Abstract
Saline–alkali soil can inhibit the growth of crops as a consequence of cellular damage through oxidation of lipids and proteins and degradation of nucleic acids, ultimately leading to cell death. The bacterial community composition and diversity in saline–alkali soil across different land uses, such as agricultural land, forest land, and grassland, were evaluated using high-throughput sequencing of the bacterial 16S rRNA gene. Significant differences in the soil physicochemical characteristics and bacterial community among different land uses were observed in this study. The soil pH value and electrical conductivity were much higher in grassland soil than in agricultural and forest soils. There were high proportions of Actinobacteria and Proteobacteria (phyla) in agricultural and forest soils, while Actinobacteria, Bacteroidetes, Gemmatimonadetes were more predominant in grassland soil. The relative abundance of dominant taxa exhibited a highly significant correlation with soil pH, water content, EC, and organic matter. The percentage of species that are shared among the different soil samples ranged from 5.3 to 30.5%. The haloalkaliphilic Actinobacterial genus Nitriliruptor was detected in grassland but not in areas with other types of land use. Results of both heatmap and principal component analysis (PCA) indicated that the soil properties and bacterial communities in the areas studied have been greatly influenced by long-term land use by different management.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Soil plays a vital role in maintaining the balance of Earth’s biosphere [1]. However, soil quality has been degraded seriously because of the improper management and use of land resources, leaving the global biogeochemical cycle changed. Soil microorganisms, including protozoa, fungi, bacteria, and archaea, play an important role in biogeochemical cycle, and are an important component in soil [2]. They hold dominant position in nutrient cycles, ensuring system stability, anti-jamming capability, and sustainable development of soil [3]. As sensitive indicators, changes in bacterial diversity and community structure in soil can rapidly reflect the quality and health of soil ecosystems.
The trajectory of land use can change land cover and plant litter on the soil surface, and affect many ecological processes, such as biodiversity, surface runoff, soil erosion, and soil environment. Recently, the influence of different land uses on soil microbiological indicators has been reported both in China and abroad [4, 5]. Bossio et al. [6] and Lagerlöf et al. [7] reported on soil microbial communities in different sites by different land uses (wooded and agricultural soils) in Kenya. Li et al. [8] studied the changes of soil organic carbon and the functional diversity of microbial communities experiencing different histories of land use (agricultural and forest systems) in a subtropical zone. All of these results showed that the different types of land use have a significant impact on soil physicochemical and biological properties.
Saline–alkali soil can inhibit the growth of crops as a consequence of cellular damage through oxidation of lipids and proteins and degradation of nucleic acids, ultimately leading to cell death [9, 10]. Currently, the total area occupied by saline–alkali soil is increasing in the world [11]. Therefore, it is an urgent issue to improve the saline–alkali soil by different land uses. However, none of the researches to date have been reported on the soil biodiversity affected by different land uses in saline–alkali sites. Here, pyrosequencing techniques were used to (1) determine the bacterial community structure and diversity in saline–alkali soil, (2) evaluate the influence on bacterial diversity of different land uses in this context and compare the differences and similarities of the bacterial communities in different land use systems.
Materials and Methods
Site Description and Soil Sampling
The study site is located in Daqing, China, at 45°46′–46°55′ north between 124°19′ and 125°12′ longitude, and the elevation in 640 m, where the soil type is primarily saline–alkali soil. This area remains with the continental monsoon climate, where the mean annual rainfall is 500 mm and the mean annual temperature is 4.2 °C [12].
In the present study, the six soil sites were selected, each covered with different vegetation for more than 20 years, including (i) two forest plantations sites formed by human (ZZS, YZY); (ii) two agricultural sites (DDN, YMN); and (iii) two meadow sites (JBT1, JBT2). The locations of the sampling sites and their soil properties are shown in Figure S1 and Table 1, respectively. The soil samples were collected at 20-cm depths using the five-point sampling method of Wakelin et al. [13] in three replications. From these sites, a total of 18 composite samples were collected. Soil samples were transported to the laboratory under ice and half of each sample air-dried for soil physicochemical analysis.
Soil Physicochemical Property Determination
Soil physicochemical parameters (soil water content, soil pH, electric conductivity (EC) of soil leaching solution, total nitrogen and total phosphorus, and organic content) were measured as described by Schinner et al. [14] with three replications for each treatment.
DNA Extraction, Amplification, and Pyrosequencing of Bacterial 16S rRNA Genes
Genomic DNA was extracted in triplicate from 500 mg subsample of the composite soil samples using E.Z.N.A. Soil DNA Kit (OMEGA, USA) according to the manufacture’s protocol. The DNA was extracted in each soil sample and pooled together, and then stored in −20 °C for downstream manipulation. Bacterial 16 S rRNA genes were amplified with the following universal primer set: 28F (5′-GAGTTTGATCNTGGCTCAG-3′) and 519R (5′-GTNTTACNGCGGCKGCTG-3′). PCR reactions were performed in 20 μL volumes containing 4 μL 5× FastPfu Buffer, 2 μL 2.5 mM dNTPs, 0.8 μL primers, 10 ng pooled DNA, and 1 U polymerase. The reaction conditions were programmed to predenaturing at 94 °C for 2 min, 25 cycles of 30 s at 94 °C, 30 s at 55 °C, 1 min at 72 °C, and a final cycle of 5 min at 72 °C. The PCR amplicons were checked by electrophoresis in a 2% agarose gel and then purified using a DNA gel extraction kit (OMEGA, USA). The 200 ng purified products were subjected to 454 pyrosequencing on the Roche 454 GS FLX Titanium platform (Roche 454 Life Sciences, Branford, CT, USA) at Majorbio Bio Technology Co. Ltd., Shanghai, China.
Statistical and Bioinformatics Analysis
Data for soil physicochemical characteristics were analyzed using the SPSS 17.0 software package. All values were presented as the means ± standard deviation (mean ± SD). One-way analysis of variance (ANOVA) was used to determine the minimum significant difference (P < 0.05) among different land uses.
To improve the quality of pyrosequencing data and eliminate the effect of random sequencing errors, we deleted some defective data from the libraries according to the protocol of Liu et al. [15], including (i) reads with ambiguous bases, (ii) more than six repeated single bases, and (iii) reads shorter than 150 bp. The sequences obtained from pyrosequencing were clustered into operational taxonomic units (OTUs), which were defined as having 97% sequence similarity. Taxonomic classification down to the phylum, class, and genus level was carried out by matching the reads against sequences using the Ribosomal Database Project (RDP) [16]. Based on OTU data, we calculated rarefaction curves as well as Shannon and coverage indices. ACE and Chao1 indices were calculated according to Chao, Bunge [17]. A principal component analysis (PCA) was carried out based on weighted UniFrac distance. In addition, a heatmap and Venn diagram were generated with R-package plots (http://www.r-project.org).
Results
Physicochemical Properties of Six Soil Samples
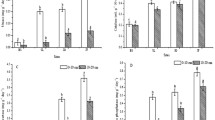
Soil physicochemical properties of the six samples are shown in Table 1. All the sampling sites were located in saline–alkali soil, formed under the influence of electrolytes resulting in a relatively high pH value (7.37–9.40) in all land use types. The soil water content ranged from 13.64 to 18.45%. EC had a huge variability with values ranging from 80.11 to 1225.17 uS/cm. Soil total N and total P varied from 0.70 to 1.98 and 0.19 to 0.31 mg/g, respectively. In addition, soil organic content showed a high variability, ranging from 0.28 to 5.34%. The ANOVA for soil properties showed that the significant differences were observed in all soil indices for comparisons between types of land use (Table 1).
Microbial Richness and Diversity
A total of 30,714 16S rRNA sequences were obtained from the six soil samples by pyrosequencing of bacterial 16S rRNA genes. The optimized sequence numbers for each sample ranged from 3529 to 8187 with an average of 5119. Among all the samples, a total of 10,559 OTUs were obtained and ranged from 852 in grassland soil (JBT1) and 2431 in forest soil (ZZS) (Table 2). The coverage of all samples was estimated and varied from 61 to 94% with the highest value in grassland soil JBT2 and lowest in forest soil (YZY). Among three kinds of land uses, the ACE and Chao1 in agricultural land (DDN, YMN) and forest land (ZZS, YZY) were higher than that in the grassland soil (JBT1, JBT2), which indicated that long-term vegetation coverage and human activity had changed the soil bacterial richness. In addition, the Shannon diversity indices showed the same trends as the ACE values. Therefore, the richness and diversity of soil bacterial communities showed strong variations among management types. Though all sequences were far from saturation based on the rarefaction curves, a distinct difference in bacterial diversity could be seen among them in OTUs (Figure S2).
Taxonomic Composition of Bacterial Communities among Different Land Uses
All valid sequences from the six sample libraries were classified from phylum to order based on RDP database. There were significant differences in bacterial community abundance at different phylogenetic levels (Tables S1–S3). Thirty-one phyla were found in the six samples, in which Acidobacteria, Actinobacteria, Bacteroidetes, Chloroflexi, Gemmatimonadetes, and Proteobacteria were the dominant bacterial taxa, accounting for over 80% of the reads in each sample (Fig. 1). Proportionally dominant sequences belonged to Acidobacteria, Actinobacteria, and Proteobacteria in agricultural and forest land. The most abundant sequences in the grassland were those related to Actinobacteria, Bacteroidetes, Gemmatimonadetes, and Proteobacteria. In addition, sequences related to Cyanobacteria, Elusimicrobia, and Deinococcus were detected with a relatively low abundance threshold (less than 1%) in all soil samples. Some groups were found only in particular land uses, e.g., Elusimicrobia and Fibrobacteres only in agricultural and forest land, Deinococcus-Thermus was found only in agricultural and grassland.
The presence of Chlorobi, Fibrobacteres, and Thermotogae were indicated in the soils of most of the systems evaluated, except for grassland soils. There was higher abundance of Gemmatimonadetes under JBT1 than in JBT2 and greater abundance in grass systems compared with the agricultural and forest systems. Nitrospira abundance was similar among the six soil samples. Besides, Planctomycetacia were comparable values in forest systems.
Soil pH and water content closely correlated with the abundance of mostly dominant bacterial phyla (Table 3). The relative abundance of Acidobacteria (r = 0.96), Chloroflexi (r = 0.94), Planctomycetes (r = 0.93), Proteobacteria (r = 0.88) in the analyzed soils significantly increased with higher water content values (P < 0.01), while Actinobacteria (r = −0.93, P < 0.01) and Gemmatimonadetes (r = −0.90, P < 0.01) were negatively correlated with soil water content. Soil pH, EC, and organic matter were also shown to be closely related to the abundance of five dominant bacterial phyla (Table 3). The abundance of Acidobacteria and Chloroflexi had a markedly negative relationship versus pH (r = −0.98, P < 0.01; r = −0.89, P < 0.01, respectively), EC (r = −0.95, P < 0.01; r = −0.89, P < 0.01, respectively), and organic matter (r = −0.98, P < 0.01; r = −0.92, P < 0.01; r = −0.90, P < 0.01, respectively); and the Gemmatimonadetes exhibited a highly significant positive correlation with pH (r = 0.96, P < 0.01), EC (r = 0.97, P < 0.01), and organic matter (r = 0.95, P < 0.01). The abundance of Bacteroidetes had no significant relationship with soil properties. Additionally, the soil TN and TP were found to have no significant correlations with the abundance of all the dominant bacterial phyla.
Relationships of the Bacterial Communities among the Different Soil Samples
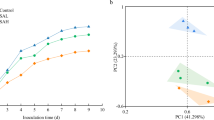
A principal component analysis was performed in relation to soil basic characteristics and bacterial community properties, using pyrosequencing data to reveal the relationships among the six samples at the genus level. The PCA score plot showed that JBT1 and JBT2 grouped to the right of the graph along the first principal component (PC1), while the other four samples closely clustered on the left of PC1, which represented 68.14% of the total variations (Fig. 2). The second principal component (PC2) accounted for 27.9% of the total variations. Overall, these two PCs were involved in 96.04% of the total variations, which could explain the most variations of all the bacterial communities.
The relationship of the bacterial community by different land uses at the genus level. Principal component 1 and 2 explained 68.14 and 27.9% of the total variations, respectively. Just based on PC1, six samples were divided into two groups, JBT1 and JBT2 were clustered together, and the other four samples were in the same group. Codes indicating the different land uses are given right of the graph. DDN, YMN: Agricultural land; ZZS, YZY: Forest land; JBT1, JBT2: Grassland
A Venn diagram was further used to find the relationships among the six samples in detail at the species level [18]. The diagram showed specific and common OTUs in all samples categorized by different land uses and management (Fig. 3). The percentage of species shared between the different soil samples ranged from 5.3 to 30.5%. The two grassland samples showed the lowest percentage of similarity in bacterial community versus the other samples (5.3–7.06%) (Fig. 3). We selectively analyzed and listed the top 20 predominant bacterial groups in the six soil samples (Table S4); 14 groups were unique to the soil from agricultural and forest systems, including Rhodospirillum, Bryobacter, Gaiella, Solirubrobacter, Patulibacter, Alcaligenes, Nitrosomonadaceae, Gemmata, Pseudonocardia, Blastococcus, Terrabacter, Luteimonas, Blastococcus, Massilia. Some bacteria were unique to grass systems: Halomonas, Echinicola, Planococcus, Nesterenkonia, Amycolicicoccus, Dietzia, Haloactinospora, Tepidamorphus. However, bacteria unique to other agricultural or forest lands are not listed due to very low content. In addition, areas with similar land uses with different vegetation possessed different bacterial abundance (Table S4).
Percentage of shared OTUs between the different soil samples at species level. The percentage is to the number of shared OTUs exiting in the two samples accounting for the ratio of all OTUs. Codes indicating the different land uses are given in the graph. DDN, YMN: Agricultural land; ZZS, YZY: Forest land; JBT1, JBT2: Grassland
A hierarchically clustered heatmap was also constructed at the genus level based on the top 100 most abundant bacterial community using R software. The result showed that the six soil samples were separated into two groups: one group contained the JBT1 and JBT2 libraries, and the other group was composed by agricultural (DDN and YMN) and forest soil samples (ZZS and YZY) (Fig. 4). The most abundant genus, Nitriliruptor in red color, was found in JBT1 and JBT2, accounting for 39.45 and 32.64% of sequences, respectively. The most abundant taxonomic groups in DDN, YMN, ZZS, and YZY libraries were Planctomycetaceae, Acidimicrobiaceae, Marmoricola, Chloracidobacterium, Arthrobacter, Solirubrobacter, Sphingomonas, Nocardioides, and Pseudomonas. The sequences belonging to Rhodococcus and Planctomycetes were detected in forest land but were absent or rarely detected in the other soil samples. The richness of the bacterial communities of the grassland was much lower than that of the agriculture and forest lands, in which the richness of JBT1 was clearly higher than that of JBT2. This factor underlies the obvious separation between JBT1 and JBT2 by Principal Component 2. The heatmap analyses (Fig. 4) agree with the results of the PCA (Fig. 2), with both analyses indicating that different land uses had a great effect on the soil characteristics and bacterial communities.
Hierarchically clustered heatmap of bacterial community distribution at the genus level categorized by different land uses (Note row represents the relative percentage of each bacterial genus, and column stands for different samples. The relative abundance for each bacterial genus was depicted by color intensity in the figure below. Codes indicating the different land uses are given below the graph. DDN, YMN: Agricultural land; ZZS, YZY: Forest land; JBT1, JBT2: Grassland)
Discussion
To the best of our knowledge, this is the first work in soil pyrosequencing to analyze the bacterial structure variation by different land management of the saline–alkali soil zone of Daqing, China. Acosta-Martinez et al. [19] found differences in bacterial diversity among the two non-disturbed grasslands and two agricultural lands affected by land management using the method of bacterial tag-encoded FLX amplicon pyrosequencing. Roesch et al. [20], using high-throughput DNA pyrosequencing, found that agricultural management could significantly influence the diversity of the soil bacteria and archaea, and the microbial diversity of the forest land was much higher than that of the agricultural land in phylum level. Thus, land use system was an important factor that affected the diversity of the soil bacterial communities.
Seven dominant bacterial phyla in all six soil samples included Acidobacteria, Actinobacteria, Bacteroidetes, Chloroflexi, Gemmatimonadetes, and Proteobacteria with the findings of Liu et al. [15]. Additionally, the relative abundance of Acidobacteria, Chloroflexi, Planctomycetes, and Proteobacteria was significantly positively correlated with soil water content, and negatively correlated with soil pH and organic matter (Table 3); Actinobacteria and Gemmatimonadetes produced contrary trends. These findings show that the distribution of bacterial communities in saline–alkali soils is affected by soil pH, soil moisture, and organic matter. Interestingly, the relative abundance of Chloroflexi, Gemmatimonadetes, and Planctomycetes from different soil samples in our study ranged from 12 to 20% (Table S1), which is ten times higher than that of the other research (<1%) [21, 22] and twice as much as that reported by Liu et al. [15] (nearly 5%). This difference could be caused by different land management despite the original soil type being saline-alkali.
Fierer, Jackson [23] reported that the soil with relatively low soil pH value might lead to lower microbial diversity than that with higher pH value. That result was the opposite of our finding that soil pH increased with the lower richness of the soil bacterial communities. This could be attributed to the fact that our sampling area includes the zonal saline–alkali soil originally with a much higher pH value, where long-term vegetation coverage and agricultural activities decreased the soil pH to nearly neutral values better supporting bacterial diversity (Tables 1, 2).
Sequences related to Bacteroidetes were detected in all soil samples, whatever land use system, thus pointing out to their vast distribution and adaptation to different conditions. This bacterial phylum was frequently encountered in agricultural systems; presumably their widespread distribution can be attributed to their ability to rapidly exploit bioavailable organic matter and colonize aggregates [19, 24]. Interestingly, we detected much higher abundance of Bacteroidetes in both agricultural lands (DDN) and grasslands (JBT1), implying some preference to degrade lignocellulose and cellulose of crops and grasses. This result agrees with studies by Chan et al. [25] and Acosta-Martinez et al. [19], who found higher abundance of Bacteroidetes in pasture and agricultural soil, respectively. Also, OTUs related to Actinobacterial order Gaiellales were the most abundant in agricultural and forest lands; this group is known to assimilate carbohydrates, organic acids, and amino acids [26, 27].
Based on the heatmap, an interesting feature of the obtained sequences was the detection of 32.64–39.45% of the total soil bacteria belonging to the Actinobacterial genus Nitriliruptor in grassland and not in the other types of land use. Species of Nitriliruptor exhibit a haloalkaliphilic capacity [28], presumably better adapted for thriving in the grasslands with high alkali environments (pH, 9.35–9.5) (Table 1 and Fig. 5). A significant finding is that the relative abundances of Nitrospira, Nocardioides, and Pseudomonas were much higher in soil of agricultural and forest lands than that of grasslands; these groups were previously reported as belonging to the nitrite-oxidizing bacterial group and key nitrifiers in natural ecosystems [29, 30]. Presumably, a large amount of added nitrogen fertilizer and intensified human activities in agricultural and forest lands would build up a selective pressure for these species. Together, these results indicated that different land use systems have greatly influenced the diversity and structure of the soil bacterial communities.
Conclusions
Overall, this study revealed significant differences in soil properties and bacterial community composition among three different land uses in the saline–alkali soil zone. The relative abundance of dominant taxa exhibited a highly significant correlation with soil pH, water content, EC, and organic matter. Additionally, the results showed that the soil bacterial communities respond to different land uses of saline–alkali soil, and the conversion of saline–alkali soil to forest or agricultural soil increased bacterial diversity.
References
Nesme T, Colomb B, Hinsinger P, Watson CA (2014) Soil phosphorus management in organic cropping systems: from current practices to avenues for a more efficient use of P resources. In: Organic farming, prototype for sustainable agricultures. Springer, Berlin, pp 23–45
Heijden MGAvd, Bardgett RD, Straalen NMv (2008) The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol Lett 11:296–310
Gyaneshwar P, Kumar GN, Parekh L, Poole P (2002) Role of soil microorganisms in improving P nutrition of plants. In: Food security in nutrient-stressed environments: exploiting plants’ genetic capabilities. Springer, Berlin, pp 133–143
Ding G-C, Piceno YM, Heuer H, Weinert N, Dohrmann AB, Carrillo A, Andersen GL, Castellanos T, Tebbe CC, Smalla K (2013) Changes of soil bacterial diversity as a consequence of agricultural land use in a semi-arid ecosystem. PLOS one 8(3):e59497
Bissett A, Richardson AE, Baker G, Thrall PH (2011) Long-term land use effects on soil microbial community structure and function. Appl Soil Ecol 51:66–78
Bossio DA, Girvan MS, Verchot L, Bullimore J, Borelli T, Albrecht A, Scow KM, Ball AS, Pretty JN, Osborn AM (2005) Soil microbial community response to land use change in an agricultural landscape of Western Kenya. Microb Ecol 49(1):50–62. doi:10.1007/s00248-003-0209-6
Lagerlöf J, Adolfsson L, Börjesson G, Ehlers K, Vinyoles GP, Sundh I (2014) Land-use intensification and agroforestry in the Kenyan highland: impacts on soil microbial community composition and functional capacity. Appl Soil Ecol 82:93–99
Li ZP, Wu XC, Chen BY (2007) Changes in transforamtion of soil organic carbon and functional diversity of soil microbial community under different land use patterns. Scienctia Agricultura Sinica 40 (8):1712–1721
Xun F, Xie B, Liu S, Guo C (2014) Effect of plant growth-promoting bacteria (PGPR) and arbuscular mycorrhizal fungi (AMF) inoculation on oats in saline-alkali soil contaminated by petroleum to enhance phytoremediation. Environ Sci Pollut Res 22(1):598–608
Kohler J, Hernández JA, Caravaca F, Roldán A (2009) Induction of antioxidant enzymes is involved in the greater effectiveness of a PGPR versus AM fungi with respect to increasing the tolerance of lettuce to severe salt stress. Environ Exp Bot 65(2–3):245–252
Wang X, Wang J, Liu H, Zou D, Zhao H (2013) Influence of natural saline-alkali stress on chlorophyll content and chloroplast ultrastructure of two contrasting rice (Oryza sativa L. japonica) cultivars. Aust J Crop Sci 7:289–292
Tang J, Wang L, Zhang S (2005) Investigating landscape pattern and its dynamics in Daqing, China. Int J Remote Sens 26(11):2259–2280
Wakelin SA, Barratt BI, Gerard E, Gregg AL, Brodie EL, Andersen GL, DeSantis TZ, Zhou J, He Z, Kowalchuk GA (2013) Shifts in the phylogenetic structure and functional capacity of soil microbial communities follow alteration of native tussock grassland ecosystems. Soil Biol Biochem 57:675–682
Schinner F, Ohlinger R, Kandeler E, Margesin R (1995) Methods in soil biology. Springer-Verlag, Berlin
Liu J, Sui Y, Yu Z, Shi Y, Chu H, Jin J, Liu X, Wang G (2014) High throughput sequencing analysis of biogeographical distribution of bacterial communities in the black soils of northeast China. Soil Biol Biochem 70:113–122
Maidak BL, Olsen GJ, Larsen N, Overbeek R, Mccaughey MJ, Woese CR (1997) The RDP (Ribosomal Database Project). Nucleic Acids Res 25(1):109–111
Chao A, Bunge J (2002) Estimating the number of species in a stochastic abundance model. Biometrics 58(3):531–539
Wu S, Wang G, Angert ER, Wang W, Li W, Zou H (2012) Composition, diversity, and origin of the bacterial community in grass carp intestine. PloS One 7(2):e30440
Acosta-Martinez V, Dowd S, Sun Y, Allen V (2008) Tag-encoded pyrosequencing analysis of bacterial diversity in a single soil type as affected by management and land use. Soil Biol Biochem 40(11):2762–2770
Roesch LF, Fulthorpe RR, Riva A, Casella G, Hadwin AK, Kent AD, Daroub SH, Canargo FA, Farmerie WG, Triplett EW (2007) Pyrosequencing enumerates and contrasts soil microbial diversity. ISME J 1(4):283–290
Lauber C, Hamady M, Knight R, Fierer N (2009) Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl Environ Microbiol 75(15):5111–5120
Chu HY, Fierer N, Lauber CL, Caporaso JG, Knight R, Grogan P (2010) Soil bacterial diversity in the Arctic is not fundamentally different from that found in other biomes. Environ Microbiol 12:2998–3006
Fierer N, Jackson RB (2006) The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci USA 103(3):626–631
Abell GCJ, Bowman JP (2005) Colonization and community dynamics of class Flavobacteria on diatom detritus in experimental mesocosms based on Southern Ocean seawater. FEMS Microbiol Ecol 53:379–391
Chan OC, Casper P, Qing SL, Li FZ, Fu Y, Dong YX, Ulrich A, Ming ZX (2008) Vegetation cover of forest, shrub and pasture strongly influences soil bacterial community structure as revealed by 16 S rRNA gene T-RFLP analysis. FEMS Microbiol Ecol 64:449–458
Albuquerque L, França L, Rainey FA, Schumann P, Nobre MF, da Costa MS (2011) Gaiella occulta gen. nov., sp. nov., a novel representative of a deep branching phylogenetic lineage within the class Actinobacteria and proposal of Gaiellaceae fam. nov. and Gaiellales ord. nov. Syst Appl Microbiol 34(8):595–599
Navarrete AA, Kuramae EE, M DH, Pijl AS, van Veen JA, Tsai SM (2012) Acidobacterial community responses to agricultural management of soybean in Amazon forest soils. FEMS Microbiol Ecol 83(3):607–621
Sorokin DY, van Pelt S, Tourova TP, Evtushenko LI (2009) Nitriliruptor alkaliphilus gen. nov., sp. nov., a deep-lineage haloalkaliphilic actinobacterium from soda lakes capable of growth on aliphatic nitriles, and proposal of Nitriliruptoraceae fam. nov. and Nitriliruptorales ord. nov. Int J Syst Evol Microbiol 59(2):248–253
Pester M, Maixner F, Berry D, Rattei T, Koch H, Lücker S, Nowka B, Richter A, Spieck E, Lebedeva E (2014) NxrB encoding the beta subunit of nitrite oxidoreductase as functional and phylogenetic marker for nitrite-oxidizing Nitrospira. Environ Microbiol 16(10):3055–3071
Gruber-Dorninger C, Pester M, Kitzinger K, Savio DF, Loy A, Rattei T, Wagner M, Daims H (2014) Functionally relevant diversity of closely related Nitrospira in activated sludge. ISME J 9(3):643–655
Acknowledgements
The authors are grateful for the support by the Project “948” of the Chinese National Forestry Bureau, Grant (No. 2008-4-34) and the Fundamental Research Funds for the Central Universities (2572014CA22).
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Peng, M., Jia, H. & Wang, Q. The Effect of Land Use on Bacterial Communities in Saline–Alkali Soil. Curr Microbiol 74, 325–333 (2017). https://doi.org/10.1007/s00284-017-1195-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-017-1195-0