Abstract
Sickle cell anemia (SCA) is an autosomal recessive monogenic disease with significant clinical variability. Cerebrovascular disease, particularly ischemic stroke, is one of the most severe complications of SCA in children. This study aimed to investigate the influence of genetic variants on the levels of fetal hemoglobin (Hb F) and biochemical parameters related with chronic hemolysis, as well as on ischemic stroke risk, in ninety-one unrelated SCA patients, children of sub-Saharan progenitors. Our results show that a higher Hb F level has an inverse relationship with the occurrence of stroke, since the group of patients who suffered stroke presents a significantly lower mean Hb F level (5.34 ± 4.57% versus 9.36 ± 6.48%; p = 0.024). Furthermore, the co-inheritance of alpha-thalassemia improves the chronic hemolytic pattern, evidenced by a decreased reticulocyte count (8.61 ± 3.58% versus 12.85 ± 4.71%; p < 0.001). In addition, our findings have confirmed the importance of HBG2 and BCL11A loci in the regulation of Hb F expression in sub-Saharan African SCA patients, as rs7482144_A, rs11886868_C, and rs4671393_A alleles are significantly associated with a considerable increase in Hb F levels (p = 0.019, p = 0.026, and p = 0.028, respectively). Concerning KLF1, twelve different variants were identified, two of them novel. Seventy-three patients (80.2%) presented at least one variant in this gene. However, no correlation was observed between the presence of these variants and Hb F level, severity of hemolysis, or stroke occurrence, which is consistent with their in silico-predicted minor functional consequences. Thus, we conclude that the prevalence of functional KLF1 variants in a sub-Saharan African background does not seem to be relevant to SCA clinical modulation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hemoglobinopathies are inherited anemia characterized by abnormal structure, function, or amount of the globin chains of hemoglobin. One of the most common hemoglobinopathies worldwide, sickle cell anemia (SCA), is due to homozygosity for the c.20A>T mutation in the beta-globin gene (HBB) that gives rise to hemoglobin S (Hb S). Although SCA is a monogenic disease, it is clinically very heterogeneous due to the influence of environmental and genetic modifiers, such as an increased expression of fetal hemoglobin (Hb F) during adult life [1,2,3]. The beneficial effect of higher levels of Hb F results from the corresponding reduction of Hb S concentration inside the red blood cells thus hindering deoxy Hb S polymerization [3]. Another known SCA clinical modifier is alpha-thalassemia, a hemoglobinopathy due to reduced production of alpha-globin chains. Its co-inheritance with SCA results in a decreased intracellular concentration of Hb S and reduces the likelihood and rate of Hb S polymerization [4, 5].
The Hb F level in normal adults is usually less than 1% of the total hemoglobin. However, a variable level, ranging from approximately 1 to 30%, is frequently observed among SCA patients. A specific single nucleotide polymorphism (SNP) within the beta-globin gene cluster on chromosome 11p15.5 partly accounts for this variation. It is the rs7482144 (the Xmn I site) located in the promoter of the Gγ-globin gene (HBG2) [6]. Furthermore, genome-wide association studies have identified two additional quantitative trait loci that affect baseline Hb F levels in patients with hemoglobin disorders, which involve the genetic determinants of trans-acting factors not linked to the beta-globin cluster. They comprise the HBS1L-MYB intergenic region on 6q22–23 and the BCL11A gene on 2p16.1 [7,8,9,10,11,12]. More recently, the trans-acting erythroid-specific Krüppel-like factor 1 (KLF1) was also implicated in Hb F expression modulation. Additionally, KLF1 plays a multifunctional role during erythropoiesis and is involved in the activation of numerous erythroid genes, namely those encoding globins, red cell membrane and structural proteins, heme synthesis enzymes, and many other proteins involved in red cell metabolism [13,14,15,16]. KLF1 modulates the fetal to adult switching of hemoglobin by two ways: stimulating the HBB gene expression through direct interaction with its promoter, and repressing HBG gene expression indirectly through the activation of its repressor BCL11A [17]. Therefore, it seems that weakening the activity of KLF1 would stimulate Hb F synthesis through the upregulation of HBG gene expression. Accordingly, some KLF1 gene variants associated with KLF1 haploinsufficiency have been related to higher Hb F levels, which may be particularly relevant to some hemoglobinopathy clinical outcomes [14, 16, 18, 19]. Thus, it has been hypothesized that KLF1 variants may have been under positive selection by malaria and this may justify their polymorphic frequencies in populations where some type of hemoglobinopathies, such as beta-thalassemia and Hb E disorder, are commonly found, e.g., in Southern China and South East Asia [19,20,21]. In contrast, the possible interaction of KLF1 variants and SCA remains unclear. A study carried out in a group of 44 SCA patients from India and the Middle East did not replicate the previously described association between KLF1 SNPs and Hb F level [22]. In addition, studies performed on African Americans with SCA failed to detect association between KLF1 SNPs and Hb F levels [23, 24]. In this regard, the lack of knowledge about the prevalence of KLF1 variants and its contribution to Hb F levels and SCA modulation is even greater when considering the African context. A genome-wide association study performed in SCA patients from an East African country (Tanzania) did not detect an association between KLF1 SNPs and Hb F levels [25]. In accordance with those results, in a UK study where 55 SCA patients were enrolled, only one Nigerian SCA patient presented a KLF1 functional variant [26]. Therefore, due to the paucity of such data in African SCA populations, we screened a group of sub-Saharan African SCA pediatric patients for KLF1 gene variants. Apart from confirming the role of the other known SCA genetic modifiers, we aimed to examine the effect of KLF1 genetic variants on the phenotypic expression of Hb F in these patients, as well as in their hematological/biochemical chronic hemolysis biomarkers and cerebral vasculopathy outcome.
Materials and methods
Study population and data collection
Patients eligible for this study were unrelated children with SCA, ≥ 3 years old, with a direct sub-Saharan African ancestry, who had been followed up in general hospitals from the Greater Lisbon area. Their genetic background is African; however, many of these patients were born in Portugal.
After written informed consent given by the patients’ legal representatives, ninety-one SCA children (3 to 16 years old) were enrolled in this study (53 males and 38 females). All participants are children of sub-Saharan African couples. Eighty-three patients have both progenitors from the same country of origin: Angola (n = 55), Guinea-Bissau (n = 10), S. Tomé (n = 10), Cape Verde (n = 6), Guinea-Conakry (n = 1), and Nigeria (n = 1). Eight participants have progenitors with different African origins (Angola/Democratic Republic of Congo (n = 3), Angola/Cape Verde (n = 2), Angola/Guinea-Bissau (n = 1), Cape Verde/S. Tomé (n = 1), Cape Verde/Guinea-Bissau (n = 1)).
Data obtained from their interviews, which included demographic characteristics (age, gender, and parents’ birthplace), were used to create a database. The database was then filled up with data retrospectively collected from the patients’ hospital records regarding biochemical and hematological hemolysis biomarkers, such as serum lactate dehydrogenase (LDH), total bilirubin, and reticulocyte count, as well as hemoglobin profiling. These parameters were obtained by standard procedures, and Hb F levels were measured by high-performance liquid chromatography (HPLC). The collected data for each patient were the result of, at least, three different time point analyses performed in steady state, before starting treatment with hydroxyurea and more than 3 months after receiving a blood transfusion. At present, 43 out of the 91 SCA patients are under hydroxyurea treatment; however, their Hb F values registered in the database were the ones observed previously to HU treatment.
Cerebrovascular disease (CVD) is one of the most severe complications in SCA, and ischemic stroke is the most severe type of CVD, which may occur in SCA pediatric patients. Transcranial Doppler (TCD) ultrasonography is currently the standard tool for risk screening of stroke. Children who present at least two measurements of time-averaged mean of the maximum velocity (TAMMX) in the middle cerebral artery ≥ 200 cm/s were classified as high risk for stroke and, usually, entered a regular blood transfusion therapy for reducing stroke risk. HU is another successful therapeutic option, alternatively or in conjunction with blood transfusion therapy.
In this work, overt ischemic stroke occurrence was the parameter used as CVD outcome. Fifteen out of the 91 enrolled SCA patients had already experienced at least one episode of overt ischemic stroke, confirmed by magnetic resonance imaging. Four of them are currently under chronic blood transfusion therapy. The remaining eleven entered a chronic transfusion program in conjunction with HU treatment.
An aliquot of peripheral blood sample was collected from each participant for DNA extraction and candidate gene genotyping.
The study protocol was approved by INSA’s ethics committee and is in accordance with the Helsinki declaration of 1975.
Genetic testing for known variants
Genomic DNA was isolated from peripheral leukocytes using the MagNA Pure LC instrument (Roche Diagnostics GmbH, Mannheim, Germany). The homozygous status for the SCA mutation in HBB gene (c.20A>T) was confirmed by polymerase chain reaction, followed by restriction fragment length polymorphism analysis (PCR-RFLP) with the endonuclease Bsu36 I. Beta-globin cluster haplotypes were assigned after examining six restriction endonucleases sites within the cluster: Xmn I (5′ to HBG2), Hind III (within the HBG2 and HBG1), Hinc II (within and 3′ to ψHBB), and Hinf I (3′ to HBB). Aliquots of the amplified products were then digested with the appropriate restriction enzymes under the conditions recommended by the manufacturers. Haplotypes were inferred by determining the presence or absence of cleavage at each site, and by compiling the results into one known pattern [27].
The 3.7-kb alpha-thalassemia deletion was assessed by gap-PCR [28]. Concerning the two SNPs analyzed in the BCL11A gene, rs11886868 and rs4671393, the former was characterized by PCR-RFLP using the Mbo II endonuclease, while the latter was analyzed by PCR followed by Sanger sequencing in a 3130XL Genetic Analyser, ABI PRISM (Applied Biosystems, Foster City, CA, USA). All genotype distributions were tested positively for adherence to the Hardy-Weinberg equilibrium.
Screening for KLF1 variants by NGS
KLF1 gene analysis was carried out by next-generation sequencing (NGS) of a long PCR fragment (3.2 kb) including the KLF1 gene and its flanking regions. PCR was obtained by using the primers: Fw5′-CTGATAGCAGCCTCCAACGTCTGGG-3′ and Rv5′-ACCTTCAGGAGCCGCTTTCT-3′ and the FailSafe PCR enzyme with the PreMix D (Epicentre, Illumina). PCR conditions consisted of 94 °C for 2 min, followed by 35 cycles of 94 °C for 45 s, 63 °C for 45 s, and 72 °C for 4 min. This was followed by incubation at 72 °C for 5 min.
NGS workflow consisted of five steps: (1) PCR product purification using Agencourt AMPure XP (Beckman Coulter); (2) double-stranded DNA quantification in Qubit fluorometer (Thermo Fisher Scientific); (3) library preparation using the Nextera XT DNA Library Preparation Kit (Illumina) following manufacturer’s instructions; (4) sequencing on a MiSeq benchtop sequencer (Illumina); and (5) data analysis using MiSeq Reporter v2.6.2 (Illumina). Variants were inspected and validated using the Integrative Genomics Viewer v2.3.86 [29]. KLF1 variants found by NGS were confirmed by Sanger sequencing.
To further assess the effect of the novel KLF1 missense variant on protein function, bioinformatic tools were used: SIFT (http://sift.jcvi.org) and PolyPhen-2 (http://genetics.bwh.harvard.edu/pph2).
Statistical analysis
Statistical analyses were performed using the SPSS version 23 software. Normal distribution was assessed using Kolmogorov-Smirnov test. Continuous variables were evaluated as mean ± SD when the distribution was normal and median with interquartile range when the distribution was not normal. The analysis of Student’s t test was used to compare differences among means and Mann-Whitney test was used as non-parametric test of significance. A p value of < 0.05 was considered statistically significant.
Results
Genotyping was performed to (i) confirm homozygosity for the c.20A>T mutation in the HBB gene; (ii) identify beta-globin cluster haplotypes and the presence of the 3.7-kb alpha-thalassemia deletion; (iii) characterize SNPs in the HBG2 rs7482144, BCL11A rs11886868, and rs4671393; and (iv) search for KLF1 variants. The effect of genetic variants on levels of Hb F, biochemical parameters related to the severity of hemolysis, and stroke occurrence was also investigated.
Relationship between Hb F level, biochemical parameters related to chronic hemolysis severity, and stroke occurrence
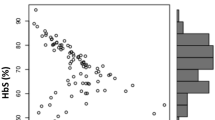
The Hb F is known as the principal SCA genetic modifier, and a concentration of at least 10% in SCA patients was reported as necessary to reduce the clinical severity [30]. The Hb F mean ± SD presented by our SCA patients was 8.9 ± 6.4% (median 7.4%, interquartile range 9.2). Using a cut-off value of 10%, patients with Hb F levels lower than this value were categorized in the class of “low Hb F,” while those with values ≥ 10% were grouped in the class “high Hb F.” Comparing the levels of hemolysis-related biomarkers between these two subgroups, the higher Hb F group presented a trend to lower values of two of these parameters, although not reaching statistical significance (reticulocyte count mean, 10.17 ± 4.47% vs 11.36 ± 4.93%, p = 0.271; and LDH mean 689.96 ± 324.73 U/L vs 853.35 ± 366.93 U/L; p = 0.053, respectively). On the other hand, the group of patients who had suffered at least one stroke event presented a significantly lower level of Hb F (Hb F mean 5.34 ± 4.57% vs 9.36 ± 6.48%, respectively; p = 0.024).
Relationship between common polymorphisms in (alpha and beta) globin clusters and BCL11A, Hb F level and biochemical parameters related to hemolysis severity, and stroke occurrence
The beta-globin cluster haplotypes found in our patients were consistent with parental geographic origin. Taking into consideration the three most representative geographic subgroups, they presented the following allele frequencies: (i) the Angolan group (n = 55): Bantu 91.8%, Benin 4.5%, Senegal 1.8%, and atypical 1.8%; (ii) the Guinean group (n = 10): Benin 10% and Senegal 90%, and (iii) the Santomean group (n = 10): Bantu 30%, Benin 65%, and atypical 5%. Considering genotypes, 50 patients were homozygous for the Bantu haplotype, 12 were homozygous for the Senegal haplotype, 7 were homozygous for the Benin haplotype, and the remaining 22 individuals were compound heterozygotes.
We observed that patients homozygous for the Bantu haplotype (n = 46) had lower Hb F (mean of 7.2 ± 4.9%) than the homozygous for the Senegal haplotype (n = 11) who had a Hb F mean of 12.8 ± 5.8% (p = 0.005). One fundamental difference between these two haplotypes is the SNP rs7482144 located at −158 of HBG2, where the allele A is detected as positive. In our patients, a frequency of 18.7% of the A allele was found, always in association with the Senegal haplotype. It was found largely in children whose parents originated from Cape Verde and Guinea-Bissau. The group of children presenting at least one allele A (GA+AA; dominant model of inheritance) had higher Hb F in contrast with the GG group (11.5 ± 6.35% vs 8.08 ± 6.25%; p = 0.019) (Table 1). Conversely, SCA patients belonging to the “high Hb F” group were three times more likely to present the rs7482144_allele A than those belonging to the “low Hb F” group (odds ratio = 3.0435; 95%CI = 1.1771–7.8692; p = 0.022).
Due to the African ancestry of our SCA patients, a high prevalence of alpha-thalassemia determinants was expected and confirmed. Fifty patients presented the normal four α-globin genes (αα/αα), thirty-one were heterozygous for the deletion (−α3.7kb/αα), six were homozygous (−α3.7kb/−α3.7kb), one had the triple α-globin gene (αααanti3.7kb/αα), and for the remaining three, no laboratory result was obtained. Thus, in our series, 37 out of 88 patients had an alpha-thalassemia determinant, corresponding to a prevalence of 42%, and an allele frequency of the −α3.7-kb deletion of 24.4%. However, considering only the larger subgroup of Angolan SCA patients, 24 out of these 55 patients presented alpha-thalassemia, corresponding to a higher prevalence of 45.5% and an allele frequency of the −α3.7kb deletion of 27.3%.
We observed that the SCA patients who presented at least one alpha-thalassemia allele (genotypes: −α3.7kb/αα or −α3.7kb/−α3.7kb; n = 37) had a lower reticulocyte percentage than the other group without alpha-thalassemia (genotypes: αα/αα or ααα/αα; n = 51). The first group presented a reticulocyte count mean of 8.61 ± 3.58% while the latter presented 12.85 ± 4.71% (p < 0.001). Considering the reticulocyte count in absolute values, the same conclusion was obtained: the first group having alpha-thalassemia presented a mean of 268,244 ± 98,453 cells/μL while the group without alpha-thalassemia had 358,243 ± 154,259 cells/μL (p = 0.002). Therefore, the co-inheritance of at least one alpha-thalassemia allele seems to improve the hemolytic status of SCA.
Concerning BCL11A SNPs, we observed a minor allele frequency of the rs11886868_C allele of 25.8% and the rs4671393_A allele of 27.5%. The presence of these alleles, considering the inheritance dominant model, seems to introduce a considerable increase on Hb F level (p = 0.026 and p = 0.028, respectively) (Table 1).
Searching for genetic variants in the KLF1 gene and their relationship with Hb F level
To investigate any possible relationship between KLF1 variants, Hb F level, and stroke occurrence in SCA, we conducted a KLF1 gene analysis by NGS. Twelve different variants in the KLF1 gene orin its regulatory regions were identified by NGS and validated by Sanger sequencing (Table 2). Seventy-three patients (80.2%) presented, at least, one variant in the KLF1 gene. However, almost all consist of alterations previously reported as belonging to functional class 1, which comprises those with no or minor functional consequences [16]. Six variants were considered as common SNPs with minor allele frequency (MAF) ≥ 5%: (i) one is located at the 5′UTR, c.−251C>G; (ii) three are located at exon 2 and are missense variants, c.115A>C, c.304 T > C, and c.544 T > C; and (iii) two are located at the 3′UTR, c.*277C>T and c.*296G>A. The most frequent variant found was the rs2072597, c.304T>C, with a MAF of 42.86%. It is responsible for an amino acid change in the N-terminal domain of the protein p.(Ser102Pro). However, this change was predicted as benign in terms of function by in silico analysis. The same result is observed in ClinVar. The second most frequent variant found was the rs3817621, c.−251C>G, with a MAF of the C allele of 18.68%. These two SNPs, besides being the most frequent in our series, are reported in the 1000 Genomes Project Phase 3 as widely dispersed in the worldwide population, including populations in the African continent where their MAFs are 44% and 19%, respectively.
In this study, two novel KLF1 variants were detected, both in the heterozygous state, in two patients: the c. −149(C/−) located in the promoter region, and the c.1026G>C, in exon 3 (Fig. 1). The first is a deletion of one nucleotide within an Sp1 binding site and probably affects the binding of this transcription factor. It was found in compound heterozygosity with two other KLF1 variants, the c.115A>C and c.304T>C, in an Angolan patient with 3.4% of Hb F. This patient did not have the other Hb F–related variants at HBG2 and BCL11A. The other novel variant found in KLF1 changes the corresponding amino acid, p.(Gln342His), that is located in the DNA-binding domain zinc finger 3 of the KLF1 protein. However, in silico analysis of this variant predicted a benign consequence at the protein level (Polyphen: HumDiv score = 0.013 and HumVar score = 0.063; Sorting Intolerant from Tolerant (SIFT) = 1.000). It was found in compound heterozygosity with another variant of KLF1, the c.304T>C, in a patient from Guinea-Bissau with 14.6% of Hb F. However, it was also observed that this patient was homozygous for the A allele at rs7482144 in the HBG2 gene, which may justify this phenotype. Thus, in general, we found no association between the presence of KLF1 variants and the level of Hb F or serum hemolytic parameters or stroke events.
Discussion
Several studies have been conducted to find the main risk and protective factors involved in the development of specific SCA subphenotypes. Among those factors, Hb F level has been of great interest due to its substantial protective effect on the course (timing and severity) of SCA subphenotypes. In our study, SCA children of sub-Saharan African couples (mainly Angola, Guinea-Bissau, S. Tomé, or Cape Verde) were evaluated. Their genetic background is African; however, some of these patients were born and live in Portugal. This means that their environmental and to a certain extent their behavioral determinants of health are southern European.
We evaluated the influence of genetic factors on Hb F level, on the level of the biochemical parameters related to hemolysis, and the occurrence of overt ischemic stroke. The participants in this study showed a mean level of Hb F of 8.9 ± 6.4% which is similar to values reported by other studies on African SCA children, e.g., from Nigeria, Uganda, or Congo Republic, who presented 9.9 ± 6.0% [31], 9.0 ± 5.6% [32], and 8.8 ± 5.8% [33], respectively. Nevertheless, in other studies, lower values of Hb F have been reported for African SCA populations [34,35,36,37]. This discrepancy may be the result of several factors, such as the method used for the Hb F estimation, the age of the SCA patients, or due to environmental, cellular, and genetic factor modulators of phenotypic expression of Hb F.
A level of Hb F above 10% has been suggested as necessary for reduced SCA clinical severity [30]. Accordingly, we have found a negative association between the occurrence of ischemic stroke and the level of Hb F. In fact, SCA patients who have experienced stroke presented significantly lower values of mean Hb F in contrast to the group without stroke (Hb F mean 5.34 ± 4.57% vs 9.36 ± 6.48%, respectively; p = 0.024). Other authors, when studying Nigerian SCA patients, obtained similar results: the group presenting stroke had a mean of Hb F level of 5.6 ± 1.5% in contrast to the group without stroke, which presented Hb F of 10.0 ± 6.1% [31].
Besides the role of Hb F in modulating SCA morbidity, the co-inheritance of alpha-thalassemia has also been considered an important genetic modulator. It seems to play a role in reducing the intensity of chronic hemolysis by decreasing the tendency of deoxy Hb S to polymerize through lowering the mean cell hemoglobin concentration and erythrocyte density [5, 38]. Moreover, the damaging role of hemolysis is evident in the pathophysiology of SCA since free hemoglobin is a well-known scavenger of nitric oxide, which maintains vascular patency through a cascade of biological events that culminate in the relaxation of smooth muscle cells of blood vessel walls. Our findings are in line with this theory, as we found a significant association between the presence of alpha-thalassemia and decreased reticulocyte count, one of the hemolysis biomarkers analyzed. Thus, the co-inheritance of alpha-thalassemia with SCA seems to act beneficially in two ways: by lowering reticulocyte count and consequently their harmful adhesion to endothelium and by decreasing the severity of hemolysis, and consequently free hemoglobin levels, which in turn preserves the protective effect of higher nitric oxide bioavailability.
Our results confirm the importance of HBG2 and BCL11A loci in the regulation of Hb F expression in sub-Saharan African SCA descents. On the other hand, our results did not show an association between the presence of KLF1 variants and levels of Hb F, serum hemolytic parameters, or stroke events. This fact may be due to the nature of the KLF1 variants found in this population, which is consistent with the absence of functional consequences, as predicted by in silico analysis.
The KLF1 protein contains at its C-terminal region three zinc finger domains that enable its binding to DNA at specific sites of the genome, in order to control gene expression by working primarily as a transcriptional activator. Genetic variants located at critical residues of those zinc finger domains have functional consequences and are classified as belonging to functional classes 2 to 4 [16]. On the other hand, variants located outside those DNA-binding domains are categorized in class 1, since they present minor or no functional consequences [16]. It is expected that KLF1 variants belonging to classes 2 or 3 are associated with reduced levels of KLF1 factor (and consequently of the HBG repressor BCL11A), originating derepression of the fetal globin genes and consequently increased Hb F levels [17, 18]. This is in line with some studies where KLF1 classes 2 and 3 variants were observed at high frequency in populations where thalassemias are endemic and where they were associated with high Hb F levels and/or with thalassemia milder phenotypes, e.g., in Southern China [19, 39], South East Asia [20, 21], and Sardinia [40]. However, SCA presents a different worldwide distribution, with a high prevalence in African and Arab-Indian populations. In those populations, the prevalence of KLF1 variants and their possible association with Hb F levels have been poorly investigated. Two genome-wide association studies performed in SCA patients of African descent failed to identify common KLF1 variants influencing F-cell count [41] or associated with higher Hb F levels [25]. Another study addressed the hypothesis of loss-of-function mutations in KLF1 being involved with increased Hb F levels in SCA patients of diverse origins residing in the UK. This investigation concluded that only one (Nigerian) patient out of the 55 SCA patients studied presented a KLF1 mutation and increased Hb F [26]. Similarly, research on 44 SCA patients with high Hb F levels and an Arab-Indian origin (presenting the Arab-Indian haplotype in the beta-globin cluster) did not find the variants in the KLF1 zinc finger domains that were previously associated with increased Hb F. Therefore, the authors concluded that the KLF1 SNPs detected were neutral in determining Hb F level in those patients [22]. Our study demonstrates that variants in KLF1 are prevalent in sub-Saharan populations with SCA (≈ 80% of the patients present at least one variant in this gene). However, due to their non-deleterious effect on gene expression (mainly class 1 variants), they do not show association with any peculiar subphenotype and do not seem to be a relevant modifier of Hb F expression.
On the other hand, rare variants located at specific sites of the KLF1 promoter may disturb the normal gene transcription level. This is the case of the reported c.−148G>A variant for which in vitro studies revealed a consequent ablation of the SP1 transcription factor binding and a reduction of KLF1 expression [42]. Similarly, we found a nearby novel heterozygous single nucleotide deletion c.−149(C/−), in a SCA Angolan patient presenting with a mean Hb F of 3.4% and absence of the high Hb F–related alleles at HBG2 or BCL11A. This variant may negatively affect the SP1 binding, consequently reducing KLF1 expression in this individual but, due to its low frequency in the studied population (1/182 alleles), we can conclude that its impact in sub-Saharan SCA patients’ Hb F level is not relevant.
Three previously reported KLF1 missense variants, p.(Met39Leu), p.(Ser102Pro), and p.(Phe182Leu), were found in this study. The most frequent of them, the p.(Ser102Pro), was observed with a remarkably high allele frequency (42.9%). Its widely dispersed geographical distribution is known, with a minor allele frequency > 40% [43]. In an early study, this variant in homozygosity showed a 15% reduction in KLF1 expression, when compared with the normal wild-type genotype [42]. Nevertheless, similarly to our study, other authors did not find an association of this variant with any peculiar subphenotype [44]. The other two frequent missense variants we found, the p. (Met39Leu) and p.(Phe182Leu), showed allele frequencies of 6.6 and 4.9%, respectively, which is in accordance with published data for the African population in the “1000 Genomes Project.”
Regarding the novel missense variant located in KLF1 exon 3, the c.1026G>C, p.(Gln342His), in silico analysis indicates it probably will have no effect on KLF1 function. In our study, it was found in heterozygosity in only one SCA patient, a child of a Guinea-Bissau couple. This patient had a high mean Hb F (14.6%). However, this may be just a coincidental finding, as he also presented homozygosity for the A allele at SNP rs7482144, the −158 Xmn I site of HBG2, which per se may account for the observed high Hb F phenotype.
Conclusion
We conclude that higher Hb F levels have an inverse relationship with sub-Saharan SCA disease burden, namely on the most severe vascular outcome in children—the occurrence of ischemic stroke. In addition, there is a significant effect of alpha-thalassemia co-inheritance in the intensity of chronic hemolysis in these patients. These genetic factors have the potential to be used as early prognosis biomarkers, suggesting a closer follow-up and prioritizing access to Hb F–inducing pharmacological agents. We also conclude that, although some genetic modification of KLF1 expression may result in increased Hb F levels, and consequently in ameliorating disease phenotype, the low prevalence of functional KLF1 variants in the sub-Saharan background seems to be non-significant in the SCA clinical modulation context.
References
Sebastiani P, Solovieff N, Hartley SW, Milton JN, Riva A, Dworkis DA et al (2010) Genetic modifiers of the severity of sickle cell anemia identified through a genome-wide association study. Am J Haematol 85(1):29–35
Thein SL (2008) Genetic modifiers of the beta-hemoglobinopathies. Br J Haematol 141(3):357–366
Platt OS, Brambilla DJ, Rosse WF, Milner PF, Castro O, Steinberg MH, Klug PP (1994) Mortality in sickle cell disease: life expectancy and risk factors for early death. N Engl J Med 330(23):1639–1644
Embury SE, Clark MR, Monroy G, Mohandas N (1984) Concurrent sickle cell anemia and alpha-thalassemia. Effect on pathological properties of sickle erythrocytes. J Clin Invest 73(1):116–123
Rumaney MB, Bitoungui VJN, Vorster AA, Ramesar R, Kengne AP, Ngogang J, Wonkam A (2014) The co-inheritance of alpha-thalassemia and sickle cell anemia is associated with better hematological indices and lower consultations rate in Cameroonian patients and could improve their survival. PLoS One 9(6):e100516
Labie D, Dunda-Belkhodja O, Rouabhi F, Pagnier J, Ragusa A, Nagel RL (1985) The -158 site 5’ to the Gγ gene and Gγ expression. Blood 66(6):1463–1465
Lettre G, Sankaran VG, Bezerra MAC, Araújo AS, Uda M, Sanna S (2008) DNA polymorphisms at the BCL11A, HBS1L-MYB, and β-globin loci associate with fetal hemoglobin levels and pain crises in sickle cell disease. Proc Natl Acad Sci U S A 105(33):11869–11874
Uda M, Galanello R, Sanna S, Lettre G, Sankaran VG, Chen W (2008) Genome-wide association study shows BCL11A associated with persistent fetal hemoglobin and amelioration of the phenotype of beta-thalassemia. Proc Natl Acad Sci U S A 105:1620–1625
Sankaran VG, Menne TF, Xu J, Akie TE, Lettre G, Van Handel B et al (2008) Human fetal hemoglobin expression is regulated by the development stage specific repressor BCL11A. Science 322(5909):1839–1842
Thein SL, Menzel S, Lathrop M, Garner C (2009) Control of fetal hemoglobin: new insights emerging from genomics and clinical implications. Hum Mol Genet 18(R2):R216–R223
Creary LE, Ulug P, Menzel S, McKenzie CA, Hanchard NA, Taylor V et al (2009) Genetic variation on chromosome 6 influences F cells in healthy individuals of African descent and HbF levels in sickle cell patients. PLoS One 4(1):e4218
Wahlberg K, Jiang J, Rooks H, Jawaid K, Matsuda F, Yamaguchi M, Lathrop M, Thein SL, Best S (2009) The HBS1L-MYB intergenic interval associated with elevated HbF levels shows characteristics of a distal regulatory region in erythroid cells. Blood 114(6):1254–1262
Bieker JJ (2010) Putting a finger on the switch. Nat Genet 42(9):733–734
Borg J, Patrinos GP, Felice AE, Philipsen E (2011) Erythroid phenotypes associated with KLF1 mutations. Haematologica 96(5):635–638
Siatecka M, Bieker JJ (2011) The multifunctional role of EKLF/KLF1 during erythropoiesis. Blood 118(8):2044–2054
Perkins A, Xu X, Higgs D, Patrinos GP, Arnaud L, Bieker JJ, Philipsen S, the KLF1 Consensus Worgroup (2016) Krüppeling erythropoiesis: an unexpected broad spectrum of human red blood cell disorders due to KLF1 variants. Blood 127(15):1856–1862
Zhou D, Liu K, Sun CW, Pawlik KM, Townes TM (2010) KLF1 regulates BCL11A expression and gamma- to beta-globin gene switching. Nat Genet 42(9):742–774
Borg J, Papadopoulos P, Georgitsi M, Gutiérrez L, Grech G, Fanis P, Phylactides M, Verkerk AJMH, van der Spek PJ, Scerri CA, Cassar W, Galdies R, van IJcken W, Özgür Z, Gillemans N, Hou J, Bugeja M, Grosveld FG, von Lindern M, Felice AE, Patrinos GP, Philipsen S (2010) Haploinsufficiency for the erythroid transcription factor KLF1 causes hereditary persistence of fetal hemoglobin. Nat Genet 42(9):801–805
Liu D, Zhang X, Yu L, Cai R, Ma X, Zheng C (2014) KLF1 mutations are relatively more common in a thalassemia endemic region and ameliorate the severity of beta-thalassemia. Blood 124(5):803–811
Tepakhan W, Yamsri S, Sanchaisuriya K, Fucharoen G, Xu X, Fucharoen S (2016) Nine known and five novel mutations in the erythroid transcription factor KLF1 gene and phenotypic expression of fetal hemoglobin in hemoglobin E disorder. Blood Cells Mol Dis 59:85–91
Khamphikham P, Sripichai O, Munkongdee T, Fucharoen S, Tongsima S, Smith DR (2018) Genetic variation of Krüppel-like factor 1 (KLF1) and fetal hemoglobin (HbF) levels in β0-thalassemia/HbE disease. Int J Hematol 107(3):297–310
Ngo D, Bae H, Steinberg MH, Sebastiani P, Solovieff N, Baldwin CT, Melista E, Safaya S, Farrer LA, al-Suliman AM, Albuali WH, al Bagshi MH, Naserullah Z, Akinsheye I, Gallagher P, Luo HY, Chui DHK, Farrell JJ, al-Ali AK, Alsultan A (2013) Fetal hemoglobin in sickle cell anemia: genetic studies of Arab-Indian haplotype. Blood Cells Mol Dis 51(1):22–26
Bae HT, Baldwin CT, Sebastiani P, Telen MJ, Ashley-Koch A, Garrett M, Hooper WC, Bean CJ, DeBaun MR, Arking DE, Bhatnagar P, Casella JF, Keefer JR, Barron-Casella E, Gordeuk V, Kato GJ, Minniti C, Taylor J, Campbell A, Luchtman-Jones L, Hoppe C, Gladwin MT, Zhang Y, Steinberg MH (2012) Meta-analysis of 2040 sickle cell anemia patients: BCL1A, and HBS1L-MYB are the major modifiers of Hb F in African Americans. Blood 120(9):1961–1962
Liu L, Pertsemlidis A, Ding LH, Story MD, Steinberg MH (2016) A case-control genome-wide association study identifies genetic modifiers of fetal hemoglobin in sickle cell disease. Exp Biol Med 241(7):706–718
Mtatiro SK, Singh T, Rooks H, Mgaya J, Mariki H, Soka D et al (2014) Genome wide association study of fetal hemoglobin in sickle cell anemia in Tanzania. PLoS One 9(11):e111464
Gallienne AE, Dréau HM, Schuh A, Old JM, Henderson S (2012) Ten novel mutations in the erythroid transcription factor klf1 gene associated with increased fetal hemoglobin levels in adults. Haematologica 97(3):340–343
Orkin SH, Kazazian HH, Antonarakis SE, Goff SC, Boehm CD, Sexton JP, Waber PG, Giardina PJV (1982) Linkage of β-thalassaemia mutations and β-globin gene polymorphisms with DNA polymorphism in human β-globin gene cluster. Nature 296(5858):627–631
Dodé C, Krishnamoorthy R, Lamb J, Rochette J (1993) Rapid analysis of α3.7 thalassaemia and αααanti3.7 triplication by enzymatic amplification analysis. Br J Haematol 83(1):105–111
Robinson JT, Thorvaldsdóttir H, Wenger AM, Zehir A, Mesirov JP (2017) Variant review with the Integrative Genomics Viewer. Cancer Res 77(21):e31–e34
Akinsheye I, Alsultan A, Solovieff N, Ngo D, Baldwin CT, Sebastiani P, Chui DHK, Steinberg MH (2011) Fetal hemoglobin in sickle cell anemia. Blood 118(1):19–27
Adeodu OO, Akinlosotu MA, Adegoke SA, Saheed BA, Oseni SBA (2017) Fetal hemoglobin and disease severity in Nigerian children with sickle cell anemia. Mediterr J Hematol Infect Dis 9(1):e2017063
Mpalampa L, Ndugwa CM, Ddungu H, Idro R (2012) Fetal hemoglobin and disease severity in sickle cell anemia patients in Kampala Uganda. BMC Blood Disord 12:11
Mouélé R, Galactéros F, Feingold J (1999) Hemoglobin F (Hb F) levels in sickle-cell anemia patients homozygous for the Bantu haplotype. Eur J Haematol 63(2):136–137
Makani J, Menzel S, Nkya S, Cox SE, Drasar E, Soka D, Komba AN, Mgaya J, Rooks H, Vasavda N, Fegan G, Newton CR, Farrall M, Lay Thein S (2011) Genetics of fetal hemoglobin in Tanzanian and British patients with sickle cell anemia. Blood 117(4):1390–1392
Tshilolo L, Summa V, Gregorj C, Kinsiama C, Bazeboso JA, Avvisati G, Labie D (2012) Foetal hemoglobin, erythrocytes containing fetal hemoglobin, and hematological features in Congolese patients with sickle cell anemia. Anemia 2012:105349
Isah IZ, Udomah FP, Erhabor O, Aghedo F, Uko EK, Okwesili AN et al (2013) Fetal hemoglobin levels in sickle cell disease (SCD) patients in Sokoto, Nigeria. Br J Med Health Sci 1(6):36–47
Adeyemo TA, Ojewunmi OO, Oyetunji IA, Rooks H, Rees DC, Akinsulie AO, Akanmu AS, Thein SL, Menzel S (2018) A survey of genetic fetal-hemoglobin modifiers in Nigerian patients with sickle cell anemia. PLoS One 13(6):e0197927
Kato GJ, Steinberg MH, Gladwin MT (2017) Intravascular hemolysis and the pathophysiology of sickle cell disease. J Clin Invest 127(3):750–760
Yu LH, Liu D, Cai R, Shang X, Zhang XH, Ma XX, Yan SH, Fang P, Zheng CG, Wei XF, Liu YH, Zhou TB, Xu XM (2015) Changes in hematological parameters in alpha-thalassemia individuals co-inherited with erythroid Krüppel-like factor mutations. Clin Genet 88(1):56–61
Satta S, Paglietti ME, Sollaino MC, Barella S, Moi P, Desogus MF, Demartis FR, Manunza L, Origa R (2017) Changes in HbA2 and Hb F in alpha thalassemia carriers with KLF1 mutation. Blood Cells Mol Dis 64:30–32
Bhatnagar P, Purvis S, Barron-Casella E, Debaun MR, Casella JF, Arking DE, Keefer JR (2011) Genome-wide association study identifies genetic variants influencing F-cell levels in sickle-cell patients. J Hum Genet 56(4):316–323
Radmilovic M, Zukic B, Petrovic MS, Bartsakoulia M, Stankovic B, Kotur N, Dokmanovic L, Georgitsi M, Patrinos GP, Pavlovic S (2013) Functional analysis of a novel KLF1 gene promoter variation associated with hereditary persistence of fetal hemoglobin. Ann Hematol 92(1):53–58
Gnanapragasam MN, Crispino JD, Ali AM, Weinberg R, Hoffman R, Raza A, Bieker JJ (2018) Survey and evaluation of mutations in the human klf1 transcription unit. Sci Rep 8(1):6587
Hariharan P, Colah R, Ghosh K, Nadkarni A (2018) Differential role of Kruppel like factor 1 (KLF1) gene in red blood cell disorders. Genomics pii: S0888–7543(18)30542–1
Acknowledgments
The authors wish to thank the SCA patients and their parents for their participation in this study.
Funding
This work was partially funded by Fundação para a Ciência e a Tecnologia (FCT) grant PIC/IC/83084/2007, ISAMB, and INSA project 2012DGH720. Additionally, it is a result of the GenomePT project (POCI-01-0145-FEDER-022184), supported by COMPETE 2020 - Operational Programme for Competitiveness and Internationalisation (POCI), Lisboa Portugal Regional Operational Programme (Lisboa2020), Algarve Portugal Regional Operational Programme (CRESC Algarve2020), under the PORTUGAL 2020 Partnership Agreement, through the European Regional Development Fund (ERDF), FCT.
Author information
Authors and Affiliations
Contributions
MN, SV, MS, AC, and EF performed the molecular work and analyzed results; SV and AC also collected data and populating database; JM and LV performed NGS experiments and analyzed data; PK, RM, RS, AD, TF, AM, and IMS participated in clinical enrolling/work-up of patients. JL acquired funding and performed a critical revision of the manuscript. PF conceived and designed the experiments, supervised the molecular analysis, and drafted the manuscript. All authors (except AM†) revised and approved the manuscript final version.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This study was conducted in accordance with the ethical standards of the institutional review board and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. Informed consent was obtained from all patients’ legal representatives for being included in the study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Anabela Morais: deceased co-author
Rights and permissions
About this article
Cite this article
Nicolau, M., Vargas, S., Silva, M. et al. Genetic modulators of fetal hemoglobin expression and ischemic stroke occurrence in African descendant children with sickle cell anemia. Ann Hematol 98, 2673–2681 (2019). https://doi.org/10.1007/s00277-019-03783-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00277-019-03783-y