Abstract
A thermophilic anaerobic bacterium (strain TH7C1T) was isolated from the hydrothermal hot spring of Guelma in the northeast of Algeria. Strain TH7C1T stained Gram-positive, was a non-motile rod appearing singly, in pairs, or as long chains (0.7–1 × 2–6 μm2). Spores were never observed. It grew at temperatures between 55 and 75°C (optimum 65°C) and at pH between 6.2 and 8.3 (optimum 6.9). It did not require NaCl for growth, but tolerated it up to 5 g l−1. Strain TH7C1T is an obligatory heterotroph fermenting sugars including glucose, galactose, lactose, raffinose, fructose, ribose, xylose, arabinose, maltose, mannitol, cellobiose, mannose, melibiose, saccharose, but also xylan, and pyruvate. Fermentation of sugars only occurred in the presence of yeast extract (0.1%). The end-products from glucose fermentation were acetate, lactate, ethanol, CO2, and H2. Nitrate, nitrite, thiosulfate, elemental sulfur, sulfate, and sulfite were not used as electron acceptors. The G+C content of the genomic DNA was 44.7 mol% (HPLC techniques). Phylogenetic analysis of the small-subunit ribosomal RNA (rRNA) gene sequence indicated that strain TH7C1T was affiliated to Firmicutes, order Clostridiales, family Caldicoprobacteraceae, with Caldicoprobacter oshimai (98.5%) being its closest relative. Based on phenotypic, phylogenetic, and genetic characteristics, strain TH7C1T is proposed as a novel species of genus Caldicoprobacter, Caldicoprobacter algeriensis, sp. nov. (strain TH7C1T = DSM 22661T = JCM 16184T).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Several thermophilic members of orders Thermoanaerobacterales [31] and Clostridiales [24], class Clostridia are considered as common inhabitants of terrestrial hot springs where they most probably participate to fermentative processes of organic matter [28]. They are considered as strict anaerobes with representatives of the order Thermoanaerobacterales (e.g., Thermoanaerobacter and Thermoanaerobacterium spp.) being frequently isolated from such extreme ecosystems and having the ability to use thiosulfate as terminal electron acceptor and reduce it to sulfide or elemental sulfur [21, 22]. Within the order Clostridiales, most of bacterial species isolated from terrestrial hot springs pertain to family Syntrophomonadaceae (e.g., Carboxydocella, Caldicellulosiruptor, and Anaerobranca spp.) and Peptococcaceae (e.g., Desulfotomaculum, Carboxythermus, and Thermincola spp.) with few representatives pertaining to the families Acidaminococcaceae (e.g., Thermosinus carboxydivorans), Heliobacteriaceae (e.g., Heliobacterium modesticaldum), and Clostridiaceae [2, 8–10, 23, 24, 26, 27, 32] (For more information, see review of Wagner and Wiegel [28]).
It is only recently that hydrothermal ecosystems have been exploited in Algeria (eastern Algeria) to look for novel microorganisms inhabiting local hot springs. Hammam E’Dbagh (Guelma) is the hottest spring in this country, with temperatures rising up to 98°C.
Here we report on the isolation of a novel thermophilic anaerobic bacterium from this Algerian hot spring which belongs to the Firmicutes, order Clostridiales and presents significant phenotypic, genetic, and phylogenetic traits with its closest relative Caldicoprobacter oshimai [34], family Caldicoprobacteraceae to be proposed as a novel species within this genus.
Materials and Methods
Source of Sampling
Samples were collected from a terrestrial hot spring, located at 20 km from Guelma, northeast of Algeria (70°25′ East, 36°27′ North), at an altitude of 320 m. The water bearing zone was near the surface. Water contained chloride (370 mg l−1), sulfate (385 mg l−1), sodium (240 mg l−1), calcium (130 mg l−1), and sulfide. The temperature at the sampling site (Hammam D’bagh or Meskhoutine) was 98°C, and pH was 7.3. The samples were collected under anaerobic conditions and were transported to the laboratory at ambient temperature.
Isolation and Culture Techniques
Strict anaerobic procedures were followed for isolation and culture of microorganisms as previously reported by Hungate [16]. Selective medium for isolation included (g l−1): NH4Cl (1.0), K2HPO4 (0.3), KH2PO4 (0.3), KCl (0.1), MgCl2·6H2O (0.5), CaCl2·2H2O (0.1), NaCl (0.5), yeast extract (2.0), biotrypcase (2.0), Cysteine–HCl (0.5), together with sodium acetate (2 mM) and Balch trace element solution (10 ml) [3].
The pH was adjusted to 7.2 with 10 M KOH solution, and the medium was boiled and cooled to room temperature under a stream of O2-free N2 gas. Aliquots of 5 ml were dispensed into Hungate tubes, degassed under N2–CO2 (80:20 v/v), and subsequently sterilized by autoclaving at 120°C for 20 min. Before inoculation, 0.1 ml of 10% (w/v) NaHCO3, 0.1 ml of 2% (w/v) Na2S·9H2O, and 20 mM glucose were injected from sterile stock solutions into the tubes.
Enrichments were performed in Hungate tubes or serum bottles inoculated with 10% of sample and incubated at 70°C. The culture was purified by repeated use of the Hungate roll tube method, using gelrite solid medium and transferred into liquid medium as previously described [12].
Optimum Growth Conditions
The pH, temperature, and NaCl concentration ranges for growth were determined using basal medium supplemented with 20 mM glucose. The different pH (5–9) of the medium were adjusted by injecting in Hungate tubes aliquots of anaerobic stock solution of 0.1 M HCl (acidic pH), 10% NaHCO3, or 8% Na2CO3 (basic pH). Water baths were used for incubating bacterial cultures from 45 to 90°C. NaCl requirement was determined by directly weighing NaCl in Hungate tubes before dispensing medium. Cultures were subcultured at least twice under the same experimental conditions before determination of growth rates and use of substrates.
Morphological Studies
The Gram reaction was determined with heat fixed liquid cultures stained with Difco kit reagents. For electron microscopy, exponentially growth cells were negatively stained with 1% sodium phosphotungstic acid (pH 7.2). Whole cells were observed with a Hitachi model H 600 electron microscope at an accelerating voltage of 75 kV. The presence of spores was analyzed by phase-contrast microscopic observations of young and old cultures and pasteurization tests, performed at 80, 90, and 100°C for 10 and 20 min.
Substrates Utilization Tests
Substrates (glucose, ribose, sucrose, fructose, galactose, lactose, raffinose, melibiose, xylose, arabinose, maltose, melibiose, mannose, mannitol, saccharose, cellobiose, gelatine, casaminoacids, pyruvate, fumarate, succinate, ethanol, with the exception of H2/CO2 (2 bars), formate (40 mM), xylan (mix of birchwood and oats spelt) (10 g l−1), starch (10 g l−1), and peptone (10 g l−1) were tested at a final concentration of 20 mM in growth medium that lacked glucose. To test for electron acceptors, sodium thiosulfate (20 mM), sodium sulfate (20 mM), sodium sulfite (2 mM), elemental sulfur (10 g l−1), sodium nitrate (20 mM), and sodium nitrite (2 mM) were added to the medium.
Analytical Methods
The determination of fatty acids composition of strain TH7C1T was performed by DSMZ (Deutsche Sammlung von Mikroorganismen und Zellkulturen, Braunschweig, Germany) services. To test for antibiotic susceptibility, ampicillin, chloramphenicol from filter sterilized stock solutions were each added at a final concentration of 50 and 100 μg ml−1.
Enzyme Assays
For xylanolytic activity measurements, cells were harvested in the late exponential or early stationary phase. Reducing sugars were quantified with dinitrosalicylic acid [20]. Xylanolytic activity was assayed in the supernatant and in re-suspended cells by measuring the release of reducing sugars from xylan. Each assay mixture consisted of 0.5% xylan supplemented with 100 mM acetate buffer (pH 6.5) and enzyme so that the final volume was 0.2 ml. It was incubated for 30 min at 70°C. The assay was stopped by adding dinitrosalicylic acid, and xylose released from xylan was measured at 540 nm. Controls with substrate and no enzyme were included.
Determination of G+C Content and DNA–DNA Hybridization
The G+C content of DNA was determined at DSMZ. The DNA was isolated and purified by chromatography on hydroxyapatite, and the G+C content was determined by HPLC as described by Mesbah et al. [19]. The hybridization was also done at DSMZ services.
16S rRNA Sequence Studies
Methods for purification of the DNA, PCR amplification, and sequencing of the 16S rRNA gene were performed as described previously [4, 6, 11, 30]. The partial sequences generated were assembled using BioEdit v. 5.0.9. [14], and the consensus sequence of 1,525 nucleotides was corrected manually for errors. The most closely related sequences in GenBank (version 178), the Ribosomal Database Project (release 10) [5] identified using BLAST [1], and the Sequence Match program [7] were extracted and aligned. The consensus sequence of 1,179 nucleotides was then manually adjusted to conform to the 16S rRNA secondary structure model [33].
Nucleotide ambiguities were omitted and evolutionary distances were calculated using the Jukes and Cantor option [17]. Dendrograms were constructed with TREECON program using the neighbor-joining method [25]. Tree topology was re-examined by the bootstrap method (1,000 replications) of resampling [13]. Its topology was also supported using the maximum-parsimony and maximum-likelihood algorithms. The 16S rRNA sequence of strain TH7C1T has been deposited in the GenBank database under accession number GU216701.
Results
Enrichment and Isolation
A 0.5 ml aliquot of sample was inoculated into Hungate tubes containing 5 ml of basal medium and glucose as energy source. The tubes were then incubated at 70°C. To obtain pure cultures, the enrichment was subcultured several times under the same growth conditions prior the isolation. For isolation, the culture was serially diluted tenfold in roll tubes. Several colonies that developed were picked separately. The process of serial dilution was repeated several times until the isolates were deemed to be axenic. Several strains similar in morphology and producing the same end-products from glucose metabolism were isolated: strains C52, Cb-EL, CUE-CA, DC-GUE, TG-C2r, TG-C44, and strain TH7C1. Comparative 16S rRNA gene sequence similarity analysis revealed that the similarities among strains TH7C1, C52, Cb-EL, CUE-CA, DC-GUE, TG-C2r, and TG-C44 were more than 99.5%.
The strain designated TH7C1T was selected and used for further characterization.
Colony and Cell Morphology
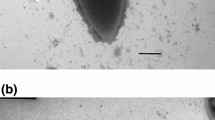
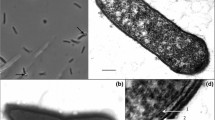
The colonies obtained in roll tubes were round, creamy, and pale yellow. They were 2–3 mm in diameter after 3 days of incubation at 70°C. Strain TH7C1T was a non-motile rod-shaped bacterium occurring singly, in pairs or occasionally as long chains with size ranging from 0.7–1.0 to 2.0–6.0 μm (Fig. 1a). Spores were not observed. Cells stained Gram-positive. The cell wall structure of strain TH7C1T was a single layer (Gram-type positive) (Fig. 1b).
Optimum Growth Conditions
Strain was strictly anaerobic and thermophilic. The optimal growth temperature was 65°C (range 55–75°C; no growth at 50 and 80°C). The isolate did not require NaCl for growth. The optimum pH range for growth was 6.9 (range 6.2–8.3).
Metabolic Properties
The isolate required yeast extract (0.1%) for growth which could not be replaced by peptides, biotrypcase, or vitamins. Sulfate, thiosulfate, elemental sulfur, sulfite, nitrate, and nitrite were not used as terminal electron acceptors. Strain TH7C1T grew on glucose, galactose, lactose, raffinose, fructose, ribose, xylose, arabinose, maltose, mannitol, cellobiose, mannose, melibiose, saccharose, xylan, and pyruvate, but not on casaminoacids, peptone, gelatine, starch, ethanol, succinate, formate, and fumarate. The main end-products resulting from glucose fermentation were acetate, lactate, ethanol, CO2 and H2. In optimal growth conditions, generation time was around 2 h. Strain TH7C1T grew with xylan as energy source. Xylanolytic activity was found extracellular. Strain TH7C1T was susceptible to ampicillin (150 μg ml−1) and to chloramphenicol (50 μg ml−1).
Cellular Fatty Acids
The major membrane fatty acids present in strain TH7C1T were C17:0 (iso) (25% of total fatty acids) and anteiso (13%), C16:0 (15%), C15:0 (13%) acids whereas minor fatty acids were C15:0 anteiso, C17:0 (iso 3OH and 2OH), and C18:0 (w9c and iso) acids.
G+C Content of DNA, Phylogeny, and DNA–DNA Hybridization Studies
The G+C content of strain TH7C1T was 44.7 mol%. Sequence alignment and subsequent comparisons with sequences of representative members of the domain Bacteria consistently placed strain TH7C1T within the phylum Firmicutes, class Clostridia, order Clostridiales, family Caldicoprobacteraceae, with Caldicoprobacter oshimai JW/HY-331T being its closest phylogenetic (98.5% similarity) (Fig. 2). DNA–DNA hybridization studies revealed low homology (45.5 ± 2%) between strain TH7C1T and C. oshimai.
Neighbor-joining phylogenetic dendrogram based on 1,179 unambiguous nucleotides of 16S rRNA gene sequences showing the relationship between strain TH7C1T pertaining to the new family Caldicoprobacteraceae and selected organisms belonging to the families Thermoanaerobacteraceae and Clostridiaceae. The dendrogram was constructed proceeding from 16S rRNA gene sequences of the type strains of the type species; GenBank accession numbers are shown in parentheses. Numbers at branch points specify the reliability of the branching order determined for 1,000 resamplings; only bootstrap values above 80% are shown. The tree was rooted using the sequence of Halanaerobium praevalens (GenBank accession no. M 59123) as an out group (not shown). Bar 0.05 substitutions per nucleotide position
Discussion
There are few studies on thermophilic anaerobes inhabiting the numerous terrestrial hot springs located in Algeria so far. To our knowledge, it is only recently that a hyperthermophilic archaeon pertaining to the genus Pyrococcus has been isolated from northeast of this country [18]. The latter microorganism was described as obligatory anaerobic, heterotrophic utilizing proteinaceous compounds and reducing elemental sulfur to sulfide [18]. Here we report on a novel thermophilic anaerobic bacterium (strain TH7C1T), also isolated from northeast Algeria, unable to use sulfur compounds as terminal electron acceptors, which pertains to the order Clostridiales, family Caldicoprobacteraceae, consisting of a single genus and single species Caldicoprobacter oshimai [34]. Not only phylogenetic, but also chemotaxonomic and metabolic characteristics are in agreement with strain TH7C1T as being a member of Caldicoprobacter. Indeed, similarly to C. oshimai, the major cellular fatty acids of strain TH7C1T were branched, saturated fatty acids with odd numbers of carbon atoms (e.g., iso- and anteiso-C17:0, and iso-C15:0). Moreover, for both bacteria, lactate, acetate, ethanol, CO2, and H2 were the end-products of glucose metabolism. However, strain TH7C1T differed markedly from C. oshimai by the absence of spores, its lower tolerance to NaCl, and the use of mannitol (Table 1). In addition DNA–DNA homology between these microorganisms is sufficiently low (45.5%) to insure strain TH7C1T as being a novel species [29] of Caldicoprobacter within the family Caldicoprobacteraceae. Interestingly, C. oshimai was isolated from sheep feces and had clone OTU4 (99.5% similarity) retrieved from cow feces enrichment cultures as its closest phylogenetic relative, thus demonstrating that similar microorganisms, despite being most probably dormant, might prevail in herbivore feces [34]. Based on molecular approaches, two unidentified clones having similarities of 97.7% (Hb), and 95.7% (LNE) with C. oshimai have been detected in thermophilic enrichment cultures inoculated with bioreactor sludge and a hot spring sample, respectively [15, 34]. In this respect, we may expect that beside strain TH7C1T, other closely phylogenetic relatives of C. oshimai pertaining to family Caldicoprobacteraceae also inhabit springs thermal where in contrast to feces, their metabolic contribution may be considered. However, further experiments are needed to ascertain that such microorganisms might be of ecological significance in these hot natural environments.
Finally, the isolation of strain TH7C1T as a member of order Clostridiales, family Caldicoprobacteraceae, from an Algerian hot spring extends our knowledge on the microbial diversity inhabiting such extreme ecosystems. Based on its phenotypic, genetic, and phylogenetic characteristics, we propose strain TH7C1T to be assigned to as a novel species within the Firmicutes, Caldicoprobacter algeriensis, sp. nov.
Emended Description of the Genus Caldicoprobacter
The genus description is the same as that given by Yokoyama et al. [34] except that abilities to form spores and to reduce mannitol are variable.
Description of Caldicoprobacter algeriensis sp. nov. (al.ge.ri.en’sis. M.L. fem. N. referring to Algeria, the country where the bacterium was first recovered). Cells stain Gram-positive. They have a Gram-positive type of cell wall. They are non-motile rods appearing singly (0.7–1 to 2–6 μm), in pairs, or occasionally as long chains. Spores are not observed. Strictly anaerobic and fermenting sugars uses glucose, galactose, lactose, raffinose, fructose, ribose, xylose, arabinose, maltose, mannitol, cellobiose, mannose, melibiose, saccharose, xylan, and pyruvate, but not casaminoacids, peptone, gelatine, starch, ethanol, succinate, formate, and fumarate. Sulfate, thiosulfate, elemental sulfur, sulfite, nitrate, and nitrite are not used as terminal electron acceptors. Optimum temperature is 65°C (range 55–75°C; no growth at 50 and 80°C). Optimum pH at 6.9 (no growth at 6.3 and 8.6). Growth occurs at NaCl concentration ranging from 0 to 0.5% (w/v). Under optimal growth conditions, the generation time is 2 h. The major membrane fatty acids are C17:0 (iso and anteiso), C16:0, and C15:0 acids.
The G+C content of genomic DNA is 44.7%. The type strain is strain TH7C1T (= DSM 22661T = JCM 16184T) which was isolated from a thermal hot spring located at 20 km from Guelma in the northeast of Algeria (70°25′ East, 36°27′ North), at an altitude of 320 m.
References
Altschul SF, Madden TL, Schaffer AA et al (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Asao M, Madigan MT (2009) Family IV. Heliobacteriaceae. In: De Vos P, Garrity GM, Jones D, Krieg NR, Ludwig W, Rainey FA, Schleifer KH, Whitman B (eds) The Firmicutes, vol 3, 2nd edn. Springer, Dordrecht, pp 913–916
Balch WE, Fox GE, Magrum LJ, Woese CR et al (1979) Methanogens: reevaluation of a unique biological group. Microbiol Rev 43:260–296
Ben Dhia Thabet O, Fardeau ML, Joulian C et al (2004) Clostridium tunisiense sp. nov., a new proteolytic, sulphur-reducing bacterium isolated from an olive mill wastewater contaminated by phosphogypse. Anaerobe 10:185–190
Benson DA, Karsch-Mizrachi I, Lipman DJ et al. (2008) GenBank. Nucleic Acids Res 36(Database issue): D25–D30. doi:10.1093/nar/gkn979
Cashion P, Holder-Franklin MA, McCully J et al (1977) A rapid method for the base ratio determination of bacterial DNA. Anal Biochem 81:461–466
Cole JR, Wang Q, Cardenas E, Fish J et al. (2009) The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res 37(Database issue): D141–D145. doi:10.1093/nar/gkn879
Collins MD, Lawson PA, Willems A et al (1994) The phylogeny of the genus Clostridium: proposal of five new genera and eleven new species combinations. Int J Syst Bacteriol 44:812–826
Engle M, Li Y, Rainey F et al (1996) Thermobrachium celere gen. nov., sp. nov., a rapidly growing thermophilic, alkalitolerant, and proteolytic obligate anaerobe. Int J Syst Bacteriol 46:1025–1033
Esaki T (2009) Family VI. Peptococcaceae. In: De Vos P, Garrity GM, Jones D, Krieg NR, Ludwig W, Rainey FA, Schleifer KH, Whitman B (eds) The Firmicutes, vol 3, 2nd edn. Springer, Dordrecht, pp 969–971
Escara JF, Hutton JR (1980) Thermal stability and renaturation of DNA in dimethylsulphoxide solutions: acceleration of renaturation rate. Biopolymers 19:1315–1327
Fardeau ML, Ollivier B, Cayol JL (2009) Genus III. Caldanaerobacter. In: De Vos P, Garrity GM, Jones D, Krieg NR, Ludwig W, Rainey FA, Schleifer KH, Whitman B (eds) The Firmicutes, vol 3, 2nd edn. Springer, Dordrecht, pp 1241–1244
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hobel CFV, Marteinsson VT, Hauksdóttir S et al (2004) Use of low nutrient enrichments to access novel amylase genes in silent diversity of thermophiles. World J Microbiol Biotechnol 20:801–809
Hungate RE (1969) A roll tube method for the cultivation of strict anaerobes. In: Norris JR, Ribbons DW (eds) Methods in microbiology, vol 3B. Academic Press, New York, pp 117–132
Jukes TH, Cantor CR (1969) Evolution of protein molecules. In: Munro HN (ed) Mammalian protein metabolism. Academic Press, New York, pp 21–132
Kecha M, Benallaoua S, Touzel JP et al (2007) Biochemical and phylogenetic characterization of a novel terrestrial hyperthermophilic archaeon pertaining to the genus Pyrococcus from an Algerian hydrothermal hot spring. Extremophiles 11:65–73
Mesbah M, Premachandran U, Whitman WB (1989) Precise measurement of the G+C content of deoxyribonucleic acid by high-performance liquid chromatography. Int J Syst Bacteriol 39:159–167
Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugars. Anal Chem 31:426–442
Onyenwoke RU, Wiegel J (2009) Genus II. Thermoanaerobacterium. In: De Vos P, Garrity GM, Jones D, Krieg NR, Ludwig W, Rainey FA, Schleifer KH, Whitman B (eds) The Firmicutes, vol 3, 2nd edn. Springer, Dordrecht, pp 1279–1287
Onyenwoke RU, Wiegel J (2009) Genus I. Thermoanaerobacter. In: De Vos P, Garrity GM, Jones D, Krieg NR, Ludwig W, Rainey FA, Schleifer KH, Whitman B (eds) The Firmicutes, vol 3, 2nd edn. Springer, Dordrecht, pp 1225–1239
Patel BKC, Monk C, Littleworth H et al (1987) Clostridium fervidus sp. nov., a new chemoorganotrophic acetogenic thermophile. Int J Syst Bacteriol 37:123–126
Rayney FA (2009) Order I. Clostridiales. In: De Vos P, Garrity GM, Jones D, Krieg NR, Ludwig W, Rainey FA, Schleifer KH, Whitman B (eds) The Firmicutes, vol 3, 2nd edn. Springer, Dordrecht, p 736
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:405–425
Sekiguchi Y (2009) Family Syntrophomonadaceae. In: De Vos P, Garrity GM, Jones D, Krieg NR, Ludwig W, Rainey FA, Schleifer KH, Whitman B (eds) The Firmicutes, vol 3, 2nd edn. Springer, Dordrecht, pp 1044–1045
Slobodkin AI, Tourova TP, Kostrikina NA et al (2006) Tepidimicrobium ferriphilum gen. nov., sp. nov., a novel moderately thermophilic, Fe(III)-reducing bacterium of the order Clostridiales. Int J Syst Evol Microbiol 56:369–372
Wagner ID, Wiegel J (2008) Diversity of thermophilic anaerobes. Ann N Y Acad Sci 1125:1–43
Wayne LG, Brenner DJ, Colwell RR et al (1987) Report of the Ad Hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464
Weisburg WG, Barns SM, Pelletier DA et al (1991) 16S Ribosomal DNA Amplification for phylogenetic study. J Bacteriol 173:697–703
Wiegel J (2009) Order III. Thermoanaerobacterales. In: De Vos P, Garrity GM, Jones D, Krieg NR, Ludwig W, Rainey FA, Schleifer KH, Whitman B (eds) The Firmicutes, vol 3, 2nd edn. Springer, Dordrecht, p 1224
Wiegel J (2009) Family I. Clostridiaceae. In: De Vos P, Garrity GM, Jones D, Krieg NR, Ludwig W, Rainey FA, Schleifer KH, Whitman B (eds) The Firmicutes, vol 3, 2nd edn. Springer, Dordrecht, pp 736–738
Winker S, Woese CR (1991) A definition of the domains, Archaea, Bacteria and Eucarya in terms of small subunit ribosomal RNA characteristics. Syst Appl Microbiol 13:161–165
Yokoyama H, Wagner ID, Wiegel J (2010) Caldicoprobacter oshimai gen. nov., sp. nov., an anaerobic, xylanolytic, extremely thermophilic bacterium isolated from sheep faeces, and proposal of Caldicoprobacteraceae fam. nov. Int J Syst Evol Microbiol 60:67–71
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bouanane-Darenfed, A., Fardeau, ML., Grégoire, P. et al. Caldicoprobacteralgeriensis sp. nov. a New Thermophilic Anaerobic, Xylanolytic Bacterium Isolated from an Algerian Hot Spring. Curr Microbiol 62, 826–832 (2011). https://doi.org/10.1007/s00284-010-9789-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-010-9789-9