Abstract
The effects of berberine on growth, arylamine N-acetyltransferase (NAT) activity, and gene expression in Salmonella Typhi (Typhi) were described. The growth inhibition of Typhi was determined by measuring absorbance by optical density (OD at 650 nm). The NAT activity was determined by measuring the levels of 2-aminofluorene (AF) and N-acetyl-2-aminofluorene (AAF) by high-performance liquid chromatography. The results demonstrated that 24-h berberine treatment decreased bacteria growth and amounts of AAF in Typhi. Western blotting and flow cytometry were used for examining the levels of NAT after bacteria were cotreated with or without various concentrations of berberine, and results indicated that berberine decreased the levels of NAT in Typhi. Polymerase chain reaction was used for examining the gene expression of NAT (mRNA NAT), and results indicated that berberine affects mRNA NAT1 expression in Typhi.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Arylamine N-acetyltransferases (NATs) are widely distributed in many species including eukaryotes and prokaryotes, with more than 20 NATs identified [11, 15, 18]. Many arylamine chemicals require N-acetylation to form metabolites before leading to carcinogenesis. N-Acetylation is the first step for arylamine metabolites, and it is catalyzed by cytosolic NAT using acetyl coenzyme A as a cofactor [26]. Individuals can be classified into slow and rapid acetylator phenotypes based on liver cytosolic NAT activity [9]. After individuals were exposed to these arylamines, the slow or rapid acetylator phenotype can influence both therapeutic response and toxicity [26]. NAT polymorphism has pharmacogenetic implications, leading to different rates of inactivation of drugs, including the anti-tubercular agent isoniazid and the anti-hypertensive drug hydralazine [22, 23].
Berberine, an isoquinoline alkaloid, has a wide range of pharmacological effects, including anti-inflammation [17], with a long history of being used as a tonic remedy for liver dysfunction in some populations [25]. Many studies have demonstrated that berberine, a yellow benzylisoquinoline alkaloid, is conventionally used as an antidiarrhetic, bitter stomachic, and antimalaria drug in many countries [27]. They often show strong cytotoxicity to prokaryotic and eukaryotic cells; for example, vincristine inhibits microtubule formation, and berberine inhibits DNA and protein synthesis [10]. Because of these activities, alkaloids are presumed to play an important role as a biological barrier to protect the plant tissue from pathogens. Indeed, berberine shows strong antimicrobial activity to both Gram-positive and Gram-negative bacteria as well as other microorganisms [15]. NAT activities had been demonstrated to exist in some human gastrointestinal flora [18]. There is no available information that addresses the effects of berberine on NAT activity (N-acetylation of 2-AF) and gene expression (mRNA NAT) in Typhi. We report herein whether berberine could affect growth in Typhi and in vitro evaluation of the activity as inhibitor of bacterial NAT.
Materials and Methods
The Bio-Rad protein assay kit was obtained from Bio-Rad Laboratories (Hercules, CA), and all other chemicals and reagents were purchased from Sigma Aldrich (St. Louis, MO) and Merck Co. (Darmstadt, Germany).
Berberine affects the growth of Salmonella Typhi
Typhi was collected in the Department of Clinical Laboratory, China Medical University Hospital, in mid-Taiwan. The isolate was identified on the basis of routine microbiologic methods and was confirmed using the VITEK system (BioMerieux Vitek Inc., Hazelwood, MO). About 1 × 108 bacteria were placed in individual tubes containing 1 mL tryptic soy broth with or without different concentrations of berberine (0.5, 1, 5, 25, and 50 μM). The control reactions had 10 μL DMSO in place of berberine. The culture tubes were incubated at 35°C with 5% CO2 and checked for growth after 24 h. The determination of the effects of berberine on the Typhi was described previously. Growth inhibition (%) was determined using the equation described previously [19].
Preparation of Salmonella Typhi cell lysates
An amount of 1 × 1010 colony-forming units (cfu) of Typhi was washed twice in cold phosphate buffered saline (PBS) and was then immediately placed in 1 mL of lysis buffer [20 mM Tris-HCl, pH 7.5 (at 4°C), 1 mM DTT, 1 mM EDTA, 50 μM PMSF, and 10 μM leupeptin]. The cell suspensions were disrupted in a sonicator (Hert Systems, Inc., Farmingdale, NY) and centrifuged for 30 min at 10,000g; the supernatant was kept on ice until assayed for NAT activity (n-acetylation of AF) and protein determination [19].
NAT activity determination
The determination of total amounts of acetylated and nonacetylated AF were determined by high-pressure liquid chromatography (HPLC) as described by Chung et al. in 1998 [5].
Protein determination
Protein concentrations of the bacterial cell lysates were determined by the method of Bradford with bovine serum albumin as the standard [2]. All of the samples were assayed in triplicate.
Berberine affects NAT activity in Salmonella Typhi cell lysates
Berberine was dissolved in DMSO at various concentrations (0.5, 1, 5, 25, and 50 μM). The reaction mixtures consisted of 50 μL cell lysates (Typhi), 20 μL of recycling mixture containing 22.5 μM AF, and 10 μL of berberine. The reactions were started by the addition of Ac-CoA. The control reactions had 20 μL of distilled water in place of Ac-CoA. After these steps, the NAT activity was determined as described above [19].
Berberine affects NAT activity in intact bacterial cells
For the intact cell studies, 1 × 109 cfu of bacteria in 1 mL trypticase soy broth were incubated with 22.5 μM AF with or without various concentrations (0.5, 1, 5, 25, and 50 μM) of berberine for 24 h. After incubation, the media suspensions were removed and centrifuged. The supernatant was immediately extracted with ethylacetate/methanol (95:5), the solvent evaporated under speed vacuum, and the residue dissolved in methanol and assayed by HPLC [19]. All samples were run in triplicate.
Berberine affects kinetic constants of NAT from Salmonella Typhi cell lysates
Cell lysates of Typhi cotreated with or without 25 μM berberine and selected concentrations (5.625, 11.25, 22.5, 45, and 90 μM) of AF were performed for NAT activity as described above [19].
Berberine effect on NAT protein levels and was examined by SDS-PAGE and Western blot
About 1 × 109 cfu of bacteria were cotreated with or without various concentrations of berberine for 24 h. Typhi were then harvested and centrifuged to remove and discard the medium, while the cells were washed with PBS. The cells were lysed in 100 μL of triple detergent buffer (50 mM Tris-Cl [pH 8.0], 150 mM NaCl, 0.02% NaN3, 0.1% SDS, 1% NP-40, 0.5% sodium deoxycholate, 100 μg/mL phenylmethylsulfonyl fluoride, 1 μg/mL aprotinin, 1 μg/mL leupeptin, and 1 μg/mL pepstatin A). The samples were then sonicated, incubated on ice for 20 min, and centrifuged at 12,000g for 10 min at 4°C. The supernatants were collected and the Bradford assay was performed to determine the protein concentration. Proteins (50 μg/lane) were separated on 12% SDS–polyacrylamide gels and blotted onto polyvinyliene difluoride membranes. The membrane was incubated with 5% bovine serum albumin and primary antibody (anti-NAT) at 4°C overnight. The blots were washed three times in PBS with 0.04% Tween-20 (PBST) for 5 min, then incubated with horseradish peroxidase (HRP)-conjugated secondary antibody at 1:1000 dilution in PBST containing 5% milk for 2 h at room temperature. The membranes were washed with PBST and visualized using an ECL detection system (Amersham, Piscataway, NJ) and quantitated by densitometry using ImageQuant image analysis.
Berberine affects NAT gene expression and was examined by PCR
The total RNA was extracted from Typhi by using Qiagen RNeasy Mini Kit at 24 h afterwards with or without co-treatment of 25 μM berberine. Total RNA (1.5 μg), 0.5 μg of oligo-dT primer, and DEPC (diethyl pyrocarbonate)-treated water were combined into a microcentrifuge tube to a final volume of 12.5 μL. The entire mixture was heated at 70°C for 10 min and chilled on ice for at least 1 min. The subsequent procedures for conducting reverse transcription were exactly the same as those in the instruction manual (First-strand cDNA synthesis kit, Novagen) [14].
The reverse transcription products from total RNA served as a template for PCR. When amplifying target cDNA, components in 50 μL of solution were as follows: 1.5 mM MgCl2, 0.2 mM dNTP mix, 20 pmol of each primer (B-MDIEA-NAT1 & VPKHGD-X-NAT1 for NAT1, FP1-NAT2 & RP1-NAT2 for NAT2, Act b1 & Act b2 for beta-actin), cDNA template corresponding to the amount synthesized from 50 ng of total RNA and 2 units of DyNAzyme DNA polymerase. The sequence of primers as follows: B-MDIEA-NAT1, 5′-CACCCG GATCCGGGATCATGGACATTGAAGC-3′, nt 435–454, GenBank accession number X17059; VPKHGD-X-NAT1, 5′-GGTCCTCG AGTCAATCACCATGTTTGGGCAC-3′, nt 1295–1278, GENBANK accession number X17059; FP1-NAT2, 5′-CTAGTTCCTGGT TGCTGGCC-3′, nt 79–98, GenBank accession number NM-000015; RP1-NAT2, 5′-TAACGTGAGGGTAGAGAGGA-3′, nt 1073–1054, GenBank accession number NM-000015; Act b1, 5′-GCTCGTCGTC GACAACGGCTC-3′, nt 94–114, GenBank accession number NM-001101; Act2 b2, 5′-CAAACATGATCTGGGTCATCTTCTC-3′, nt 446–422, GenBank accession number NM-001101 as described previously [14].
Statistical analysis
Statistical analysis of the data was performed with an unpaired Student’s t-test and ANOVA tests. The kinetic constants were calculated with the Cleland HYPER Program [6] that performs linear regression using a least-squares method.
Results
Berberine affects the growth of Salmonella Typhi
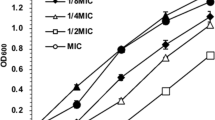
The data from intact bacteria cells culture cotreated with or without various concentrations of berberine on the growth of inhibition from Typhi is given in Fig. 1. The growth of Typhi was inhibited by berberine in a dose-dependent manner. Therefore, the concentration of berberine for IC50 value is 18 μM for Typhi.
Berberine affects the NAT activity of Salmonella Typhi
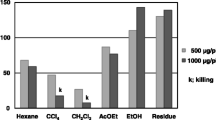
The effects of berberine on the NAT activity in Typhi in cell lysates and intact bacteria were examined by HPLC, assessing the percentage of N-acetylation of AF (AAF production). Cell lysates or suspensions of Typhi with or without various concentrations (0, 0.5, 1, 5, 25, and 50 μM) of berberine cotreatment showed different percentages of AF acetylation (Fig. 2). The data indicated that berberine induced dose-dependent effect in both cell lysates and intact bacteria.
The effect of various concentrations of berberine on NAT activity in Typhi. The intact cell incubated 24 h in the presence of 0, 0.5, 1, 5, 25, and 50 μM berberine. AAF was measured by HPLC assay. Each point represents the mean of triplicate assays of three incubations of cells. (A) AF production. (B) AAF production. *M significant differences between control and treated groups (P < 0.05).
Berberine affects the kinetic constants of NAT of Salmonella Typhi
NAT activity was found in the cell lysates of the bacterial Typhi. A double reciprocal plot of the data for 25 μM berberine inhibition to NAT activity in Typhi was prepared and given in Table 1. Clearly, when 25 μM berberine was added to the reaction mixtures, the apparent values of K m and Vmax were decreased 46.6% and 52.7% for cell lysates of Typhi.
Berberine affects the enzyme expression of NAT from Salmonella Typhi
The expression of NAT protein was measured by the ability of NAT antibody to form an antigen–antibody complex. The result from SDS-PAGE and Western blot experiment are given in Fig. 3A and 3B. The results demonstrated that 25 μM berberine inhibited NAT enzyme expression in Typhi.
Berberine affects expression of NAT1 enzyme in bacteria strain Typhi. The bacteria were incubated with 25 μM berberine for 24 h followed by evaluation of NAT expression. NAT expression was estimated by Western blotting as described in Materials and Methods. (A) The stain of NAT; (B) the ratio of NAT1/β-actin.
Berberine affects the gene expression of NAT (mRNA NAT) from Salmonella Typhi
To investigate whether berberine affects the gene expression of NAT, we performed reverse transcription–polymerase chain reaction (PCR) on Typhi RNA using specific primers for NAT1, NAT2, and β-actin. The result from the PCR experiment is given in Fig. 4A. The mRNA levels of NAT and β-actin on gel electrophoresis were quantified by densitometric analysis of gel photographs and expressed as NAT/β-actin (Fig. 4B). The results demonstrated that 25 μM berberine did inhibit NAT mRNA expression in Typhi.
Berberine affects expression of NAT mRNA in bacteria strain Typhi. The bacteria were incubated with 25 μM berberine for 24 h. The bacteria were collected to extract RNA. The extracted RNA was subjected to primers for NAT and β-actin, and then PCR-amplified cDNA derived from mRNA was applied to agarose gel-electrophoresis. (A) The mRNA NAT; (B) the ratio of NAT1/β-actin.
Discussion
In human populations, arylamine-induced neoplasms within target organs may indicate differential risks in NAT activities that are genetically mediated variations [11]. However, it is reported that enteric bacteria contain enzymes that are known to contribute to the metabolic activation of chemical carcinogens in animal studies [16]. The intestinal microflora have been revealed to contain NATs which acetylated arylamines from the animal studies [18]. The first three-dimensional NAT structure was resolved for NAT from Salmonella typhimurium, and subsequently the structure of NAT from Mycobacterium smegmatis has been elucidated [22]. In addition to the identification of NAT in prokaryotes, there has been considerable interest in the identification of homologous NAT sequences in eukaryotes. Our previous studies have shown that many kinds of enteric bacteria such as K. pneumoniae, Salmonella group B, Staphylococcus aureus, Shigella sonnei, and Helicobacter pylori exhibit NAT activity [5, 19, 24]. Therefore, the present study was focused on the effect of berberine on growth and NAT activity of Typhi.
The results clearly demonstrated that berberine inhibited growth, NAT activity, protein levels, and gene expression (mRNA NAT1) on Typhi. These effects occurred in a dose-dependent manner. Berberine can be used as a bacerticidal to Typhi. Based on the decreased value of kinetic constant of NAT, it was suggested that berberine might act as an uncompetitive inhibitor. It is well known that the enzyme inhibitor can be divided into competitive, uncompetitive, and noncompetitive inhibitor. A competitive inhibitor competes with the substrate for binding at the same site on the active enzyme. An uncompetitive inhibitor binds to a different site but blocks the conversion of the substrate to the products that lead to change Vmax while K m maintains the same. A noncompetitive inhibitor binds only to the enzyme–substrate complex, which leads to changing both values of K m and Vmax [28]. Apparently, further investigations are needed for the mechanism of berberine effects on the growth of Typhi.
The reason for our selection of AF for N-acetylation of Typhi is that 1) human intestinal bacteria exhibit different N-acetylation activity for AF; 2) AF is the common substrate for NATs [12]; 3) AF is a kind of arylamine carcinogen [13]; 4) it is more convenient for the determination by HPLC of the quantity of AF and AAF. The intestinal microflora in mice play an important role in the metabolic activation and absorption of 1-nitropyrene, and human microflora also can metabolize 1-nitropyrene to form 1-aminopyrene and N-formyl-1-aminopyrene [23]. NAT enzyme of human enteric bacteria may contribute to the metabolic activation of 2-amino-3-methylimidazo [4,5-fluroquinoline [20, 22].
Our results indicate that Typhi may be involved in the metabolic activation or the detoxification of the arylamine carcinogens in humans. Although we discovered the reducing of NAT mRNA in these bacteria after cotreatment with berberine, it is still unclear how these compounds get into the bacteria cells, how bacteria react to this chemical, and what kind of enzymes are involved in this mechanism. Apparently, further work is required to explain the mechanism of berberine inhibition of AF N-acetylation. However, our findings may offer some information on the possibility of decreasing arylamine carcinogens in induced carcinogenesis, because the increased levels of NAT activity may be associated with increased sensitivity to the mutagenic effects of many arylamines [9]. Furthermore, breast and bladder cancers have been demonstrated to be associated with the attenuation of NAT activity in the liver [13, 26]. The differences in NAT activity may be dependent on interindividual genetic variations and are potential sources of toxicological/pathological susceptibility among human populations [7, 8]. However, substrate-dependent downregulation [1, 3] and oxidative-dependent inactivation [1, 3, 4] may also regulate NAT activity. Apparently, it appears that the overall NAT activity in cells or tissues may depend on both genetic and environmental factors.
Literature Cited
Atmane N, Dairou J, Paul A, Dupret JM, Rodrigues-Lima F (2003) Redox regulation of the human xenobiotic metabolizing enzyme arylamine N-acetyltransferase 1 (NAT1). Reversible inactivation by hydrogen peroxide J Biol Chem 278:35086–35092
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding Anal Biochem 72:248–254
Butcher NJ, Arulpragasam A, Minchin RF (2004) Proteasomal degradation of N-acetyltransferase 1 is prevented by acetylation of the active site cysteine: a mechanism for the slow acetylator phenotype and substrate-dependent down-regulation J Biol Chem 279:22131–22137
Butcher NJ, Ilett KF, Minchin RF (2000) Substrate-dependent regulation of human arylamine N-acetyltransferase-1 in cultured cells Mol Pharmacol 57:468–473
Chung JG, Chen GW, Wu LT, Chang HL, Lin JG, Yeh CC, Wang TF (1998) Effects of garlic compounds diallyl sulfide and diallyl disulfide on arylamine N-acetyltransferase activity in strains of Helicobacter pylori from peptic ulcer patients Am J Chin Med 26:353–364
Cleland WW (1967) The statistical analysis of enzyme kinetic data Adv Enzymol Relat Areas Mol Biol 29:1–32
Dairou J, Atmane N, Dupret JM, Rodrigues-Lima F (2003) Reversible inhibition of the human xenobiotic-metabolizing enzyme arylamine N-acetyltransferase 1 by S-nitrosothiols Biochem Biophys Res Commun 307:1059–1065
Dairou J, Atmane N, Rodrigues-Lima F, Dupret JM (2004) Peroxynitrite irreversibly inactivates the human xenobiotic-metabolizing enzyme arylamine N-acetyltransferase 1 (NAT1) in human breast cancer cells: a cellular and mechanistic study J Biol Chem 279:7708–7714
Einisto P, Watanabe M, Ishidate M Jr, Nohmi T (1991) Mutagenicity of 30 chemicals in Salmonella typhimurium strains possessing different nitroreductase or O-acetyltransferase activities Mutat Res 259:95–102
Ghosh AK, Bhattacharyya FK, Ghosh DK (1985) Leishmania donovani: amastigote inhibition and mode of action of berberine Exp Parasitol 60:404–413
Grant DM, Blum M, Beer M, Meyer UA (1991) Monomorphic and polymorphic human arylamine N-acetyltransferases: a comparison of liver isozymes and expressed products of two cloned genes Mol Pharmacol 39:184–191
Hein DW (2002) Molecular genetics and function of NAT1 and NAT2: role in aromatic amine metabolism and carcinogenesis Mutat Res 506–507:65–77
Hein DW, Rustan TD, Bucher KD, Furman EJ, Martin WJ (1991) Extrahepatic expression of the N-acetylation polymorphism toward arylamine carcinogens in tumor target organs of an inbred rat model J Pharmacol Exp Ther 258:232–236
Hsia TC, Chung JG, Lu HF, Ho HC, Yang CC, Lu KH, Hung CF (2002) The effect of paclitaxel on 2-aminofluorene-DNA adducts formation and arylamine N-acetyltransferase activity and gene expression in human lung tumor cells (A549). Food Chem Toxicol 40:697–703
Iwasa K, Lee DU, Kang SI, Wiegrebe W (1998) Antimicrobial activity of 8-alkyl- and 8-phenyl-substituted berberines and their 12-bromo derivatives J Nat Prod 61:1150–1153
Kinouchi T, Kataoka K, Miyanishi K, Akimoto S, Ohnishi Y (1993) Biological activities of the intestinal microflora in mice treated with antibiotics or untreated and the effects of the microflora on absorption and metabolic activation of orally administered glutathione conjugates of K-region epoxides of 1-nitropyrene Carcinogenesis 14:869–874
Kuo CL, Chi CW, Liu TY (2004) The anti-inflammatory potential of berberine in vitro and in vivo Cancer Lett 203:127–137
Larse GL (1988) Deconjugation of biliary metabolites by microfloral β-glucronidase, sulphatase and cysteine conjugate β-lyase and their subsequent enterohepatic circulation. In: Rowland I (ed) Role of gut flora in toxicity and cancer. London: Academic Press, pp 79–107
Lo HH, Chung JG (1999) The effects of plant phenolics, caffeic acid, chlorogenic acid and ferulic acid on arylamine N-acetyltransferase activities in human gastrointestinal microflora Anticancer Res 19:133–139
Muckel E, Frandsen H, Glatt HR (2002) Heterologous expression of human N-acetyltransferases 1 and 2 and sulfotransferase 1A1 in Salmonella typhimurium for mutagenicity testing of heterocyclic amines Food Chem Toxicol 40:1063–1068
Payton M, Mushtaq A, Yu TW, Wu LJ, Sinclair J, Sim E (2001) Eubacterial arylamine N-acetyltransferases—identification and comparison of 18 members of the protein family with conserved active site cysteine, histidine and aspartate residues Microbiology (Reading, England) 147:1137–1147
Sim E, Pinter K, Mushtaq A, Upton A, Sandy J, Bhakta S, Noble M (2003) Arylamine N-acetyltransferases: a pharmacogenomic approach to drug metabolism and endogenous function Biochem Soc Trans 31:615–619
Steffensen IL, Fretland AJ, Paulsen JE, Feng Y, Eide TJ, Devanaboyina US, Hein DW, Alexander J (2000) DNA adduct levels and intestinal lesions in congenic rapid and slow acetylator syrian hamsters admi food mutagens 2-amino-1-methyl-6-phenylimidazo[4,5-b]pyridine (PhIP) or 2-amino-3-methylimidazo[4,5-f]quinoline (IQ) Pharmacol Toxicol 86:257–263
Tsou MF, Hung CF, Lu HF, Wu LT, Chang SH, Chang HL, Chen GW, Chung JG (2000) Effects of caffeic acid, chlorogenic acid and ferulic acid on growth and arylamine N-acetyltransferase activity in Shigella sonnei (group D) Microbios 101:37–46
Virtanen P, Lassila V, Njimi T, Ekotto MD (1988) Regeneration of D-galactosamine-traumatized rat liver with natural protoberberine alkaloids from Enantia chlorantha Acta Anat (Basel) 132:159–163
Weber WW, Hein DW (1985) N-acetylation pharmacogenetics Pharmacol Rev 37:25–79
Yamamoto K, Takase H, Abe K, Saito Y, Suzuki A (1993) Pharmacological studies on antidiarrheal effects of a preparation containing berberine and geranii herba [in Japanese] Nippon Yakurigaku Zasshi 101:169–175
Zubay GLWWPaDEV (1995) Enzyme kinetics. In: Principles of biochemistry. Wm. C. Brown, pp 133–149
Acknowledgments
This work was supported by grant CMU92-M-26 from China Medical University.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wu, LT., Tsou, MF., Ho, CC. et al. Berberine Inhibits Arylamine N-Acetyltransferase Activity and Gene Expression in Salmonella Typhi. Curr Microbiol 51, 255–261 (2005). https://doi.org/10.1007/s00284-005-4569-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-005-4569-7