Abstract
Purpose
To provide a holistic and complete comparison of the five most advanced AI models in the augmentation of low-dose 18F-FDG PET data over the entire dose reduction spectrum.
Methods
In this multicenter study, five AI models were investigated for restoring low-count whole-body PET/MRI, covering convolutional benchmarks — U-Net, enhanced deep super-resolution network (EDSR), generative adversarial network (GAN) — and the most cutting-edge image reconstruction transformer models in computer vision to date — Swin transformer image restoration network (SwinIR) and EDSR-ViT (vision transformer). The models were evaluated against six groups of count levels representing the simulated 75%, 50%, 25%, 12.5%, 6.25%, and 1% (extremely ultra-low-count) of the clinical standard 3 MBq/kg 18F-FDG dose. The comparisons were performed upon two independent cohorts — (1) a primary cohort from Stanford University and (2) a cross-continental external validation cohort from Tübingen University — in order to ensure the findings are generalizable. A total of 476 original count and simulated low-count whole-body PET/MRI scans were incorporated into this analysis.
Results
For low-count PET restoration on the primary cohort, the mean structural similarity index (SSIM) scores for dose 6.25% were 0.898 (95% CI, 0.887–0.910) for EDSR, 0.893 (0.881–0.905) for EDSR-ViT, 0.873 (0.859–0.887) for GAN, 0.885 (0.873–0.898) for U-Net, and 0.910 (0.900–0.920) for SwinIR. In continuation, SwinIR and U-Net’s performances were also discreetly evaluated at each simulated radiotracer dose levels. Using the primary Stanford cohort, the mean diagnostic image quality (DIQ; 5-point Likert scale) scores of SwinIR restoration were 5 (SD, 0) for dose 75%, 4.50 (0.535) for dose 50%, 3.75 (0.463) for dose 25%, 3.25 (0.463) for dose 12.5%, 4 (0.926) for dose 6.25%, and 2.5 (0.534) for dose 1%.
Conclusion
Compared to low-count PET images, with near-to or nondiagnostic images at higher dose reduction levels (up to 6.25%), both SwinIR and U-Net significantly improve the diagnostic quality of PET images. A radiotracer dose reduction to 1% of the current clinical standard radiotracer dose is out of scope for current AI techniques.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The use of artificial intelligence (AI) technology for medical image restoration has accelerated rapidly in the past decade. AI-powered deep learning neural networks are increasingly being used to augment low-count medical images, such as those acquired by positron emission tomography (PET) [1]. PET has been considered as the gold standard for staging and treatment monitoring of patients with solid cancers [2,3,4]. However, the disadvantages of PET imaging as compared to magnetic resonance imaging (MRI) are its high cost and ionizing radiation exposure [5,6,7]. Reductions in radiotracer dosage could minimize radiation exposure, and reductions in scan time could enhance patient throughput and reduce scan costs. However, reductions in radiotracer dosage and scan times lower the detection of PET annihilation events, resulting in low-count PET scans with reduced diagnostic image quality (DIQ) [8]. Based on a comprehensive literature review, the restoration of a standard full-count PET imaging from this reduced DIQ cannot be achieved by simple postprocessing operations such as denoising, since lowering the number of coincidence events in the PET detector introduces both noise and local uptake value changes [9]. Hence, sophisticated AI-powered deep learning techniques for image restoration became increasingly more popularized to facilitate PET image restoration [10,11,12].
Multiple AI models have emerged in recent years to enhance low-count PET scans [13,14,15], with some convolutional neural network (CNN) methods approved by the U.S. Food and Drug Administration (FDA) [16]. However, the FDA does not recommend which specific FDA-approved software should be used for a given medical problem. Most available AI-powered PET restoration publications feature a single AI model. As such, the literature currently lacks an unbiased, systematic evaluation comparing multiple state-of-the-art AI models in this context. Moreover, the rapid rate of progress in AI and deep learning research has given way to transformer-based models with innate global self-attention mechanisms capable of outperforming CNN-based benchmarks in a variety of imaging-related tasks including image reconstruction [17,18,19,20]. To our knowledge, transformers have not yet been well-adapted and utilized for whole-body PET restoration, nor have they been directly compared against the state-of-the-art CNNs. Thus, we herein seek to fulfill an unmet need by performing a comprehensive comparison of state-of-the-art AI models for low-count whole-body (WB) PET imaging restoration.
Reducing the 18F-FDG dose increases image artifacts, because the image quality is proportional to the number of coincidence events in the PET detector following radiopharmaceutical positron annihilation [1]. Such significant artifacts and noise introduce challenges for the recovery of true radiotracer signal by AI models. Three recent studies have explored AI-based augmentation in WB PET images at 50% [16, 21], 25% [1], and 6.25% [22] of the clinical standard doses. To date, few efforts have been reported on conducting a comprehensive assessment across the dosage reduction spectrum [16]. There is also a lack of PET databases containing list-mode data that can be used to generate a wide array of dose-reduced images for direct comparison [23]. A key question that has not yet been addressed in low-count PET image augmentation is that of model limitation (i.e., what is the lowest reduction percentage that a given AI model can enhance with acceptable clinical utility).
To close the gaps on the aforementioned challenges, our study aimed to compare five different AI models in the augmentation of low-dose 18F-FDG PET data. Using two cross-continental independent PET/MRI datasets, we examined six PET dose level percentages ranging from 75 to 1% against the five most advanced models — spanning the CNN and transformer categories. The five models include three CNN benchmarks: U-Net [24], enhanced deep super-resolution network EDSR [22], generative adversarial network (GAN) [25], and two transformer models: SwinIR [17] and EDSR-ViT [18]. Notably, the recent advancement — Swin transformer — was leveraged for whole-body PET restoration for the first time in this study.
To integrate these AI-powered low-count PET restorations in a clinical setting, a comprehensive investigation is critical. Hence, we considered different anatomical regions for the training of our model, which has been underexplored in previous studies. This study is pertinent for implementers developing AI models optimized for achieving PET imaging that preserves the best image quality with the lowest possible radiation exposure to patients. To promote the continued advancement of this domain, we have open-sourced the code underpinning the five AI models tailored for PET/MRI restoration.
Materials and methods
Participants and dose reduction spectrum
In this multicenter, restrospective evaluation of data from the Health Insurance Portability and Accountability (HIPAA)-compliant clinical trials, two participating centers (University of Tübingen, Germany, and Stanford University, CA, USA) obtained approval from their institutional review board (IRB). Written informed consent was obtained from all adult patients and parents of pediatric patients. Stanford cohort: Between July 2015 and June 2019, we collected 48 whole-body PET/MRI scans (Supplementary pp 1–2) from 22 children and young adults (13 females, 9 males) with lymphoma and a mean age (standard deviation; range) of 17 years (7; 6–30 years). Tumor histology consisted of 14 patients with Hodgkin lymphoma, six with non-Hodgkin lymphoma, and two patients with posttransplant lymphoproliferative disorder (PTLD). Tübingen cohort: 20 whole-body PET/MRI scans (Supplementary pp 1–2) from 10 patients (5 females, 5 males) with a mean age (standard deviation; range) of 14 years (5; 3–18 years) were collected. The distribution of tumor histologies was eight with Hodgkin lymphoma and two with non-Hodgkin lymphoma.
Radiotracer input data were used to generate images. Full-dose (3 MBq/kg) PET data were acquired in list mode, which helps detect coincidence events across the entire duration of the PET bed time (3 min 30 s). Low-dose PET images were retrospectively simulated by unlisting the PET list-mode data and reconstructing them based on the percentage of coincidence events [26]. List-mode PET input data were collected over time periods: the first block of 3 min 30 s, 2 min 38 s, 1 min 45 s, 53 s, 26 s, 13 s, and 2 s. These were used to simulate 100%, 75%, 50%, 25%, 12.5%, 6.25%, and 1% 18F-FDG PET dose levels, respectively. This resulted in 476 original count standard-dose and simulated low-count PET/MRI images (336 from the Stanford cohort and 140 from the Tübingen cohort) included in this study.
Study design
Five different AI models were trained and tested separately over six dose reduction percentages ranging from 75 to 1% (of the clinical standard-dose) on the primary Stanford PET/MRI images. This resulted in 30 AI models in total. All of the 30 AI models were further tested on the Tübingen external validation cohort. The Tübingen cohort was not included in the training of each model, making it a true external test set. The same image pre-processing steps (Supplementary p 2) were applied to all PET/MRI images from each cohort. Using an approach which aimed to alleviate additional burden on the network learning methods to find patterns between images for final restoration, the top 0.1% of the pixels in PET images were clipped, i.e., the intensity values of the top 0.1% pixels were mapped to the intensity of the top 0.1% pixel. This operation was critical for model convergence and training stability, as these pixels possessed high noise and were therefore outliers of the distribution.
The 3D whole-body volume was predicted in a slice-by-slice fashion and the predicted 2D slices were stacked together to reconstruct the final 3D PET prediction. We adopted 2.5D input scheme to ensure vertical spatial consistency. Five consecutive axial slices from both PET and MRI modalities were fed into the model as combined inputs, resulting in ten input slices in total for one evaluation. Fivefold cross-validation was applied to ensure generalization in model performance. A combination of mean square error (MSE) and the structural similarity index measure (SSIM) loss was used to train the model (Supplementary pp 2–3).
Five AI models evaluated
The framework illustrating the five AI models in low-count PET restoration is shown in Fig. 1. We investigated three CNN benchmarks (U-Net, EDSR, and GAN) and two transformer models (EDSR-ViT and SwinIR). Below, we detailed the models and their advantages.
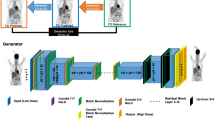
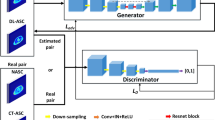
Schematic overviews of AI model frameworks for low-count PET reconstruction. a The classic U-Net model. b The adapted EDSR (enhanced deep super-resolution network) model. c The GAN (generative adversarial network) model. d The EDSR-ViT model. EDSR-ViT takes the feature encoder part from the adapted EDSR (b) directly, and makes use of the ViT (visual transformer) block to obtain global self-attention within the image. e The SwinIR model, consisting of Swin transformer blocks. The main difference of Swin transformer and ViT transformer is where the self-attention operation applies. For Swin transformer block, the self-attention is applied within each of the local windows, including the regular window partitions (Layer l) and the following shifted windows (Layer l + 1, etc.). For ViT, the self-attention is applied within the global image, which is equally partitioned into fixed size patches
U-Net
Proposed in 2015 [24], the U-Net was first invented for biomedical image segmentation and has rapidly become the most well-recognized and classic AI model in the medical imaging community. Previous studies [1, 16] have utilized U-Net in low-count PET restoration. The name “U-Net” borrows intuitively from the U-shaped structure of the model diagram, as shown in Fig. 1a. It consists of (1) the left side encoder, where convolution layers intercalate with max-pooling layers that gradually reduce the dimensions of the image, and (2) the right side decoder, where a set of convolution operations and upscaling brings the feature map back to the original dimensions. This architecture is well-suited for middle-level segmentation tasks, as the semantic information extracted from the encoder, along with the spatial information kept from the skip connection and decoder, provides almost everything needed for semantic segmentation in biomedical images. The limitation of the classic U-Net model is that it cannot sufficiently model the global and long-range semantic information due to the restriction of localized convolutional operations [27, 28].
EDSR
Investigated on 6.25%-low-count PET/MRI restoration in 2021 [22], the adapted EDSR is inspired by the classic enhanced deep super-resolution network [29] model in computer vision. The main innovation of EDSR is the organization and optimization of the building block, with only two convolutions, a rectified linear unit (ReLU) activation in between, and an add residual — as shown in Fig. 1b. The unnecessary modules — batch normalization and follow-up ReLU activation — in conventional residual networks, ResNet [30] and SRResNet [31], are removed.
GAN
First proposed in 2014 [32] and now widely used in image generation, GANs originated from the notion of having two neural networks, a generator and a discriminator, pitted against one other as adversaries in order to generate new, synthetic instances of data that can pass for real data (Fig. 1c); in short, the generator’s goal is to fool the system by trying to produce images that the discriminator cannot distinguish from real-world ones [33]. Several studies [25, 34] have explored GANs in PET restoration. However, most of the superior performance has been achieved by introducing additional clinical data — e.g., amyloid status within the brain [25] — which are not always available in real practice.
EDSR-ViT
Originally designed for sequence-to-sequence prediction in natural language processing (NLP) [35], transformer applications had been expanded to image processing very recently and soon became a game-changing technique in computer vision [36]. As opposed to FCN, where the receptive fields are gradually expanded through a series of convolution operations, the self-attention operations inherited in transformers allow full coverage of the entire input space at the beginning, demonstrating exceptional representation power. Vision transformer (ViT) — a transformer adapted for image processing — has shown impressive performance on high-level vision tasks [37, 38], but few efforts have been made to explore its role in image restoration. In order to examine its performance on PET/MRI restoration, we tailored the original ViT by adding an EDSR CNN encoder on top of the transformer block, as shown in Fig. 1d. The rationale for this is that the global long-range dependency from ViT and the precise localization from CNN encoder are complimentary for low-level vision tasks [39].
SwinIR
Proposed in 2021 [17], SwinIR is among the pioneering efforts in transformer utilization for image restoration, showing surperior performance over a variety of state-of-the-art methods spanning image super-resolution, image denoising, and JPEG compression artifact reduction. The highlight of SwinIR is the adoption of Swin transformer [19]. Swin transformer is a hierarchical transformer whose representation is computed with shifted windows, reducing the border artifacts in ViT — as ViT usually divides the input image into patches with fixed size (e.g., \(48*48\)). This brings greater efficiency by limiting self-attention computation to these local shifted windows and allowing cross-window connection to capture global dependency (Fig. 1e). According to a recent study [19], Swin transformer outperformed ViT in high-level tasks including image classification, object detection, and semantic segmentation. In this study, we adopted the backbone of SwinIR [17], which consists of 24 Swin transformer blocks for PET restoration.
Evaluation framework
We adopted three quantitative metrics to measure the quality of the restored PET images: SSIM (the structural similarity index), PSNR (peak signal-to-noise ratio), and VIF (visual information fidelity). SSIM is the most widely used metric in radiology imaging reconstruction [40] (which are a combination of luminance, contrast, and structural comparison functions). Specifically, the SSIM score was derived by comparing the AI-restored PET to the original standard-dose PET sequences and quantifying similarity on a scale of 0 (no similarity) to 1 (perfect similarity). PSNR is most commonly used to measure the reconstruction quality of a lossy transformation [41]. The higher the PSNR, the better the degraded image has been restored to match the original image. SSIM and PSNR mainly focus on pixel-wise similarity; thus, we introduce VIF, which uses natural statistics models to evaluate psychovisual features of the human visual system [42]. The code for calculating the performance was written with Python using SciPy and Scikit-image toolkits (script; Supplementary p 3).
Two board-certified radiologists (with 6 and 10 years of experience) independently reviewed the standard-dose PET, low-count PET, and the AI-restored PET by SwinIR and U-Net (SwinIR and U-Net were chosen to represent the transformer and CNN family, respectively) across the whole dose reduction spectrum (75%, 50%, 25%, 12.5%, 6.25%, and 1%). The subjects were anonymized and the order of the scans were randomized. The DIQ was assessed with 5-point Likert scale. The 5-point Likert scale for DIQ was 1. Nondiagnostic, 2. Poor, 3. Acceptable, 4. Good, and 5. Excellent image quality. In addition, the radiologists rated the lesions in 8 anatomical regions including CNS (central nervous system), paraspinal, neck, arms, chest, abdomen, pelvis, and legs, and determined the false-positive and false-negative errors using the standard-dose PET as a reference for the evaluation.
To investigate the utility of AI-restored PET scans in providing quantitative measures of tumor metabolism required for clinical PET interpretations, we measured standardized uptake values (SUVs) for the tumors and used liver as an internal reference standard. SUVs are the most widely used metric in clinical oncologic imaging and play a germane role in assessing tumor glucose metabolism on FDG-PET [43, 44]. The SUVmax of target lesions and SUVmax of liver were measured by placing separate three-dimensional volumes of interest over tumor lesions and the liver. SUVs were measured using OsiriX version 12.5.1. (OsiriX software; Supplementary p 3). SUV values were calculated based on patient body weight and injected dose by using the equation in Supplementary p 3.
Statistical analysis
We used Wilcoxon signed-rank t test as implemented in R software (V4.0.3) to assess the significance of the difference between two models. The ratings of two radiologists were used to test the difference between the AI-restored PET images and the corresponding original low-count PET images using Wilcoxon signed-rank tests. We used a predefined P < 0.01 for significance. The performance tables show the mean, standard deviation (SD), and the first (25%) and third (75%) quartiles of the data. The evaluation metrics are provided with two-sided 95% confidence intervals (CIs). All models were written in Python3, with model training and testing performed using the Pytorch package (version 1.10).
Results
Both baseline and follow-up WB PET/MRI scans of 32 children and young adult lymphoma patients were collected and six dose levels (75%, 50%, 25%, 12.5%, 6.25%, and 1%) were simulated, resulting in 476 PET/MRI scans (336 from the primary Stanford cohort and 140 from the Tübingen external cohort). The cross-continental PET/MRI cohorts were used to examine the generalization of our findings. To the best of our knowledge, large pooled PET/MRI databases containing PET list-mode data amenable to simulate low-dose PET for AI model evaluation do not exist. As such, our collected cohort is unique in that it is among the first PET/MRI databases for AI-enabled dose reduction studies.
Five AI models on six dose reduction percentages
To provide a holistic comparison of the five AI models, all models were evaluated in the restoration of low-count whole-body PET images at six reduction percentages (75%, 50%, 25%, 12.5%, 6.25%, and 1% of the clinical standard 3 MBq/kg 18F-FDG dose). The quantitative performance metrics of all AI models over the entire dose reduction spectrum are shown in Fig. 3. Model comparisons at doses 25% and 12.5% revealed that SSIM scores were highest for SwinIR on the Stanford internal test set. At dose 6.25%, SSIM scores were highest for SwinIR and lowest for U-Net. Differences in SSIM score became apparent between models at dose 6.25%, ranging from 0.883 (U-Net) to 0.914 (SwinIR). At dose 1%, SSIM scores were highest for SwinIR and U-Net, and lowest for EDSR and EDSR-ViT. Differences in SSIM score between models were the least appreciable at dose 1%, ranging from 0.842 (SwinIR and U-Net) to 0.848 (EDSR and EDSR-ViT). For the Tübingen cohort, SwinIR also achieved the best performance in the SSIM metric with doses below 50% (Fig. 3). More detailed performance metrics for 6.25% low-count PET restoration are shown in Table 1. Dose 6.25% was the lowest dosage with around 40 dB PSNR for the AI restoration and thereby became our dose of choice for further investigation. The systematic evaluation presented herein is rendered in summary form, with mean and median quantitative values over the fourfold cross-validations on the two cohorts of interest (Table 1). SwinIR achieved the best quantitative results, with the highest SSIM score of 0.910 (95% CI 0.900–0.920), PSNR score of 39.9 (39.1–40.6), and VIF score of 0.485 (0.469–0.501) on the primary Stanford test set. It was also generalized to the external Tübingen test set with the highest SSIM score of 0.950 (0.942–0.958) and VIF score of 0.483 (0.464–0.502), demonstrating model generalization across different institutions and scanner types.
The qualitative comparisons between the five AI models on 6.25% low-count restoration are shown in Fig. 2. The PET images restored from the SwinIR model were superior in reflecting some of the underlying anatomic patterns of the tracer uptake (the basal ganglia; Fig. 2A) when compared to the images generated from the other four models. Meanwhile, though lesions could be detected on all AI-restored scans (Fig. 2B-D), lesion-to-background contrast and confidence for lesion detection were improved on SwinIR (Fig. 2C). Compared to the standard-dose 18F-FDG PET scans, the simulated 6.25% low-count PET images had significantly higher SUVmax values of the liver as a result of increased image noise. All five AI models managed to recover SUVmax values of the liver similar to the values in standard-dose PET, demonstrating good denoising capability (Supplementary Table 1). All tumors had SUV values above that of the liver on all AI-restored PET images. Table 2 provides the comparison of five models in low-count PET restoration.
PET image comparison across five state-of-the-art AI models on 6.25% low-count PET reconstruction. A Representative 18F-FDG PET scan of a 29-year-old female patient with Hodgkin lymphoma (HL). The enlarged patches are shown on the second panel (yellow arrows: basal ganglia). The structural similarity index (SSIM) and visual information fidelity (VIF) metrics are presented under each PET image. B Representative 18F-FDG PET scan of a 14-year-old male patient with HL. The SUVmax of the lesion (delineated by red circle) and liver for this patient are shown under each PET image. C The same patient as B. The small lesion (less than 1.5 cm3; 5 mm < width < 10 mm; height > 10 mm; red arrow) is enhanced by SwinIR with the lesion-to-liver contrast of SUVmax retained. The lesions (black arrow) are also clearly depicted by SwinIR, in contrast with being blurred and mixed together by the other reconstructions. D Representative 18F-FDG PET scan of a 17-year-old female patient from the external Tübingen testing cohort. All AI models successfully denoise the 6.25% low-count images and provide similar diagnostic conspicuity of the lesion (red circle; red arrows) as the standard-dose PET, demonstrating the model is generalizable across different institutions for all AI models. SwinIR shows superiority in retaining lesion-to-liver contrast and structural fidelity
Restoration across the dose reduction spectrum
Next, we examine the AI-powered PET restoration through the lens of dose reduction spectrum. AI-restored PET images consistently achieved improved SSIM, VIF, and PSNR over original low-count PET images at dose 25%, 12.5%, 6.25%, and 1% (Fig. 3). Among the six dose reduction percentages, the improvement from AI restoration was largest at dose 6.25%. The average improvement scores for the five AI models were 0.106 (95% CI 0.102–0.110) in SSIM, 3.97 dB (3.78–4.16) in PSNR, and 0.183 (0.178–0.188) in VIF on Stanford internal test cohort; 0.211 (0.208–0.215) in SSIM, 3.54 dB (3.20–3.88) in PSNR, and 0.196 (0.190–0.202) in VIF on Tübingen external test set. Pair-wise t-tests between the AI-restored PET images and the low-count PET images revealed p-values consistently less than 0.001, suggesting that all AI models possessed statistically significant capacities for restoration and generalization. Figure 4 provides the detailed qualitive PET image comparisons between different dosages. With reduction in simulated radiotracer dose, PET images exhibited higher noise and information loss, leading to increased SUVmax values in the liver and tumors (Fig. 4B). The AI models tested herein reduced artifacts for the low-count PET images and recovered the SUVmax values of liver and tumors to values commensurate with those derived from standard-dose PET (Fig. 4C, D).
Quantitative metrics over the dose reduction spectrum. The five AI models were adapted for the low-count PET reconstruction task. The AI models were trained on 75%, 50%, 25%, 12.5%, 6.25%, and 1% of the clinical standard 18F-FDG dose PET/MRI images from the primary Stanford cohort. One round of cross-validation was adopted. The trained models were then evaluated on the corresponding low-count PET/MRI test set. The performance on the Stanford internal test set is shown on the top panel, and the performance on the external Tubingen test cohort is shown on the bottom panel. Measures of performance include structural similarity index (SSIM), peak signal-to-noise ratio (PSNR), and visual information fidelity (VIF). For all three metrics, higher represents better reconstruction. All comparisons are made against the ground-truth standard-count PET images. The blue line presents the original low-count PET images without AI enhancement and serves as the baseline for direct comparisons
PET image comparisons across the dose reduction spectrum from 75 to 1% (of the clinical standard 3 MBq/kg 18F-FDG dose). Representative 18F-FDG PET scan of 13-year-old male patient with diffuse large B cell lymphoma (DLBCL). The SUVmax of two tumors and liver were measured for each PET image. SwinIR and U-Net are our demonstration models of choice, representing the transformer and CNN categories, respectively. A The coronal slice of the standard-dose PET, showing the chest region. SUVmax of two tumors and liver were measured for direct comparison. B The original low-count PET images with SUVmax measured under the same regions of tumors and liver as in A. C U-Net restored low-count PET images. The red arrows point to corrupted reconstruction in mediastinum and erroneous upstaging in liver. Red rectangle: enlargement of false upstaging in the liver area. U-Net-75p = U-Net restored 75% low-count PET image. D SwinIR restored low-count PET images. The red arrows point to the erroneous upstaging. Red rectangle: enlargement of the degraded reconstruction in liver. SwinIR-75p, SwinIR restored 75% low-count PET image
For doses 75% and 50%, there were discrepancies between quantitative metrics and visual appearances. All AI models have enhanced the 75% and 50% low-count PETs visually with reduced image noise (Fig. 4), but the improvements were not reflected quantitatively (Fig. 3). A possible explanation is that 75% and 50% low-count PET images are sufficiently similar to standard-dose PET. Their PSNR values are greater than the threshold — 40 dB — which corresponds to nearly undiscernable differences, and thus passes the considerations for good image quality [45, 46]. Therefore, the quantitative metrics might not be able to reasonably depict improvements above this threshold.
In general, the quantative metrics — SSIM, VIF, and PSNR — of both original low-count PET and AI-restored PET images decreased over the dose reduction spectrum. However, AI restorations (powered by SwinIR, EDSR, and EDSR-ViT) between doses 12.5% and 6.25% achieved similar performance in the three metrics (Fig. 3). This is partly owing to the smoothing effect of 6.25% low-count restoration (the liver area in SwinIR-6.25p; Fig. 4D). The AI models in 6.25% low-count restoration converged on an approach that smoothed particular regions with significantly decreased noise.
From doses ranging from 6.25 to 1%, there was a steep drop (Fig. 3) in SSIM, PSNR, and VIF across both Stanford and Tübingen cohorts, indicating the challenge of extreme-low-count PET restoration. Indeed, AI restoration introduced hallucinated signals and erroneous upstaging in 1% low-count PET restoration (Fig. 4C, D; far right column). The extreme-low-count scenario degraded PET images with substantial artifacts and information loss that were difficult for the current AI techniques to handle without the incorporation of additional information. Supplementary Fig. 2 shows the whole-body PET restoration from the coronal view, across the dose reduction spectrum powered by SwinIR.
Model training strategy
Figure 5 demonstrates an interesting observation when training SwinIR in 6.25% low-count PET images. In epoch 24, the trained model was able to reconstruct the shape and contrast of the basal ganglia in the brain, but failed to clearly depict a small lesion (less than 1.5 cm3) in the liver. Meanwhile in epoch 4, the brain structure was not well-restored, but the diagnostic conspicuity of the small lesion was preserved. Our experiment suggested that the discrepancies in restoration quality between different anatomical regions were agnostic to specific model architectures. The possible reasons may be twofold: (1) the commonly used loss functions — mean square loss (MSE loss) and the structural similarity index loss (SSIM loss) — were originally proposed for natural image reconstruction and not specifically designed for diagnostic radiology images, thus limiting their ability to guide model training for these specific clinical needs; and (2) whole-body PET images have large intra-patient uptake variation. The metabolic activities of the brain and bladder are greater than other anatomical locations, shown as hyperintensities in PET images. As the training progresses, the focus of model optimization can shift to these hyperintense regions easily, as they can possess larger absolute loss penality values; this can in turn cause over-smoothing of other relatively low-intensity regions (e.g., the liver).
Representative discrepancy in reconstruction quality between different anatomical regions over the course of model training. SwinIR is the model of choice for this demonstration. The performance is based on the primary Stanford PET/MRI cohort. The line chart shows the SSIM metric of the Stanford validation set over models at different training epochs. PET images illustrate cases from the Stanford testing set. The patches (top panel) are enlarged crops of a, b, and c, respectively. As the training progresses from epoch 4 to epoch 24, the structure of the basal ganglia within the brain becomes better restored, while the small lesion (less than 1 cm3) within the liver gets over-smoothed
Clinical diagnostic evaluation
Compared to the original low-count PET images, with near-to or nondiagnostic images at higher dose reduction levels, both SwinIR and U-Net can significantly improve the diagnostic quality of PET images (Table 3; Supplementary Fig. 3). From the radiologists’ assessment, it became apparent that both SwinIR and U-Net were able to significantly reduce the number of false-negative/false-positive lesions compared to the original low-count PET images (the overall rate of false negatives/false positives in Table 4).
When comparing SwinIR and U-Net, there was a total of 3 false-positive and 1 false-negative lesions for SwinIR and 3 false-positive lesions for U-Net, indicating a similar diagnostic performance for both models. There were more false-positive lesions for the U-Net architecture at 50% and 25% (3 false positives for U-Net compared to 1 false positive for SwinIR), while there were more false positives for SwinIR at 12.5% and 6.25% (2 false positives versus 0 false positives). Even though the 6.25% simulation shows a notable increase in image quality with smoothed organ borders for SwinIR with minimal noise (Fig. 4), some original image information is lost through the strong smoothing effect, which is underscored by the presence of a false-positive lesion in contrast to the U-Net. The occurrence of this false-positive finding may have been favored by the fact that SwinIR image quality at the 6.25% level appears visually high with minimal noise, which carries a certain risk of false diagnostic confidence.
Overall, there was no noticeable difference in diagnostic performance between SwinIR and U-Net. For both architectures, some image information was lost at higher dose reduction levels, e.g., loss of myocardial uptake or decreased definition of the spine or the ribs (Fig. 4; Supplementary Fig. 2). While diagnostic image quality (DIQ) was slightly higher for SwinIR compared to U-Net, especially at 6.25%, this did not lead to significant differences in lesion detection rates. The assessment of the radiologists adds to the information from SSIM, PSNR, and VIF metrics, confirming that both SwinIR and U-Net bring a significant gain in diagnostic image quality compared to the original low-count PET, with comparable improvement of lesion detection (false negatives/positives). Including the information from SSIM, PSNR, and VIF metrics in the assessment (Table 1), SwinIR might possibly have a slight advantage over U-Net, even though this did not result in a significant difference in diagnostic performance for specialist-based lesion assessment.
Discussion
In this study, we provide the first unbiased and comprehensive investigation of AI-enabled low-count whole-body PET restoration from two perspectives: the restoration models and the dose reduction percentages. Six reduction percentages covering the entire dose spectrum — 75%, 50%, 25%, 12.5%, 6.25%, and 1% (extreme low count) of the clinical standard 18F-FDG dose — were investigated. In addition, we adapted five state-of-the-art AI models for this task, including the classic CNN benchmarks and the most advanced transformer models. Two cross-continental PET/MRI cohorts were used to examine the generalization of our findings.
All five AI models possess PET restoration capability. From the algorithmic perspective, the advantage of SwinIR in low-count PET/MRI restoration is that no pre-training is needed according to our experiments; the transformer approach complemented the conventional CNN approaches in that the innate global self-attention mechanism provided long-range dependency that is otherwise lacking in CNNs due to the limited receptive field of convolution operations. The Swin transformer model (SwinIR) with its shifted window mechanism further improved the depiction of structural details and small lesions that could be missed if the fixed partition operations of ViT transformer alone were used. A major drawback, however, is the large number of operations required in SwinIR — resulting in training and testing times that were 10 × longer compared to other state-of-the-art models. A few studies to date applying transformers on PET restoration are mainly focused on low-count brain images [47, 48]. This is one of the first studies utilizing Swin transformer for whole-body PET restoration.
While AI deep learning architectures are essential in low-count PET restoration, equally important is the model training strategy, i.e., the procedure used to carry out the learning process; this includes specific considerations such as the loss function and when to stop training. To date, few efforts have been made to reconcile these considerations. We made an oberservation about the discrepancy in restoration quality among different anatomical regions on PET restoration over the course of training. This observation underscores the role of training strategy in building up the optimal model for low-count whole-body PET restoration. Our findings suggest that engaging radiologists in the model development loop is imperative so that the PET restoration training process can be effectively and efficiently guided by domain experts in a task-specific fashion. Another possible direction is region-based restoration that takes the regional difference priors into consideration for effectively designing WB PET restoration models. Future work is needed on optimizing PET restoration training process that can be effectively guided in a clinic-task-specific fashion.
Another key contribution of this study is the examination of AI-powered PET restoration over six groups of count levels, representing 75%, 50%, 25%, 12.5%, and 6.25%, to extremely ultra-low-count 1% (of the clinical standard 3 MBq/kg 18F-FDG dose). In order to perform a holistic assessment of low-count PET restoration, we adapted multiple AI models upon the complete dose reduction spectrum. The most relevant work to our study, published in 2021 [16], evaluated the FDA-approved U-Net software across various dosages. This commercially available software was trained only on 25% low-count PET images and was tested at other percentages. In contrast, our study takes the approach of training and testing images in a manner consistent with the relevant reduced dosage. To our best knowledge, this study is the first complete investigation of AI-powered whole-body PET restoration over the entire dose reducing spectrum. Note that the lowest possible reduction in radiotracer dose is dependent on many factors, such as tumor type (e.g., pediatric lymphomas and sarcomas typically present with strong radiotracer uptake at the time of the diagnosis), the timing of the scan with regard to the treatment schedule (baseline images typically show strong 18F-FDG tumor uptake, follow-up scans after chemotherapy demonstrate markedly reduced 18F-FDG tumor signal if the tumor responds to therapy), the sensitivity of the PET detector, and the duration of the PET data acquisition time. The relation between image quality and dose is not binary, but continuous. In this study, the most cutting-edge AI models enabled low-count PET restoration of doses above 6.25% with acceptable DIQ, which is consistent with another recent study on 6.25% low-count PET restoration [49], while dose 1% without additional clincal information was out of scope for the AI techniques evaluated herein.
This study has the following limitations. Simulated low-dose PET images were used instead of injecting multiple different PET tracer doses in a single patient, considering ethically feasiblity. Though previous data have shown that simulated low-dose images have characteristics similar to those of actual low-dose images [50], evidence of AI restoration in true injected low-dose cases is needed. In addition, this study only included patients scanned with FDG, due to its clinical prevalency. The use of the deep-learning approaches to reconstruct images obtained with non-FDG radiotracers may entail different performances dependent upon signal-to-noise ratios, and the uptake dynamics and locations. The model generalizability across different diseases and a wide range of patients should be investigated in future work.
In conclusion, the findings from this study hold important implications for implementers developing the optimal AI model in order to achieve PET imaging with the lowest radiation exposure to patients and non-inferior DIQ. Mitigation of ionizing radiation exposure from medical imaging procedures holds critically important potential for clinical impact, as reducing such exposure could minimize the potential risk of secondary cancer development later in life [5, 51, 52]. This is especially important for pediatric patients or patients receiving therapies that require repeat imaging with reoccurring radiation exposure. Toward further advancement of this domain, we open-sourced the five AI models specifically tailored for low-count PET/MRI restoration. Of note, our code may easily be applied to other medical imaging modalities (e.g., MRI and CT) and could thereby potentially serve as a common foundation for medical image restoration.
Data Availability
The data from this study are not publicly available in accordance to institutional requirements governing human subject privacy considerations. The data may be made available from the authors upon reasonable request.
References
Chaudhari AS, Mittra E, Davidzon GA, Gulaka P, Gandhi H, Brown A, et al. Low-count whole-body PET with deep learning in a multicenter and externally validated study. NPJ Digit Med. 2021;4:1–11.
Baum SH, Frühwald M, Rahbar K, Wessling J, Schober O, Weckesser M. Contribution of PET/CT to prediction of outcome in children and young adults with rhabdomyosarcoma. J Nucl Med. 2011;52:1535–40.
Kleis M, Daldrup-Link H, Matthay K, Goldsby R, Lu Y, Schuster T, et al. Diagnostic value of PET/CT for the staging and restaging of pediatric tumors. Eur J Nucl Med Mol Imaging. 2009;36:23–36.
Baratto L, Hawk KE, Qi J, Gatidis S, Kiru L, Daldrup-Link HE. PET/MRI improves management of children with cancer. J Nucl Med. 2021;62:1334–40.
Huang B, Law MW-M, Khong P-L. Whole-body PET/CT scanning: estimation of radiation dose and cancer risk. Radiology. 2009;251:166–74.
Meulepas JM, Ronckers CM, Smets AM, Nievelstein RA, Gradowska P, Lee C, et al. Radiation exposure from pediatric CT scans and subsequent cancer risk in the Netherlands. JNCI J Natl Cancer Institute. 2019;111:256–63.
Brenner DJ, Doll R, Goodhead DT, Hall EJ, Land CE, Little JB, et al. Cancer risks attributable to low doses of ionizing radiation: assessing what we really know. Proc Natl Acad Sci. 2003;100:13761–6.
Townsend D. Physical principles and technology of clinical PET imaging. Ann Acad Med Singap. 2004;33:133–45.
Wang T, Lei Y, Fu Y, Curran WJ, Liu T, Nye JA, et al. Machine learning in quantitative PET: a review of attenuation correction and low-count image reconstruction methods. Phys Med. 2020;76:294–306.
Wang G, Ye JC, Mueller K, Fessler JA. Image reconstruction is a new frontier of machine learning. IEEE Trans Med Imaging. 2018;37:1289–96.
Reader AJ, Corda G, Mehranian A, da Costa-Luis C, Ellis S, Schnabel JA. Deep learning for PET image reconstruction. IEEE Trans Radiat Plasma Med Sci. 2020;5:1–25.
Raj A, Bresler Y, Li B. Improving robustness of deep-learning-based image reconstruction. International Conference on Machine Learning. 2020:7932–42.
Häggström I, Schmidtlein CR, Campanella G, Fuchs TJ. DeepPET: a deep encoder–decoder network for directly solving the PET image reconstruction inverse problem. Med Image Anal. 2019;54:253–62.
Gong K, Catana C, Qi J, Li Q. PET image reconstruction using deep image prior. IEEE Trans Med Imaging. 2018;38:1655–65.
Feng Q, Liu H. Rethinking PET image reconstruction: ultra-low-dose, sinogram and deep learning. International Conference on Medical Image Computing and Computer-Assisted Intervention: Springer. 2020:783–92.
Theruvath AJ, Siedek F, Yerneni K, Muehe AM, Spunt SL, Pribnow A, et al. Validation of deep learning–based augmentation for reduced 18F-FDG dose for PET/MRI in children and young adults with lymphoma. Radiology: Artificial Intelligence. 2021;3:e200232.
Liang J, Cao J, Sun G, Zhang K, Gool LV, Timofte R. SwinIR: image restoration using Swin transformer. IEEE/CVF International Conference on Computer Vision Workshops (ICCVW). 2021:1833–44.
Dosovitskiy A, Beyer L, Kolesnikov A, Weissenborn D, Zhai X, Unterthiner T, et al. An image is worth 16x16 words: transformers for image recognition at scale. The International Conference on Learning Representations (ICLR). 2021.
Liu Z, Lin Y, Cao Y, Hu H, Wei Y, Zhang Z, et al. Swin transformer: Hierarchical vision transformer using shifted windows. Proceedings of the IEEE/CVF International Conference on Computer Vision. 2021:10012–22.
Khan S, Naseer M, Hayat M, Zamir SW, Khan FS, Shah M. Transformers in vision: A survey. ACM computing surveys (CSUR). 2022;54:1–41.
Whiteley W, Luk WK, Gregor J. DirectPET: full-size neural network PET reconstruction from sinogram data. J Med Imaging. 2020;7: 032503.
Wang Y-RJ, Baratto L, Hawk KE, Theruvath AJ, Pribnow A, Thakor AS, et al. Artificial intelligence enables whole-body positron emission tomography scans with minimal radiation exposure. Eur J Nucl Med Mol Imaging. 2021:1–11.
Schramm G, Rigie D, Vahle T, Rezaei A, Van Laere K, Shepherd T, et al. Approximating anatomically-guided PET reconstruction in image space using a convolutional neural network. Neuroimage. 2021;224: 117399.
Ronneberger O, Fischer P, Brox T. U-net: convolutional networks for biomedical image segmentation. International Conference on Medical image computing and computer-assisted intervention. 2015:234–41.
Ouyang J, Chen KT, Gong E, Pauly J, Zaharchuk G. Ultra-low-dose PET reconstruction using generative adversarial network with feature matching and task-specific perceptual loss. Med Phys. 2019;46:3555–64.
Sekine T, Delso G, Zeimpekis KG, de Galiza BF, Ter Voert EE, Huellner M, et al. Reduction of 18F-FDG dose in clinical PET/MR imaging by using silicon photomultiplier detectors. Radiology. 2018;286:249–59.
Esser P, Rombach R, Ommer B. Taming transformers for high-resolution image synthesis. Proceedings of the IEEE/CVF conference on computer vision and pattern recognition. 2021:2873–83.
Zheng S, Lu J, Zhao H, Zhu X, Luo Z, Wang Y, et al. Rethinking semantic segmentation from a sequence-to-sequence perspective with transformers. Proceedings of the IEEE/CVF conference on computer vision and pattern recognition. 2021;6881–90.
Lim B, Son S, Kim H, Nah S, Mu Lee K. Enhanced deep residual networks for single image super-resolution. Proceedings of the IEEE conference on computer vision and pattern recognition workshops. 2017:136–44.
He K, Zhang X, Ren S, Sun J. Deep residual learning for image recognition. Proceedings of the IEEE conference on computer vision and pattern recognition. 2016:770–8.
Ledig C, Theis L, Huszár F, Caballero J, Cunningham A, Acosta A, et al. Photo-realistic single image super-resolution using a generative adversarial network. Proceedings of the IEEE conference on computer vision and pattern recognition. 2017:4681–90.
Goodfellow I, Pouget-Abadie J, Mirza M, Xu B, Warde-Farley D, Ozair S, et al. Generative adversarial nets. Advances in neural information processing systems. 2014:27.
Lucas A, Iliadis M, Molina R, Katsaggelos AK. Using deep neural networks for inverse problems in imaging: beyond analytical methods. IEEE Signal Process Mag. 2018;35:20–36.
Islam J, Zhang Y. GAN-based synthetic brain PET image generation. Brain Inform. 2020;7:1–12.
Wolf T, Chaumond J, Debut L, Sanh V, Delangue C, Moi A, et al. Transformers: state-of-the-art natural language processing. Proceedings of the 2020 Conference on Empirical Methods in Natural Language Processing: System Demonstrations. 2020:38–45.
Tang C, Zhao Y, Wang G, Luo C, Xie W, Zeng W. Sparse MLP for image recognition: Is self-attention really necessary? Proceedings of the AAAI Conference on Artificial Intelligence. 2022:2344–51.
Carion N, Massa F, Synnaeve G, Usunier N, Kirillov A, Zagoruyko S. End-to-end object detection with transformers. European conference on computer vision. 2020:213–29.
Touvron H, Cord M, Douze M, Massa F, Sablayrolles A, Jégou H. Training data-efficient image transformers & distillation through attention. International Conference on Machine Learning. 2021:10347–57.
Zhang Y, Liu H, Hu Q. Transfuse: Fusing transformers and cnns for medical image segmentation. International Conference on Medical Image Computing and Computer-Assisted Intervention: Springer; 2021. p. 14-24.
Preetha CJ, Meredig H, Brugnara G, Mahmutoglu MA, Foltyn M, Isensee F, et al. Deep-learning-based synthesis of post-contrast T1-weighted MRI for tumour response assessment in neuro-oncology: a multicentre, retrospective cohort study. Lancet Digit Health. 2021;3:e784–94.
Hore A, Ziou D. Image quality metrics: PSNR vs. SSIM. 20th international conference on pattern recognition. 2010:2366–9.
Sheikh HR, Bovik AC. Image information and visual quality. IEEE Trans Image Process. 2006;15:430–44.
Thie JA. Understanding the standardized uptake value, its methods, and implications for usage. J Nucl Med. 2004;45:1431–4.
Fletcher J, Kinahan P. PET/CT standardized uptake values (SUVs) in clinical practice and assessing response to therapy. NIH Public Access. 2010;31:496–505.
Yang Q, Tan K-H, Ahuja N. Real-time O (1) bilateral filtering. 2009 IEEE Conference on Computer Vision and Pattern Recognition. 2009:557-64
Paris S, Durand F. A fast approximation of the bilateral filter using a signal processing approach. European conference on computer vision. 2006:568–80.
Luo Y, Wang Y, Zu C, Zhan B, Wu X, Zhou J, et al. 3D Transformer-GAN for high-quality PET reconstruction. International conference on medical image computing and computer-assisted intervention: Springer; 2021. p. 276–85.
Hu R, Liu H. TransEM: residual Swin-transformer based regularized PET image reconstruction. arXiv preprint arXiv:220504204. 2022.
Wang Y-RJ, Baratto L, Hawk KE, Theruvath AJ, Pribnow A, Thakor AS, et al. Artificial intelligence enables whole-body positron emission tomography scans with minimal radiation exposure. Eur J Nucl Med Mol Imaging. 2021;48:2771–81.
MD Dipl-math SG, Seith F, Schäfer JF, Christian la Fougère M, Nikolaou K, Schwenzer NF. Towards tracer dose reduction in PET studies: simulation of dose reduction by retrospective randomized undersampling of list-mode data. Hell J Nucl Med. 2016;19:15–8.
Brenner DJ, Hall EJ. Computed tomography—an increasing source of radiation exposure. N Engl J Med. 2007;357:2277–84.
Chawla SC, Federman N, Zhang D, Nagata K, Nuthakki S, McNitt-Gray M, et al. Estimated cumulative radiation dose from PET/CT in children with malignancies: a 5-year retrospective review. Pediatr Radiol. 2010;40:681–6.
Funding
This study was supported by a grant from the National Cancer Institute of the US National Institutes of Health, grant number R01CA269231, and the Andrew McDonough B + Foundation.
Author information
Authors and Affiliations
Contributions
Guarantors of integrity of entire study: Y-R (J) W, DR, HED-L
Study concepts/study design: Y-R (J) W, HED-L
Primary drafting of paper and figures: Y-R (J) W, PW
Clinical evaluations: LCA, AHS
Data acquisition: NDS, AJT, SG, AP, AST
Critical revision of the manuscript for important intellectual content: HED-L, TH, NDS
Data analysis/interpretation: Y-R (J) W, PW, LCA, LQ, QZ
Final approval of the paper: All authors
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Disclaimer
The funders had no role in the study design, data collection and analysis, decision to publish, and preparation of the manuscript.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Daniel Rubin and Heike E. Daldrup-Link are senior authors.
This article is part of the Topical Collection on Advanced Image Analyses (Radiomics and Artificial Intelligence)
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, YR.(., Wang, P., Adams, L.C. et al. Low-count whole-body PET/MRI restoration: an evaluation of dose reduction spectrum and five state-of-the-art artificial intelligence models. Eur J Nucl Med Mol Imaging 50, 1337–1350 (2023). https://doi.org/10.1007/s00259-022-06097-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00259-022-06097-w