Abstract
Endophytes are fungi and bacteria that inhabit plant tissues without causing disease. Endophytes have characteristics that are important for the health of the plant and have been isolated from several plants of economic and medicinal interest but rarely from ornamental plants. The current study isolates and identifies endophytic fungi from the leaves of Pachystachys lutea and evaluates the antagonistic activity of these endophytes as well as cellulase production by the endophytes. Fungi were isolated by fragmentation from surface-disinfected leaves and were identified by the sequencing of the ITS gene and the genes coding for EF 1-α and β-tubulin followed by multilocus sequence analysis. Molecular taxonomic analysis revealed that 78% of the identified fungi belonged to the genus Diaporthe. We also identified strains belonging to the genera Colletotrichum, Phyllosticta, Xylaria, Nemania, and Alternaria. Most of the strains tested were able to inhibit the growth of pathogenic fungi, especially PL09 (Diaporthe sp.), which inhibited the growth of Colletotrichum sp., and PL03 (Diaporthe sp.), which inhibited the growth of Fusarium oxysporum. The production of cellulase ranged from 0.87 to 1.60 μmol/min. Foliar endophytic fungal isolates from P. lutea showed promising results for the in vitro control of plant pathogens and for cellulase production. This paper is the first report on culturable endophytic fungi isolated from the ornamental plant P. lutea.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Society’s concerns about the impact of agriculture on the environment have led to the use of biological control as one of the most discussed alternatives to reduce the intensive use of pesticides to control plant diseases, pests, and weeds. Furthermore, endophytic microorganisms have proved to be effective in blocking the growth of various groups of plant pathogens, similar to biological control agents [52].
By occupying inter- and intra-cellular spaces [13, 19], endophytes are capable of colonizing plant tissues without causing apparent damage [16]. Endophytic colonization may be beneficial for host plants in many ways, including promotion of plant growth, production of phytohormones, nitrogen fixation [16, 47], biological pest control [39], increased resistance of plants to stress conditions [2, 39], and inhibition or reduction of phytopathogen growth by mycoparasitism, antibiosis, production of metabolites, competition for nutrients, or resistance induction in plants [39].
Most plants have endophytic microorganisms, which include fungi and bacteria. In general, there are dominant species, which are fairly frequent in a particular host, and secondary species, which are rarer. Consequently, plants have a characteristic endophytic microbiota that is likely to be important for plant protection and maintenance. Since host specificity requires co-adaptation between the host plant and its fungal partner, a mutual influence arising from an ancient cohabitation and co-evolution may be suggested [34]. Factors shaping plant–endophyte interactions include transmission mode, infection pattern, plant age, environmental conditions, and genetic background [2].
In fact, these microorganisms have been widely studied due to the interactions between endophytes, plants, and other microorganisms, which requires a great variety of substances, including enzymes [11] and a wide range of bioactive secondary metabolites [32, 42]. Endophytes have developed special mechanisms to penetrate host tissues and to reside in close association with each other in the host tissues. Endophytes possess exoenzymes necessary for host colonization, while they grow well in the apoplastic washing fluid of the host [10]. To compete with pathogenic fungi, endophytes also release hydrolytic enzymes, such as proteases, glucanases, and chitinases, which are capable of degrading the cell walls of the fungal hyphae [16].
There have been reports of cellulase, amylase, phenoloxidase, pectinase, xylanase, tyrosinase, gelatinase, and lipase production by endophytic fungi isolated from various plants [11, 16]. Enzymes obtained from microorganisms are used in the detergent, starch, fuel, food, beverage, textile, paper, leather, and several other industries. Indeed, endophytes are a potential source of enzymes of interest for industrial use [11].
Cellulases are the third most produced group of industrial enzymes worldwide due to their applications in cotton processing, paper recycling, and juice extraction and as enzymatic detergents and animal food additives [31]. Cellulases are classified into three groups: endoglucanases, which cleave the internal bonds of cellulosic fiber, generating oligosaccharides of different lengths and, therefore, new chain ends; exoglucanases, which are divided into cellobiohydrolases, which release cellobiose (glucose dimer) from the ends of cellulose, and glucanohydrolases, which are capable of directly releasing the glucose polymer; and β-glucosidase, which hydrolyses soluble oligosaccharides to glucose [22]. Endoglucanases are involved in plant colonization by endophytes [12].
Pachystachys lutea (Acanthaceae), popularly known as golden shrimp plant, is a subtropical shrub that is 90–120 cm tall and is commonly used as an ornamental plant [36]. This species is native to South America and was collected for the first time in the Amazon region; more precisely, this plant was first collected in the state of Acre, Brazil [56]. Studies of endophytes from ornamental plants are still rare and have mainly been conducted on Orchidaceae. In this family, endophytic fungi have been isolated from Lepanthes [5], Bletilla ochracea [51], and 54 other species; some of these endophytic fungi exhibited antimicrobial activity [54]. Fungi and endophytic bacteria have been isolated from Acanthaceae plants [33, 43].
These findings demonstrate the versatility of endophytic microorganisms isolated from different hosts as sources of information about host–plant interactions and biomolecule production. Therefore, the diverse endophytic fungi in P. lutea may be a rich source for the discovery of new, potentially bioactive compounds generated by these microorganisms. Current study isolates and identifies endophytic fungi in the leaves of P. lutea, and evaluates the antagonistic activity and cellulose production for them.
Materials and Methods
Isolation of Endophytic Fungi
P. lutea leaves were collected randomly from two specimens at the plant nursery of the Universidade Estadual de Maringá (23°24′S; 51°56′W). The rainfall during the month of collection was 151.3 mm, the average temperature was 24.1 °C, and the relative humidity was 63%. For the isolation of endophytes, the surfaces of 50 leaves were sterilized by immersion in 70% ethanol for 1 min, in 3% sodium hypochlorite for 4 min, and in 70% ethanol again for 30 s; then, the leaves were rinsed twice in autoclaved distilled water. The effectiveness of this method was verified by spreading 100 μL of the water from the final rinse on Petri dishes containing potato dextrose agar (PDA) medium (HiMedia®, Mumbai, India), pH 6.6, supplemented with tetracycline (Sigma, St. Louis, MO) (50 μg/mL in 50% ethanol) to prevent bacterial growth.
Then, 50 disinfected leaves were cut into small fragments measuring approximately 2 mm2, which were then deposited (five fragments per plate, on a total of 100 plates) on plates containing PDA supplemented with tetracycline. The plates were incubated at 28 °C for 7 days. The colonization frequency (CF) (%) was determined as the ratio of the number of fragments colonized by fungi and the total number of fragments × 100. For fungal purification, the fungal isolates were transferred to PDA plates and grown for 7 days. Then, the fragments (5 mm2) were crushed in 1 mL of an aqueous solution of 0.01% Tween 80, and a 100-μL aliquot of this solution was then spread on plates containing PDA and incubated for 24 h. Single colonies were immediately transferred to new plates with PDA and incubated for 7 days. If necessary, the process was repeated until pure colonies were obtained.
All strains were preserved according to Castellani’s method [9] and deposited in the fungal collection of the Laboratory of Microbial Biotechnology (LBIOMIC) at the Universidade Estadual de Maringá, Brazil.
Multilocus Sequence Analysis (MLSA)
Genomic DNA extraction, amplification, and phylogenetic analysis were performed as described by Polonio et al. [38]. For MLSA, partial sequences of the region ITS1–5.8S–ITS2 and of the genes coding for elongation factor 1-α (EF1α) and β-tubulin (TUB) were used. Primers used for amplification and the PCR (polymerase chain reaction) conditions are presented in Supplementary Table 1.
Sequences of the isolates were compared to the sequences deposited in GenBank (http://www.ncbi.nlm.nih.gov) using BLASTn and limiting the alignment with type–strain sequences, and the isolates were identified based on percentage identity and sequence coverage. Based on the available phylogenetic data on the TreeBASE database, other species were also selected (Study S13943; S15707; S14141; S14146; S141147; SN1525; ID:14671; http://www.treebase.org). The sequences were then rescued and aligned with the online interface MAFFT [25] (http://mafft.cbrc.jp/alignment/server/). After alignment, multigene assembly of sequences was performed using SequenceMatrix [53] (http://gaurav.github.io/taxondna/).
For phylogenetic analysis based on the maximum likelihood and Bayesian inference, MrModelTest v. 2.3 [30] was used to choose the best evolutionary model. The phylogenetic tree was constructed using MrBayes v. 3.2.5 [44], taking into consideration the parameters generated by MrModelTest, with Markov chain Monte Carlo (MCMC), which lasted until the average standard deviation of the split frequencies was below 0.01 (100.000 generations). The Bayesian probability was demonstrated on the nodes between each individual. The tree was edited with FigTree v. 1.4.2 [41].
All sequences in the current study were deposited in GenBank and are available under the Accession Numbers listed in Supplementary Table 2.
In Vitro Antagonist Activity of Endophytic Fungi
The antagonistic activity of the endophytic fungi was evaluated against the pathogenic fungi Fusarium oxysporum (ATCC2163, André Tosello Foundation, Campinas, SP, Brazil) and Colletotrichum sp. (CNPUV378, Embrapa Grape and Wine, Bento Gonçalves, RS, Brazil) using a paired-culture technique described by Campanile et al. [8], with modifications described by Polonio et al. [37].
Competitive interactions between the endophytes and pathogens were analyzed in vitro on the scale described by Badalyan et al. [4], with modifications, based on four types of interactions: A, B, C, and D, with C and D being divided into subcategories. The interaction types are as follows: A = inhibition of mycelial growth with contact; B = inhibition from a distance; C = endophytic growth on the pathogen without initial inhibition; CA1 and CA2 = partial and complete endophytic growth, respectively, on the pathogen after initial inhibition with mycelial contact; CB1 and CB2 = partial and complete endophytic growth, respectively, on the pathogen after initial inhibition from a distance; D = pathogen growth on the endophyte without initial inhibition; DA1 and DA2 = partial and complete pathogen growth on the endophyte after initial inhibition with mycelial contact, respectively.
Detection of Cellulase Production
To select endophytes with cellulolytic activity in the extracellular medium, all isolates were inoculated in minimal medium (6 g/L NaNO3, 5 g/L KCl, 1.5 g/L KH2PO4, 0.5 g/L MgSO4·7H2O, 0.01 g/L ZnSO4, 0.01 g/L FeSO4, and 15 g/L agar; pH 5.0) supplemented with 5 g/L yeast extract and 1% carboxymethylcellulose (CMC) (Sigma-Aldrich®, São Paulo, Brazil). Plates were incubated for 7 days at 28 °C and then visualized with 0.1% Congo red dye. Enzyme production was indicated by halo formation.
The fungi with the best results were studied by the cup–plate method as described by Souza et al. [48], with modifications.
Three mycelial discs were cultured in Mancini solution (2 g/L KH2PO4, 1 g/L (NH4)2SO4, 0.1 g/L MgSO4·7H2O, 0.9 g/L Na2HPO4·2H2O, and 1 g/L yeast extract in 1000 mL of distilled water; pH 5.0) with 0.5% CMC. The cultures were incubated for 7 days at 28 °C. Then, fungal mycelia were separated from the liquid media using sterilized gauze. The filtrate (enzyme crude extract) (50 µL) was inoculated into 6-mm-diameter holes removed from the central part of the culture media, CMC-agar (18 g/L agar, 10 g/L CMC, 0.1 M sodium acetate buffer; pH 5.0). A positive control experiment was performed using a commercially available enzyme (cellulase from Aspergillus niger, Sigma-Aldrich) at 1 mg/mL. The plates were incubated at 28 °C for 24 h and then stained with 0.1% Congo red dye. Tests were performed in triplicate. Enzyme production was indicated by halo formation; diameters were measured in mm. Mean rates were compared by the Scott–Knott test (p < 0.05) using the statistical program Sisvar 5.5 [14].
Determination of Endoglucanase Activity
Endoglucanase activity was determined as described by Ghose [19], adapted for CMC analysis. A 0.5 mL aliquot of crude enzymatic extract was pipetted into test tubes containing 0.5 mL of CMC solution (1% w/v) in sodium citrate buffer (50 mM; pH 4.8). Then, 1 mL of DNS (3,5-dinitrosalicylic acid) was added into the tubes after 15 and 30 min of incubation at 40 °C.
To prepare the DNS solution, 4.0 g of sodium hydroxide (Panreac®, Barcelona, Spain) was dissolved in 50 mL of distilled water, to which 2.5 g of DNS reagent (Dinâmica®, Diadema, Brazil) was added, and the mixture was homogenized. Meanwhile, 75 g of sodium potassium tartrate (KNaC4H4O6·4H2O) (Nuclear®, Diadema, Brazil) was dissolved in 125 mL of distilled water under constant agitation. The solutions were mixed with heating until completely dissolved. After cooling, the volume of the mixture was adjusted to 250 mL, and the mixture was stored at room temperature and protected from light.
After homogenization of the contents of the tubes, the tubes were heated for 5 min and then cooled. Then, 3 mL of distilled water was added to each tube, and the contents of each tube were subsequently homogenized by vortexing for 15 s. The absorbance of the reaction was measured at a wavelength of 540 nm by a spectrophotometer [29].
A blank comprising crude enzymatic extract, CMC solution (1% w/v) and DNS, which were immediately mixed, was measured for each sample. The absorbance of the blank of each sample was subtracted from the measurement of the corresponding test sample. The absorbance values were compared to a standard glucose curve.
Each sample was assayed in triplicate. One unit of endoglucanase activity was defined as the amount of enzyme required to liberate 1 µmol/min/mL glucose. Average rates were compared by the Scott–Knott test (p < 0.05) using the statistical program Sisvar 5.5 [14].
Results
Isolation
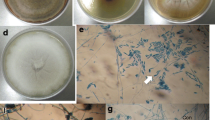
The isolation rate was 26.6%. From the obtained isolates, 85 were randomly selected and grouped into 23 morpho-groups based on macroscopic characteristics, including morphology and characteristics observed when the isolates were grown on PDA culture media: sporulation, mycelial properties, mycelial coloration, coloration of the reverse side of the Petri dish, pigmentation of the culture medium, and average diameter of the colony. One of the fungi from each morpho-group was randomly selected for molecular identification based on DNA sequencing (Supplementary Fig. 1).
Multilocus Sequence Analyses
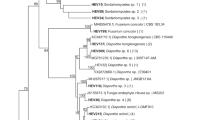
Multilocus sequence analyses (MLSAs) revealed that 78% of the isolates identified belonged to the genus Diaporthe. These isolates were categorized into four main groups (Supplementary Fig. 2).
The endophytic isolates PL01, PL64, and PL43, belonging to group A, were sub-clustered into sub-group 1A, exhibiting 100% Bayesian probability (BP) with Diaporthe anacardii CBS 720.97.
In sub-group 2B, with 100% BP, the isolates PL39, PL74, PL63, PL73, and PL66 were clustered with another three Diaporthe infecunda strains taking into account that only the ITS and EF1-α genes were sequenced for strains PL74, PL63, and PL66.
In sub-group 1C, with 87% BP, the isolates PL03 and PL09 were clustered with different species of Diaporthe; however, the identity of these isolates was confirmed to be Diaporthe only at the genus level.
Sub-group 2C, with isolates PL40, PL53, and PL47, was clustered with Diaporthe schini CBS 133181 and presented 100% BP. The strains were grouped based on the sequences of the ITS and EF1-α genes.
In group D, the isolates PL18, PL50, PL61, PL67, and PL71 were clustered with Diaporthe sp. 2 LGMF932 and Diaporthe mayteni CBS133185 with 100% BP. These isolates were classified taxonomically as Diaporthe sp.
For the remaining endophytic isolates, sequence comparison against the NCBI database did not yield DNA sequences with close identities to the DNA sequences of those endophytes; this observation was true when considering all three genes (ITS, EF1α, and TUB) together as well as when only considering two of the three sequenced genes. Therefore, these endophytic strains were clustered into four groups of different genera based on the partial information obtained from the NCBI database (Supplementary Fig. 3).
In sub-group 1.1, the endophyte PL75 was clustered with different species of Alternaria (based on the analysis of ITS and EF1-α genes). Consequently, molecular taxonomical classification was confirmed only at the genus level as Alternaria.
In sub-group 1.2, the endophyte PL49 was clustered with five strains of Phyllosticta capitalensis (1908) [synonyms: Phyllostictina pyriformis (1955), Guignardia mangiferae (1968), and Guignardia endophyllicola (2001)] and confirmed as P. capitalensis by analysis of ITS and EF1-α genes.
In sub-group 2.2, the endophyte PL36 was clustered with three strains of Xylaria berteroi with 99% BP based on the analysis of the ITS and TUB genes. When these two genes were considered, the endophyte PL27 (sub-group 2.4) was clustered with Nemania sp. FL0031 with 100% BP, while strain PL45 (sub-group 3.2) was clustered with three strains of Colletotrichum fructicola. Taxonomy was confirmed at the species level.
In Vitro Antagonist Activity of Endophytic Fungi
The analysis of variance of antagonistic activities measured for the endophytic fungi showed statistically significant differences between the endophytic strains tested against phytopathogens. These data are shown in Table 1 along with the percentages of inhibition and types of competitive interactions.
The analysis of the antagonism of 20 endophytic fungi against F. oxysporum yielded four statistical groups. The best performance was observed for strain PL03 (Diaporthe sp.), which exhibited a 64.62% inhibition rate (Table 1; Supplementary Fig. 4). Three types of interactions were reported: type CA1 (partial growth of the endophyte on the pathogen after initial inhibition with mycelial contact), 45%; type A (inhibition of mycelial growth with contact), 40%; and type DA1, 15% (Table 1).
The antagonistic activity of 20 endophytic fungi against Colletotrichum sp. was also evaluated. Four statistical groups were obtained for Colletotrichum sp., with the best inhibition rate obtained for endophyte PL09 (59.54%), identified as Diaporthe sp. Again, three types of interactions were reported: type A, 50%; type DA1, 40%; and type CA1, 10% (Table 1, Supplementary Fig. 4).
Enzyme Production
The statistical analysis of the results from the cup–plate test of cellulase production revealed significant differences between the control and the endophytes tested, with the formation of four separate groups (Table 2). The best results were reported for the endophyte PL01 (Diaporthe anacardii), with halos measuring 15.02 mm, followed by PL67 (Diaporthe sp.), with halos of 12.89 mm; PL36 (Xylaria berteroi); PL03 (Diaporthe sp.) and PL35, with halos ranging between 11.62 and 11.07 mm (Table 2, Supplementary Fig. 5).
Spectrophotometric analysis of cellulase production revealed results ranging between 0.87 and 1.60 μmol/min of endoglucanase with no significant difference (Table 2).
Discussion
The colonization frequency of P. lutea leaves was 26.6%. The colonization may even be as high as 100% in plants from tropical environments and between 1 and 40% in plants from northern and Arctic ecosystems [34, 39, 42].
In this study, the endophytic genus Diaporthe was the one most abundant (78%) in the foliar tissue of P. lutea. Similarly, several authors showed that one or two species of endophytes appear in high abundance in different host. Bogner et al. [6] reported that Fusarium oxysporum and F. solani were the most frequently observed endophytes in tomato roots from Kenya. Pamphile and Azevedo [34] showed that the most prevalent endophyte in close association with maize seeds from different genotypes was Fusarium verticillioides (= Fusarium moniliforme). These results are also consistent with other reports in which the genus Diaporthe is commonly isolated from several plants, such as Sapindus saponaria [18], Eichhornia azurea [1], Vitis labrusca [13], and Mikania glomerata [37], and from agricultural crops with high economic value, such as cocoa [45], coffee [55], soybean [26], and common bean [21]. Rhoden et al. [42] observed an 84.3% abundance of Diaporthe sp. endophytic isolates in the leaves of Trichilia elegans. Ferreira et al. [15] analyzed the diversity of endophytic fungi associated with Vellozia gigantean, and Diaporthe was seen to be the most abundant genus, with 70 endophytic isolates recovered from the leaves and roots.
The genus Diaporthe comprises close to 800 species and is distributed worldwide, with a great variety of hosts. Species of this genus may be phytopathogens, saprophytes, or endophytic symbionts [20, 46]. Some species of Diaporthe may be either pathogenic or harmless endophytes, depending on the type of host and the health of the host [20].
Other species found in the leaves of P. lutea were Alternaria sp., Phyllosticta capitalensis, Xylaria berteroi, Nemania sp., and Colletotrichum fructicola. The genera found in the current study may be associated with the isolation approach used, which generally excludes the detection of non-culturable species, as highlighted by Bogner et al. [6]. Consequently, the number of genera observed in P. lutea leaves in the current study may represent only a fraction of the total fungal diversity present. A major limitation of crop-dependent studies for unraveling the diversity of endophytes is the prevalence of fast-growing ubiquitous species. On the other hand, rare species with minor competitive strength and more specialized requirements may remain undiscovered [2]. It should be noted that the time between plant collection and endophyte isolation, the culture medium selected, the size of the plant fragment, and the growth conditions could also affect infection frequency, diversity, and species composition [3].
The endophytes that inhabit leaves are under a different set of selective pressures than those inhabiting other parts of the plant. Leaves have a short life span compared to stems and roots, but leaves are more biochemically dynamic, more sensitive to changes in environmental conditions, and more prone to damage caused by herbivores. The implications of these differences have not yet been explored, and they can be used to elucidate general patterns of the evolution of the life histories and specificity between the host and the endophyte [3]. Therefore, different environmental pressures can select endophytes with different physiological abilities, such as enzyme production and antagonistic activity against phytopathogens of economically important plant cultivars.
The current study demonstrated the ability of several endophytic fungi of P. lutea to reduce the mycelial growth of two phytopathogenic fungi. The endophytic fungi tested were efficient against F. oxysporum and Colletotrichum sp. Diseases caused by F. oxysporum, especially fusarium wilt, and crown and root rot in tomato plants (Solanum lycopersicum L., formerly, Lycopersicon esculentum Mill.), have been and continue to be among the most intensively studied plant diseases. These pathogens cause extensive damage to this important vegetable crop in fields and in greenhouses and continue to be the major limiting factors in tomato production [28]. Colletotrichum spp., an ascomycete, is a pathogenic fungus that causes anthracnose, a highly destructive plant disease, in a wide range of plants, including vegetables, fruits, legumes, and perennial trees, worldwide, with severe economic repercussions [23].
Endophytic fungi with the highest antagonistic activity against F. oxysporum and Colletotrichum sp. in the current study belong to the genus Diaporthe. This genus is known as a prolific source for the production of natural products, many of which possess antifungal activity. Tanneye et al. [50] characterized three dihydropyrones, phomopsolides A, B, and C, and a stable alpha-pyrone obtained from Diaporthe maritima with potent in vitro antifungal and antibiotic activities. Specian et al. [49] investigated secondary metabolites from endophytic D. helianthi by column chromatography and 1H and 13C nuclear magnetic resonance and identified the phenolic compound 2(-4 hydroxyphenyl)-ethanol (Tyrosol). These bioactive compounds, including Tyrosol, have antimicrobial effects against human and phytopathogenic bacteria. Several studies have demonstrated the ability of Diaporthe to inhibit other fungi and bacteria. The antifungal activity of D. citri, isolated from Mikania glomerata, was verified by in vitro tests against Fusarium solani and Didymella bryoniae [37]. Diaporthe phaseolorum isolated from Espeletia sp. demonstrated significant activity against the plant pathogen Phytophthora infestans; this organism contains a gene encoding an amylase, which was differentially expressed during the interaction and is possibly involved in this antagonism [40]. Compounds obtained from Diaporthe sp. have also demonstrated potent antifungal and antibacterial activity against gram-positive and gram-negative bacteria [27, 37, 50].
There has been increasing interest in endophytic microorganisms as biological control agents because they are capable of inhibiting plant pathogens via several mechanisms of direct or indirect inhibition. Endophytic organisms produce substances (antibiotics and enzymes) that directly inhibit pathogens or induce systemic resistance in the host; merely occupying space and mobilizing plant nutrients prevent infection by pathogens [17], with the added benefit of decreasing the levels of xenophobic chemical products that may damage the environment [24, 43]. The current study further highlights the well-documented role of endophytes as biocontrol agents and reinforces the observations from other studies that use the same methodologies employed in this study. Orlandelli et al. [32] evaluated the antifungal activity of endophytic fungi isolated from Piper hispidum against plant pathogens Alternaria alternata, Colletotrichum sp., Phyllosticta citricarpa, and Moniliophthora perniciosa and observed that, in some cases, the endophytes were more effective than commercial fungicides. This result highlights the capacity of these microorganisms to act as biological control agents.
One of the mechanisms by which endophytes compete with pathogenic fungi is by releasing hydrolytic enzymes. Panka et al. [35] confirmed that colonized plants invoke a faster defensive reaction against pathogens than non-colonized plants, thus increasing plant resistance against infection. Furthermore, microorganisms produce enzymes that facilitate their penetration into plants. Corrêa et al. [11] highlight endophytes as a new source of many interesting industrial enzymes, such as lipases, phytases, amylases, proteases, and cellulases. Moreover, the ability of endophytes to degrade the complex structure of lignocellulose makes them potentially useful in the exploitation of lignocellulosic biomass to produce ethanol and other value-added products.
Cellulase is involved in plant colonization by endophytes, and there is great interest in cellulase for industrial application [11, 12]. Cellulose is the most abundant renewable source of carbon on the earth’s crust; however, in nature, few microorganisms are able to degrade cellulose by cellulase production; filamentous fungi are the most efficient degraders of cellulose [22].
The most studied models of cellulase production are Trichoderma reesei, Aspergillus niger, Aspergillus nidulans, and Neurospora crassa. Endoglucanase production by T. reesei may vary between 0.2 and 137 μmol/min/mg, depending on test conditions and substrate used. Endoglucanase production by A. niger and A. nidulans varies between 1.0 and 59 μmol/min/mg and 65.66 μmol/min/mg, respectively [22]. Endoglucanase production by endophytes of P. lutea may be compared to endoglucanase production by T. reesei and A. niger, taking into account differences in test conditions and substrates. Laboratory production of cellulase may be optimized to increase the efficiency of P. lutea endophytes.
The current analysis reported the relationship between antifungal and enzymatic activities. The strains PL01 (Diaporthe anacardii), PL03 (Diaporthe sp.), PL36 (Xylaria berteroi), and PL35 (unidentified) showed significant antifungal activity against at least one of the pathogenic fungi tested, in addition to producing cellulase. These observations are not unexpected since the same fungal endophyte has often exhibited a positive behavior in different in vitro tests [32, 37]. The species Xylaria berteroi has been isolated as an endophyte from grapevines (Vitis labrusca L.) and has shown significant activity against F. oxysporum [7].
Ornamental plants have rarely been studied for their endophytic biodiversity. The current analysis shows, for the first time, that endophytic fungi from P. lutea are promising agents for the biological control of diseases of agronomic interest and are useful for cellulase production.
References
Almeida TT, Orlandelli RC, Azevedo JL, Pamphile JA (2015) Molecular characterization of the endophytic fungal community associated with Eichhornia azurea (Kunth) and Eichhornia crassipes (Mart.) (Pontederiaceae) native to the Upper Parana River floodplain, Brazil. Genet Mol Res 14(2):4920–4931. https://doi.org/10.4238/2015.May.11.25
Aly AH, Debbab A, Proksch P (2011) Fungal endophytes: unique plant inhabitants with great promises. Appl Microbiol Biotechnol 90(6):1829–1845. https://doi.org/10.1007/s00253-011-3270-y
Arnold AE (2007) Understanding the diversity of foliar endophytic fungi: progress, challenges, and frontiers. Fungal Biol 21:51–66. https://doi.org/10.1016/j.fbr.2007.05.003
Badalyan SM, Innocenti G, Garibyan NG (2002) Antagonistic activity of xylotrophic mushrooms against pathogenic fungi of cereals in dual culture. Phytopathol Mediterr 41:200–225
Bayman P, Lebro LL, Tremblay RL, Lodge DJ (1997) Variation in endophytic fungi from roots and leaves of Lepenthes (Orchidaceae). New Phytol 135:143–149
Bogner CW, Kariuki GM, Elashry A, Sichtermann G, Buch AK, Mishra B, Thines M, Grundler FMW, Schouten A (2016) Fungal root endophytes of tomato from Kenya and their nematode biocontrol potential. Mycol Progress 15:30. https://doi.org/10.1007/s11557-016-1169-9
Brum MCP, Araújo WL, Maki CS, Azevedo JL (2012) Endophytic fungi from Vitis labrusca L. (Niagara Rosada) and its potential for the biological control of Fusarium oxysporum. Genet Mol Res 11(4):4187–4197. https://doi.org/10.4238/2012
Campanile G, Ruscelli A, Luisi N (2007) Antagonistc activity of endophytic fungi towards Diplodia corticola assessed by in vitro and in planta test. Eur J Plant Pathol 117:237–246. https://doi.org/10.1007/s10658-006-9089-1
Castellani A (1967) Maintenance and cultivation of common pathogenic fungi in distilled water. J Trop Med Hyg 42:181–184
Chandra S (2012) Endophytic fungi: novel sources of anticancer lead molecules. Appl Microbiol Biotechnol 95:47–59. https://doi.org/10.1007/s00253-012-4128-7
Corrêa RCG, Rhoden SA, Mota TR, Azevedo JL, Pamphile JA, Souza CGM et al (2014) Endophytic fungi: expanding the arsenal of industrial enzyme producers. J Ind Microbiol Biotechnol 41(10):1467–1478. https://doi.org/10.1007/s10295-014-1496-2
Fan X, Yang R, Qiu S, Cai X, Zou H, Hu F (2016) The endo-β-1,4-glucanase of Bacillus amyloliquefaciens Is required for optimum endophytic colonization of plants. J Microbiol Biotechnol 26(5):946–952. https://doi.org/10.4014/jmb.1512.12055
Felber AC, Orlandelli RC, Rhoden SA, Garcia A, Costa AT, Azevedo JL et al (2015) Bioprospecting foliar endophytic fungi of Vitis labrusca Linnaeus, Bordô and Concord cv. Ann Microbiol 66:765–775. https://doi.org/10.1007/s13213-015-1162-6
Ferreira DF (2008) SISVAR: um programa para análise e ensino de estatística. Rev Científica Symp 6:36–41
Ferreira MC, Cantrell CL, Wedge DE, Gonçalves VN, Jacob MR, Khan S et al (2017) Diversity of the endophytic fungi associated with the ancient and narrowly endemic neotropical plant Vellozia gigantean from the endangered Brazilian rupestrian grasslands. Biochem Syst Ecol 71:163–169. https://doi.org/10.1016/j.bse.2017.02.006
Fouda AH, Hassan SE, Eid AM, Ewais EE (2015) Biotechnological applications of fungal endophytes associated with medicinal plant Asclepias sinaica (Bioss.). Ann Agric Sci 60:95–104. https://doi.org/10.1016/j.aoas.2015.04.001
Gao F, Dai C, Liu X (2010) Mechanisms of fungal endophytes in plant protection against pathogens. Afr J Microbiol Res 4(13):1346–1351
Garcia A, Rhoden SA, Rubin Filho CJ, Nakamura CV, Pamphile JA (2012) Diversity of foliar endophytic fungi from the medicinal plant Sapindus saponaria L. and their localization by scanning electron microscopy. Biol Res 45:139–148. https://doi.org/10.4067/S0716-97602012000200006
Ghose TK (1987) Measurement of cellulase activities. Pure Appl Chem 59:257–268
Gomes RR, Glienke C, Videira SIR, Lombard L, Groenewald JZ, Crous PW (2013) Diaporthe: a genus of endophytic, saprobic and plant pathogenic fungi. Persoonia 31:1–41. https://doi.org/10.3767/003158513X666844
Gonzaga LL, Costa LE, Santos TT, Araujo EF, Queiroz MV (2014) Endophytic fungi from the genus Colletotrichum are abundant in the Phaseolus vulgaris and have high genetic diversity. J Appl Microbiol 118:485–496. https://doi.org/10.1111/jam.12696
Gutiérrez-Rojas I, Moreno-Sarmientoa N, Montoya D (2015) Mecanismos y regulación de la hidrólisis enzimática de celulosa en hongos filamentosos: casos clásicos y nuevos modelos. Rev Iberoam Micol 32:1–12. https://doi.org/10.1016/j.riam.2013.10.009
Han JH, Chon JK, Ahn JH, Choi IY, Lee YH, Kim KS (2016) Whole genome sequence and genome annotation of Colletotrichum acutatum, causal agent of anthracnose in pepper plants in South Korea. Genom Data 8:45–46. https://doi.org/10.1016/j.gdata.2016.03.007
Heydari A, Pessarakli M (2010) A review on biological control of fungal plant pathogens using microbial antagonists. J Biol Sci 10:273–290. https://doi.org/10.3923/jbs.2010.273.290
Katoh K, Toh H (2008) Recent developments in the MAFFT multiple sequence alignment program. Brief Bioinform 9(4):286–298. https://doi.org/10.1093/bib/bbn013
Leite TS, Cnossen-Fassoni A, Pereira OL, Mizubuti ESG, Araujo EF, Queiroz MV (2013) Novel and highly diverse fungal endophytes in soybean revealed by the consortium of two different techniques. J Microbiol 51:56–69. https://doi.org/10.1007/s12275-013-2356-x
Li G, Kusari S, Kusari P, Kayser O, Spiteller M (2015) Endophytic Diaporthe sp. LG23 produces a potent antibacterial tetracyclic triterpenoid. J Nat Prod 78(8):2128 – 2132. https://doi.org/10.1021/acs.jnatprod.5b00170
McGovern RJ (2015) Management of tomato diseases caused by Fusarium oxysporum. Crop Prot 73:78–92. https://doi.org/10.1016/j.cropro.2015.02.021
Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal Chem 31:426–428. https://doi.org/10.1021/ac60147a030
Nylander JAA (2004) MrModeltest v2. Program distributed by the author. Evolutionary Biology Centre, Uppsala University, Uppsala
Onofre SB, Mattiello SP, Silva GC, Groth D, Malagi I (2013) Production of cellulases by the endophytic fungus Fusarium oxysporum.. Journal of Microbiology Research 3(4):131–134. https://doi.org/10.5923/j.microbiology.20130304.01
Orlandelli RC, Almeida TT, Alberto RN, Polonio JC, Azevedo JL, Pamphile J (2015) Antifungal and proteolytic activities of endophytic fungi isolated from Piper hispidum Sw. Braz J Microbiol 46(2):359–366. https://doi.org/10.1590/S1517-838246220131042
Pal A, Paul AK (2013) Bacterial endophytes of the medicinal herb Hygrophila spinosa T. Anders and their antimicrobial activity. Br J Pharm Res 3:795–806
Pamphile JA, Azevedo JL (2002) Molecular characterization of endophytic strains of Fusarium verticillioides (= Fusarium moniliforme) from maize (Zea mays. L). World J Microb Biotechnol 18:391–396. https://doi.org/10.1023/A:1015507008786
Panka D, Piesik D, Jeske M, Baturo-Ciesniewska A (2013) Production of phenolics and the emission of volatile organic compounds by perennial ryegrass (Lolium perenne L.)/Neotyphodium lolii association as a response to infection by Fusarium poae. J Plant Physiol 170(11):1010–1019. https://doi.org/10.1016/j.jplph.2013.02.009
Paulsen E, Andersen SL, Andersen KE (2009) Occupational contact dermatitis from golden shrimp plant (Pachystachys lutea). Contact Dermatitis 60(5):293–294. https://doi.org/10.1111/j.1600-0536.2009.01531.x
Polonio JC, Almeida TT, Garcia A, Mariucci GE, Azevedo JL, Rhoden SA et al (2015) Biotechnological prospecting of foliar endophytic fungi of guaco (Mikania glomerata Spreng.) with antibacterial and antagonistic activity against phytopathogens. Genet Mol Res 14(3):7297–7309. https://doi.org/10.4238/2015.July.3.5
Polonio JC, Ribeiro MAS, Rhoden SA, Sarragiotto MH, Azevedo JL, Pamphile JA (2016) 3-Nitropropionic acid production by the endophytic Diaporthe citri: Molecular taxonomy, chemical characterization, and quantification under pH variation. Fungal Biol 120:1600–1608. https://doi.org/10.1016/j.funbio.2016.08.006
Porras-Alfaro A, Bayman P (2011) Hidden fungi, emergent properties: endophytes and microbiomes. Annu Rev Phytopathol 49:291–315. https://doi.org/10.1146/annurev-phyto-080508-081831
Prada H, Ávila L, Sierra R, Bernal A, Restrepo S (2009) Caracterización morfológica y molecular del antagonismo entre el endofito Diaporthe sp. aislado de frailejón (Espeletia sp.) y el fitopatógeno Phytophthora infestans. Rev Iberoam Micol 26:198–201. https://doi.org/10.1016/j.riam.2009.01.002
Rambaut A (2009) FigTree v1. 3.1: Tree figure drawing tool. http://tree.bio.ed.ac.uk/software/figtree
Rhoden SA, Garcia A, Rubin Filho CJ, Azevedo JL, Pamphile JA (2012) Phylogenetic diversity of endophytic leaf fungus isolates from the medicinal tree Trichilia elegans (Meliaceae). Genet Mol Res 11(3):2513–2522. https://doi.org/10.4238/2012.June.15.8
Romeralo C, Santamaria O, Pando V, Diez JJ (2015) Fungal endophytes reduce necrosis length produced by Gremmeniella abietina in Pinus halepensis seedlings. Biol Control 80:30–39. https://doi.org/10.1016/j.biocontrol.2014.09.010
Ronquist F, Teslenko M, Van Der Mark P, Ayres DL, Darling A, Höhna S, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61(3):539–542. https://doi.org/10.1093/sysbio/sys029
Rubini MR, Silva-Ribeiro RT, Pomella AWV, Maki CS, Araujo WL, Santos DR, Azevedo JL (2005) Diversity of endophytic fungal community of cacao (Theobroma cacao L.) and biological control of Crinipellis perniciosa, causal agent of Witches’ Broom Disease. Int J Biol Sci 1:24–33
Santos TT, Leite TS, Queiroz CB, Araújo EF, Pereira OL, Queiroz MV (2016) High genetic variability in endophytic fungi from the genus Diaporthe isolated from common bean (Phaseolus vulgaris L.). J Appl Microbiol 120:388–401. https://doi.org/10.1111/jam.12985
Santoyoa G, Moreno-Hagelsieb G, Orozco-Mosqueda MC, Glick BR (2016) Plant growth-promoting bacterial endophytes. Microbiol Res 183:92–99. https://doi.org/10.1016/j.micres.2015.11.008
Souza HQ, Oliveira LA, Andrade JS (2008) Seleção de Basidiomycetes da Amazônia para produção de enzimas de interesse biotecnológico. Ciência Tecnol Alimentos 28:116–124
Specian V, Sarragiotto MH, Pamphile JA, Clemente E (2012) Chemical characterization of bioactive compounds from the endophytic fungus Diaporthe helianthi isolated from Luehea divaricata. Braz J Microbiol 43(3):1174–1182. https://doi.org/10.1590/S1517-838220120003000045
Tanney JB, McMullin DR, Green BD, Miller JD, Seifert KA (2016) Production of antifungal and antiinsectan metabolites by the Picea endophyte Diaporthe maritima sp. nov. Fungal Biol. https://doi.org/10.1016/j.funbio.2016.05.007
Tao G, Liu ZY, Hyde KD, Liu XZ, Yu ZN (2008) Whole rDNA analysis reveals novel and endophytic fungi in Bletilla ochracea (Orchidaceae). Fungal Divers 33:101–122
Terhonen E, Sipari N, Asiegbu FO (2016) Inhibition of phytopathogens by fungal root endophytes of Norway Spruce. Biol Control 99:53–63. https://doi.org/10.1016/j.biocontrol.2016.04.006
Vaidya G, Lohman DJ, Meier R (2011) SequenceMatrix: concatenation software for the fast assembly of multi-gene datasets with character set and codon information. Cladistics 27:171–180. https://doi.org/10.1111/j.1096-0031.2010.00329.x
Vaz AB, Mota RC, Bomfim MR, Vieira ML, Zani CL, Rosa CA et al (2009) Antimicrobial activity of endophytic fungi associated with Orchidaceae in Brazil. Can J Microbiol 55(12):1381–1391. https://doi.org/10.1139/W09-101
Vega FE, Simpkins A, Aime MC, Posada F, Peterson SW, Rehner SA, Infante F, Castillo A et al (2010) Fungal endophyte diversity in coffee plants from Colombia, Hawaii, Mexico and Puerto Rico. Fungal Ecol 3:122–138. https://doi.org/10.1016/j.funeco.2009.07.002
Wasshausen DC (1986) The systematics of the genus Pachystachys (Acanthaceae) Proc. Biol Soc Washington 99:162–163
Acknowledgements
The authors would like to thank CAPES (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior) for the Master’s scholarship and CAPES/PNPD-UEM post-doctoral scholarship and CNPq—Conselho Nacional de Desenvolvimento Científico e Tecnológico—(311534/2014-7; 447265/2014-8) and the Fundação Araucária (276/2014) for financial support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
da Silva Ribeiro, A., Polonio, J.C., Costa, A.T. et al. Bioprospection of Culturable Endophytic Fungi Associated with the Ornamental Plant Pachystachys lutea. Curr Microbiol 75, 588–596 (2018). https://doi.org/10.1007/s00284-017-1421-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-017-1421-9