Abstract
Purpose
Three-dimensional fibrous scaffolds provide an environment that enhances transplanted stem cell survival in vivo and facilitates imaging their localization, viability, and growth in vivo. To assess transplanted stem cell viability on biocompatible polymer scaffolds in vivo, we developed in vivo imaging systems for evaluation of implanted viable neural stem cells (NSC) and mesenchymal stem cells (MSC) on scaffolds using luciferase or sodium/iodide symporter (NIS) genes.
Methods
Firefly luciferase stably expressing-C6 cell was established (C6-Fluc). The human neural stem cell, F3, was infected with adenoviral vector carrying luciferase gene (F3-Fluc) and MSC expressing NIS controlled by ubiquitin C promoter using lentiviral vector was established by treating blasticidine for 2 weeks (MSC-NIS). Chitosan and poly l-lactic acid (PLLA) scaffolds were used for in vivo image. In vivo expression of luciferase and human NIS was examined by bioluminescence image or 99mTc-pertechnetate gamma camera image, respectively. The cell/scaffold complex was implanted into subcutaneous or abdominal area of BALB/C nude mouse. For quantitative evaluation of cell viability, regions of interest were drawn on 99mTc-pertechnetate scintigraphy by manual.
Results
The gradual increase of luciferase activity was observed in C6-Fluc seeded with chitosan according to the increase in the number of cells. C6-Fluc/chitosan complex subcutaneously implanted into nude mice showed longitudinal bioluminescence image until 34 days. Luciferase image of abdominal-injected C6-Fluc/PLLA complex was saturated in only 14 days, showing great cell growth due to abundant nutrients. F3 cells showed well-incorporated pattern with fibrous chitosan scaffold using scanning electron microscopy. F3 infected with Ad-Fluc showed >100-fold higher luciferase activity than luciferase activity in F3. Cell-number-dependent increase of luciferase activity was shown in F3-Fluc seeded on chitosan. F3-Fluc incorporation into chitosan after abdominal injection was clearly visible on bioluminescence image up to 11 days. Radionuclide imaging showed higher uptake by MSC-NIS on PLLA scaffolds than by MSC-NIS not seeded on a scaffold. Quantitative data showed significantly better survival of MSC-NIS on PLLA scaffolds than without scaffold at 72 h post-implantation, which concurred with histologic findings.
Conclusion
These results suggest that NSC-Fluc and MSC-NIS cells incorporated within polymer scaffolds can be monitored on a long-term basis by serial in vivo imaging. We believe that a biocompatible scaffold-based imaging system could be used to assess stem cell viabilities in a non-invasive way to aid the development of regenerative therapeutics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tissue engineering technology will in the future probably enable treatment of a wide spectrum of diseases that are associated with significant tissue losses in the human body. Two major strategies are used to achieve this goal: One involves the direct introduction of cell suspensions into tissues of interest and the other involves the transplantation of pre-cultured cells on scaffolds serving as templates for tissue reconstitution [1, 2]. Polymer scaffolds that provide biomechanical support for stem cells have been widely used in regenerative therapeutics to restock lost tissue space with appropriate numbers of cells [3–6]. Recent reports have suggested that neural stem cells on nanofibrous scaffolds are biologically and mechanically stable and that their differentiations are well supported, thus, demonstrating the suitability of the scaffold approach for tissue engineering [7, 8]. However, we need to examine how many grafted stem cells survive on transplanted scaffolds and whether these cells proliferate and/or differentiate in vivo after transplanting stem cell–scaffold complexes. Molecular imaging techniques might serve this purpose by enabling the repetitive monitoring of implanted stem cells attached to scaffolds.
Recent advances in molecular imaging techniques enable us to investigate the localization, survival, and proliferation of grafted stem cell non-invasively in vivo. However, unlike cancer cells, stem cells do not always form masses after implantation but rather spread to their favorite niches. Moreover, if they remain in situ, growing stem cell masses may become necrotic internally before cell masses are excessive. On the other hand, if stem cells are transplanted as cell–scaffold complex, the extended surface areas of scaffolds should help the stem cells settle down in vivo in a scaffold cage preventing internal necrosis. Cell–scaffold complexes, thus, might aid molecular stem cell imaging of cell proliferation or differentiation in vivo in the long term.
Poly l-lactic acid (PLLA) is an FDA-approved polymer that is easily hydrolyzed in vivo, and a wide variety of tissue engineering applications have demonstrated the ability of PLLA to form cell–scaffold complexes in vivo [9, 10]. Chitosan is a derivative of chitin with excellent biocompatibility properties and forms stable and porous crystalline structures and, thus, is suitable for cell transplantation/regeneration studies [11, 12]. Moreover, fibrous polymer forms of these materials can be infiltrated easily by transplanted cells. In the present study, we chose chitosan and PLLA, which are both biocompatible and biodegradable, to produce cell/scaffold complexes for the in vivo monitoring of transplanted stem cells.
Luciferase is a good reporter transgene to monitor stem cell behavior in vivo in small animals due to its high sensitivity and low background activity despite its relatively low tissue penetration. Sodium iodide symporter (NIS), a transmembrane protein, which selectively transports iodide anion into thyroid cells, can also be used as a reporter gene and had the advantages of easier imaging with 99mTc-pertechnetate or 123I and better quantification [13, 14]. Two cell lines, human neural stem cells (F3) and rat mesenchymal stem cells, were used in this study to investigate in vivo behaviors as the potential bases of regenerative therapies [15]. We examined the viabilities of these two stem cells incorporated into biocompatible chitosan or PLLA scaffolds in nude mice using NIS and luciferase reporter genes. In addition, we proposed the feasibility of using cell–scaffold complex for imaging the long-term survivals of these cells based on observation of viable cells inside three-dimensional scaffolds.
Materials and methods
Construction of pCMV-Fluc plasmid and Ad-Fluc viral vector
The coding region of firefly luciferase (Fluc) released by HindIII and XhoI restriction enzymes was inserted into multi-cloning site of the pcDNA3.1 vector (Invitrogen, Carlsbad, CA, USA) regulated by CMV (cytomegalovirus) promoter. Also, recombinant adenoviral vector expressing firefly luciferase gene was cloned and purified by CsCl ultracentrifugation (Ad-Fluc).
Lentiviral cloning
Human NIS gene was cloned into pNenti6/UbC/V5-DEST (Invitrogen, Carlsbad, CA, USA) under the control of ubiquitin C promoter. Lentivirus containing NIS gene and packaging plasmid (gag, pol, vsv-g, rev; Invitrogen, Carlsbad, CA, USA) were transfected into a packaging cell line (293FT) using lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA). Supernatant containing Lenti-Fluc was collected by centrifugation (3,000 rpm, 15 min) at 72 h post-transfection. HT1080 human fibrosarcoma cells were infected with lentivirus with 10 μg/ml polybrene to check NIS activity.
Cell culture and the establishments of C6-Fluc, F3-Fluc, and MSC-NIS cell lines
C6, a rat glioma cell line, was grown in RPMI medium (Jeil Biotechservices Inc., Korea) containing 10% fetal bovine serum (Invitrogen, Carlsbad, CA, USA), 10 U/ml penicillin (Invitrogen, Carlsbad, CA, USA), and 10 μg/ml streptomycin (Invitrogen, Carlsbad, CA, USA).
HB1.F3 (a human neural stem cell line; provided by S.U. Kim, Ajou University, Suwon, Korea) was cultured in DMEM medium (Invitrogen, Carlsbad, CA, USA) supplemented with 5% fetal bovine serum, 5% horse serum (Invitrogen, Carlsbad, CA, USA), 10 U/ml penicillin, and 10 μg/ml streptomycin. Rat bone-marrow-derived mesenchymal stem cells (MSCs; kindly donated by Dr. Kim, Asan Medical Center, Seoul) was maintained in McCoy medium with each supplement.
CMV-driven luciferase recombinant plasmid was transfected into C6 using lipofectamine. C6 expressing stably luciferase gene was established by selection with hygromycin (Invitrogen, Carlsbad, CA, USA) for 2 weeks (C6-Fluc). F3 cells were also infected with Ad-Fluc (F3-Fluc). To select stable mesenchymal stem cells expressing the human NIS (hNIS) gene, MSC cells were infected with lentivirus containing the hNIS gene. Twenty-four hours after infection, cells were treated with blasticidine for 2 weeks. The stable cell line was named MSC-NIS.
Preparation of PLLA and chitosan microfibrous matrix
PLLA were dissolved in methylene chloride and acetone (9:1 v/v) at a concentration of 10 w/v %. The polymer solution was loaded into a glass syringe, placed in a syringe pump, and connected the syringe to an 18-gauge needle. Only blunt-tipped needles were used because the slope on sharp needles would cause problems during extrusion. The extrusion flow rate was from 0.10 ml/min. The needle tip was immersed in a small stainless container full of methanol as the coagulating bath fluid. Fibrous matrices were shaped to have pore size of 150–300 μm. The fibrous matrices were exposed to organic solvent vapor for 30 min to bond the fibers. Chitosan fibers were prepared by extruding 4% chitosan solution in 4% acetic acid into a pH 13 aqueous solution. An acid-treated fiber-bonding technique for forming three-dimensional scaffolds was developed. Chitosan disks were fabricated by pressing dried chitosan fibers. The chitosan disks were equilibrated in water for pore generation and were subjected to a pH 5.6 acetate buffer for 15 s, utilizing the fact that chitosan is partially soluble in this solution. After the matrices were placed quickly in a 1 N NaOH solution for inhibiting solubilizing chitosan, nonwoven chitosan fibrous matrix was washed in distilled water and freeze-dried. This process yielded a scaffold of chitosan fibers bonded together by the acid treatment (fiber diameter, 25 μm; fiber density, 0.0139 g/cm3) [16, 17].
Preparation and cell seeding of chitosan or PLLA scaffold
Three-dimensional scaffolds were prepared from poly l-lactic acid microfibers spun using a wet spinning technique. All scaffolds were provided by one of the authors (S. J. Lee, Ewha Womans’ University College of Pharmacy). Prior to cell seeding, all scaffolds were sterilized by wetting in 70% isopropyl alcohol over 24 h and were then washed with phosphate-buffered saline (PBS) three times. MSC culture medium was used to pre-wet scaffolds for 12 h before cell seeding.
125I uptake assay of hNIS transgene activity
MSC-NIS cells were incubated for 30 min in Hanks balanced salt solution (HBSS) containing 0.5% bovine serum albumin, 3.7 kBq of carrier-free Na125I, and 10 μM NaI. These cells were rapidly rinsed with cold HBSS twice, and cells were removed using 0.2% SDS solution [18]. Radioactivities were counted using a gamma counter (CobraII, Packard Instrument Co, Meridien, CT, USA) for iodine uptake testing. For inhibition testing, radioactivities were counted over 30 min after adding 50 μM KClO4.
In vitro bioluminescence assay
Cells were washed with PBS and lysed in lysis solution at 48 h after transfection. Cell homogenates were transferred to a 96-well microplate after collection by scraping. A microplate luminometer (TR717; Applied Biosystems) was used to measure bioluminescence, expressed as relative light units.
In vivo bioluminescence imaging
After C6-Fluc or F3-Fluc cells had been seeded onto polymer scaffolds, cells/scaffold complexes were implanted intraperitoneally. Ten-week BALB/c nude mice (n = 4) were anesthetized with 50 μl of ketamine–xylazine (2:1) solution, and bioluminescence images were acquired after administering 3 mg/0.1 ml/mouse luciferase substrate luciferin by intraperitoneal injection. Animals were placed in an IVIS-200 equipped with a cooled CCD camera (Xenogen, CA, USA). Bioluminescence images were acquired by integration for 3 min.
In vivo image acquisition and analyses
99mTc-pertechnetate 15.2 MBq/0.1 ml were injected intraperitoneally in seven mice. Scintigraphy was acquired 30 min after injections using a single-head gamma camera with pin-hole collimator (LEM, Siemens, Germany). All seven mice were under ketamine sulfate (5 mg/100 g of body weight) anesthesia. For quantification, maximum counts per pixel in regions of interests that were drawn on scintigraphic images were measured using an imaging tool, M-view 5.3 (Marosys, Seoul, South Korea). To draw regions of interest (ROIs) more accurately, all scintigraphic images were overlapped with real photographic images. Corrected maximum counts (CMC) were calculated using the following formula:
Histopathologic examination
All seven mice were sacrificed by cervical dislocation 80 h post-implantation. Subcutaneous injection sites were excised with skin, and scaffold insertion sites were widely excised. All tissues were fixed in 10% formaldehyde solution for 24 h, then paraffin blocks were made. For histopathological examination, hematoxylin–eosin-stained slides were reviewed by a pathologist.
Statistical analyses
All data were processed using SPSS Ver. 12.0 k (SPSS Korea DataSolution, Seoul, South Korea) and Excel 2002 (Microsoft Korea, Seoul, South Korea). To compare time-corrected maximum uptakes by scaffold, repeated measurement analysis of variance (ANOVA) and the paired t test were used. p values of <0.05 were considered to be significant.
Results
Production of cell–scaffold complexes in vitro using C6 tumor cells and the feasibility of in vivo bioluminescence imaging
The stable expression of luciferase in C6 (C6-Fluc) was confirmed using luminometer assays in vitro. Luciferase activities of C6-Fluc were >100 higher than those of C6 parental cells (Fig. S1). To examine the relation between C6 adhesion to chitosan scaffolds versus administered cell numbers, increasing numbers of C6-Fluc were seeded onto chitosan scaffolds. Twenty-four hours after mixing cells with scaffold, luciferase activity increased gradually for C6-Fluc cells on chitosan scaffolds, and this was similar to that of luciferase activity in C6-Fluc seeded in plates. When C6-Fluc/chitosan complex was removed from wells, little luciferase activity remained, indicating that few cells overflowed cell–scaffold complexes regardless of the number of cells inoculated (Fig. 1a).
In vitro and in vivo time course of luciferase activities in C6-Fluc cells incorporated into chitosan scaffold. a C6-Fluc cells were incubated in a chitosan scaffold for 24 h. A gradual increase in luciferase activity was observed in C6-Fluc–chitosan complex as cell numbers increased, showing similar pattern in C6-Fluc on plate. b C6-Fluc on chitosan scaffold was implanted subcutaneously in nude mice [regions 2 (back), 3 (left flank), 4 (right flank)]. Bioluminescence images were acquired at 1, 7, 26, and 34 days post-implantation. C6-Fluc without scaffold [positive control, region 1 (shoulder)] was shielded due to its oversaturated luciferase activities at later images (26 and 34 days post-implantation). c The 2 × 106 C6-Fluc cells seeded on a PLLA scaffold were implanted intraperitoneally (region B). The 1 × 105 C6-Fluc cells were also injected subcutaneously as a control without scaffold (region A). Rapid growth of C6-Fluc cells spread the bioluminescence throughout the whole peritoneum on the day 14 post-implantation. Total luciferase activities were summed up and depicted in Supplementary Fig. 2. The data are the means ± SD of tetraplicates
To confirm the survival of C6-Fluc glioma cells in scaffolds in vivo, C6-Fluc–scaffold complex was subcutaneously grafted into nude mice using C6-Fluc seeded onto chitosan scaffold incubated for 24 h in vitro (regions 2 (back), 3 (left flank), and 4 (right flank) in Fig. 1b). The 2 × 105 C6-Fluc without scaffold was grafted into left shoulders as a control (region 1 in Fig. 1b), and this C6-Fluc showed much higher luciferase activity due to rapid cell growth. Though survivals of 5 × 105 C6-Fluc on scaffold were confirmed up to 34 days (region 3 in Fig. 1b), individual variations were high, and background luminescence was also observed along surgical wounds (day 7 of Fig. 1b).
Because peritoneal cavity appeared suitable for implanting cells in terms of space and nutrient supply, we implanted 2 × 106 C6-Fluc cells seeded on a PLLA scaffold intraperitoneally (region B of Fig. 1c). The 1 × 105 C6-Fluc cells were also injected subcutaneously as a control without scaffold (region A of Fig. 1c). Spread intense bioluminescence were observed in the peritoneal cavities after C6-Fluc scaffold implantation on day 1, and this activity became oversaturated on day 14 (Fig. 1c). Luciferase activities in C6-Fluc–scaffold complexes gradually increased with time, showing rapid cell growth in peritoneal cavities (days 6 and 14 of Fig. 1c). Bioluminescences for each region were quantified and are shown in Fig. S2, which showed similar total sum of luminescence activities of region B between individuals.
In vitro validation of the production of cell–scaffold complexes using neural stem cells (F3-Fluc) and fibrous chitosan scaffolds
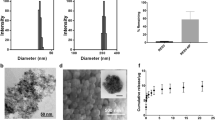
The 1 × 106 F3 cells were seeded onto chitosan scaffolds to examine F3 cell attachment. Six days after cell seeding, F3 cells were found to have attached well onto fibrous chitosan scaffolds by scanning electron microscopy (Fig. 2a). Ad-Fluc vector was infected into F3 at a multiplicity of infection of 0.75 (F3-Fluc). Luciferase activity increased significantly in F3 infected with Ad-Fluc as compared with parental F3 (Fig. S3). When F3-Fluc were seeded onto chitosan at increasing cell numbers, luciferase activities increased linearly up to 5 × 105 cells/well, whereas activities showed a bell-shaped response according to the increase of the number of cells and decreased when more than 4 × 105 cells were seeded on a plate. This was probably due to a confluence effect (Fig. 2b). The linear relationship between increases in the numbers of F3-Fluc cells up to 1 × 106 cells in chitosan scaffolds was confirmed in a separate experiment (Fig. S4).
Adherent effect of F3-Fluc to the chitosan scaffolds in vitro and their in vivo visualization of F3-Fluc scaffolds on optical imaging in peritoneal cavities. a F3 adherence to chitosan scaffolds was shown by SEM imaging at 6 days after seeding at a cell concentration of 1 × 106 cells/μl. a Chitosan scaffold (×100), b F3, neural stem cells on a chitosan scaffold (×100). b Luciferase activity in F3-Fluc on chitosan scaffolds was found to be dependent on cell number. Bell-shaped number activity curve in F3-Fluc cells seeded on plates was due to cell overgrowth. c In vivo monitoring of F3-Fluc incorporated into chitosan scaffold. Two clones of Ad-Fluc vector infected F3 cells were implanted with scaffolds into peritoneal cavities (region B). On day 3, bioluminescence in the abdomen of these mice showed well-incorporated stem cells within the chitosan scaffolds. Follow-up images of days 6 and 11; the activities of the grafted cells of group 1 were sustained until 11 days, but those of group 2 vanished probably due to transgene silencing. C6-Fluc cells without scaffold were used as a control on the shoulders (region A). Data are expressed as means ± SD in tetraplicates
In vivo bioluminescence imaging of F3-fluc–scaffold complexes
The 1 × 106 F3-Fluc cells (two clones, groups 1 and 2) and 1 × 105 C6-Fluc cells were seeded onto chitosan scaffolds. F3-Fluc–scaffolds were implanted into peritoneal cavities and C6-Fluc–scaffolds into right shoulders as a positive control. On day 3, luciferase activity was evident in peritoneal cavities (F3-Fluc–scaffold) and in right shoulders (C6-Fluc–scaffold) in all mice. On day 11, luciferase activity in the peritoneal cavities of group 1 mice had increased or at least persisted, but in group 2, luciferase activities disappeared probably due to the transient expression of this clone infected with adenovirus carrying luciferase gene (Fig. 2c). In right shoulders, luciferase activity increased gradually in C6-Fluc–scaffolds up to 11 days in Groups 1 and 2 (Fig. 2c). In another group of mice (negative controls) injected with F3-Fluc alone (without scaffold) in the peritoneal cavity, no luciferase activity was observed (data not shown), probably because F3-Fluc cells could not settle in the peritoneal cavity without scaffold support, which resulted in no bioluminescence.
In vitro validation of mesenchymal stem cells (MSC-NIS) on PLLA scaffold using iodine uptake assays
Lentivirus-mediated NIS-transfected MSCs stably expressed NIS genes and, thus, took up radioactive iodine up to 18.0 times more than MSCs without NIS (415.5 ± 4.1 pmol/105 cells). After treatment with KClO4, radioactive iodine uptake was completely inhibited (Fig. S5). Chitosan or PLLA scaffolds were pre-wetted for 24 h in serum-free medium, and 1 × 106 MSC-NIS and MSC were seeded onto these pre-wetted scaffolds. MSC-NIS–chitosan or MSC-NIS–PLLA complex showed approximately five-fold higher iodine uptake as compared with MSC–chitosan, MSC–PLLA, chitosan, or PLLA. Glass pipette suction was enough to remove iodine inside scaffolds, which yielded no background activities both in MSC and in scaffolds (Fig. 3a).
In vitro and in vivo activity of MSC-NIS cells with scaffold on radioiodine uptake assay and 99mTc-pertechnetate imaging. a MSC-NIS cells adherent to chitosan or PLLA scaffolds showed comparable (about five-fold increase) radioiodine uptake in vitro. b The 3 × 106 MSC-NIS cells and c 9 × 106 MSC-NIS cells were seeded onto PLLA scaffold, and this cell/scaffold complex was transplanted into mice. Radionuclide image in MSC-NIS/PLLA scaffold region was higher than that of MSC-NIS without scaffold. ROI analysis for MSC-NIS cells on PLLA scaffold 72 h after implantation (3 × 106 cells on PLLA scaffolds, 9 × 106 cells on PLLA scaffolds)
In vivo 99mTc-pertechnetate imaging of MSC-NIS–scaffolds and the histopathologic validation of the proliferation of implanted cells on scaffolds
Cells and cell–scaffold complexes were subcutaneously injected into each mouse (Fig. S6. A: MSC-NIS–PLLA scaffold, B: MSC-NIS without scaffold, C: PLLA scaffold alone). MSC-NIS and MSC-NIS–scaffold showed specific 99mTc-pertechnetate uptake. Serial scintigraphic images showed changes in the uptake of 99mTc-pertechnetate with time (Fig. S7).
MSC-NIS incorporated with PLLA scaffold showed higher radionuclide image than MSC-NIS without PLLA scaffold (Fig. 3b: 3 × 106, 9 × 106). For quantitative evaluation, regions of interest were defined on co-registered images, and CMCs were determined (Fig. 3b: 3 × 106, 9 × 106). CMCs on MSC-NIS with scaffolds and without scaffolds (Table S1) are shown in Fig. 4a–d, and these figures show the time courses of specific 99mTc-pertechnetate uptake in implanted areas in each group. Repeated measure ANOVA of the MSC-NIS–scaffold versus MSC-NIS groups (between A and B ROIs in Fig. S6) disclosed the effect of scaffold on 99mTc-pertechnetate uptake.
Quantified data for CMC curves of MSC-NIS on PLLA scaffolds and without a scaffold. The first three graphs represent average CMCs of mice treated with the same number of cells. The last graph is of the CMC of mouse 4. Overall CMCs of MSC-NIS on a PLLA scaffold were higher than those of MSC-NIS without a scaffold. In subject 4, two scaffolds were implanted; CMC at 72 h post-implantation was higher than the mouse with one scaffold. Each figure was represented from Supplementary Fig. 16 data (a mice 1a, 1b; b mice 2a, 2b; c mice 3a, 3b; d mouse 4)
No difference in CMCs was observed between bare MSC and non-specific background activity (Table S2). No difference in CMC was observed between scaffold without cells and non-specific background area (Table S3).
Implanted MSC-NIS–scaffold, MSC-NIS without scaffold, and scaffold-alone excised after sacrifice revealed increased cellular density by H&E stain in MSC-NIS–scaffold and MSC-NIS areas but not in the site of scaffold-alone (Fig. 5). PLLA scaffolds with or without cells showed inflammatory cell infiltration of macrophages around PLLA scaffolds (Table 1). MSC injection sites with lower numbers of cells (3 × 106 cells) showed no stem cells, and MSC masses without scaffold sometimes showed internal necrosis (Fig. 5c).
Discussion
Many trials on stem cell imaging have been carried out to track grafted stem cell behaviors in vivo for cell localization, survival, and proliferation. To monitor grafted stem cells on a long-term bases, three-dimensional meshes that provide spaces and surfaces for cell growth and that ‘cage’ grafted cells are required. This space support by scaffolds eases contact inhibition between proliferating grafted cells, improves grafted cell survival, and enables grafted cells to form a supported mass in vivo. For this purpose, scaffolds should have high porosity, large surfaces, and adequate porosity. Biocompatible polymer scaffolds help defective tissues and organs regenerate their original masses. Cell–scaffold complexes attached with cells grown in vitro have become important recently in the field of cell therapy [19, 20].
Scaffolds should be easily absorbed as soon as cells have filled defective spaces to minimize foreign body reactions. Biocompatible polymers like chitosan or PLLA are widely used as biomaterials for tissue reconstruction because of their biodegradabilities, mechanical stabilities, and non-toxic natures [21–23]. These three-dimensional polymer scaffolds offer mechanical templates that support cell survival and growth in vivo. In this study, we developed a non-invasive monitoring system for stem cells within polymer scaffolds in vivo that allow investigations of grafted stem cell survival and growth by optical or radionuclide imaging.
We first tested the feasibility of in vivo polymer scaffold-cell imaging using C6 glioma cells by implanting C6-Fluc–scaffold complex subcutaneously and intraperitoneally. As was expected for tumor cells, the luminescence of subcutaneously injected C6-Fluc cells without scaffold was easily saturated with huge size of tumor mass. After shielding this hot area with black paper, we could confirm the survivals of tumor cells within scaffolds for up to 34 days. However, surgical regions implanted with scaffolds did not heal well and showed non-specific background bioluminescence, though not so hot. And C6-Fluc cells within scaffolds in subcutaneous tissues yielded much less luminescence, probably reflecting a poorer blood and nutrient supply via scaffolds than ‘uncaged’ C6-Fluc tumor cells without scaffolds. However, when we changed the implantation site to the peritoneal cavity where nutrient supply is ample for cell growth and survival, marked cell growth in peritoneal cavities with tumor cells with scaffolds was observed much the same as in subcutaneous C6-Fluc cells without scaffold on shoulders. Peritoneal cavities looked much favored by ‘caged’ tumor cells within scaffolds than subcutaneous location in that the tumor cells had spread into surroundings in peritoneal cavities outside of scaffold cages due to their cellular overgrowth (Fig. 1c).
We chose neural stem cells and mesenchymal stem cells because these cells may be suitable for the treatment of neurodegenerative diseases and bone disorders, respectively. The viability of neural stem cells (F3-Fluc) seeded on PLLA scaffold was maintained for up to 11 days, but unlike C6 cells, F3-Fluc cells should have been well-entrapped and organized onto fibrous polymer scaffolds, which were observed on bioluminescence images (Fig. 2c). Abdominal activities of F3–scaffold complexes in mice were more confined than tumor–scaffold complexes (Fig. 1c). For this experiment, we used an adenoviral system associated with non-integrative expression of transgenes and found one cell line with persistent expression of the Fluc reporter transgene (group 1) and another cell line showing rapid silencing of the Fluc transgene (group 2 of Fig. 2c). In group 2, lack of bioluminescence on later days might be either due to cell loss or due to transgene silencing, which obscured the differentiation between cell loss and transgene loss in viable cells. In this respect, retroviral or lentiviral system might be a better alternative in stem cell–scaffold imaging, especially for longer term studies. In this experiment, the activities were not quantified in vivo because of their variable attenuation according to the depth of location. This was the main shortcoming of optical imaging. F3-Fluc cells injected without scaffolds in the peritoneal cavity were not observed on site because F3-Fluc might have spread out to the entire intraperitoneal cavity or remote tissues as soon as these cells were injected.
Radionuclide imaging enables not only tomography but also quantitative analysis in small animals and in humans. In this experiment, a mesenchymal stem cell line expressing NIS gene (MSC-NIS) regulated by ubiquitin C promoter was established using a lentiviral system. Visual and quantitative analyses using grafted MSC-NIS on serial images revealed temporal changes of 99mTc-pertechnetate uptake by MSC-NIS with or without scaffolds. By visual inspection, we confirmed that MSC-NIS took up 99mTc-pertechnetate, but ROI quantification (after removing the confounding effects of background) using corrected maximum counts was required to prove the beneficial effects of scaffolds on stem cell survival. Individual variations were prominent as shown in Fig. 4; the activities of MSC-NIS–scaffold persisted in a certain mouse probably because implanted stem cells survived owing to better scaffold support and nutrient supply of that mouse. We found that reporter NIS activities of MSC-NIS cells on PLLA scaffolds were higher than those without a scaffold, while these reporter activities were the same between cells with or without scaffolds at the start in every mouse (3 h after implantation). We think that the scaffolds allow the required nutrition to be supplied and create spaces for stem cells. Now, we believe that we have a tool to examine the suitabilities of scaffolds and in vivo environments. For example, a comparison between chitosan and PLLA or any other polymer system can be performed in terms of their in vivo suitability as supporting scaffolds for each cell.
In the present study, histopathologic examinations showed mild inflammatory responses probably due to PLLA scaffold (Fig. 5). However, no internal necrosis was found in cell–scaffold implants, but stem cell masses without scaffolds showed internal necrosis.
Noninvasive in vivo molecular imaging using stem cells carrying reporter genes attached to polymer scaffolds provides a real benefit in terms of developing stem cell therapies using cell–scaffold complexes. Dynamic imaging and the real-time monitoring of stem cell behaviors in vivo such as survival, distribution, and proliferation are mandatory if the beneficial effects of newly devised scaffold supports for cells are to be understood by elucidating benefits on survival or the proliferation of grafted stem cells in vivo. Recently, a variety of modified polymer scaffolds such as polymer surface modifications were proposed to improve biocompatibility or survival effect in vivo [24–28]. In vivo molecular imaging might help us circumvent the effects of individual variations in small animal experiments as this technique allows examinations of single animals repetitively over several weeks. Exploiting this advantage, this technique will allow us to evaluate the superiority of a novel biocompatible polymer scaffold over another. We also can evaluate cellular differentiation patterns or regenerative induction of grafted stem cells by modifying scaffold surfaces with molecular attachment that induce differentiation and promote regeneration responses [7, 29, 30].
This study represents the first successful monitoring of grafted stem cells incorporated into biocompatible scaffolds using luciferase or nuclear reporters for the development of cell/scaffold-based engraftment. Application of in vivo molecular imaging to tissue engineering using cell-polymer scaffolds as a bio-bridge provides an approach closer to clinical applications for the assessment of regenerative and cell-replacement therapies.
References
Langer R, Vacanti JP. Tissue engineering. Science 1993;260:920–6.
Griffith LG, Naughton G. Tissue engineering: current challenges and expanding opportunities. Science 2002;295:1009–14.
Shi C, Zhu Y, Ran X, Wang M, Su Y, Cheng T. Therapeutic potential of chitosan and its derivatives in regenerative medicine. J Surg Res 2006;15:185–92.
Boo JS, Yamada Y, Okazaki Y, Hibino Y, Okada K, Hata K, et al. Tissue-engineered bone using mesenchymal stem cells and a biodegradable scaffold. J Craniofac Surg 2002;13:231–9.
Hammond JS, Beckingham IJ, Shakesheff KM. Scaffolds for liver tissue engineering. Expert Rev Med Devices 2006;3:21–7.
Liu W, Cui L, Cao Y. Bone reconstruction with bone marrow stromal cells. Methods Enzymol 2006;420:362–80.
Silva GA, Czeisler C, Niece KL, Beniash E, Harrington DA, Kessler JA, et al. Selective differentiation of neural progenitor cells by high-epitope density nanofibers. Science 2004;303:1352–5.
Park KI, Teng YD, Snyder EY. The injured brain interacts reciprocally with neural stem cells supported by scaffolds to reconstitute lost tissue. Nat Biotechnol 2002;20:1111–7.
Evans GR, Brandt K, Niederbichler AD, Chauvin P, Herrman S, Bogle M, et al. Clinical long-term in vivo evaluation of poly(l-lactic acid) porous conduits for peripheral nerve regeneration. J Biomater Sci Polym Ed 2000;11:869–78.
Yang F, Murugan R, Wang S, Ramakrishna S. Electrospinning of nano/micro scale poly(l-lactic acid) aligned fibers and their potential in neural tissue engineering. Biomaterials 2005;26:2603–10.
Lee JY, Nam SH, Im SY, Park YJ, Lee YM, Seol YJ, et al. Enhanced bone formation by controlled growth factor delivery from chitosan-based biomaterials. J Control Release 2002;78:187–97.
Suh JK, Matthew HW. Application of chitosan-based polysaccharide biomaterials in cartilage tissue engineering: a review. Biomaterials 2000;21:2589–98.
Chung JK. Sodium iodide symporter: its role in nuclear medicine. J Nucl Med 2002;43:1188–200.
Kang JH, Lee DS, Paeng JC, Lee JS, Kim YH, Lee YJ, et al. Development of a sodium/iodide symporter (NIS)-transgenic mouse for imaging of cardiomyocyte-specific reporter gene expression. J Nucl Med 2005;46:479–83.
Ourednik V, Ourednik J, Flax JD, Zawada WM, Hutt C, Yang C, et al. Segregation of human neural stem cells in the developing primate forebrain. Science 2001;293:1820–4.
Scoof H, Apel J, Heschel I, Rau G. Control of pore structure and size in freeze dried collagen sponges. J Biomed Mater Res 2001;58:352–7.
Mikos AG, Thorsen AJ, Czwerwonka LA, Bao Y, Langer R. Preparation and characterization of poly(l-lactic acid) foams. Polymer 1994;35:1068–77.
Kosugi S, Sasaki N, Hai N, Sugawa H, Aoki N, Shigemasa C, et al. Establishment and characterization of a Chinese hamster ovary cell line, CHO-4J, stably expressing a number of Na+/I− symporters. Biochem Biophys Res Commun 1996;227:94–101.
Bensaïd W, Triffitt JT, Blanchat C, Oudina K, Sedel L, Petite H. A biodegradable fibrin scaffold for mesenchymal stem cell transplantation. Biomaterials 2003;24:2497–502.
Elçin YM, Dixit V, Lewin K, Gitnick G. Xenotransplantation of fetal porcine hepatocytes in rats using a tissue engineering approach. Artif Organs 1999;23:146–52.
Hutmacher DW, Goh JC, Teoh SH. An introduction to biodegradable materials for tissue engineering applications. Ann Acad Med Singapore 2001;30:183–91.
Madihally SV, Matthew HW. Porous chitosan scaffolds for tissue engineering. Biomaterials 1999;20:1133–42.
Chen VJ, Ma PX. Nano-fibrous poly(l-lactic acid) scaffolds with interconnected spherical macropores. Biomaterials 2004;25:2065–73.
Yamaguchi M, Shinbo T, Kanamori T. Surface modification of poly(l-lactic acid) affects initial cell attachment, cell morphology, and cell growth. J Artif Organs 2004;7:187–93.
Liu X, Won Y, Ma PX. Porogen-induced surface modification of nano-fibrous poly(l-lactic acid) scaffolds for tissue engineering. Biomaterials 2006;27:3980–7.
Ma Z, Gao C, Gong Y, Shen J. Cartilage tissue engineering PLLA scaffold with surface immobilized collagen and basic fibroblast growth factor. Biomaterials 2005;26:1253–9.
Li J, Pan J, Zhang L, Yu Y. Culture of hepatocytes on fructose-modified chitosan scaffolds. Biomaterials 2003;24:2317–22.
Ho MH, Hou LT, Tu CY, Hsieh HJ, Lai JY, Chen WJ, et al. Promotion of cell affinity of porous PLLA scaffolds by immobilization of RGD peptides via plasma treatment. Macromol Biosci 2006;6:90–8.
Levenberg S, Huang NF, Lavik E, Rogers AB, Itskovitz-Eldor J, Langer R. Differentiation of human embryonic stem cells on three-dimensional polymer scaffolds. Proc Natl Acad Sci USA 2003;100:12741–6.
Taqvi S, Roy K. Influence of scaffold physical properties and stromal cell coculture on hematopoietic differentiation of mouse embryonic stem cells. Biomaterials 2006;27:6024–31.
Acknowledgments
We thank Dr. Sang Hee Kim of the Asan Medical Center, Seoul, South Korea for generously donating the rat mesenchymal stem cells. This work was supported by Nano Bio Regenomics Project of Korean Science and Engineering Foundation and by Innovation Cluster for Advanced Medical Imaging Technology. This study was made easier using KREONET, Korean Research Network, a nation-wide Giga-bps network system.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Soonhag Kim and Dong Soo Lee contributed equally to this investigation as corresponding authors and Do Won Hwang and Sung June Jang equally contributed as co-first author.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table 1
CMCs of 99mTc-pertechnetate uptake at the implantation sites of MSC-NIS with scaffold and MSC-NIS without scaffold. Repeated measurement ANOVA revealed a significant difference between CMCs of MSC-NIS with scaffold and MSC-NIS without scaffold (p = 0.034) (DOC 50.0 KB)
Supplementary Table 2
Ratio of corrected maximum counts on MSC injection site and control area. The paired t test showed no significant difference between groups (DOC 28.5 KB)
Supplementary Table 3
Ratio of corrected maximum counts on scaffold alone and control area. The paired t test showed no significant difference between groups (p > 0.05) (DOC 28.0 KB)
Supplementary Fig. 1
C6 glioma cells stably expressing luciferase showed much higher luciferase activity than parental C6 cells. The values are mean ± SD of tetraplicate wells (GIF 67.2 KB)
Supplementary Fig. 2
The 2 × 106 C6-Fluc cells seeded on a PLLA scaffold were implanted intraperitoneally. Summed activities in C6-Fluc/PLLA complex of peritoneal cavities (region B of Fig. 1c) were quantified, and their time courses were shown (GIF 76.4 KB)
Supplementary Fig. 3
F3-Fluc cells showed markedly higher luciferase activity than in parental F3 cells at 48 h after infecting F3 with Ad-Fluc vector (GIF 70.8 KB)
Supplementary Fig. 4
On these microplate images, luciferase activities gradually increased in F3-Fluc/chitosan scaffolds as cell numbers were increased. Initially administered number of cells was comparable to the final activities in cell–scaffold complexes in vitro. Linearity was preserved up to the cell density of 1 × 106 cells/well (GIF 105 KB)
Supplementary Fig. 5
NIS activity in MSC infected with NIS using a lentiviral system was higher than in parental MSC cells (black bar), and KClO4 completely inhibited NIS expression (white bar) (GIF 70.1 KB)
Supplementary Fig. 6
MSC-NIS was implanted to three or four subcutaneous areas of these mice. These are co-registered images showing ROIs used for quantitative analyses. A: MSC-NIS–PLLA scaffold complex was implanted surgically in right flanks. B: MSC-NIS cells without scaffold were injected subcutaneously using a 22 G needle. C: PLLA scaffold alone without cells was implanted subcutaneously. In the mice 1, 3, and 5, on the left scapular area opposite to region C, MSC without NIS was injected subcutaneously as a negative control (GIF 119 KB)
Supplementary Fig. 7a
To the seven mice, MSC-NIS–scaffold complex and MSC-NIS cells without scaffold were implanted. The number of implanted cells was varied for each mouse (1a, 1b: 3 × 106; 2a, 2b: 6 × 106; 3a, 3b: 9 × 106; 4: two scaffolds with 3 × 106cells. In addition to normal thyroid, stomach, and occasional bladder uptakes, the 99mTc-pertechnetate uptake on the implanted sites by MSC-NIS were shown repeatedly imaged from 3 to 72 h post-implantation. MSCs without NIS were implanted in the left scapular area of subjects 1a, 2a, and 3a (GIF 309 KB)
Supplementary Fig. 7b
(GIF 256 KB)
Rights and permissions
About this article
Cite this article
Hwang, D.W., Jang, S.J., Kim, Y.H. et al. Real-time in vivo monitoring of viable stem cells implanted on biocompatible scaffolds. Eur J Nucl Med Mol Imaging 35, 1887–1898 (2008). https://doi.org/10.1007/s00259-008-0751-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00259-008-0751-z