Abstract
Genetic engineering of Lactococcus lactis to produce a heterologous protein may cause potential risks to the environment despite the industrial usefulness of engineered strains. To reduce the risks, we generated three auxotrophic recombinant L. lactis subsp. lactis IL1403 strains expressing a heterologous protein, BmpB, using thyA- and alr-targeting integration vectors: ITD (thyA − alr + bmpB +), IAD (thyA + alr − bmpB +), and ITDAD (thyA − alr − bmpB +). After construction of integration vectors, each vector was introduced into IL1403 genome. Integration of BmpB expression cassette, deletion of thyA, and inactivation of alr were verified by using PCR reaction. All heterologous DNA fragments except bmpB were eliminated from those recombinants during double crossover events. By using five selective agar plates, we also showed thymidine auxotrophy of ITD and ITDAD and d-alanine auxotrophy of IAD and ITDAD. In M17G and skim milk (SYG) media, the growth of the three recombinants was limited. In MRS media, the growth of IAD and ITDAD was limited, but ITD showed a normal growth pattern as compared with the wild-type strain (WT). All the recombinants showed maximal BmpB expression at an early stationary phase when they were cultivated in M17G supplemented with thymidine and d-alanine. These results suggest that auxotrophic recombinant L. lactis expressing a heterologous protein could be generated to reduce the ecological risks of a recombinant L. lactis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lactic acid bacteria (LAB) are food-grade microorganisms widely used in the food industry. For example, Lactococcus lactis is one species of LAB used for fermented dairy products such as cheese and fermented milk [16]. Genetic engineering of L. lactis has been used to enhance the probiotic potential of wild-type (WT) strains [18]. They have been genetically engineered to express enzymes or antigens, and to produce functional molecules.
Although the genetically engineered LAB (GE-LAB) are useful in the food industry, they may pose potential risks to the ecosystem including human [5]. The well-known problems with GE-LAB are the predominance of themselves in unwanted places and the dissemination of antibiotic resistance genes derived from them [15]. To overcome those problems, the survival of GE-LAB released into the environment can be limited by introducing a mechanism to kill them. Unnecessary DNA elements, including antibiotic resistance genes, should also be eliminated from them.
Two useful biological mechanisms to kill LAB cells have been reported. The first mechanism is the active containment based on cell lysis by cell wall-degrading enzymes such as enterolysin A [6] and autolysin [3]. However, the expression control of killing enzymes is challenging when GE-LAB are released into the environment, and the killing effect may be inactivated by spontaneous mutations in the killing genes or controlling promoters for the genes. The second mechanism is the passive containment, which is based on auxotrophy for specific nutrients such as thymidine [7, 14] and d-alanine [2, 13]. However, auxotrophic GE-LAB for a single specific nutrient can grow and survive where the nutrient is present. To overcome the shortcomings associated with the above strategies, dual lethal systems have been reported [11, 17].
In this study, we combined two kinds of auxotrophy, thymidine and d-alanine, to introduce a dual lethal system into recombinant L. lactis expressing a heterologous protein, BmpB, which has been reported to be a vaccine candidate against swine dysentery [9]. Growth in LAB media and BmpB expression of the recombinant L. lactis strain were evaluated and compared with those of WT and single auxotrophic strains.
Materials and Methods
Bacterial Strains and Plasmids
The bacterial strains and plasmids used in this study are listed in Table 1. Escherichia coli Top10F′ was used as a host bacterium for the construction of each integration vector targeting IL1403 thyA or alr.
Culture Media and Cultivation
Escherichia coli and L. lactis were cultivated in Luria–Bertani broth and M17 broth containing 0.5% (w/v) glucose (M17G), respectively. The antibiotics used were ampicillin (100 μg/ml) for E. coli and erythromycin (5 μg/ml) for L. lactis. When indicated, 40 μM of thymidine or 200 μg/ml of d-alanine was added to the media. E. coli was cultivated at 37°C with agitation, and L. lactis was cultivated at 30°C without agitation.
Construction of Integration Vectors
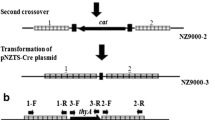
As shown in Fig. 1a, integration vectors were constructed using the following DNA fragments; thyA flanking regions (thyA F1 and thyA F2), alr flanking regions (alr F1 and alr F3), ermAM with a promoter (Pro.ermAM), and modified bmpB with the IL1403 pfkA promoter (Ppfk.bmpB). These fragments were amplified by PCR using the primers shown in Table 2 and cloned into the pGEM-T easy plasmid. IL1403 genomic DNA was used as a template to clone thyA F1, thyA F2, alr F1, alr F3, and the pfkA promoter (Ppfk). The pILPpfk.Mb plasmid was used as a template to clone Pro.ermAM and modified bmpB, which encodes the partial cytosolic BmpB protein. Two thyA flanking regions were cut with XbaI, and they were ligated to each other by T4 DNA ligase. The ligated fragment (thyAF1.F2) was amplified by PCR and cloned into the pGEM-T easy plasmid to generate the pT.thyAF1.F2 plasmid. Pro.ermAM was cut with SalI and SacI, and inserted into the SalI and SacI site of pT.thyAF1.F2 to generate a thyA-targeting integration backbone pT.I2 plasmid. Similarly, the alr-targeting integration backbone pA.I2 plasmid was generated using the pGEM-T easy, alrF1.F3, and Pro.ermAM. After construction of the two integration backbone plasmids, Ppfk.bmpB was cut with SpeI and XhoI, and inserted into the XbaI and XhoI site of pT.I2 and pA.I2, respectively, to generate the final integration vectors, pT.Ppfk.MbI2 and pA.Ppfk.MbI2.
Analysis of Nucleotide Sequences
The nucleotide sequences of all fragments were identified by sequencing based on Sanger’s dideoxy sequencing method and the absence of mutations was confirmed. DNA sequencing was carried out by using an Applied Biosystems 3730xl sequencer of the National Instrumental Center for Environmental Management (NICEM, Seoul National University, South Korea).
Integration of Foreign DNA
Each integration vector was introduced into IL1403 competent cells by electroporation [1]. Transformants were then incubated at 30°C for 8 h in 50 ml of M17G supplemented with thymidine and d-alanine (M17G.TD). After incubation, cells, harvested from 4 ml of the culture, were spread on M17G.TD plates supplemented with erythromycin (M17G.TD.Em5) and the cells were incubated at 30°C for 48 h. Colonies formed after incubation were assumed to be single crossover integrant candidates. To induce re-crossover at a different flanking region, single crossover cells were continuously sub-cultivated using M17G.TD without erythromycin. After approximately 100 generations, erythromycin-sensitive colonies were isolated and assumed to be WT revertants or double crossover integrant strains.
Verification of Integration
Integration of the vectors into the IL1403 genome was verified by PCR using i-MAX II DNA polymerase (iNtRON Biotechnology, South Korea). Genomic DNA extracted from IL1403-derived integrant strains was used as a template. The primers used and their locations are indicated in Fig. 1a and b and Table 2.
Test of Auxotrophy and Erythromycin-Sensitivity Using Selective Agar Plates
Each test strain was cultivated overnight with M17G.TD broth. Each overnight culture was rubbed on five selective agar plates and the plates were incubated for 36 h. After incubation, the plates were scanned.
Growth Analysis in Growth Media
For growth analysis of the three recombinant strains, M17G, MRS, and SYG [10% (w/v) skim milk, 0.5% (w/v) yeast extract, and 0.5% (w/v) glucose] were used as growth media. An overnight culture of each strain was inoculated into 10 ml of growth media and the cells were incubated at 30°C until specified time points. During incubation, the optical density of the culture at 600 nm (OD600) or the number of colony-forming units per ml (CFU/ml) was measured. In the SYG medium, we could not detect the OD600 value due to the high turbidity of the medium.
Bacterial Cell Harvest for Protein Extraction
Single colony of each strain was inoculated into 10 ml of fresh M17G.TD broth, and the cells were incubated overnight at 30°C without agitation. The overnight culture (500 μl) was subsequently inoculated into 50 ml of fresh M17G.TD broth, and the cells were incubated at 30°C without agitation. The OD600 was measured to draw a growth curve. At specified time points, 5 ml of culture was collected and centrifuged at 5,000g for 10 min at 4°C to harvest cells. The harvested cells were stored at −80°C until protein extraction.
BmpB Detection by SDS-PAGE and Western Blot Assay
To extract proteins from IL1403-derived recombinant cells, the harvested cells were washed twice with phosphate buffered saline (PBS) and resuspended in 200 μl of PBS. Glass beads (212–300 μm, Sigma) were added, and the cells were incubated on ice and shaken intermittently for 1 h by vortex. Cell-debris was removed from the cell extract by centrifugation at 14,000g for 10 min. The protein-containing supernatant was stored at −80°C until analysis. Quantitative analysis of protein was carried out with Protein Assay (Bio-Rad, USA, #500-0006) and bovine serum albumin (BSA) according to manufacturer’s instructions. SDS-PAGE was carried out with a 12% poly-acrylamide gel. Each well was loaded with 40 μg protein. BmpB was detected with polyclonal anti-BmpB mouse serum (1:4,000, Aprogen, South Korea) and anti-mouse IgG HRP-linked antibody (1:20,000, Cell Signaling Technology, USA) [8].
Results
Generation of Three Double Crossover Integrant Strains
We constructed thyA- and alr-targeting integration vectors (pT.Ppfk.MbI2 and pA.Ppfk.MbI2) using the pGEM-T easy plasmid, which does not replicate in LAB. Each vector was introduced into L. lactis IL1403 by electro-transformation and expected to be inserted into the cognate target locus of chromosome via homologous recombination. After transformation erythromycin-resistant colonies were isolated and identified as single crossover integrant strains (ITS and IAS) by PCR (data not shown). These single crossover strains are still having the full ORF sequences of thyA and alr within their genomes. The single crossover thyA-integrant, ITS, did not show thymidine auxotrophy in M17G, but the single crossover alr-integrant, IAS, showed d-alanine auxotrophy (data not shown). During continuous sub-cultivation of these strains using M17G.TD, double crossover strains (ITD and IAD) were isolated. In a similar way, a thymidine/d-alanine double auxotrophic strain (ITDAD) was isolated from sub-cultivation of an ITD-derived alr-integrant (ITDAS).
The genetic structure of the recombinant strains was confirmed by PCR (Fig. 1b, Table 2). In ITD and ITDAD, crossover at thyA F1 and F2 was identified by 2,377-bp and 1,993-bp bands, respectively. In IAD and ITDAD, crossover at alr F1 and F3 was identified by 2,334-bp and 2,022-bp bands, respectively. These four bands were made between IL1403 genomic DNA and an integration vector, which are separate DNA molecules if the crossover did not occur. In ITD and ITDAD, the absence of thyA was identified by the absence of the 685-bp band shown in WT and IAD. In IAD, insertion of bmpB and truncation of alr were identified by the 1,852-bp band, which is longer than the 1,028-bp band shown in WT and ITD. In ITDAD, truncation of alr was identified by the 902-bp band, which is shorter than the 1,028-bp band of WT and ITD. In all the four strains, the absence of Pro.ermAM was identified by the absence of the 1,232-bp band, which is shown in a Pro.ermAM-containing strain (P). All recombinant and wild-type strains could not grow when erythromycin was supplemented, which means that they have no active ermAM gene (Fig. 2). These results suggest that all three recombinant strains are in double crossover at thyA or alr locus and they did not have any heterologous DNA elements except the BmpB expression cassette.
Growth of Recombinant L. lactis Strains in M17G, MRS, and SYG
The growth of thyA-deficient LAB strains is dependent on the presence of thymidine or thymine, because these strains cannot produce dTTP without supplementation [7, 14]. Likewise, the growth of alr-deficient LAB strains is dependent on the presence of d-alanine, which is required to synthesize normal cell walls [2, 13]. To show the thymidine or/and d-alanine dependency of the three recombinant strains (ITD, IAD, and ITDAD), we checked auxotrophy of each recombinant using five different M17G-derived selective agar plates (Fig. 2). ITD or IAD could grow normally only when thymidine or d-alanine was supplemented, respectively. ITDAD could grow normally only when both thymidine and d-alanine were supplemented.
A growth curve of each recombinant strain was compared with that of WT IL1403 using conventional LAB media such as M17G and MRS. In addition, pH changes were also measured because LAB produce lactic acid, thereby lowering the pH of the medium. When M17G was used (Fig. 3a), the growth of the three recombinants was limited as compared with the WT strain. The growth limitation of two alr-integrants (IAD and ITDAD) was affected to a greater extent than that of ITD. The pH value of M17G changed from 6.90 at 0 h to 5.85 (WT), 6.34 (ITD), and 6.85 (IAD and ITDAD) after 24 h of incubation. When MRS was used (Fig. 3b), the growth of only the two alr-integrants was limited, while the ITD strain showed a normal growth pattern identical to that of the WT strain. The pH value of MRS changed from 6.73 at 0 h to 4.58 (WT), 4.60 (ITD), and 5.28 (IAD and ITDAD) after 24 h of incubation. In contrast, when the two media were supplemented with thymidine and d-alanine, all recombinants were able to grow and change the pH of media in a manner identical to that of the WT strain, even though the growth of ITD was slightly retarded in M17G.TD.
Milk has been recognized as a nutrient-rich niches for LAB [10], but it has insufficient pyrimidines to support the growth of some LAB strains. The pyrimidine source present in milk is orotate, an intermediate compound in pyrimidine biosynthesis. However, only some LAB strains can utilize orotate to synthesize pyrimidines and grow in milk [7]. To examine the growth and thymidine or/and d-alanine dependency of the auxotrophic recombinants in milk, a skim milk-based SYG medium was used. When SYG was used (Fig. 3c), all the recombinants showed similar levels of growth limitation and cell death, and only the WT strain could grow normally and lower the pH of SYG thereby inducing curd formation (data not shown). The pH value of SYG changed from 6.54 at 0 h to 4.72 (WT), 6.15 (ITD), and 6.56 (IAD and ITDAD) after 24 h of incubation. On the contrary, when thymidine and d-alanine were added to SYG, normal growth and curd formation were observed in all strains (data not shown).
BmpB Expression in Recombinant L. lactis Strains
A heterologous gene, modified bmpB, was inserted into the genome of each recombinant. When they are expected to be used as LAB-based oral vaccines against the swine dysentery, the BmpB expression is an important issue because expressed BmpB antigen should be delivered to the intestinal immune system. Therefore, in this study, the BmpB expression in the recombinants was examined. All the recombinant strains were able to express BmpB and the expression was growth phase-dependent. The maximal BmpB expression was observed at an early stationary phase regardless of the strain type (Fig. 4a, b).
Discussion
We generated three recombinant L. lactis strains expressing BmpB. They do not have any antibiotic resistance gene and vector-derived DNA fragments, and the only heterologous genetic element inserted into their chromosome is the gene of interest (modified bmpB). The methods we used to engineer L. lactis seem to be safer than conventional methods using plasmid-based expression vectors, because expression vectors generally have selection markers such as antibiotic resistance genes and contain DNA elements to replicate and maintain themselves in host bacteria. These antibiotic resistance genes can be transferred to other bacteria, including pathogens. The transfer is one of the major problems of genetically modified organisms [15].
The growth of auxotrophic strains in M17G, MRS, and SYG was limited. In particular, the two d-alanine auxotrophs (IAD and ITDAD) experienced not only growth limitation but also cell death (Fig. 3c). But the thymidine auxotroph, ITD, showed a normal growth pattern in MRS medium without additional thymidine. MRS is a representative LAB growth medium and widely used in LAB cultivation. Due to both normal growth of ITD in MRS and absence of antibiotic resistance gene, we may misjudge that the recombinant strain is a WT strain. However, the defect of a single auxotrophy of ITD could be overcome by a dual lethal system based on thymidine and d-alanine auxotrophy (ITDAD). Nevertheless, the mortality of ITDAD was not greater than that of the single lethal system based on d-alanine auxotrophy (IAD).
Gene expression in LAB has been reported to be affected by the growth phase [4]. In this study, BmpB expression was also growth phase-dependent. The same BmpB expression pattern was observed when the BmpB expression cassette (Ppfk.bmpB) was used in a plasmid expression vector [8]. Therefore, the growth phase-dependency of BmpB expression is likely to be a result of Ppfk. Maximal BmpB expression was observed in the early stationary phase. This time point is optimal for cell harvesting if the recombinants are to be used as oral vaccines, because both cell yield and BmpB expression are maximal at this time point.
In conclusion, we generated recombinant L. lactis strains using genetic engineering based on gene replacement and auxotrophy. The resulting recombinant strains were able to express a heterologous protein, BmpB. In addition, the recombinants showed growth limitation and cell death in growth media. The approach we took can be used to generate safe GE-LAB. However, further study is needed to examine their in vivo mortality and effects as LAB-based oral vaccines.
References
Alegre MT, Rodríguez MC, Mesas JM (2004) Transformation of Lactobacillus plantarum by electroporation with in vitro modified plasmid DNA. FEMS Microbiol Lett 241:73–77
Bron PA, Benchimol MG, Lambert J et al (2002) Use of the alr gene as a food-grade selection marker in lactic acid bacteria. Appl Environ Microbiol 68:5663–5670
Buist G, Karsens H, Nauta A et al (1997) Autolysis of Lactococcus lactis caused by induced overproduction of its major autolysin, AcmA. Appl Environ Microbiol 63:2722–2728
Cohen DPA, Renes J, Bouwman FG et al (2006) Proteomic analysis of log to stationary growth phase Lactobacillus plantarum cells and a 2-DE database. Proteomics 6:6485–6493
Cummins J, Ho M-W (2005) Genetically modified probiotics should be banned. Microb Ecol Health Dis 17:66–68
Hickey RM, Ross RP, Hill C (2004) Controlled autolysis and enzyme release in a recombinant lactococcal strain expressing the metalloendopeptidase enterolysin A. Appl Environ Microbiol 70:1744–1748
Kilstrup M, Hammer K, Jensen PR et al (2005) Nucleotide metabolism and its control in lactic acid bacteria. FEMS Microbiol Rev 29:555–590
Kim EB, Piao DC, Son JS et al (2009) Cloning and characterization of a novel tuf promoter from Lactococcus lactis subsp. lactis IL1403. Curr Microbiol 59:425–431
Lee BJ, La T, Mikosza ASJ et al (2000) Identification of the gene encoding BmpB, a 30 kDa outer envelope lipoprotein of Brachyspira (Serpulina) hyodysenteriae, and immunogenicity of recombinant BmpB in mice and pigs. Vet Microbiol 76:245–257
Pfeiler EA, Klaenhammer TR (2007) The genomics of lactic acid bacteria. Trends Microbiol 15:546–553
Ronchel MC, Ramos JL (2001) Dual system to reinforce biological containment of recombinant bacteria designed for rhizoremediation. Appl Environ Microbiol 67:2649–2656
Simon D, Chopin A (1988) Construction of a vector plasmid family and its use for molecular cloning in Streptococcus lactis. Biochimie 70:559–566
Steen A, Palumbo E, Deghorain M et al (2005) Autolysis of Lactococcus lactis is increased upon d-alanine depletion of peptidoglycan and lipoteichoic acids. J Bacteriol 187:114–124
Steidler L, Neirynck S, Huyghebaert N et al (2003) Biological containment of genetically modified Lactococcus lactis for intestinal delivery of human interleukin 10. Nat Biotechnol 21:785–789
Sybesma W, Hugenholtz J, Vos WMd et al (2006) Safe use of genetically modified lactic acid bacteria in food. Bridging the gap between consumers, green groups, and industry. Electron J Biotechnol 9:424–448
Teuber M, Geis A (2006) The genus lactococcus. In: The prokaryotes. Springer, New York, pp 205–228
Torres B, Jaenecke S, Timmis KN et al (2003) A dual lethal system to enhance containment of recombinant micro-organisms. Microbiology 149:3595–3601
Wells JM, Mercenier A (2008) Mucosal delivery of therapeutic and prophylactic molecules using lactic acid bacteria. Nat Rev Microbiol 6:349–362
Acknowledgments
This study was supported by a graduate fellowship from the BK 21 project and the Research Institute for Agriculture and Life Sciences, Seoul National University.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kim, E.B., Son, J.S., Zhang, Q.K. et al. Generation and Characterization of Thymidine/d-Alanine Auxotrophic Recombinant Lactococcus lactis subsp. lactis IL1403 Expressing BmpB. Curr Microbiol 61, 29–36 (2010). https://doi.org/10.1007/s00284-009-9572-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-009-9572-y