Abstract
Enteric viruses are shed in the feces and may be present in environmental waters. Their detection in wastewater, even at low concentration, is a major challenge. In this study, recoveries of Echovirus 7 (EV7), virions and RNA in wastewater, using virus concentration methods were determined to evaluate the detection of infectious viruses and the possibility of recovering viral genomes. Two virus concentration methods, PEG precipitation method and two-phase separation method, were applied to recovery experiments of EV7–virions from wastewater, in parallel with recovery experiments of EV7 RNA. The titration of EV7 virions was carried out by cell culture using human rhabdomyosarcoma tumor tissue and the EV7 RNA quantification was performed by real-time PCR. The mean recovery yields of EV7 virions using the PEG precipitation method and the two-phase separation method were 78.5 ± 10.99 and 83.1 ± 0.28 %, respectively. Besides, EV7 RNA recoveries obtained using the PEG precipitation method were four times higher than those using the two-phase separation method. According to our results, the two methods enable to concentrate both infectious viruses and viral genomes. Moreover, considering the protocol time and cost together with the ratio of the EV7 virion recovery to the EV7 RNA recovery, the two-phase separation method (83.1/2.71 %, or 30.6) seems to be more appropriate for selective concentration of viral virions than the PEG precipitation method (78.5/10.33 %, or 7.6).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Viruses are a major cause of water-related diseases [4]. They are constantly found in raw sewage of large populations and in environmental waters affected by fecal contamination, sometimes leading to waterborne outbreaks [28]. According to public health data, the most important enteric viruses are the enteroviruses (EV), adenoviruses (HAdV), noroviruses, rotaviruses, astroviruses as well as hepatitis A (HAV) and E viruses. More than 100 virus types can colonize the gastrointestinal tracts and cause a wide variety of illnesses, inducing variable epidemiological features and some water-related diseases [36].
Enteric viruses are excreted in the feces of infected individuals in high concentrations and transmitted mainly by the fecal–oral route via contaminated water [30]. Indeed, they have been detected in various water samples [8, 9, 13, 19, 26, 33]. The detection of viruses in wastewater, even at low concentrations, is a major challenge. Because usually only a few viral particles are present in wastewater samples, it is necessary to concentrate the viruses from a large volume of water, commonly from 13 ml to 2 l [13, 23]. Hence, many virus concentration methods have been developed: the two-phase separation [11], the polyethylene glycol (PEG) precipitation [20, 21], the ultracentrifugation [15, 34], and methods using an electropositive filter [16] and an electronegative filter [10]. Furthermore, two virus concentration methods from wastewater are advised by WHO for environmental poliovirus surveillance [37]. The first one uses bags with sorbent macroporous glass to trap viruses and the second uses the two-phase separation, which is applied in this study. Regarding the comparison of virus concentration methods from water, several studies have been conducted using quantitative polymerase chain reaction (qPCR) [1, 7, 31]. Until now, to our knowledge, there is only one published study in which the recovery of naked viral genomes from wastewater, using two virus concentration methods, have been evaluated [10].
During the virus replication in the host cell, many proteins and viral genomes are produced from the mRNA separately and assembled into the infectious virions at the end of this step. A large number of viral genomes are released into the feces of patients and are discarded in wastewater. PCR amplification detects genomes of both infectious and non-infectious viruses even though the infectious virus ratio in water remains unknown in the absence of cell culture. Thus, it is important to know the recovery of both infectious and non-infectious viruses in the sample for the concentration method used.
The aim of this study was to compare two virus concentration methods, the PEG precipitation and the two-phase separation, to evaluate the recovery of infectious viruses and the possibility to recover viral genomes of Echovirus 7 (EV7) that were artificially seeded in wastewater. The titer of EV7 was quantified by cell culture and the EV7 RNA was quantitatively determined by qRT-PCR. Wastewater was spiked with EV7 virions and RNA, to provide information about which method is more suitable, taking into consideration time and cost.
Materials and Methods
Spiking Experiments and Wastewater Samples
Raw wastewater samples used for spiking experiments were collected from a wastewater treatment plant (WWTP), located at 70 km from Casablanca in Morocco. One single pool of 26 l was collected in plastic bottles. Eighteen liters were used for EV7 virions experiments and 8 l for EV7 RNA. In order to damage viruses and nucleic acids which could be present in wastewater, samples were first autoclaved at 121 °C for 20 min. Then, they were divided into 52 volumes of 500 ml and stored at +4 °C until use.
For each virus concentration method, two EV7 virions concentrations 1.00E+06 TCID50/100 μl and 1.00E+03 TCID50/100 μl (tissue culture infectious dose 50 %) were seeded in nine replicates in 500 ml of autoclaved wastewater. The EV7 RNA was also seeded in four replicates at two final RNA concentrations: 1.00E+04 and 1.00E+02 RT-PCR Unit (RT-PCR U). In all experiments, negative control without virus or RNA spiking was included. Each spiked sample was vigorously shaken before concentration proceeding. The titer of EV7 virions stock used in spiking experiments was 1.00E+08 TCID50/100 μl and the concentration of EV7 RNA stock used was 1.00E+06 RT-PCR U. The titers were calculated as follows: Virions volume seeded in wastewater = (Concentration of seeded virions × 500 ml)/1.00E+08 TCID50/100 μl and RNA volume seeded in wastewater = (Concentration of seeded RNA × 500 ml)/1.00E+06 RT-PCR U.
Concentration Methods
The seeded wastewater samples were concentrated using two methods. The first one “PEG precipitation” was carried out as described elsewhere [20] with modifications. Five hundred milliliters of sample were clarified by centrifugation (30 min, 1,000×g) and the pellet was resuspended in 10 ml of the supernatant. The remaining portion of supernatant was saved. Chloroform was added to the resuspended sample to a concentration of 10 % (v/v, Sigma, USA) and mixed, and the mixture was centrifuged again (5 min, 1,000×g). The first and second supernatants were combined. The volume was measured and the pH was adjusted to 7.5. The PEG–NaCl technique as previously described [22] was used with modification. The combined supernatants were supplemented with NaCl and PEG until a final concentration of 2.2 % (w) NaCl (Sigma, St Louis, USA) and 7 % (w) PEG 6000 (Fluka, Steinheim, Germany) was obtained. The mixture was stirred 4 h at 4 °C and then centrifuged for 2 h at 2,000×g at 4 °C. Supernatant was discarded and the pellet was suspended in 5 ml of phosphate buffer saline (PBS).
In the second method “two-phase separation” [37], 500 ml of sample was centrifuged for 10 min at 1,000×g. The pellet was stored and the pH of the supernatant was adjusted to neutral (pH 7–7.5). Usually only a few milliliters of NaOH 1 N is needed. The supernatant was mixed with 39.5 ml of 22 % dextran (Sigma Aldrich, St Louis, USA), 287 ml of 29 % PEG 6000 (Fluka, Steinheim, Germany), and 35 ml of 5 N NaCl (Sigma, St Louis, USA) and kept in constant agitation for 1 h at 4 °C using a magnetic stirrer. After overnight incubation at 4 °C in a separation funnel, the entire lower layer and the interphase were collected. The pellet from the first centrifugation was resuspended in this concentrate and the suspension was extracted with 20 % volume of chloroform by shaking vigorously, followed by centrifugation for 5 min at 1,000×g. The supernatant (10 ml) was recovered and decontaminated by antibiotic (e.g., penicillin G and streptomycin to final concentrations of 100 IU/ml and 100 μg/ml, respectively).
Titration of EV7 Virions and Samples
EV7 strain used in this study was obtained from patient fecal samples and maintained in our laboratory. The human rhabdomyosarcoma tumor tissue (RD) cell lines were cultivated in minimum essential medium (MEM, Gibco) with 2 % fetal bovine serum (FBS), penicillin and streptomycin, and were incubated at 37 °C with 5 % CO2 for 5 days.
To determine the viral titer of EV7, the TCID50 values were calculated by the Reed and Müench equation [7, 27]. The titer of EV7 stock used (1.00E+08 TCID50/100 μl) was determined using serial tenfold dilutions in a 96-well plate. Cells were plated at a density of 2 × 104 cells/ml in MEM with 10 % (v/v) FBS. After 24 h of incubation at 37 °C with 5 % of CO2, 100 μl of tenfold serial dilutions of virus were added per well with 8 replicates for each dilution. The plates with MEM with 2 % (v/v) FBS were incubated for 7 days at 37 °C with 5 % CO2. Virus replication was monitored daily under an inverted microscope. Before the titration with the same protocol, wastewater concentrates were filtered though filters with 0.22 μm pore size (Millipore, Billerica, USA). Negative and positive controls were included in each plate.
RNA Extraction
The High Pure Viral Nucleic Acid Kit (Roche, California, USA) was used to extract the RNA from the samples according to manufacturer’s instructions. RNA extracts obtained were performed on 200 μl of sample concentrate and eluted in 50 μl.
Quantification of EV7 RNA
The quantitative reverse transcription-PCR (qRT-PCR) was performed using Superscript III Platinum One-Step Quantitative RT-PCR System (Invitrogen, California, USA) with an ABI PRISM 7500 Fast Real-Time PCR System (Applied Biosystems, California, USA). In brief, five microliters out of 50 μl of the extracted RNA were mixed with 20 μl of a reaction mixture containing 12.5 μl of 2× buffer, 4 mM of MgSO4, 50 nM of ROX Reference Dye, 0.5 μl of Superscript III RT/Platinum Taq Mix, 0.4 μM of forward primer (5′-CCCTGAATGCGGCTAATCC-3′), 0.4 μM of reverse primer (5-ATTGTCACCATAAGCAGCCA-3′), and 0.2 μl of 10 μM TaqMan probe (5′-FAM-AACCGACTACTTTGGGTGTCCGTGTTTC-TAMRA-3′) [25]. Subsequently, the mixture was added to 96-well plate (Applied Biosystems, California, USA) and was incubated at 50 °C for 30 min and 95 °C for 5 min followed by 45 cycles at 95 °C for 15 s and at 60 °C for 1 min.
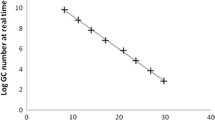
In order to draw the standard curve, EV7 RNA was diluted by serial tenfold dilution. The samples and the standard samples were applied to RT-PCR at the same time followed by analysis using the SDS software (Version 2.0.3, Applied Biosystems), to obtain quantitative data on EV7 RNA RT-PCR U in the well. One RT-PCR U is defined as the concentration of the last positive dilution with Ct value less than 40 cycles. Three wells were used for each sample, negative and positive controls were included in each run. The average was used for the subsequent calculation.
Inhibition Test
To assess the potential presence of PCR inhibitors, the RNA of mengovirus strain MC0 (ATCC VR-1957) (kindly provided by Albert Bosch, Department of Microbiology, University of Barcelona, Barcelona, Spain) was used as an internal control (IC). In each EV7 RNA spiking experiment, a 1.00E+03 RT-PCR U of mengovirus RNA was tenfold diluted in each RNA sample directly after nucleic acid extraction (sample with IC), and compared with positive control which includes tenfold diluted mengovirus RNA in RNase-free water. Negative control was also included.
The quantification of mengovirus was carried out through a real-time procedure following the same conditions as for EV7 RNA with the reverse primer Mengo209 (5′-GAAGTAACATATAGACAGACGCACAC-3′), the forward primer Mengo110 (5′-GCGGGTCCTGCCGAAAGT-3′), and the FAM-MGB probe Mengo147 (5′-ATCACATTACTGGCCGAAGC-3′) [24].
Statistical Analysis
Percent recoveries of EV7 virions and RNA were calculated using the following equation: \( {\text{percent recovery}}\; = \; {\text{the number of recovered viruses / the number of seeded viruses}} \; \times \; 100 \).
The Mann–Whitney U (MWU) test was used for EV7 virions tests, to examine mean recoveries differences from nine trials. The MWU test is a nonparametric version of the independent samples and does not require a normal distribution of the data. The independent variable is the percent recovery. All statistical analyses were carried out with GraphPad Prism software version 5.03 (GraphPad Software, www.graphpad.com). Differences were considered significant if P was < 0.05.
Results
Inhibition Test
Data obtained from the EV7 RNA experiments demonstrated that all samples with IC were found positive for the presence of mengovirus RNA. The comparison between the Ct value mean obtained by the PEG precipitation method and that obtained for positive control showed 30.42 ± 0.37 % (Ct value mean ± SD), while a mean Ct value of 30.62 ± 0.11 % was obtained for the two-phase separation method. That explains the absence of PCR inhibition in all RNA quantification tests.
EV7 Virions/RNA Recoveries
The yields of both the virus concentration methods were determined by spiking EV7 virions and RNA in wastewater using cell culture and qRT-PCR. Results showed that the two methods were able to concentrate both seeded infectious EV7 virions and RNA, as reported in Table 1. The EV7 virions were recovered at a relatively high level, 78.5 ± 10.99 % (Ct mean ± standard deviation) for the PEG precipitation and 83.1 ± 0.28 % for the two-phase separation method. The statistical analysis of EV7 virions indicates that spiked wastewater with 1.00E+06 TCID50/100 μl showed a mean recovery of 70.72 ± 19.37 % for the PEG precipitation, whereas the two-phase separation showed 83.3 ± 11.23 %. However, the difference of the average was not statistically significant (n = 9, P = 0.195, MWU test).
Similar results were obtained for spiked wastewater with 1.00E+03 TCID50/100 μl of EV7 virions. The mean recovery was 86.27 ± 9.88 % for the PEG precipitation, while the two-phase separation showed a mean of 82.9 ± 8.48 %. However, the difference of the average was not statistically significant (n = 9, P = 0.453, MWU test).
Recoveries of EV7 RNA obtained for the PEG precipitation and the two-phase separation were 10.33 ± 2.85 and 2.71 ± 1.95 %, respectively. The mean recovery of EV7 RNA, resulting from four repeated experiments was four times higher for the PEG precipitation than that obtained for the two-phase separation method.
Discussion
Enteroviruses are recommended as a virologic marker for water quality in different studies, because they are environmental resistant, abundant, and easily detected by molecular methods and are of great importance in public health. In the WHO guidelines for environmental surveillance, the poliovirus was used as a model to determine the recovery of two-phase separation concentration method. In this study, EV7 which is an enterovirus was chosen as a model to represent enteric viruses because in our previous study, when the PEG precipitation method was used, no EV7 have been detected in wastewater samples [2]. We suggest that perhaps the PEG precipitation did not concentrate EV7.
PCR problems are commonly related to the presence of enzyme inhibitors which are also concentrated with viral particles [2, 5]. The absence of PCR inhibition in this study showed that the autoclaving may also remove inhibitors from wastewater samples.
The difference between the two methods to concentrate the seeded EV7 virions was not significant, which explains that the two methods were able to concentrate virions similarly for both 1.00E+06 TCID50/100 μl and 1.00E+03 TCID50/100 μl concentrations. For the RNA experiments, the mean recovery obtained using the PEG precipitation was four times higher than that obtained using the two-phase separation method. That may result from the centrifugation time of 2 h used in the PEG precipitation protocol which increased the RNA precipitation. The detection of virions with higher recoveries than those obtained for RNA is in agreement with a previous study [10], reporting the yields of spiked Poliovirus 1 (PV1) virions and RNA, (50 ± 8.9 %) and (3.4 ± 1.6 %) using Mg-method, (22 ± 9.3 %) and (5.2 ± 4.9 %) using Al-method, respectively.
We made some modifications in the PEG precipitation protocol. The elution of pellet was performed using PBS instead of lysis buffer, to be able to amplify the viruses by cell culture after the concentration step. The time of stirring was also reduced to 4 h to decrease the concentration time. Comparing the time and cost of the protocols, the PEG precipitation requires less than 8 h while the two-phase separation requires two half days with overnight incubation. The latter protocol needs more reagent than the PEG precipitation, such as dextran, and twice the quantity of PEG and NaCl, which increases the cost of the method.
Traditionally, the PEG has been used as a virus precipitating agent in water in several studies [14, 17, 21, 32] but it can also precipitate DNA and RNA [3, 18, 35]. Therefore, the ability of PEG to precipitate RNA may provide an increase in the recovery for PCR (e.g., when detecting noroviruses from foods). The purpose of using the EV7 RNA experiments was to project the state of genome in wastewater which could become naked by treatment or transport. In this study, the free RNA was tested to demonstrate that both the virus concentration methods can concentrate RNA, even if the presence of viral genome has not been reported as a causative agent of disease. We suggest that if RNA is not recovered as efficiently as virions by a virus concentration method, the method is expected to have selectivity in detecting virions, although this hypothesis depends on the ratio of infectious virions and RNA (defective virions), as published previously [10].
Although, the presence of viral genome has not been reported as a causative agent of disease. However, the recovery of viral genome should be determined for the virus concentration method used. In previous studies, viruses have been detected from water using cell culture, PCR, or qPCR. Most of them were optimized by spiking different infectious viruses such as: HAV and HAdV [30], HAV [7], Influenza A virus [6], PV1 [29], and Echovirus 1 and Coxsackievirus B5 [12]. The cell culture is useful to detect infectious viruses. Moreover, wastewater concentrate may also be analyzed by molecular biology techniques (frequently PCR or qPCR). The use of PCR is relatively rapid and specific and may detect a small quantity of genome but PCR fails to discern between infectious and non-infectious particles which can be defective virions or naked viral genomes.
In conclusion, our study revealed that the two methods were able to concentrate EV7 virions similarly. The PEG precipitation method resulted in a better recovery of RNA when compared to the two-phase separation. Moreover, considering the protocol time and cost together with the ratio of the EV7 virion recovery to the EV7 RNA recovery, the two-phase separation method (83.1/2.71 %, or 30.6) seems to be more appropriate for selective concentration of virions than the PEG precipitation method (78.5/10.33 %, or 7.6). However, a logical follow-up would be to study the performance of both the methods using wastewater sampling in quest for some enteric viruses to assess the most relevant method for all enteric viruses.
References
Albinana-Gimenez N, Clemente-Casares P, Calgua B, Huguet JM, Courtois S, Girones R (2009) Comparison of methods for concentrating human adenoviruses, polyomavirus JC and noroviruses in source waters and drinking water using quantitative PCR. J Virol Methods 158:104–109
Amdiouni H, Faouzi A, Fariat N, Hassar M, Soukri A, Nourlil J (2012) Detection and molecular identification of human adenoviruses and enteroviruses in wastewater from Morocco. Lett Appl Microbiol 54(4):359–366. doi:10.1111/j.1472-765X.2012.03220.x
Bartram A, Poon C, Neufeld J (2009) Nucleic acid contamination of glycogen used in nucleic acid precipitation and assessment of linear polyacrylamide as an alternative co-precipitant. Biotechniques 47:1019–1022
Bosch A, Guix S, Sano D, Pinto RM (2008) New tools for the study and direct surveillance of viral pathogens in water. Curr Opin Biotechnol 19:295–301
Chapron CD, Ballester NA, Fontaine JH, Frades CN, Margolin AB (2000) Detection of astroviruses, enteroviruses, and adenovirus type 40 and 41 in surface waters collected and evaluated by the information Collection rule and an integrated cell culture-nested PCR procedure. Appl Environ Microbiol 66:2520–2525
Deboosere N, Horm SV, Pinon A, Gachet J, Coldefy C, Buchy P, Vialette M (2011) Development and validation of a concentration method for the detection of influenza a viruses from large volumes of surface water. Appl Environ Microbiol 77:3802–3808
Di Pasquale S, Paniconi M, Auricchio B, Orefice L, Schultz AC, De Medici D (2010) Comparison of different concentration methods for the detection of hepatitis A virus and calicivirus from bottled natural mineral waters. J Virol Methods 165:57–63
Gibbons CD, Rodríguez RA, Tallon L, Sobsey MD (2010) Evaluation of positively charged alumina nanofibre cartridge filters for the primary concentration of noroviruses, adenoviruses and male-specific coliphages from seawater. J Appl Microbiol 109:635–641
Gibson KE, Opryszko MC, Schissler JT, Guo Y, Schwab KJ (2011) Evaluation of human enteric viruses in surface water and drinking water resources in southern Ghana. Am J Trop Med Hyg 84:20–29
Haramoto E, Katayama H, Oguma K, Ohgaki S (2007) Recovery of naked viral genomes in water by virus concentration methods. J Virol Methods 142:169–173
Hovi T, Stenvik M, Partanen H, Kangas A (2001) Poliovirus surveillance by examining sewage specimens: quantitative recovery of virus after introduction into sewerage at remote upstream location. Epidemiol Infect 127:101–106
Ikner LA, Soto-Beltran M, Bright KR (2011) New method using a positively charged microporous filter and ultrafiltration for concentration of viruses from tap water. Appl Environ Microbiol 77:3500–3506
Kamel AH, Ali MA, El-Nady HG, Aho S, Pothier P, Belliot G (2010) Evidence of the co-circulation of enteric viruses in sewage and in the population of greater Cairo. J Appl Microbiol 108:1620–1629
Kiulia NM, Netshikweta R, Page NA, Van Zyl WB, Kiraithe MM, Nyachieo A, Mwenda JM, Taylor MB (2010) The detection of enteric viruses in selected urban and rural river water and sewage in Kenya, with special reference to rotaviruses. J Appl Microbiol 109:818–828
Kokkinos K, Filippidou S, Karlou K, Vantarakis A (2010) Molecular typing of enteroviruses, adenoviruses, and hepatitis A viruses in untreated and treated sewage of a biological treatment plant in Greece. Food Environ Virol 2:89–96
Lee H, Kim M, Paik SY, Lee CH, Jheong WH, Kim J, Ko G (2011) Evaluation of electropositive filtration for recovering norovirus in water. J Water Health 9:27–36
Liang S, Xie GC, Xu ZQ, Li JS, Li DT, Feng SL, Duan ZJ (2011) Study on concentration of nuorovirus gene group II from environmental water. Bing Du Xue Bao 27:58–63
Lu C, Meyers BC, Green PJ (2007) Construction of small RNA cDNA libraries for deep sequencing. Methods 43:110–117
Maunula L, Miettinen IT, von Bonsdorff CH (2005) Norovirus outbreaks from drinking water. Emerg Infect Dis 11:1716–1721
Meleg E, Jakab F, Kocsis B, Banyai K, Melegh B, Szucs G (2006) Human astroviruses in raw sewage samples in Hungary. J Appl Microbiol 101:1123–1129
Melegh B, Szucs G (2008) Detection and quantification of group C rotaviruses in communal sewage. Appl Environ Microbiol 74:3394–3399
Minor PD (1985) Growth, assay and purification of picornaviruses. Mahy BW (ed) Virology, a practical approach, IRL Press, Oxford/Washington, pp 25–41
Myrmel M, Berg EM, Grinde B, Rimstad E (2006) Enteric viruses in inlet and outlet samples from sewage treatment plants. J Water Health 4:197–209
Pintó RM, Costafreda MI, Bosch A (2009) Risk assessment in shellfish-borne outbreaks of hepatitis A. Appl Environ Microbiol 75:7350–7355
Piqueur MA, Verstrepen WA, Bruynseels P, Mertens AH (2009) Improvement of a real-time RT-PCR assay for the detection of enterovirus RNA. Virol J 6:95
Prado T, Silva DM, Guilayn WC, Rose TL, Gaspar AM, Miagostovich MP (2011) Quantification and molecular characterization of enteric viruses detected in effluents from two hospital wastewater treatment plants. Water Res 45:1287–1297
Reed JL, Muench H (1938) A simple method of estimating per cent endpoint. Am J Hyg 27:493–497
Reynolds KA, Mena KD, Gerba CP (2008) Risk of waterborne illness via drinking water in the United States. Rev Environ Contam Toxicol 192:117–158
Rhodes ER, Hamilton DW, See MJ, Wymer L (2011) Evaluation of hollow-fiber ultrafiltration primary concentration of pathogens and secondary concentration of viruses from water. J Virol Methods 176:38–45
Rigotto C, Kolesnikovas CK, Moresco V, Simões CM, Barardi CR (2009) Evaluation of HA negatively charged membranes in the recovery of human adenoviruses and hepatitis A virus in different water matrices. Mem Inst Oswaldo Cruz 104:970–974
Schultz AC, Perelle S, Di Pasquale S, Kovac K, De Medici D, Fach P, Sommer HM, Hoorfar J (2011) Collaborative validation of a rapid method for efficient virus concentration in bottled water. Int J Food Microbiol 145:158–166
Shieh YC, Wong CI, Krantz JA, Hsub FC (2007) Detection naturally occurring enteroviruses in waters using direct RT-PCR integrated cell culture-RT-PCR. J Virol Methods 149:184–189
Tong HI, Lu Y (2011) Effective detection of human adenovirus in Hawaiian waters using enhanced PCR methods. Virol J 8:57
Vantarakis A, Papapetropoulou M (1998) Detection of enteroviruses, adenoviruses in sea water in southern Greece. Water Res 32:2365–2372
Wang XW, Xiong AS, Yao QH, Zhang Z, Qiao YS (2010) Direct isolation of high-quality low molecular weight RNA of pear peel from the extraction mixture containing nucleic acid. Mol Biotechnol 44:61–65
WHO (2001) Waterborne zoonoses. http://www.who.int/water_sanitation_health/diseases/zoonoses.pdf. Accessed 15 Sep 2011
WHO (2003) Guidelines for environmental surveillance of poliovirus circulation. Geneva, Switzerland. http://whqlibdoc.who.int/hq/2003/WHO_V&B_03.03.pdf. Accessed 20 Feb 2009
Acknowledgments
The authors thank Prof Albert Bosch from the University of Barcelona (Barcelona, Spain) for providing us with the internal control.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Amdiouni, H., Maunula, L., Hajjami, K. et al. Recovery Comparison of Two Virus Concentration Methods from Wastewater Using Cell Culture and Real-Time PCR. Curr Microbiol 65, 432–437 (2012). https://doi.org/10.1007/s00284-012-0174-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-012-0174-8