Abstract
In this study, we have investigated intrinsic salt tolerance of Astragalus cicer microsymbionts (USDA3350, ACMP18) and the role of exogenous glycine betaine in osmoprotection in these bacteria. Salt stress was imposed by NaCl concentrations ranging from 0.5 to 2 %. A. cicer mesorhizobia were capable of tolerating up to 2 % sodium chloride with a population count that was inversely proportional to the salt content. When the extracellular concentration of NaCl was raised to 2 %, the generation time of the UDSA3350 strain in the mid-exponential phase of growth was 3.9-times greater than that in the no-salt control medium, whereas the ACMP18 strain survived under the same conditions but did not multiply. Application of 1 mM glycine betaine into the salt-stressed rhizobium cultures increased the number of culturable bacteria, pointing out that this molecule was involved in restoration of osmotic balance. The decline in A. cicer symbiont viability in the medium with sodium chloride and the osmoprotective role of glycine betaine for these bacteria was confirmed in the experiment using the live/dead Bac Light Bacterial Vibility Kit. Data presented in this study showed the presence of proU-like genes in the genomes of A. cicer rhizobia with high sequence similarity to the genes of the ProU-like system in Sinorhizobium meliloti and the proU operon of Escherichia coli.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Soil bacteria called rhizobia are frequently exposed to osmotic stress that adversely affects their metabolic activity and, in consequence, the growth of bacteria is generally strongly reduced. Rhizobia exhibit significant differences in salt tolerance [14–16, 47]. While strains of the genera Rhizobium and Bradyrhizobium are inhibited by 100–300 mM NaCl, Sinorhizobium meliloti strains grow at salt concentrations higher than 300 mM. The studies of salt tolerance in bacteria of the Mesorhizobium genus showed that most of these strains were more sensitive to salt stress than Rhizobium and Sinorhizobium strains but more resistant than bacteria of the Bradyrhizobium genus. The growth of most mesorhizobial strains was inhibited by 100 mM NaCl with some exceptions, e.g., Mesorhizobium thiogangenicum and Mesorhizobium ciceri (they tolerated up to 200 mM NaCl) [8, 23]. To maintain the osmotic equilibrium between the cytoplasm and the saline habitat, rhizobia accumulate high levels of small, highly water-soluble organic compounds termed compatible solutes, i.e., proline, glutamate, glycine betaine, and trehalose [7, 9, 11, 18, 29, 38, 46]. Such solutes, which do not have a detrimental effect on vital processes in stressed cells and function as osmoprotectants, stabilize enzymes, DNA, membranes, and whole cells against different stresses [10]. Compatible solutes are released into the ecosystem by primary microbial producers upon dilution stress, decaying plant and animal cells, and as excretion fluids of mammals (urine) [34]. A very important compatible solute for most rhizobia is glycine betaine [3]. In the best osmotically characterized rhizobia, S. meliloti, glycine betaine can be taken up from the environment or synthesized from choline or choline-O-sulfate [27, 35]. Glycine betaine transport systems in S. meliloti are poorly understood; however, it was found that their activity is strongly stimulated when bacteria are grown at elevated osmolarity [4, 6]. One of these transporters is a constitutively expressed BetS system, rapidly posttranslationally activated by high medium osmolarity, which transports glycine betaine and proline betaine into stressed cells [1, 6]. In S. meliloti, ProU-like systems Cho and Hut were identified. It is interesting that HutXVW proteins showed higher similarity to ProU transporter proteins than to the Escherichia coli histidine transport protein, although the Hut system is involved in histidine but not glycine betaine uptake. The Cho system showed homology to the proU glycine betaine transport system but it is engaged in choline rather than glycine betaine uptake [1, 5, 13, 17]. In E. coli, two transporter systems, ProP and ProU, are primarily responsible for the import of glycine betaine [10, 22, 26, 32, 45]. ProP is a single-component transport system, regulated mainly at the activity level, which functions as an H+ symporter and imports a wide variety of compatible solutes related to proline and glycine betaine as well as ectoine. ProU is a high-affinity ATP-binding transport system regulated at the level of both transcription and activity. A sudden osmotic upshock results in a rapid increase in proU operon transcription which is proportional to the osmolarity level of the growth medium, and expression of proU operon genes is kept at an elevated level as long as the osmotic stimulus exists. The ProU transporter, a member of the ABC superfamily, consists of three proteins: ProV, ProW, and ProX. ProV is a protein associated with the cytoplasmic side of the membrane and shares considerable sequence identity with the ATP-binding protein. ProW is a transmembrane component of the transport system, whereas ProX is the periplasmic protein binding glycine betaine.
Bacterial cells subjected to a number adverse environmental stresses (high salt concentration, O2 limitation, temperature and pH variations, nutrient deficiencies) can die, exist in a metabolically active state, and proliferate or survive as “dormant” cells called a viable but nonculturable (VBNC) with reduced metabolism and suppressed reproduction [2, 28, 39, 42]. Viable but nonculturable bacteria cannot be cultured on conventional laboratory growth media but it is possible to demonstrate whether they still alive using the live/dead BacLight bacterial viability kit (Life Technologies).
In the present study, we examined the tolerance of Astragalus cicer symbionts to salt stress and the possible role of glycine betaine in NaCl tolerance. We also analyzed whether hyperosmolarity induces the VBNC state in A. cicer-specific rhizobia.
Materials and Methods
Bacterial Strains and Media
Mesorhizobium sp. (A. cicer) strain USDA3350 from the culture collection of the United States Department of Agriculture (USDA) and strain ACMP18 from our collection [44] were used throughout this study. The rhizobia were isolated from A. cicer root nodules which were surface sterilized in 70 % ethanol (2 min) and next in 0.1 % HgCl2 solution (2 min), and were rinsed in sterile water several times. The nodules were individually crushed, and the nodule suspension was streaked onto yeast mannitol agar (YEM) plates. The plates were incubated at 28 °C for 3–5 days. The purity of the bacteria was checked by repeated streaking on the YEM medium. The phenotypic characterization of ACMP18 and USDA3350 strains was presented in the paper by Wdowiak et al. [44]. The bacteria were maintained at 4 °C on the YEM rich medium [41]. Their osmotolerance phenotype and glycine betaine osmoprotective properties were determined in a modified YEM medium, in which mannitol was substituted by 1 % lactate and the yeast extract by 1 % NH4NO3. The osmotic strength of the medium was increased by NaCl addition to achieve final concentrations of 0.5, 1, 1.5, and 2 %. Glycine betaine used in these studies was prepared as a 0.5 M solution and sterilized by filtration before incorporation into the medium. Its final concentration in the medium was 1 mM for osmoprotection assays and 10 mM when used as sole carbon and nitrogen sources.
Culture Conditions for NaCl Tolerance Studies and Analysis of glycine Betaine Osmoprotective Properties
5 ml volumes of a modified YEM medium supplemented with different NaCl concentrations and with NaCl + 1 mM glycine betaine were prepared and inoculated with 0.5 ml of a late exponential phase culture of the bacteria studied. Cells were grown at 28 °C in an orbital shaker (160 rev min−1). Growth was monitored for 72 h spectrophotometrically by optical density at 600 nm and by plate counts of cell numbers as colony-forming units (CFU) on the YEM medium at 24 h intervals. The generation time from the log phase of the bacterial growth was also determined. Additionally, the relative number of dead and living bacterial cells growing under the above-mentioned conditions was estimated using the staining live/dead BacLight Bacterial Viability Kit (Life Technologies). This kit consists of two nucleic acid stains: SYTO 9 which penetrates most membranes freely and propidium iodide which is highly charged and permeates only damaged membranes. Simultaneous application of both these dyes results in green fluorescence of viable cells with intact membranes, whereas dead cells with an injured membrane show intense red fluorescence. The rapid visualization and evaluation of the viability of cells were performed by the live/dead BacLight bacterial viability kit and a confocal laser scanning microscope (Zeiss KLM150) in accordance with the manufacturer’s instruction.
The growth of Rhizobia with Glycine Betaine as the Sole Carbon and Nitrogen Source
To determine the growth of rhizobia with glycine betaine as the sole carbon or nitrogen source, precultures from overnight bacterial cultures on the modified YEM medium were pelleted, resuspended in the YEM medium without carbon or nitrogen source, and used to inoculate (3 %) the same medium containing glycine betaine as a sole carbon or nitrogen source. The rhizobial growth was determined by monitoring the optical density at 600 nm for 72 h at 24 h intervals.
DNA Isolation
Genomic DNAs from 30 ml of 72 h bacterial liquid cultures in the YEM medium were prepared according to the method described by Pitcher et al. [31]. The purities and concentrations of DNAs were determined with the spectrophotometer (Bio-Rad, SmartSpecTM3000).
PCR Amplification and Sequencing of Amplicons
PCRs were performed with the ReadyMixTM Taq kit and bacterial genomic DNA according to the manufacturer’s instructions (Sigma-Aldrich). The 640-bp fragment of proV gene was amplified and sequenced with primer proVf (5’-TCATCATCATGGGCCTGTCGG-3’) and reverse primer proVr (5’-CGGTCGACGTTGCGGGTGAAC-3’). To amplify and sequence the 450-bp fragment of the proW gene, primers proWf (5’-GCGCACAAGCCGAAGGTCTAC -3’ and proWr (5’-ACGATGGCCAGCACGACGAT-3’) were used. The 718-bp fragment of the proX gene was amplified and sequenced using primers proXf (5’-GCGGCTCTGTTGTCCACGAG-3’) and proXr (5’-TTGAACTTCAGGTTGGCGATG-3’). The PCR protocols for the proV, proW, and proX genes consisted of an initial denaturation step at 95 °C for 2 min followed by 35 cycles of denaturing at 94 °C for 1 min, annealing at 52 °C for 1 min, extension at 72 °C for 2 min, and final extension at 72 °C for 5 min. The amplification products were purified with the PCR purification Clean up kit (A&A Biotechnology) as recommended by the manufacturer and next they were sequenced using the ABI Prism BigDye Terminator Cycle sequence kit (Applied Biosystems model 310 DNA sequencer).
The nucleotide sequences determined in this study have been deposited in the GenBank database under the following accession numbers: KC155290 and KC155291 for the proV genes of USDA3350 and ACMP18, respectively; KC155293 and KC155292 for the proW genes of USDA3350 and ACMP18, respectively; and KC155294 and KC155295 for the proX genes of USDA3350 and ACMP18, respectively.
Results
Growth Characteristics Under Saline Conditions and Osmoprotection By Glycine Betaine
The intrinstic osmotolerance in two Mesorhizobium sp. (A. cicer) strains USDA3350 and ACMP18 was evaluated by their growth in the medium with five different NaCl concentrations (0, 0.5, 1, 1.5, and 2 %). This was done by measuring absorbance at 600 nm and the cell titer for 72 h at 12 and 24 h intervals. In the absence of NaCl (control medium), the strains studied showed different growth characteristics, i.e., the generation time of the ACMP18 strain (in the middle of the exponential growth phase) was about half an hour longer (5.20 h) than that of the USDA3350 strain (4.70 h), and the final bacterial yield was higher in the case of USDA3350 rhizobia (Table 1). However, both strains reached the stationary phase at the same time i.e., after 48 h. The A. cicer microsymbionts survived the entire period of cultivation in the medium with all four sodium chloride concentrations. It was found that the recovered viable count of the USDA3350 bacteria growing for 72 h in the presence of 0.5, 1, 1.5, and 2 % increased about 52-, 22-, 11-, and 5-fold, respectively, compared to the ~70-fold increase in the bacterial count in the no-salt control YEM medium. In the case of the ACMP18 strain growing over 72 h period in the presence of 0.5, 1, and 1.5 % NaCl, the culturable cell number increased ~37-,~15-, and ~7-fold, respectively. In turn, the number of CFU ml−1 recovered after 72 h of bacterium incubation in the medium with 2 % salt remained at the same level as at the beginning of the experiment. It is noteworthy that the increase in the number of CFU ml−1 of the ACMP18 strain growing in the no-salt control medium from time 0 to 72 h was only 58-fold. Furthermore, the doubling times for both studied strains growing in the presence of sodium chloride was significantly longer compared to that in the no-salt control medium. In the case of the USDA3350 strain, the generation time of bacteria cultivated in the medium with 1, 1.5, and 2 % NaCl was 1.8-, 2.7-, and 3.9-times greater, respectively, than in the control medium. The reduction in the growth rate under conditions of salt stress was much more pronounced for the ACMP18 strain. Its generation time in the YEM medium without salt was 5.20 h and increased to 10.20 h at 1 % NaCl and 22.9 h in the medium with 2 % sodium chloride (Table 1). In the presence of 2 % NaCl, the bacteria survived but did not multiply. To determine the effect of glycine betaine in relieving salt stress in Mesorhizobium sp. (A. cicer) strains USDA3350 and ACMP18, the bacteria were cultivated in the modified YEM medium containing 0, 0.5, 1, 1.5, and 2 % NaCl as well as under the same conditions plus the osmoprotectant supplied at a final concentration of 1 mM (Table 1). Addition of exogenous GB to the salt-containing medium had a beneficial effect on the growth of both strains and led to an increase in the growth rate and culturable cell number (data not shown) in comparison to the medium without the osmoprotectant; however, their growth was only partially recovered. For example, the addition of 1 mM of GB to the USDA3350 culture growing in the presence of 1.5 and 2 % NaCl reduced the doubling time of bacteria from 12.6 to 10.9 h and from 18.5 to 15.6 h, respectively. Incorporation of 1 mM GB to the culture medium with 1.5 % NaCl reduced the generation time of the ACMP18 strain from 16.0 to 13.3 h. The addition of 1 mM GB to the medium with 2 % NaCl decreased the doubling time of this strain from 22.9 to 18.5 h. The beneficial effect of GB on the growth rate of USDA3350 and ACMP18 under elevated salinity was less pronounced in the presence of 0.5 and 1 % NaCl.
Live/Dead Analysis
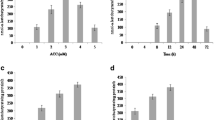
The live/dead Bac Light Bacterial Viability Kit was used to determine the relative proportions of viable and dead USDA3350 and ACMP18 bacterial cells under salt stress conditions (0.5, 1, 1.5, and 2 % NaCl) and in the salt solution plus glycine betaine as an osmoprotectant. In this assay containing two fluorescent nucleic acid stains, bacteria with intact membranes stain fluorescent green, whereas cells with damaged membranes (dead bacteria) stain fluorescent red. The percentage of live USDA3350 cells recovered after 72 h incubation at 0.5, 1, and 1.5 % NaCl in the medium with and without 1 mM glycine betaine remained at similar levels. However, the number of CFU/mL in the medium with NaCl but without GB in relation to the control clearly decreased and significantly increased after GB addition (data not shown). At the strong growth-inhibiting 2 % NaCl concentration, the percentage of dead cells clearly increased. The presence of GB in the medium with 2 % sodium chloride significantly elevated the percentage of viable cells in comparison to the percentage of live cells in the medium with 2 % NaCl but without glycine betaine (Fig. 1). The fluorescent dye treatment of the ACMP18 strain growing for 72 h in the presence of 1, 1.5, and 2 % NaCl showed that the percentage of viable cells declined gradually with the increasing concentrations of salt. Incorporation of GB to these growth media clearly alleviated viability inhibition by NaCl and clear improvement of the viable cell fraction in the presence of this osmoprotectant (5–11 %) was found. In the presence of 0.5 % NaCl and the non-salt control, the percentage of live ACMP18 cells (fluorescent green) was similar to that of the USDA3350 strain (Fig. 1).
Percentage of live/dead cells of Mesorhizobium sp. (A. cicer) ACMP18 and USDA3350 strains during growth on the modified YEM medium without and with NaCl and 1 mM glycine betaine (GB) determined by microscopic enumeration with live/dead BacLight Bacterial Viability Kit. Standard deviations never exceeded 10 %
Glycine Betaine as Sole Carbon and/or Nitrogen Sources
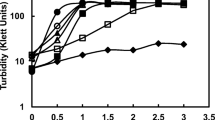
To test whether the Mesorhizobium sp. (A. cicer) strains USDA3350 and ACMP18 can catabolize GB and use it as potential carbon and nitrogen sources at low and elevated osmolarities, the growth of bacteria was monitored in the presence of 10 mM GB added to the carbon- and nitrogen-free YEM medium with and without NaCl. As shown in Fig. 2, both strains were able to assimilate GB at low osmolarity and grow in such a medium at a rate comparable to that in the YEM medium. In osmotically stressed A. cicer mesorhizobia, glycine betaine was not used as a carbon and nitrogen nutrient and the growth of the USDA3350 as well as ACMP18 strains was not supported by this compatible solute (data not shown).
Identification of proU Operon Genes
The PCR strategy was used to identify proU operon genes encoding a high-affinity glycine betaine transporter of E. coli in the Mesorhizobium sp. (A. cicer) strains USDA3350 and ACMP18. PCR amplification and sequencing of DNA with primers specific for the proX gene provided, in the case of two A. cicer microsymbionts, reliable 718-bp gene sequences sharing 41–42 % identical sequences with the E. coli proX gene and 92–91 % identical nucleotides with the proX-like gene of Mesorhizobium loti MAFF303099 (Table 2). The proX-like gene hutX of S. meliloti 1021 showed 68–69 % sequences identical to the proX sequences of A. cicer nodulators. PCR amplification and sequencing of DNA with specific primers was also used to identify proV- and proW-like genes of E. coli in A. cicer symbionts. 643-bp proV gene amplicons of the USDA3350 and ACMP18 strains, with 95 % sequence identity to each other, shared 58–60 % identical nucleotides with the E. coli proV gene, 90–92 % sequence identity with the proV-putative gene of M. loti MAFF303099, and 68 % sequence similarity with the proV-like hutV gene of S. meliloti 1021 (Table 2). It was also found that the 450-bp fragments of proW genes of the A. cicer rhizobia, with 98 % sequence identity to each other, displayed 54–55 % sequence similarity to the proW gene of E. coli, 73 % identical nucleotides with the proW-like hutW gene of S. meliloti 1021, and many more, i.e., 89–90 %, identical nucleotides with the proW-like gene of M. loti MAFF303099 (Table 2).
Discussion
As soil bacteria, Rhizobia frequently cope with high osmolarity of the environment. As the osmolarity of the surrounding environment increases, the turgor pressure drops because of cytoplasm dehydration, metabolic processes are inhibited, and bacterial growth slows down or is halted [28, 40, 43, 48]. Rhizobia differ in their tolerance to salinity. Generally, fast-growing rhizobia are more salt-tolerant than slow-growing Bradyrhizobium strains [15, 24, 33, 36, 49]. To present day, A. cicer microsymbionts have not been studied in terms of NaCl stress, although it is known that in nature elevated osmolarity affects both plant and bacterium growth as well as legume-rhizobium symbiosis. To characterize the intrinsic NaCl tolerance of A. cicer symbionts, two strains, USDA3350 from Canada and ACMP18 from Poland, were chosen based on our previous work, and studied in detail for their growth rate in the YEM medium with NaCl concentrations ranging from 0.5 to 2 %. Both strains were able to withstand even 2 % sodium chloride over 72 h. The ACMP18 strain was more sensitive to salt than USDA3350 (Table 1). It only survived in the YEM medium with 2 % NaCl and the titer of these bacteria was maintained under these conditions at a similar level within 72 h. The USDA3350 strain withstands the presence of NaCl in the culture medium better and its growth parameters (the generation time, growth rate) were less affected by sodium chloride. It was found that multiplication of USDA3350 bacteria occurred even in the presence of 2 % NaCl. Evaluation of salinity resistance of the Mesorhizobium USDA3350 and ACMP18 strains forming symbiosis with A. cicer allowed us to classify them as moderately NaCl-tolerant bacteria.
Rhizobia exposed to increased salinity defend themselves against the outflow of water from the cells by accumulation of organic osmolytes called compatible compounds [7, 18, 19, 24]. These highly water-soluble osmoprotectants can be accumulated to very high levels in stressed cells without disturbing vital cellular functions [3, 4, 25, 34]. It has been shown that glycine betaine is a very potent osmoprotectant in many bacteria including rhizobia [4, 29]. Its accumulation not only allows the cells to withstand a given osmolarity but also permits the cell to proliferate under unfavourable conditions. The growth of A. cicer mesorhizobia inhibited in the presence of 0.5–2 % NaCl was clearly stimulated by the addition of 1 mM glycine betaine to the culture medium (Table 1). The beneficial effect of glycine betaine on the growth of A. cicer rhizobia in the elevated osmolarity medium was particularly pronounced in the 48–72 h stationary phase culture.
The rapid decline in the culturability (CFU) of the ACPM18 and USDA3350 strains after 72 h incubation in the salted medium was also supported by the live/dead Bac Light Bacterial Viability Kit (Fig. 1). This assay with two fluorescent nucleic acid dyes differentiated fluorescent green viable cells from dead bacteria which stain with red fluorescent dye. The decrease in the culturability of the A. cicer symbionts that occurred in response to the addition of NaCl to the medium can be due to cell death or entry of the bacteria into the viable but nonculturable state (VBNC). In the medium with sodium chloride and glycine betaine, the number of metabolically active ACMP18 cells able to produce colonies and the proportion of viable to dead cells estimated as a percentage of green versus red fluorescent cells increased in comparison to these parameters in the salt-supplemented medium without the osmoprotectant. We conclude from these data that the compatible solute glycine betaine really protects ACMP18 strain cells from the detrimental effect of salt stress. It is interesting that the live/dead viability staining of USDA3350 cells growing at 0.5, 1, and 1.5 % NaCl with and without glycine betaine showed a comparable ratio of live to dead cells under both conditions. However, the number of colony-forming cells in the salt medium with the osmoprotectant was larger than that in the medium without glycine betaine (data not shown). We suppose that in the salt medium part of live USDA3350 bacterial cells are culturable and form colonies on a stable medium but part of this population enter the viable but nonculturable state. In the presence of the osmoprotectant glycine betaine, metabolically active but nonculturable cells of the USDA3350 strain affected by salt stress resuscitate to bacterial colony formation. Since elevated NaCl concentrations in the culture medium can induce the viable but nonculturable state in rhizobia, we can suppose that such bacteria exist under soil adverse conditions. The capability of rhizobia to enter the VBNC state can be an advantage for their survival in soil under stress conditions and further studies are needed to elucidate the effect of this metabolic shift on rhizobium-legume symbiosis.
Depending on their growth medium osmolarity, A. cicer rhizobia, like S. meliloti, use glycine betaine as an osmolite or carbon and nitrogen source [3, 20, 29, 30, 35, 37]. In unstressed cells of the ACMP18 and USDA3350 strains, glycine betaine was catabolized and used as a nitrogen or carbon nutrient (Fig. 2). Under osmotic stress, glycine betaine was an effective osmoprotectant but not a nitrogen and carbon source (data not shown). The osmotic strength of the environment can modulate the pathway of glycine betaine metabolism [35]. It was found that in low-salt grown S. meliloti cells, glycine betaine stimulated the enzymes involved in the catabolism of glycine betaine into serine and then to pyruvate. Under high-osmolarity growth conditions, the activities of these enzymes are inhibited and glycine betaine is accumulated in the cytoplasm as an osmoprotectant. The ability to accumulate and catabolize glycine betaine, liberated into the environment from plant and animal residues, may be beneficial to bacteria and allows them to survive as well as thrive in natural habitats.
The mechanism of the glycine betaine transport into the A. cicer symbiont cells is not known and no genes have been attributed to this function. As shown from the fully annotated nucleotide sequences of the M. loti MAFF303099 genome, the genes encoding transporter-related proteins constitute 10.5 % of the total DNA. The data presented in this paper show that the Mesorhizobium sp. (A.cicer) USDA3350 and ACMP18 strains have proX, proW, and proV-like genes which encode a high-affinity glycine betaine ProU uptake system of E. coli [2, 12, 21, 22, 26]. The ProU transporter consists of a periplasmic substrate binding protein ProX, a transmembrane component ProW, and an inner membrane-associated ATPase protein ProV. Comparison of proX, proW, and proV gene sequences of A. cicer symbionts with those in the data libraries revealed their highest sequence similarity with genes encoding components of the ProU-like glycine betaine system of M. loti (89–92 %). Lower identity scores were noted between the nucleotide sequences of proX, proW, and proV genes of the USDA3350 and ACMP18 strains and those of S. meliloti (68–73 %) as well as E. coli proU operon sequences (41–60 %). Based on the annotated sequences in the A. cicer symbionts studied, the presence of a potential glycine betaine ProU-like transporter that allows bacteria to cope with salt stress was revealed.
References
Alloing G, Travers I, Sagot B, Le Rudulier D, Dupont L (2006) Proline betaine uptake in Sinorhizobium meliloti: characterization of Prb, an opp-like ABC transporter regulated by both proline betaine and salinity stress. J Bacteriol 188(17):6308–6317
Alexander E, Pham D, Steck TR (1999) The viable-but-nonculturable condition is induced by cooper in Agrobacterium tumefaciens and Rhizobium leguminosarum. Appl Environ Microbiol 65:3754–3756
Bernard T, Pocard JA, Perroud B, Le Rudulier D (1986) Variations in the response of salt-stressed Rhizobium strains to betaines. Arch Microbiol 143:359–364
Boncompagni E, Østreås M, Poggi MC, Le Rudulier D (1999) Occurrence of choline and glycine betaine uptake and metabolism in the family Rhizobiaceae and their roles in osmoprotection. Appl Environ Microbiol 65:2072–2077
Boncompagni E, Dupont L, Mignot T, Østeräs M, Lambert A, Poggi MC, Le Rudulier D (2000) Characterization of a Sinorhizobium meliloti ATP-binding cassette histidine transporter also involved in betaine and proline uptake. J Bacteriol 182(13):3717–3725
Boscari A, Mandon K, Dupont L, Poggi MC, Le Rudulier D (2002) BetS is a major glycine betaine/proline betaine transporter required for early osmotic adjustment in Sinorhizobium meliloti. J Bacteriol 184:2654–2663
Botsford JL, Lewis TA (1990) Osmoregulation in Rhizobium meliloti: production of glutamic acid in response to osmotic stress. Appl Environ Microbiol 56:488–494
Brígido C, Alexandre A, Oliveira S (2012) Transcriptional analysis of major chaperone genes in salt-tolerant and salt-sensitive mesorhizobia. Microbiol Res 167(10):623–629
Cleland D, Krader P, McCree C, Tang J, Emerson D (2004) Glycine betaine as a cryoprotectant for prokaryotes. J Microbiol Methods 58:31–38
Csonka LN, Epstein W (1996) Osmoregulation. In: Neidhardt FC (ed) Escherichia coli and Salmonella: Cellular and Molecular Biology. ASM Press, Washington DC, pp 1210–1223
Dardanelli MS, Gonzáles PS, Bueno MA, Ghittoni NE (2000) Synthesis, accumulation and hydrolysis of trehalose during growth of peanut rhizobia in hyperosmotic media. J Basic Microbiol 40:149–156
Dattananda CS, Gowrishankar J (1989) Osmoregulation in Escherichia coli: complementation analysis and gene-protein relationships in the proU locus. J Bacteriol 171:1915–1922
Dupont L, Garcia I, Poggi MC, Alloing G, Mandon K, Le Rudulier D (2004) The Sinorhizobium meliloti ABC transporter Chois highly specific for choline and expressed in bacteroids from Medicago sativa nodules. J Bacteriol 186(18):5988–5996
Elkan GH (1992) Taxonomy of the rhizobia. Can J Microbiol 38:446–450
Elsheikh EAE (1998) Effects of salt rhizobia and bradyrhizobia: a review. Ann Appl Biol 132:507–524
Elsheikh EAE, Wood M (1990) Rhizobia and bradyrhizobia under salt stress: possible role of trehalose in osmoregulation. Lett Appl Microbiol 10:127–129
Fox MA, Karunakaran R, Leonard ME, Mouhsine B, Williams A, East AK, Downie JA, Poole PS (2008) Rhizobia and bradyrhizobia under salt stress: possible role of trechalose in osmoregulation. FEMS Microbiol Lett 287(2):212–220
Fujihara S, Yoneyama T (1994) Response of Rhizobium fredii P220 to osmotic shock: interrelationships between K+, Mg2+, glutamate and homospermidine. Microbiol 140:1909–1916
Gloux K, Le Rudulier D (1989) Transport and catabolism of proline betaine in salt-stressed Rhizobium meliloti. Arch Microbiol 151:143–148
Goldman A, Lecoeur L, Massage B, Delarue M, Schoonejans E, Tapfer D (1994) Symbiotic plasmid genes essential to the catabolism of proline betaine, or stachydine, are also required for efficient nodulation by Rhizobium meliloti. FEMS Microbiol Lett 115:305–312
Gowrishankar J (1989) Nucleotide sequence of the osmoregulatory proU operon of Escherichia coli. J Bacteriol 171:1923–1931
Kempf B, Bremer E (1998) Uptake and synthesis of compatible solutes as microbial stress responses to high osmolality environments. Arch Microbiol 170:319–330
Laranjo M, Oliveira S (2011) Tolerance of Mesorhizobium type strains to different environmental stresses. Antonie van Leeuvenhoek 99(3):651–662
Le Rudulier D, Gloux K, Riou N (1991) Identification of an osmotically induced periplasmic glycine betaine-binding protein from Rhizobium meliloti. Biochim Biophys Acta 1061:197–205
Lippert K, Galinski EA (1992) Enzyme stabilization by ectoine-type compatible solutes: protection against heating, freezing and drying. Appl Microbiol Biotechnol 37:61–64
Lucht JM, Bremer E (1994) Adaptation of Escherichia coli to high osmolarity environments: osmoregulation of the high-affinity glycine betaine transport system ProU. FEMS Microbiol Rev 14:3–20
Mandon K, Østerås M, Boncompagni E, Trinchant JC, Spennato G, Poggi MC, Le Rudulier D (2003) The Sinorhizobium meliloti glycine betaine biosynthetic genes (betCBA) are induced by choline and highly expressed in bacteroids. Mol Plant-Microbe Interact 16:709–719
McDougald D, Rice SA, Weichart D, Kjelleberg S (1998) Nonculturability: adaption or debilitation. FEMS Microbiol Ecol 25:1–9
Miller KJ, Wood JM (1996) Osmoadaptation by rhizosphere bacteria. Ann Rev Microbiol 50:101–106
Pichereau V, Pocard JA, Hamelin J, Blanco C, Bernard T (1998) Differential effects of dimethylsulfoniopropionate, dimethylsulfonioacetate, and other S-Methylated compounds on the growth of Sinorhizobium meliloti at low and high osmolarities. Appl Environ Microbiol 64:1420–1429
Pitcher DG, Saunders NA, Owen RJ (1989) Rapid extraction of bacterial genomic DNA with guanifium thiocyanate. Lett Appl Microbiol 8:151–156
Racher KI, Voegele RT, Marshall EV (1999) Purification and reconstitution of an osmosensor: transporter ProP of Escherichia coli senses and responds to osmotic shifts. Biochemistry 38:1676–1684
Singleton PW, Elswaify SA, Bohlool BB (1982) Effect of salinity on Rhizobium growth and survival. Appl Environ Microbiol 44:884–890
Sleator RD, Hill C (2001) Bacterial osmoadaptation: the role of osmolytes in bacteria stress and virulence. FEMS Microbiol 26:49–71
Smith LT, Pocard JA, Bernard T, Le Rudulier D (1988) Osmotic control of glycine betaine biosynthesis and degradation in Rhizobium meliloti. J Bacteriol 170:3142–3149
Soussi M, Ocaña A, Lluch C (1998) Effects of salt stress on growth, photosynthesis and nitrogen fixation in chick-pea (Cicer arietinum L.). J Exper Bot 49:1329–1337
Talibart R, Jebbar M, Gouffi K, Pichereau V, Gouesbet G, Blanco C, Bernard T, Pocard JA (1997) Transient accumulation of glycine betaine and dynamics of endogenous osmolytes in salt-stressed cultures of Sinorhizobium meliloti. Appl Environ Microbiol 63:4657–4663
Tempest DW, Meers JL, Brown CM (1970) Influence of environment on the content and composition of microbial free amino acid pools. J Gen Microbiol 64:171–185
Toffanin A, Basaglia M, Ciardi C, Vian P, Povolo S, Casella S (2000) Energy content decrease and viable-not-culturable status induced by xygen limitation coupled to the presence of nitrogen oxides in Rhizobium hedysari. Biol Fertil Soils 31:484–488
Trotman AP, Weaver RW (1995) Tolerace of clover rhizobia to heat and desiccation stresses in soil. Soil Sci Soc Am J 59:466–470
Vincent J (1970) A manual for the practical study of root nodule bacteria. UK, Blackwell Scientific Publication Ltd, Oxford
Vriezen JAC, deBruijn FJ, Nüsslein K (2012) Desiccation induces viable but non-culturable cells in Sinorhizobium meliloti 1021. AMB Express 2:6
Vriezen JAC, deBruijn FJ, Nüsslein K (2007) Responses of rhizobia to desiccation in relation to osmotic stress, oxygen and temperature. Appl Environ Microbiol 73:3451–3459
Wdowiak S, Małek W (2000) Numerial analysis of Astragalus cicer microsymbionts. Curr Microbiol 41:142–148
Wood JM, Bremer E, Csonka LN, Kramer R, Poolman B, van der Heide T, Smith LT (2001) Osmosensing and osmoregulatory compatible solute accumulation by bacteria. Comp Biochem Physiol 130:437–460
Yap SF, Lim ST (1983) Response of Rhizobium sp. UMKL 20 to sodium chloride stress. Arch Microbiol 135:224–228
Yelton MM, Yang SS, Edie SA, Lim ST (1983) Characterization of an effective salt tolerant fast growing strain of Rhizobium japonicum. J Gen Microbiol 129:1537–1547
Zahran HH (1999) Rhizobium-legume symbiosis and nitrogen fixation under severe conditions and in an arid climate. Microbiol Mol Biol Rev 63:968–989
Zou N, Dart PJ, Marcar NE, Bushby HVA (1995) Interaction of salinity and rhizobial strain on growth and nitrogen fixation by Acacia ampliceps. Soil Biol Biochem 27:409–413
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wdowiak-Wróbel, S., Leszcz, A. & Małek, W. Salt Tolerance in Astragalus cicer Microsymbionts: The Role of Glycine Betaine in Osmoprotection. Curr Microbiol 66, 428–436 (2013). https://doi.org/10.1007/s00284-012-0293-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-012-0293-2