Abstract
A novel Acinetobacter strain, Ud-4, possessing a strong capacity to degrade edible, lubricating, and heavy oil was isolated from seawater in a fishing port located in Toyama, Japan. It was identified by morphological and physiological analyses and 16S rDNA sequencing. This strain could utilize five types of edible oils (canola oil, olive oil, sesame oil, soybean oil, and lard), lubricating oil, and C-heavy oil as the sole carbon source for growth in M9 medium. The strain grew well and heavily degraded edible oils in Luria–Bertani medium during a 7-day culture at 25°C; it also degraded all kinds of oils in artificial seawater medium for marine bacteria. Furthermore, this strain was capable of degrading almost all C10–C25 n-alkanes in C-heavy oil during a 4-week culture. Oligonucleotide primers specific to two catabolic genes involved in the degradation of n-alkanes (Acinetobacter sp. alkM) and triglyceride (Acinetobacter sp. lipA) allowed amplification of these genes in strain Ud-4. To our knowledge, this is the first report on the isolation of a bacterium that can efficiently degrade both edible and mineral oils.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Various kinds of oils—edible oil, lubricating oil, and heavy oil—cause environmental pollution. Lipids are major components of domestic and industrial wastewater, including edible vegetable oils and animal fats, especially from restaurants and food processing industries [27, 37]. The amount of lipid-rich wastewater increases every year due to urbanization. Such wastewater can form large aggregates that obstruct drainage lines. Lubricating oil contaminates the soil and groundwater during daily use and via leaks and spills from industrial machines [36, 41]. Petroleum oil spills are one of the most serious problems that can occur in a marine environment [10, 11], endangering birds, fish, shells, and algae. Oil slicks in coastal regions, apart from destroying scenery, cause considerable economic damage to the fishery industry. In Japan, a heavy oil spill from the Russian tanker Nakhodka in 1997 received much attention [3, 23, 24, 34]. The main fraction of lubricating oil and heavy oil is a complex mixture of hydrocarbons, linear and branched paraffins, cyclic alkanes, and aromatic hydrocarbons [7, 38].
Physical and chemical technologies have been developed for the remediation of oil-contaminated environments. However, they are expensive for large-scale application, and are rarely completely successful. Recently, bioremediation techniques using microorganisms have attracted great interest [2, 35]. For bioremediation (bioaugmentation) to succeed, it is necessary to select microorganisms that can degrade contaminated materials under various temperatures, pH, salinities, and nutrient concentrations. Many kinds of microorganisms have been isolated for improved bioremediation of contaminated environments with lipids or petroleum hydrocarbons [3, 9, 15, 21, 31, 41]. However, most studies have focused on biodegradation of only one type of oil by each microorganism.
In the present study, we isolated a novel bacterium that degrades various kinds of oils. The bacterium showed the ability to efficiently degrade edible oil, lubricating oil, and heavy oil. Furthermore, the catabolic genes of the isolated strain were investigated.
Materials and Methods
Isolation, Media, and Culture Conditions
Seawater samples containing oil film were collected from the Uozu fishing port on the coast of Toyama Bay, Toyama, Japan. The culture media used in this study were Luria–Bertani (LB) medium (1% NaCl, 0.5% yeast extract, and 1% tryptone), M9 medium (0.68% Na2HPO4, 0.3% KH2PO4, 0.05% NaCl, 0.1% NH4Cl, 2 ml of 24.6% MgSO4·7H2O, and 0.1 ml of 11.1% CaCl2 per liter), and artificial seawater (ASW) medium (artificial seawater 800 ml, NH4NO3 1 g, K2HPO4 0.02 g, yeast extract 0.5 g, and deionized water 200 ml, pH 7.8). To isolate the oil degrader, 1 ml of a sample was transferred to a glass tube containing 10 ml of ASW medium with 0.5% (w/v) C-heavy oil (Cosmo Oil Co., Tokyo, Japan), and incubated at 25°C in a rotary shaker at 120 rpm until emulsification was observed. Part of the culture solution containing the proliferated bacteria was then spread on ASW agar plates (1.5% agar) containing 0.5% (w/v) heavy oil. The colonies that formed transparent and irregular zones were isolated and transferred into the ASW liquid medium containing 0.5% (w/v) heavy oil and were shaken for 5–10 days. The bacterial clones that emulsified the oil were selected as candidate heavy-oil-degrading strains. Then, to find bacteria that could degrade other oils, the heavy-oil-degrading bacteria were examined by halo assay using agar plates with 1% (w/v) edible oil or lubricating oil. One strain, designated as Ud-4, was confirmed to degrade not only heavy oil, but also edible oils and lubricating oil. This strain was selected for further study.
Morphological and Physiological Analysis of the Microorganism
The morphology and motility of the bacteria were first observed by a phase microscope (Olympus BH-2, Tokyo, Japan). Electron microscopy samples were negatively stained with 1% phosphotungstic acid (pH 3–9), and then observed with a JEM-1200EXII electron microscope (JEOL, Tokyo, Japan). Micrographs were taken at an accelerating voltage of 80 kV. Physiological characterization of the microorganism was performed by an API 20NE system (BioMerieux, Marcy, l’Etoile, France). In addition, gram staining (FAVOR-G SET-S, Nissui Pharmaceutical, Tokyo, Japan), catalase, and oxidase tests were conducted.
16S rDNA Sequencing
Almost the full length of the bacterial 16S rDNA was amplified by PCR using the primers 27f (5′-AGAGTTTGATCCTGGCTCAG-3′) and 1525r (5′-AAAGGAGGTGATCCAGCC-3′). The amplified products were purified with a QIAquick PCR purification kit (Qiagen, Tokyo, Japan), and then sequenced directly using a BigDye Terminator v3.1 cycle sequencing kit (Applied Biosystems, Foster City, CA, USA), an ABI Prism 3130xl genetic analyzer (Applied Biosystems), and the following sequencing primers: r1L (5′-GTATTACCGCGGCTGCTGG-3′), r2L (5′-CATCGTTTACGGCGTGGAC-3′), r3L (5′-TTGCGCTCGTTGCGGGACT-3′), r4L (5′-ACGGGCGGTGTGTACAAG-3′), f1L (5′-GAGTTTGATCCTGGCTCAG-3′), f3L (5′-GTCCCGCAACGAGCGCAAC-3′), and 926f (5′-AAACTCAAAGGAATTGACGG-3′). The DNA sequence was searched by the BLAST program on the NCBI website (http://www.ncbi.nlm.nih.gov). The 16S rDNA sequence of strain Ud-4 was deposited in GenBank/EMBL/DDBJ under accession number AB488778.
Degradation of Edible Oil, Lubricating Oil, or Heavy Oil by Strain Ud-4
The oil degradation rate was examined according to previously described methods [7, 15, 24]. Bacterial cells precultured overnight at 25°C in LB medium were transferred to silicone-capped test tubes containing 6 ml of each medium with one of the edible oils (e.g., canola oil, olive oil, sesame oil, soybean oil, and lard) or lubricating oil (Noritake Cut; Noritake Co., Limited, Aichi, Japan) at a concentration of 1% (w/v), i.e. 60 mg. The cells were inoculated at a cell density of approximately 107 cells per ml and grown at 25°C by shaking at 120 rpm, and the remaining oil in each medium was extracted with n-hexane after 7 days. The extract was dehydrated with Na2SO4, and then dried at 30°C. The dry weight of extracts was measured using an electronic microbalance accurate to 0.1 mg in three sets of samples, with and without bacterial inoculation (negative control). The degradation rate of edible oils and lubricating oil was assessed from the average dry weight difference. The oil degradation rate was calculated as follows: percentage oil degradation (% per 7 days) = [(oil weight in negative control) − (oil weight in sample)/(oil weight in negative control)] × 100. For the degradation test of heavy oil, culture media with 0.5% (w/v) type C heavy oil were used. The remaining heavy oil in each medium was extracted with chloroform, and the degradation rate of heavy oil was calculated using a method similar to that used for edible oils and lubricating oil. The extraction efficiency for control was 85–100% in edible oils, 40–45% in lubricating oil, and 85–90% in heavy oil.
Gas chromatography (GC) was performed with a gas chromatograph (Shimadzu GC-14A; Tokyo, Japan) and a capillary column (OV-1, 50 m × 0.25 mm i.d.; film thickness, 1.5 μm). Helium was the carrier gas. The injector and detector temperatures were set at 280°C. The oven temperature program was 70–160°C at 10°C/min, 160–280°C at 2°C/min, and held at 280°C for 30 min.
Detection of Catabolic Genes by PCR
PCR primers were used to detect catabolic genes as follows: Pseudomonas putida GPo1 alkB, Acinetobacter sp. ADP-1 alkM and Rhodococcus sp. Q15 alkB1 and alkB2 for alkane monooxygenase and alkane degradation [40]; P. putida ATCC 33015 xylE for catechol-2,3-dioxygenase, xylene, and toluene degradation; P. putida ATCC 17484 ndoB for naphthalene dioxygenase and naphthalene degradation [39]; and Mycobacterium sp. PYR-1 nidA for pyrene dioxygenase and PAH (pyrene) degradation [20]. For Acinetobacter sp. lipA, which encodes a lipase that catalyzes the hydrolysis of triglycerides, primers Ac lipA-F (5′-TGCHMARACMAARTATCC-3′) and Ac lipA-R (5′-AYYTGATTSACTTCATC-3′) were derived from regions of high DNA sequence identity from four lipase structural gene sequences from A. calcoaceticus RAG-1 [32] (GenBank accession number AF047691), Acinetobacter sp. SY-01 [8] (AF518410), A. baumannii BD5 [26] (EU078176), and A. calcoaceticus BD413 [18] (X80800). The degenerate bases used were H = A/C/T, M = A/C, R = A/G, Y = C/T, and S = G/C. In all cases, the primers were designed toward a sequence located inside the open reading frame. The expected size of PCR products was 870 bp for alkB, 496 bp for alkM, 629 bp for alkB1, 552 bp for alkB2, 834 bp for xylE, 642 bp for ndoB, 323 bp for nidA, and approximately 770 bp for lipA. All PCR amplifications were carried out as previously described [39] for 30 cycles at 94°C for 1 min, 60°C for 1 min, and 72°C for 1 min, with final extension at 72°C for 3 min. The PCR products were analyzed by electrophoresis on 2% (w/v) agarose gel and visualized by ethidium bromide staining. Amplified PCR fragments were sequenced as described at the beginning of this section for 16S rDNA sequencing.
Results
Identification of Strain Ud-4
Morphological and physiological properties presented by strain Ud-4 were consistent with its assignment to the genus Acinetobacter. The bacterium was gram-negative, non-spore forming, and non-motile. Phase microscopy demonstrated that the cells predominantly grew in pairs at logarithmic growth phase. Observation by electron microscope revealed that the paired cells were short rods in shape, 1.0–2.0 μm long, and 0.8–1.0 μm wide; peritrichate fibrils [12] could be observed on the cells (Fig. 1). Colonies were formed as circular and white on LB agar 2–3 days after inoculation. They grew well in media supplemented with 0–3% NaCl, but not in media with salinity exceeding 5%. Growth occurred at temperatures ranging from 5 to 35°C, with an optimum at 25°C. Catalase and oxidase reactions were positive and negative, respectively. In addition to these properties, the physiological characteristics (using the API 20NE system) are summarized in Table 1. Analysis with the api computer program indicated that it belongs to the genus Acinetobacter.
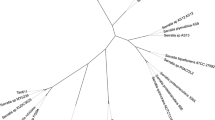
The 16S rDNA sequence of strain Ud-4, 1493 bp in length, was determined for phylogenetic analysis. It showed high sequence similarity to the Acinetobacter species, with the highest degree of similarity attained at 99% with Acinetobacter psychrotolerans (GenBank accession number AB207814), described as the Acinetobacter sp. strain Ths [43]. A phylogenic tree constructed by the neighbor joining method demonstrated that strain Ud-4 closely clusters with genus Acinetobacter (data not shown).
Degradation of Edible Oil, Lubricating Oil, and Heavy Oil by Strain Ud-4
To study the oil degradation potential in various environments, three different liquid media were used. The degradation of five different kinds of edible oils and mineral oils, such as lubricating oil and C-heavy oil, by strain Ud-4 was examined (Table 2). The results showed that strain Ud-4 could degrade all kinds of edible oils (canola oil, olive oil, sesame oil, soybean oil, and lard) in each medium after cultivation at 25°C for 7 days; however, the level of edible oil degradation varied with different oils or media. The degradation of edible oils in LB medium occurred more efficiently than those in M9 medium or ASW medium. In particular, the degradation of canola oil in LB medium reached up to 96.3%. This strain degraded lubricating oil in LB, M9, and ASW mediums up to 67.6% (LB), 34.7% (M9), and 31.7% (ASW). It also degraded C-heavy oil in each medium up to 6.5% (LB), 9.6% (M9), and 6.9% (ASW).
The GC profile of the remnant n-alkanes of C-heavy oil was compared with those of the control (Fig. 2). At the end of the incubation period (4 weeks), the n-alkanes from C10 to C25 were almost completely degraded by strain Ud-4. This finding is consistent with the presence of alkane monooxygenase gene (alk M) in this strain as described below. However, one compound was not appreciably degraded. Since the peak is found near C17 n-alkane, it is possible that the compound is pristane, which is a highly branched alkane and is biodegraded slowly [1, 22].
Detection of Catabolic Genes
A novel bacterial strain, Ud-4, possessing high capacity to degrade edible oil, lubricating oil, and heavy oil was examined for the presence of known catabolic genes (alkB, alkM, alkB1, alkB2, xylE, ndoB, nidA, and lipA) by PCR using oligonucleotide primers derived from these genes (Fig. 3). This strain showed PCR-positive for alkM (496 bp) and lipA (approximately 770 bp), involved in the degradation of n-alkanes and triglyceride, and showed PCR-negative for alkB, alkB1, alkB2, xylE, ndoB, and nidA. Sequence analysis showed that the obtained fragment of alkM gene had 85% similarity with that from Acinetobacter sp. M-1 (GenBank accession number AB049410). Meanwhile, a fragment of lipA had 77% homology with that from Acinetobacter sp. SY-01 (AF518410).
Discussion
Strain Ud-4 was identified as a new strain of the genus Acinetobacter. The 16S rDNA sequence of strain Ud-4 was 99% identical to that of Acinetobacter psychrotolerans (strain Ths), which has not yet been validated as an Acinetobacter species in the list of prokaryotic names with standing in nomenclature [6]. The results of physiological characterization of Acinetobacter sp. strain Ths [43] show that strain Ud-4 is slightly different in the utilization of citrate (Table 1) and in the hydrolysis of lipids. Therefore, strain Ud-4 may be a member of Acinetobacter psychrotolerans. Acinetobacter species are ubiquitous in geographical distribution, and are known to be involved in the bioremediation of alkanes and aromatic hydrocarbons as well as in the production of high-molecular-weight heteropolysaccharides that act as powerful emulsifiers, with many having high commercial potential [16, 25, 29]. Moreover, several Acinetobacter strains are known to produce extracellular lipase [17, 30, 33], which can degrade lipids. In this study, we observed that strain Ud-4 emulsified C-heavy oil to fine droplets during the initial 1-week period, and hence, probably produced a bioemulsifier. In preliminary experiments, we also observed that strain Ud-4 showed lipase activity in the culture supernatant.
In general, lard is persistent to biodegradation because of its high melting temperature (28–48°C) and fatty acid composition [5, 31]. Lubricating oils are also assumed to be relatively recalcitrant to biodegradation due to their high content of recalcitrant cycloalkanes and isoalkanes [13, 41]. Therefore, it is interesting that strain Ud-4 showed a strong ability to degrade lard and lubricating oil, as shown in Table 2. Interestingly, all kinds of edible oils were more efficiently degraded in LB medium than in ASW or M9 medium. LB medium contains a high nutrient content and enhances cell growth. From the results of the culture using M9 medium, we found that this strain could use edible oil, lubricating oil, or heavy oil as the sole carbon source. The culture using ASW medium showed that strain Ud-4 was able to degrade various kinds of oils in a medium containing high salinity as well as seawater. Thus, we expect that this strain certainly has the ability to cleanup oil contamination in the sea, wastewater with high salinity, and oligotrophic environments. Then, strain Ud-4 was capable of degrading almost all components of C10–C25 n-alkanes in C-heavy oil during 4-week culture. Since lubricating oil also contains n-alkanes [41], it is supposed that these components were degraded by strain Ud-4.
The use of specific primers to amplify catabolic genes is a very useful tool for characterizing microorganisms possessing a variety of biodegradative capabilities and for examining their catabolic pathways [39]. Therefore, we carried out PCR detection of the genes involved in n-alkane, aromatic, and triglyceride degradation pathways. This strain possessed Acinetobacter sp. alkM, involved in the initial oxidation of the alkane substrate to 1-alkanol [42] and Acinetobacter sp. lipA, involving the hydrolysis of triglyceride to glycerol and fatty acids [14], respectively. Thus, initial reactions involved in the degradation of n-alkane and triglyceride are different, but both metabolites are finally oxidized via beta-oxidation of fatty acids [4, 19, 28]. In addition to oil-degradation experiments, molecular detection of catabolic genes by PCR could provide additional information rapidly and accurately.
To our knowledge, this is the first report on the isolation of a bacterium that can efficiently degrade both edible and mineral oils. Therefore, this strain could be useful for application to bioremediation technology. However, further investigation is still needed to evaluate the performance in situ, because a variety of factors can affect the ability of microorganisms to degrade contaminated oils in natural environments.
References
Atlas RM (1981) Microbial degradation of petroleum hydrocarbons: an environmental perspective. Microbiol Rev 45:180–209
Brooksbank AM, Latchford JW, Mudge SM (2007) Degradation and modification of fats, oils and grease by commercial microbial supplements. World J Microbiol Biotechnol 23:977–985
Chaerun SK, Tazaki K, Asada R, Kogure K (2004) Bioremediation of coastal areas 5 years after the Nakhodka oil spill in the Sea of Japan: isolation and characterization of hydrocarbon-degrading bacteria. Environ Int 30:911–922
Chipasa KB, Medrzycka K (2008) Characterization of the fate of lipids in activated sludge. J Environ Sci 20:536–542
de Renobales M, Agud I, Lascaray JM, Múgica JC, Landeta LC, Solozábal R (1992) Hydrolysis of animal fats by lipase at temperatures below their melting points. Biotechnol Lett 14:683–688
Euzéby JP (2009) List of prokaryotic names with standing in nomenclature-genus Acinetobacter. Available at http://www.bacterio.cict.fr/. Accessed 7 Aug 2009
Goto M, Kato M, Asaumi M, Shirai K, Venkateswaren K (1994) TLC/FID method for evaluation of the crude-oil-degrading capability of marine microorganisms. J Mar Biotechnol 2:45–50
Han SJ, Back JH, Yoon MY, Shin PK, Cheong CS, Sung MH, Hong SP, Chung IY, Han YS (2003) Expression and characterization of a novel enantioselective lipase from Acinetobacter species SY-01. Biochimie 85:501–510
Hasanuzzaman M, Umadhay-Briones KM, Zsiros SM, Morita N, Nodasaka Y, Yumoto I, Okuyama H (2004) Isolation, identification, and characterization of a novel, oil-degrading bacterium, Pseudomonas aeruginosa T1. Curr Microbiol 49:108–114
Hester MW, Mendelssohn IA (2000) Long-term recovery of a Louisiana brackish marsh plant community from oil-spill impact: vegetation response and mitigating effects of marsh surface elevation. Mar Environ Res 49:233–254
Hose JE, Brown ED (1998) Field applications of the piscine anaphase aberration test: lessons from the Exxon Valdez oil spill. Mutat Res 399:167–178
Ishii S, Koki J, Unno H, Hori K (2004) Two morphological types of cell appendages on a strongly adhesive bacterium, Acinetobacter sp. strain Tol 5. Appl Environ Microbiol 70:5026–5029
Ito H, Hosokawa R, Morikawa M, Okuyama H (2008) A turbine oil-degrading bacterial consortium from soils of oil fields and its characteristics. Int Biodeterior Biodegrad 61:223–232
Jaeger KE, Ransac S, Dijkstra BW, Colson C, van Heuvel M, Misset O (1994) Bacterial lipases. FEMS Microbiol Rev 15:29–63
Jirasripongpun K (2002) The characterization of oil-degrading microorganisms from lubricating oil contaminated (scale) soil. Lett Appl Microbiol 35:296–300
Kaplan N, Rosenberg E (1982) Exopolysaccharide distribution of and bioemulsifier production by Acinetobacter calcoaceticus BD4 and BD413. Appl Environ Microbiol 44:1335–1341
Kasana RC, Kaur B, Yadav SK (2008) Isolation and identification of a psychrotrophic Acinetobacter sp. CR9 and characterization of its alkaline lipase. J Basic Microbiol 48:207–212
Kok RG, Thor JJ, Nugteren-Roodzant IM, Brouwer MBW, Egmond MR, Nudel CB, Vosman B, Hellingwerf KJ (1995) Characterization of the extracellular lipase, LipA, of Acinetobacter calcoaceticus BD413 and sequence analysis of the cloned structural gene. Mol Microbiol 15:803–818
Kunau WH, Dommes V, Schulz H (1995) β-oxidation of fatty acids in mitochondria, peroxisomes, and bacteria: a century of continued progress. Prog Lipid Res 34:267–342
Margesin R, Labbé D, Schinner F, Greer CW, Whyte LG (2003) Characterization of hydrocarbon-degrading microbial populations in contaminated and pristine Alpine soils. Appl Environ Microbiol 69:3085–3092
Matsumiya Y, Wakita D, Kimura A, Sanpa S, Kubo M (2007) Isolation and characterization of a lipid-degrading bacterium and its application to lipid-containing wastewater treatment. J Biosci Bioeng 103:325–330
Mohantya G, Mukherji S (2008) Biodegradation rate of diesel range n-alkanes by bacterial cultures Exiguobacterium aurantiacum and Burkholderia cepacia. Int Biodeterior Biodegrad 61:240–250
Morita A, Kusaka Y, Deguchi Y, Moriuchi A, Nakanaga Y, Iki M, Miyazaki S, Kawahara K (1999) Acute health problems among the people engaged in the cleanup of the Nakhodka oil spill. Environ Res 81:185–194
Nakamura S, Sakamoto Y, Ishiyama M, Tanaka D, Kunii K, Kubo K, Sato C (2007) Characterization of two oil-degrading bacterial groups in the Nakhodka oil spill. Int Biodeterior Biodegrad 60:202–207
Navon-Venezia S, Zosim Z, Gottlieb A, Legmann R, Carmeli S, Ron EZ, Rosenberg E (1995) Alasan, a new bioemulsifier from Acinetobacter radioresistens. Appl Environ Microbiol 61:3240–3244
Park IH, Kim SH, Lee YS, Lee SC, Zhou Y, Kim CM, Ahn SC, Choi YL (2009) Gene cloning, purification, and characterization of a cold-adapted lipase produced by Acinetobacter baumannii BD5. J Microbiol Biotechnol 19:128–135
Quéméneur M, Marty Y (1994) Fatty acids and sterols in domestic wastewaters. Water Res 28:1217–1226
Rehm HJ, Reiff I (1981) Mechanisms and occurrence of microbial oxidation of long-chain alkanes. Adv Biochem Eng 19:175–215
Reisfeld A, Rosenberg E, Gutnick D (1972) Microbial degradation of crude oil: factors affecting the dispersion in sea water by mixed and pure cultures. Appl Microbiol 24:363–368
Snellman EA, Sullivan ER, Colwell RR (2002) Purification and properties of the extracellular lipase, LipA, of Acinetobacter sp. RAG-1. Eur J Biochem 269:5771–5779
Sugimori D, Nakamura M, Mihara Y (2002) Microbial degradation of lipid by Acinetobacter sp. strain SOD-1. Biosci Biotechnol Biochem 66:1579–1582
Sullivan ER, Leahy JG, Colwell RR (1999) Cloning and sequence analysis of the lipase and lipase chaperone-encoding genes from Acinetobacter calcoaceticus RAG-1, and redefinition of a proteobacterial lipase family and an analogous lipase chaperone family. Gene 230:277–285
Suzuki T, Nakayama T, Kurihara T, Nishino T, Esaki N (2001) Cold-active lipolytic activity of psychrotrophic Acinetobacter sp. strain no. 6. J Biosci Bioeng 92:144–148
Tazaki K (ed) (2003) Heavy oil spilled from Russian tanker “Nakhodka” in 1997: towards eco-responsibility, earth sense. 21st Century COE Kanazawa University. Kanazawa University Press, Kanazawa, Japan
Timmis KN, Pieper DH (1999) Bacteria designed for bioremediation. Trends Biotechnol 17:201–204
Vwioko DE, Fashemi DS (2005) Growth response of Ricinus communis L. (Castor Oil) in spent lubricating oil polluted soil. J Appl Sci Environ Manage 10:127–134
Wakelin NG, Forster CF (1997) An investigation into microbial removal of fats, oils and greases. Bioresour Technol 59:37–43
Westlake DWS, Jobson A, Phillipe R, Cook FD (1974) Biodegradability and crude oil composition. Can J Microbiol 20:915–928
Whyte LG, Greer CW, Inniss WE (1996) Assessment of the biodegradation potential of psychrotrophic microorganisms. Can J Microbiol 42:99–106
Whyte LG, Schultz A, van Beilen JB, Luz AP, Pellizari V, Labbé D, Greer CW (2002) Prevalence of alkane monooxygenase genes in Arctic and Antarctic hydrocarbon-contaminated and pristine soils. FEMS Microbiol Ecol 41:141–150
Wongsa P, Tanaka M, Ueno A, Hasanuzzaman M, Yumoto I, Okuyama H (2004) Isolation and characterization of novel strains of Pseudomonas aeruginosa and Serratia marcescens possessing high efficiency to degrade gasoline, kerosene, diesel oil, and lubricating oil. Curr Microbiol 49:415–422
Wyatt TM (1984) The microbial degradation of hydrocarbons. Trends Biochem Sci 9:20–23
Yamahira K, Hirota K, Nakajima K, Morita N, Nodasaka Y, Yumoto I (2008) Acinetobacter sp. strain Ths, a novel psychrotolerant and alkalitolerant bacterium that utilizes hydrocarbon. Extremophiles 12:729–734
Acknowledgments
This work was supported in part by a Grant-in-Aid from the Ministry of Education, Culture, Sports, Science and Technology of Japan (No. 19510082).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tanaka, D., Takashima, M., Mizuta, A. et al. Acinetobacter sp. Ud-4 Efficiently Degrades Both Edible and Mineral Oils: Isolation and Characterization. Curr Microbiol 60, 203–209 (2010). https://doi.org/10.1007/s00284-009-9525-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-009-9525-5