predominant phyla were Proteobacteria and Actinobacteria, with the Proteobacteria as the most abundant bacterial group. Although the Actinobacteria dominated before, they were inhibited after the soil was treated with hydrogen, which was in agreement with many studies [14, 34]; these studies revealed that the populations of α- and β-Proteobacteria were stimulated by hydrogen exposure, but the populations of Actinobacteria were suppressed.
The bacterial diversity in the soil samples of different months was assessed using the microbial diversity index. The data suggested that the bacterial diversity in July was the highest. Studies have shown that natural and human factors [18, 19, 28] affect the composition and diversity of the soil microbial community. Excessive moisture in the soil causes an anaerobic environment, soil cooling, and nutrient loss, followed by crop rot, breathing difficulties, and negative nodulation effects. In the case of less soil moisture, the extension of plant roots is blocked, which reduces the chances of M. sativa nodule infection [20]. In July, the temperature and rainfall in Tongchuan were appropriate, which led to an increase in the number of M. sativa nodules. This resulted in the release of a considerable amount of hydrogen during nitrogen fixation and the emission of hydrogen gas obviously increased the number of hydrogen-oxidizing bacteria. The fine roots of legumes played a very important role in the underground ecology process. They were particularly sensitive to environmental factors and had a relatively short life [21]. Leguminous plant fine roots peak in the middle of July and then begin to die. The death of M. sativa fine roots may increase the number of microorganisms that can decompose fine roots. From July to September, the bacterial diversity decreases. The bacterial diversity in the rhizosphere changes during the growing stage of the crop. This change may be associated with the decrease in the root exudates. The competition among the microbial populations for root nutrition causes the disappearance of some populations [22, 25]. This in turn destroys the balance of the rhizosphere microbial populations and causes a decline of the rhizosphere microbial diversity. The microbial diversity of soil samples treated with hydrogen decreases; this is different from the result obtained by Zou [34]. The type of soil may be responsible for these different results; the soil samples used by Zou were from non-planted legumes, and the soil samples used in this study were from the soils with M. sativa. Studies have shown that soil samples after hydrogen treatment can promote the growth of some microorganisms and inhibit other populations of microorganisms [14, 34]. This result is consistent with the results obtained from the heat map, which shows that the microbial community composition of soil samples was different before and after the hydrogen treatment. Although the microbial diversity of soil samples treated with hydrogen in September was higher than that in July, the difference was not significant. In the raw soil of September, the hydrogen-oxidizing bacteria may be suppressed because of the decrease in hydrogen. This might be attributed to the fact that the expression of some genes using hydrogen is not active. When the raw soil in September was treated with hydrogen coming from the hydrogen treatment equipment, the suppressed hydrogen-oxidizing bacteria may start growing again after being stimulated by the presence of hydrogen. Zou [34] found that the level of expression of some genes in the hydrogen-treated soil samples is higher than that in the air-treated soil samples and the small subunit gene copy number of HoxK NiFe hydrogenase, the key enzyme in the hydrogen oxidation in soil, increases significantly after the hydrogen treatment.
Monthly changes in the microbial community diversity were also visualized by high-throughput sequencing. For the soil samples collected in May, July, and September, the most dominant bacterial species were Actinobacteria and Proteobacteria, which was similar to the data reported by He and other researchers [14, 34]. It is well known that soil microorganisms including Streptomyces sp.PCB7 [5], Streptomyces avermitilis [4], and Mycobacterium smegmatis [12] consume the atmospheric tropospheric trace hydrogen 0.553 ppmv [24] through high affinity [NiFe]-hydrogenase, which indicates that hydrogen peroxide is a common feature of Actinobacteria. In May, the hydrogen concentration is low (0.553 ppmv) in the rhizosphere because of the absence of root nodules. This might be the reason why the abundance of Actinobacteria was larger than that in July and September. After May, the abundance of Actinobacteria began to show a downward trend; the abundance change of Proteobacteria was opposite. Aerobic soil bacteria scavenging the hydrogen diffused from nitrogen-fixing nodules showed high km (app) (> 800 nM) [21]. For example, Paracoccus denitrificans with an enzyme having a Km of 1.1 µM may absorb relatively high concentrations of hydrogen gas mainly using the first type of membrane-bound [NiFe]-hydrogenase [1]. At the same time, Actinobacteria were unable to use hydrogen because of the high hydrogen threshold concentration. The trend of Actinobacteria and Proteobacteria appears to couple the oxidation of hydrogen to maintain the trace concentrations of atmospheric hydrogen. Whether there is a connection between the two phyla remains to be studied further.
In the cycle and transformation of ecosystem nutrients, nitrogen is one of the most active factors in the soil–plant system [8]. Nitrification is a key process in soil nitrogen conversion and plays an important role in nitrogen cycling. Nitrifier plays an irreplaceable role in the growth and development of plants and is the hub of rhizosphere micro-environment and soil material circulation and transformation. The change in the concentration of the Nitrifier and the ecological distribution of the populations affect the soil nutrient cycling, the concentration of the greenhouse gas N2O, and the function of the ecosystem directly or indirectly. Many studies have shown that soil temperature and moisture have a significant effect on nitrification [30]. In July, the mutual effects of rainfall and temperature promote the nitrification in the soil, which leads to the highest abundance of Nitrospirae.
In summary, the microbial diversity of M. sativa rhizospheric soil in July and September was higher than that of May, irrespective of the of hydrogen treatment. The structure of the soil samples changed significantly after the hydrogen treatment. This provided an appropriate amount of time for the isolation of bacteria for future research on hydrogen-oxidizing bacteria in the rhizosphere soil of M. sativa.
References
Berney M, Greening C, Hards K et al (2014) Three different [NiFe] hydrogenases confer metabolic flexibility in the obligate aerobe Mycobacterium smegmatis. Environ Microbiol 16:318–330. https://doi.org/10.1111/1462-2920.12320
Bloem J, Veninga M, Sheppard J (1995) Fully automated determination of soil bacterium numbers, cell volumes, and frequencies of dividing cells by confocal laser scanning microscopy and image analysis. Appl Environ Microbiol 61:926–936
Bossio DA, Scow KM, Gunapala N et al (1998) Determinants of soil microbial communities: effects of agricultural management, season, and soil type on phospholipid fatty acid profiles. Microbial Ecol 36:1–12. https://doi.org/10.1007/s002489900087
Constant P, Chowdhury SP, Pratscher J et al (2010) Streptomycetes contributing to atmospheric molecular hydrogen soil uptake are widespread and encode a putative high-affinity [NiFe]-hydrogenase. Environ Microbiol 12:821–829. https://doi.org/10.1111/j.1462-2920.2009.02130.x
Constant P, Poissant L, Villemur R (2008) Isolation of Streptomyces sp. PCB7, the first microorganism demonstrating high-affinity uptake of tropospheric H2. Isme J 2:1066. https://doi.org/10.1038/ismej.2008.59
Dong Z, Layzell DB (2001) H2 oxidation, O2 uptake and CO2 fixation in hydrogen treated soils. Plant Soil 229:1–12. https://doi.org/10.1023/A:1004810017490
Dong Z, Wu L, Kettlewell B et al (2003) Hydrogen fertilization of soils—is this a benefit of legumes in rotation? Plant Cell Environ 26:1875–1879. https://doi.org/10.1046/j.1365-3040.2003.01103.x
Duan Q, Liao TW (2013) A new age-based replenishment policy for supply chain inventory optimization of highly perishable products. Int J Pro Econ 145:658–671. https://doi.org/10.1016/j.ijpe.2013.05.020
Edgar RC, Haas BJ, Clemente JC et al (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. https://doi.org/10.1093/bioinformatics/btr381
Feest A, Madelin MF (1988) Seasonal population changes of myxomycetes and associated organisms in five non-woodland soils, and correlations between their numbers and soil characteristics. Fems Microbiol Lett 53:141–152. https://doi.org/10.1111/j.1574-6968.1988.tb02658.x
George E, Marschner H, Jakobsen I (1995) Role of arbuscular mycorrhizal fungi in uptake of phosphorus and nitrogen from soil. Crit Rev Biotechnol 15:257–270. https://doi.org/10.3109/07388559509147412
Greening C, Berney M, Hards K et al (2014) A soil actinobacterium scavenges atmospheric H2 using two membrane-associated, oxygen-dependent [NiFe] hydrogenases. Proc Natl Acad Sci USA 111:4257–4261. https://doi.org/10.1073/pnas.1320586111
Grice EA, Kong HH, Conlan S et al (2009) Topographical and temporal diversity of the human skin microbiome. Science 324:1190–1192. https://doi.org/10.1126/science.1171700
He X, Zou Y, Flynn B et al (2014) Assessment of soil bacterial community structure changes in response to hydrogen gas released by N2-fixing nodules. Int J Plant Soil Sci 3:47–61
Irvine P, Smith M, Dong Z (2004) Hydrogen fertilizer: bacteria or fungi? Acta Hort 631:239–242. https://doi.org/10.17660/ActaHortic.2004.631.30
Kirk JL, Beaudette LA, Hart M et al (2004) Methods for studying soil microbial diversity. J Microbiol Meth 58:169–188. https://doi.org/10.1016/j.mimet.2004.04.006
Lebbink G, Van Faassen HG, Van Ouwerkerk C et al (1994) The Dutch programme on soil ecology of arable farming systems: farm management monitoring programme and general results. Agr Ecosyst Environ 51:7–20. https://doi.org/10.1016/0167-8809(94)90032-9
Lipson DA, Schmidt SK (2004) Seasonal changes in an alpine soil bacterial community in the colorado rocky mountains. Appl Environ Micro 70:2867–2879. https://doi.org/10.1128/AEM.70.5.2867-2879.2004
Lupwayi NZ, Arshad MA, Rice WA, Clayton GW (2001) Bacterial diversity in water-stable aggregates of soils under conventional and zero tillage management. Appl Soil Ecol 16:251–261. https://doi.org/10.1016/S0929-1393(00)00123-2
Mouradi M, Farissi M, Bouizgaren A et al (2016) Effects of water deficit on growth, nodulation and physiological and biochemical processes in Medicago sativa-rhizobia symbiotic association. Arid Land Res Mana 30:193–208. https://doi.org/10.1080/15324982.2015.1073194
Noguchi K, Sakata T, Mizoguchi T et al (2005) Estimating the production and mortality of fine roots in a Japanese cedar (Cryptomeria japonica D. Don) plantation using a minirhizotron technique. J For Res Jpn 10:435–441. https://doi.org/10.1007/s10310-005-0163-x
Orlando J, Chavez M, Bravo L et al (2007) Effect of Colletia hystrix (Clos), a pioneer actinorhizal plant from the Chilean matorral, on the genetic and potential metabolic diversity of the soil bacterial community. Soil Biol Biochem 39:2769–2776. https://doi.org/10.1016/j.soilbio.2007.05.024
Osborne CA, Peoples MB, Janssen PH (2010) Detection of a reproducible, single-member shift in soil bacterial communities exposed to low levels of hydrogen. Appl Environ Microbiol 76:1471–1479. https://doi.org/10.1128/AEM.02072-09
Piche-Choquette S, Tremblay J, Tringe SG, Constant P (2016) H2-saturation of high affinity H2-oxidizing bacteria alters the ecological niche of soil microorganisms unevenly among taxonomic groups. Peer J 4:e1782. https://doi.org/10.7717/peerj.1782
Rumberger A, Merwin IA, Thies JE (2007) Microbial community development in the rhizosphere of apple trees at a replant disease site. Soil Biol Biochem 39:1645–1654. https://doi.org/10.1016/j.soilbio.2007.01.023
Schloss PD, Westcott SL, Ryabin T et al (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microb 75:7537. https://doi.org/10.1128/AEM.01541-09
Schnurer J, Clarholm M, Rosswall T (1986) Fungi, bacteria and protozoa in soil from four arable cropping systems. Biol Fert Soils 2:119–126. https://doi.org/10.1007/BF00257590
Schutter ME, Dick RP (2002) Microbial community profiles and activities among aggregates of winter fallow and cover-cropped soil. Soil Sci Soc Am J 66:142–153. https://doi.org/10.2136/sssaj2002.1420
Smit E, Leeflang P, Gommans S et al (2001) Diversity and seasonal fluctuations of the dominant members of the bacterial soil community in a wheat field as determined by cultivation and molecular methods. Appl Environ Microb 67:2284–2291. https://doi.org/10.1128/AEM.67.5.2284-2291.2001
Stark JM (1996) Modeling the temperature response of nitrification. Biogeochemistry 35:433–445. https://doi.org/10.1007/BF02183035
Stursová M, Zifčáková L, Leigh MB et al (2012) Cellulose utilization in forest litter and soil: identification of bacterial and fungal decomposers. Fems Microbiol Ecol 80:735–746
Wang Y, Sheng H, He Y et al (2012) Comparison of the levels of bacterial diversity in freshwater, intertidal wetland, and marine sediments by using millions of illumina tags. Appl Environ Microb 78:8264–8271. https://doi.org/10.1128/AEM.01821-12
Zhang X, Tian X, Ma L et al (2015) Biodiversity of the symbiotic bacteria associated with toxic marine dinoflagellate Alexandrium tamarense. Journal of Biosciences Medicines 03:23–28. https://doi.org/10.4236/jbm.2015.36004
Zou Y (2012) Effect of H2 on soil bacterial community structure and gene expression. Dissertation, Saint Mary’s University, Halifax, NS, Canada
Acknowledgements
This study is supported by the National Natural Science Foundation (Grant No. 41571243) and the project of young talents of science and technology association of Shaanxi Province (Project Number: 20160232). We thank Biomarker Technologies for their excellent technical assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Li, Z., Liu, X., Liu, R. et al. Insight into Bacterial Community Diversity and Monthly Fluctuations of Medicago sativa Rhizosphere Soil in Response to Hydrogen Gas Using Illumina High-Throughput Sequencing. Curr Microbiol 75, 1626–1633 (2018). https://doi.org/10.1007/s00284-018-1569-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-018-1569-y