Abstract
Understanding the transmission of the human microbiota from mother to child is of major importance. Although we are gaining knowledge using 16S rRNA gene analyses, the resolution of this gene is not sufficient to determine transmission patterns. We therefore developed an Illumina deep sequencing approach targeting the 16–23S rRNA Internal Transcribed Spacer (ITS) for high-resolution microbiota analyses. Using this approach, we analyzed the composition and potential mother to child transmission patterns of the microbiota (milk and stool) in a longitudinal cohort of 20 mother/child pairs. Our results show overlap in the infant stool microbiota with both mother’s milk and stool, and that the overlap with stool increases with age. We found an Operational Taxonomic Unit resembling Streptococcus gordonii as the most widespread colonizer of both mothers and their children. In conclusion, the increased resolution of 16–23S rRNA ITS deep sequencing revealed new knowledge about potential transmission patterns of human-associated bacteria.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Taxonomic assignments of complex microbial assemblages have until now mainly been performed using 16S rRNA gene analyses. A limitation with these analyses, however, is the lack of depth in taxonomic assignments due to the conserved nature of the 16S rRNA gene [14]. This limits the possibility of addressing questions about prevalence and transmission patterns of the microbiota.
The establishment of the human gut microbiota during infancy can be considered a succession process [2]. In this process, both mother’s milk and feces are proposed as important vectors for transmission of bacteria from mother to child [2, 3, 8, 9]. The transmission patterns and importance of feces and mother’s milk, however, have not yet been resolved using high-resolution analyses.
Using a Illumina 16–23S rRNA Internal Transcribed Spacer (ITS) region deep sequencing approach, the aim of our work was to determine the high-resolution overlap in the dominant microbiota in a cohort of mothers and their children [6]. The 16–23S rRNA ITS region has previously been extensively used for high-resolution analyses, utilizing a range of detection approaches [4, 5, 10, 24], including Illumina sequencing [18].
We present results suggesting Streptococcus gordonii as the most widespread colonizer of mothers and their children. Furthermore, our results support mother’s milk as an important source for bacterial transmission from mother to child.
Materials and Methods
Reference Genomes and Strains
We downloaded all 1838 bacterial genomes available in Genbank February 2014. The 16–23S ITS rRNA sequences flanked by the 16S IX-23S I primer pair [19] was exacted in silico from each genome allowing two mismatches in the both the forward and the reverse primes. These sequences were used as references for defining the 16–23S ITS rRNA OTU’s.
A collection of 89 bacterial 6 reference strains was selected to cover the main phyla and genera of the human-associated microbiota (the strains are described in Suppl Table 1). These strains were used as references for testing the coverage and reproducibility of the developed Illumina 16–23S rRNA ITS PCR. The coverage was tested on pure strains, while the reproducibly was tested on three strain mixtures. One high complex with PCR products from 83 strains mixed in equal amounts, one low complex consisting of 10 strains, and one middle complex with 10 strains in different portions.
Clinical Samples
The clinical samples represent a subset of the Pro-PACT study. This study represents an unselected cohort of healthy mothers and children from the Trondheim region, with nearly 100 % breastfeeding at 10 and 90 days [6]. From the study cohort, we selected 20 mother/child pairs delivered at term based on the criterion that they had a full set of samples (characteristics are shown in Table 1). Five samples were selected for each of the 20 mother/child pair. Milk samples were analyzed from 10 and 90 days after delivery for the mothers, in addition to stool samples from 90 days. For the children, stool samples were analyzed from 10 to 90 days after birth. All the samples were collected at home in sterile containers and immediately stored at −20 °C before transportation to centralized storage at −80 °C.
Sample Preparation and DNA Extraction
Fecal samples from the subjects were collected in Cary-Blair transport and holding medium (BD Diagnostics Sparks, MD 21152), while no buffers were added to the mother’s milk samples. The samples were frozen at −20 °C within 2 h from collection. The samples were then transferred to the lab and stored at −80 °C.
The solid material in 1 ml of the mother’s milk was precipitated by centrifugation at 20,000×g for 15 min prior to DNA extraction. The supernatants were discharged, and the solid material pellets were used for DNA extraction. DNA from both the mother’s milk and feces samples was purified with an automated protocol using DNA extraction kit based on paramagnetic particles (LGC Genomics, UK). In brief, the samples were subjected to mechanical lysis using glass beads, and the DNA was purified by eluting from the paramagnetic particles by downstream processes, as described by the manufacturer.
PCR Amplification
For the 16–23S rRNA ITS PCR, we used the 16S IX-23S I primer pair [19] with denaturation, annealing, and extension for 1 min each. The 16S rRNA gene was amplified using the general PRK primers, with the original protocol [23]. All PCR reactions were initially denatured at 95 °C for 15 min.
One µl DNA (concentration 1–10 ng/μl) was mixed with a PCR master mix consisting of the following reaction mixture: 1.25U HOT FIREpol ® DNA polymerase, 1× HOT FIREpol ® buffer B2, 2.5 mM magnesium-dichloride (MgCl2), 0.2 mM dNTP, 0.2 µM forward primer, and 0.2 µM reverse primer in the Dnase/Rnase free water to a final volume of 25 μl per reaction.
Illumina Sequencing
The Illumina adapters were added by nested PCR where the PCR product from the first round was diluted 1/100, and additional 12 PCR cycles were run with the adapter primers. The same adapters as previously described by us for the 16S rRNA gene were used [15].
The PCR products were normalized by agarose gel electrophoresis, pooled, and purified with AMPure (Beckman Coulter Life Sciences, Lakeview Parkway S. Drive, Indianapolis), following the manufacturer’s recommendations prior to sequencing. The sequencing was done using the MiSeq sequencing platform (Illumina, San Diego, California) with the Reagents Kit v3 (300 bp paired end sequencing).
Data Analysis
Operational Taxonomic Unit (OTU) Identification
We used a word-based Principle Component Analysis (PCA) approach [17] in defining OTU’s based on 16S-23S rRNA ITS sequences. The sequences for the 300 first base pairs corresponding to the forward reads were then transformed into hexamer words. The hexamer word table was further compressed to 5 latent variables using PCA. All the forward reads were subsequently projected onto the PCA model. We then defined OTU’s as cubes in the 5 dimensional space of the PCA score plot (each side having the size of one). The taxonomic assignments were done by identifying the closest genome sequenced strain to the de novo sequences (from the sequenced samples) in the PCA space.
Microbiota Analyses
The number of reads per sample was normalized to 10,000. Samples with less reads were discharged for further analyses (7 for mother’s stool, 9 for infant 10 days feces, 15 for infant 90 days feces, 6 for mother’s 10 days milk, and 7 for mother’s 90 days milk), resulting in 55 analyzed samples. The overall strain distribution was analyzed using PCA, while Venn diagrams were used to investigate the overlap in the microbiota between the different categories. The age-related development of the microbiota was analyzed using partial least square discriminant analyses (PLS-DA), which is a multivariate statistical approach to build classification models (user manual PLS toolbox, Eigenvector, Seattle, Washington). In addition, we used standard pairwise parametric and non-parametric statistical tests. All analyses were done using Matlab (MathWorks Inc., Natick, Massachusetts), with the plugin PLS toolbox for conducting the multivariate statistical analyses.
Results
OTU Identification Model for 16–23S ITS rRNA
We identified 8494 reference spacer regions from 1838 genome sequenced strains, corresponding on average 4.6 spacers per genome. PCA analyses of the hexamers using five components revealed high-resolution differentiation of the reference spacers (Suppl. Fig. 1).
Verification of the 16–23S ITS rRNA Gene Deep Sequencing
We first determined the coverage of the 16S IX-23S I primer pair by comparison to the 16S rRNA gene PRK primers. This was done by analyzing a reference strain collection of human-associated bacteria. The comparison showed that the main phyla and strains were covered by both primer pairs (Suppl. Table 1). This result was supported by probe match searches in both the RDP database and in the Silva 23S rRNA gene database. These searches revealed one or less mismatch to the 16–23S rRNA ITS primers for bacteria belonging to the main human-associated phyla Bateroidetes, Firmicutes, Proteobacteria, and Actinobacteria.
We verified the Illumina 16S-13S rRNA OTU assignment approach using three defined mixes of bacterial strains reflecting different levels of diversity. These analyses revealed a high degree of reproducibility with R 2 values >0.98 (Suppl. Fig. 2). After removing singletons, we identified 331 OTU’s for the high-complexity sample (83 strains), 88 for low complexity (10 strains), and 104 for the middle complexity sample (10 stains in different proportions). Assuming an average of 4 spacers for genome, this would correspond to approximately 83 strains identified for the high-complexity, 22 for the low-complexity, and 26 for the middle-complexity sample.
Overall Microbiota Composition in the Pro-PACT Cohort
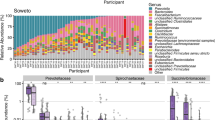
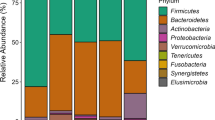
We detected in total 1356 OTU’s (corresponding to approximately 340 strains), represented by two or more sequences. Hierarchical clustering for the 40 most dominant OTU’s revealed a clustering of mother’s milk, infant feces, and mothers feces, although these did not reveal distinct groups either qualitatively (Fig. 1) or quantitatively (Fig. 2). The bacteria, on the other hand, showed distinct clustering in two main groups qualitatively (Fig. 1), but not quantitatively (Fig. 2). OTU88 (resembling Streptococcus gordonii) show a distinct colonization pattern, being both more widespread and detected in 52 out of 55 samples (Fig. 1), and quantitatively the most numerous (Fig. 2).
Prevalence of the 40 most dominant OTU’s for all samples analyzed. The dendrograms were created using average linkage clustering. Red represents detected (excluding singletons), while black represents non-detected. The following abbreviations were used: FC feces child, FM feces mother, and BM breast milk. Age is indicated with 10 and 90 days (Color figure online)
Heat map for the 40 most dominant OTU’s for all samples analyzed. The dendrograms were created using average linkage clustering. The color code represents the number of reads, with bright red color representing reads >100, while black represent no reads. The following abbreviations were used: FC feces child, FM feces mother, and BM breast milk. Age is indicated with 10 and 90 days (Color figure online)
The three most dominant OTU’s, OTU88 (S. gordonii), OTU349 (S. agalactiae) and OTU375 (S. pneumoniae), represented 9.5, 6.6 and 6.6 % of the sequences, respectively. The highest level of colonization by these OTU’s was in mother’s milk, where there seemed to be about equal amounts of all three species (Fig. 3a). However, in both mother and infant stool, the level of S. gordonii was significantly higher (P < 0.01, Mann–Whitney U test) than of both S. agalactiae and S. pneumoniae (Fig. 3b, c).
Distribution of the three most abundant species in a mother’s milk, b mothers stool and c infants stool for all samples analyzed. The number of reads per sample (out of 10,000) is indicated with red dots, while the means are indicated with the crossed dots. Significant differences are indicated with ** (P < 0.01 Mann–Whitney U test) (Color figure online)
For the observed species richness, the highest richness was detected in mother’s feces, while there was approximately the same richness in the child’s feces and mother’s milk (Fig. 4). Although not as pronounced, this was also reflected for the alpha diversity analyses (results not shown).
OTU Prevalence
The prevalence of OTU’s (detected more than once in a given sample) for the infant stool samples followed a negative geometric distribution (Fig. 5a, b). However, for the mother’s sample, we found a deviation from the negative geometric distribution for the highly prevalent OTU’s (present in more than 60 % of the samples). The most prevalent OTU’s were clearly overrepresented (P < 0.0005 t test deviation from the respective regression lines) compared to the negative geometric distribution. Furthermore, for the prevalent OTU’s, there were positive associations between the prevalence and number of OTU’s showing this prevalence (Fig. 5c–e).
Number of OTUs at different prevalence levels for child stool 10 days (a), 90 days (b), mother milk 10 days (c), 90 days (d), and stool (e). The regression lines represent fitting to a geometric distribution. Blue lines represent fitting of OTU’s with prevalence’s below 60 %, while the red lines represent fitting to OTU’s above. Since no OTU’s were identified in all samples for the children we used one to have a data point. The P values indicate the deviation of the number of the most prevalent OTU’s from the distribution below 60 % (Color figure online)
For the mother’s milk samples, the most prevalent OTU’s belonged to streptococci, while most of the highly prevalent OTU’s from the mothers stool samples belonged to the clostridia.
Temporal Changes of Microbiota
There was a consistent change of the mother’s milk microbiota composition from 10 to 90 days (Suppl. Fig. 3A). There was an association of Staphylococcus aureus for the 10 days samples, which shifted towards streptococci at 90 days. The 90 days samples were also associated with Aggregatibacter actinomycetemcomitans which has been identified as an aggressive periodontics bacterium.
Comparison of the fecal microbiota at 10 and 90 days for the children showed that S. aureus, Acetobacter woodi and an unidentified bacterium related to Clostridium botulinum were associated with the 10 days samples, while Bifidobacterium longum, Brevundimonas subvibrioides, and Escherichia coli were associated with the 90 days samples (Suppl. Figure 3B).
Overlap in Microbiota Between Mother and Child
The children’s stool microbiota composes of bacteria overlapping with both mother’s milk and mother’s stool (Fig. 6). There was an increase in the number of OTU’s with time for both mother’s milk (P = 0.02, Binomial test) and the children stool samples (P = 0.02, Binomial test). The mothers stool microbiota at 90 days showed a larger overlap with the children’s stool microbiota at 10 days than at 90 days (0.02, Binomial test), while for the mother’s milk at 10 and 90 days, there were no significant differences in overlap with mother’s stool.
Discussion
The overrepresentation of highly prevalent strains is in accordance with a core colonizers model [1, 21]. Furthermore, the apparent widespread presence of strains/species colonizing multiple body sites may indicate the presence of cosmopolitan colonizers of the human body.
We found an OTU resembling S. gordonii to be both the most abundant and prevalent colonizer in our dataset, as determined from the individual distribution and prevalence patterns. This species has not previously been reported abundant in mother’s milk, neither by culture dependent nor independent techniques [8]. Although there is a potential for taxonomic misclassification of the S. gordonii resembling OTU88, this OTU is highly widespread among individuals and body sites. Furthermore, S. gordonii is an important dental colonizer [11], with the potential to enter the blood stream and colonize the aortic valve [22]. This may indicate that OTU88 is a cosmopolitan colonizer, with the potential to colonize a wide range of body sites.
A question is the nature of the host association for the most dominant OTU (OTU88)—whether it is mutualistic or opportunistic. Pointing towards a mutualistic interaction is the fact that the related S. gordonii can produce hydrogen peroxide—potentially preventing the colonization by pathogens [25]. Furthermore, despite the high numbers and prevalence, relatively few incidences of infections have been reported for S. gordonii [11]. The fact that this bacterium together with other low-pathogenicity streptococci can colonize the aortic valve without causing instant fatal disease may also support its low virulence [7]. The low virulence is also confirmed by that its genome contain only few virulence genes (about 70) compared to more than 250 for the closely related S. pneumonia (identified from the Victors database). Although the potentially pathogenic S. agalactiae and S. pneumoniae showed about equal colonization of mother’s milk, they had a much lower colonization level of both infant and mothers stool than S. gordonii, suggesting that S. gordonii is allowed at more body sites.
Although bifidobacteria show high-relative levels in infant feces [2] and are assumed selected by mother’s milk oligosaccharides [13], we did find relative low levels of bifidobacteria in mother’s milk, but still relatively high prevalence. This indicates that the bifidobacteria are present in mother’s milk, but are not able to utilize the oligosaccharides there.
It seems like the infant stool microbiota is recruited from both the mother’s stool and milk microbiota, but that there is an expansion of the stool associated bacteria with age. The genera that showed the largest time-related shift for mother’s milk were Staphylococcus and Aggregatibacter, being linked with 10 and 90 days samples, respectively. Thus, it seems like there is a time-related shift from skin [12] to oral [20] associated bacteria for mother’s milk. For the infant stool, bacteria related to Acetobacterium were most diagnostic for the 10 days samples, while Brevundimonas showed the strongest association with the 90 days samples. Although associated with the gut [16], little is known about the potential role of these bacteria.
The mother’s stool and milk microbiota follow a distribution with an overrepresentation of highly prevalent strains. This is in line with a model, we have previously proposed [2], advocating the importance of core colonizing strains.
In conclusion, we have identified a prevalent colonizer with the potential to colonize multiple body sites, conferring potential host benefits. In addition, we show a relatively limited number of prevalent bacterial phylotypes. Taken together, these results support the importance of specific bacterial phylotypes in human/bacterial interactions [1].
References
Avershina E, Rudi K (2013) Is it who you are or what you do that is important in the human gut? Benef Microbes 4(3):219–222. doi:10.3920/BM2013.0016
Avershina E, Storro O, Oien T, Johnsen R, Pope P, Rudi K (2014) Major faecal microbiota shifts in composition and diversity with age in a geographically restricted cohort of mothers and their children. FEMS Microbiol Ecol 87(1):280–290. doi:10.1111/1574-6941.12223
Cabrera-Rubio R, Collado MC, Laitinen K, Salminen S, Isolauri E, Mira A (2012) The human milk microbiome changes over lactation and is shaped by maternal weight and mode of delivery. Am J Clin Nutr 96(3):544–551. doi:10.3945/ajcn.112.037382
D’Elia TV, Cooper CR, Johnston CG (2007) Source tracking of Escherichia coli by 16S-23S intergenic spacer region denaturing gradient gel electrophoresis (DGGE) of the rrnB ribosomal operon. Can J Microbiol 53(10):1174–1184. doi:10.1139/W07-083
Dec M, Urban-Chmiel R, Gnat S, Puchalski A, Wernicki A (2014) Identification of Lactobacillus strains of goose origin using MALDI-TOF mass spectrometry and 16S-23S rDNA intergenic spacer PCR analysis. Res Microbiol 165(3):190–201. doi:10.1016/j.resmic.2014.02.003
Dotterud CK, Storro O, Johnsen R, Oien T (2010) Probiotics in pregnant women to prevent allergic disease: a randomized, double-blind trial. Br J Dermatol 163(3):616–623. doi:10.1111/j.1365-2133.2010.09889.x
Douglas CWI, Heath J, Hampton KK, Preston FE (1993) Identity of viridans streptococci isolated from cases of infective endocarditis. J Med Microbiol 39(3):179–182. doi:10.1099/00222615-39-3-179
Jeurink PV, van Bergenhenegouwen J, Jimenez E, Knippels LM, Fernandez L, Garssen J, Knol J, Rodriguez JM, Martin R (2013) Human milk: a source of more life than we imagine. Benef Microbes 4(1):17–30. doi:10.3920/BM2012.0040
Khodayar-Pardo P, Mira-Pascual L, Collado MC, Martinez-Costa C (2014) Impact of lactation stage, gestational age and mode of delivery on breast milk microbiota. J Perinatol 34(8):599–605. doi:10.1038/jp.2014.47
Kim CM, Song ES, Jang HJ, Kim HJ, Lee S, Shin JH, Kim SJ, Jeong SH, Jeong J, Koh K, Choi GE, Lee EY, Chang CL (2010) Development and evaluation of oligonucleotide chip based on the 16S-23S rRNA gene spacer region for detection of pathogenic microorganisms associated with sepsis. J Clin Microbiol 48(5):1578–1583. doi:10.1128/JCM.01130-09
Lee SF (2003) Oral colonization and immune responses to Streptococcus gordonii: potential use as a vector to induce antibodies against respiratory pathogens. Curr Opin Infect Dis 16(3):231–235. doi:10.1097/01.qco.0000073772.11390.ae
Lindberg E, Adlerberth I, Hesselmar B, Saalman R, Strannegard I-L, Aberg N, Wold AE (2004) High rate of transfer of Staphylococcus aureus from parental skin to infant gut flora. J Clin Microbiol 42(2):530–534
LoCascio RG, Desai P, Sela DA, Weimer B, Mills DA (2010) Broad Conservation of milk utilization genes in Bifidobacterium longum subsp. infantis as revealed by comparative genomic hybridization. Appl Environ Microbiol 76(22):7373–7381
McDonald D, Price MN, Goodrich J, Nawrocki EP, DeSantis TZ, Probst A, Andersen GL, Knight R, Hugenholtz P (2012) An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J 6 (3):610–618. http://www.nature.com/ismej/journal/v6/n3/suppinfo/ismej2011139s1.html
Naseribafrouei A, Hestad K, Avershina E, Sekelja M, Linlokken A, Wilson R, Rudi K (2014) Correlation between the human fecal microbiota and depression. Neurogastroenterol Motil. doi:10.1111/nmo.12378
Rajilic-Stojanovic M, de Vos WM (2014) The first 1000 cultured species of the human gastrointestinal microbiota. FEMS Microbiol Rev 38(5):996–1047. doi:10.1111/1574-6976.12075
Rudi K, Zimonja M, Kvenshagen B, Rugtveit J, Midtvedt T, Eggesbo M (2007) Alignment-independent comparisons of human gastrointestinal tract microbial communities in a multidimensional 16S rRNA gene evolutionary space. Appl Environ Microbiol 73(8):2727–2734
Ruegger PM, Clark RT, Weger JR, Braun J, Borneman J (2014) Improved resolution of bacteria by high throughput sequence analysis of the rRNA internal transcribed spacer. J Microbiol Methods 105:82–87. doi:10.1016/j.mimet.2014.07.001
Saruta K, Matsunaga T, Kono M, Hoshina S, Ikawa S, Sakai O, Machida K (1997) Rapid identification and typing of Staphylococcus aureus by nested PCR amplified ribosomal DNA spacer region. FEMS Microbiol Lett 146(2):271–278
Schacher B, Baron F, Rossberg M, Wohlfeil M, Arndt R, Eickholz P (2007) Aggregatibacter actinomycetemcomitans as indicator for aggressive periodontitis by two analysing strategies. J Clin Periodontol 34(7):566–573. doi:10.1111/j.1600-051X.2007.01080.x
Sekelja M, Berget I, Naes T, Rudi K (2011) Unveiling an abundant core microbiota in the human adult colon by a phylogroup-independent searching approach. ISME J 5(3):519–531. doi:10.1038/ismej.2010.129
Takahashi Y, Takashima E, Shimazu K, Yagishita H, Aoba T, Konishi K (2006) Contribution of sialic acid-binding adhesin to pathogenesis of experimental endocarditis caused by Streptococcus gordonii DL1. Infect Immun 74(1):740–743. doi:10.1128/IAI.74.1.740-743.2006
Yu Y, Lee C, Kim J, Hwang S (2005) Group-specific primer and probe sets to detect methanogenic communities using quantitative real-time polymerase chain reaction. Biotechnol Bioeng 89(6):670–679. doi:10.1002/bit.20347
Zeng YH, Koblizek M, Li YX, Liu YP, Feng FY, Ji JD, Jian JC, Wu ZH (2013) Long PCR-RFLP of 16S-ITS-23S rRNA genes: a high-resolution molecular tool for bacterial genotyping. J Appl Microbiol 114(2):433–447. doi:10.1111/jam.12057
Zheng L, Itzek A, Chen Z, Kreth J (2011) Environmental influences on competitive hydrogen peroxide production in Streptococcus gordonii. Appl Environ Microbiol 77(13):4318–4328. doi:10.1128/AEM.00309-11
Acknowledgments
We wish to thank the Norwegian University of Life Sciences for financial support and the team at the MiDiv Lab for fruitful discussions.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Schanche, M., Avershina, E., Dotterud, C. et al. High-Resolution Analyses of Overlap in the Microbiota Between Mothers and Their Children. Curr Microbiol 71, 283–290 (2015). https://doi.org/10.1007/s00284-015-0843-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-015-0843-5