Abstract
Purpose
Serotonergic brain regions play a crucial role in the modulation of emotion, and serotonergic dysfunction may contribute to several neurological disorders. [123I]ADAM is a novel SPECT tracer which binds with high affinity to serotonin transporters (SERT). The objective of this study was to compare different methods for the quantification of tracer binding and to develop a simplified single-scan protocol for this tracer, as well as to investigate its potential for characterisation of the transporter occupancy versus plasma concentration curve of a selective serotonin re-uptake inhibitor (SSRI).
Methods
Dynamic SPECT scans were performed on 16 healthy volunteers after administration of ∼150 MBq [123I]ADAM. Data were acquired from the time of injection until ∼5.5 h after injection in 30- or 45-min sessions. Each subject was scanned twice: with and without pre-treatment with the SSRI citalopram in various dosage regimens. The plasma concentration of citalopram (C p) was determined from venous samples. Images were reconstructed by filtered back-projection with scatter and attenuation correction. Tracer binding was quantified for midbrain, striatum and thalamus using cerebellum as a reference region. Quantification was done by kinetic modelling, graphical analysis and multi-linear regression, as well as by the ratio method, with binding potential (BP2) as the outcome measure. The SERT occupancy by citalopram was determined relative to the baseline scan for each subject, and the occupancy versus C p curve was fitted with the E max model.
Results
The highest binding of [123I]ADAM was in midbrain (mean baseline BP2±SD=1.31±0.29), with lower binding in thalamus (0.79±0.16) and striatum (0.66±0.13). There was good agreement between BP2 values obtained by different quantification methods. Using the ratio method, the best agreement with kinetic modelling was obtained with data from the time interval [200,260] min after injection. The fitting of the midbrain occupancy curve yielded a maximum occupancy of 84% and a plasma concentration required to reach 50% of the maximum of 2.5 ng/ml, with a goodness-of-fit variability of 13% (SD).
Conclusion
Binding of [123I]ADAM to SERT in midbrain can be quantified with a single scan starting 200 min after injection. However, the variability of estimated occupancy values may be too high for critical assessment of occupancy of SERT by SSRI.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Serotonergic dysfunction may contribute to neurological disorders such as depression [1, 2], obsessive compulsive disorder [3], alcoholism [4], Parkinson’s disease [5], Huntington’s disease, Pick’s disease and dementia of the Alzheimer type [6, 7]. Serotonergic brain regions also play a crucial role in the modulation of normal emotions. Therefore non-invasive studies of this system in living humans are scientifically important as well as clinically relevant, and there is a great demand for highly selective SERT radioligands for positron emission tomography (PET) and single-photon emission computed tomography (SPECT) [8].
[123I]ADAM (2-((2-((dimethylamino)methyl)phenyl)thio)-5-iodophenylamine) is a novel tracer that binds to serotonin transporters (SERT), with high affinity for SERT (Ki=0.013 nM) and very high selectivity over the noradrenaline and dopamine transporter [9]. [123I]ADAM is a safe and effective radiotracer for imaging serotonin transporters in humans [10, 11]. The biodistribution of [123I]ADAM in animals and human studies appears to be consistent with the known distribution of SERT binding sites in the brain [9, 11, 12]. In autoradiographic studies in rat and mouse brains, the highest initial uptake of [123I]ADAM was found in the dorsal raphe nucleus within the midbrain [13]. Human studies consistently showed highest binding of [123I]ADAM in midbrain regions [10, 11, 14, 15]. High binding levels were also found in thalamus [14, 15] and striatum [11, 14, 15]. In healthy volunteers, pre-treatment with citalopram [a selective serotonin reuptake inhibitor (SSRI)] resulted in reduced [123I]ADAM binding in midbrain regions [16]. Studies in non-human primates showed that quantification of [123I]ADAM binding is possible without arterial sampling, using cerebellum as a reference region [17]. This was confirmed in a human study, in which blocking with the SSRI citalopram led to displacement of [123I]ADAM in midbrain but not in cerebellum [18].
[123I]ADAM may have the potential to be used to investigate diseases that affect the serotonergic system or drugs that act on SERT, either directly or indirectly. Our goal was to contribute to the development of [123I]ADAM as an imaging agent for quantifying serotonin transporter blockade by drugs [19, 20, 21]. The specific objectives of this study were to compare different methods for quantification of tracer binding, to develop a simplified scanning protocol and to investigate whether [123I]ADAM can be used effectively to characterise the transporter occupancy versus plasma concentration curve for the SSRI citalopram.
Materials and methods
Subjects
A total of 16 healthy male volunteers entered the study. Fourteen subjects were Caucasian, one was Afro-Caribbean and one was South American in origin. Their mean (range) age was 32 (25–52) years. Fifteen subjects consumed alcohol (1–14 units per week) and seven subjects smoked tobacco (2–5 cigarettes per day). All serology (for HIV, Hep B and Hep C), urinary drug screen and alcohol breath test results were negative at screening. Routine biochemical and haematological screening tests and urinalysis were done within 21 days before the first scan, and all results were acceptable. All subjects gave written informed consent, and ethics committee approval was obtained.
Drug treatment
Each subject who completed the study had two SPECT scans, separated by at least 7 days. The first scan was a baseline measurement, and the second was done after pre-treatment with the SSRI citalopram. One subject (#8) was withdrawn after his baseline scan owing to an accidental injury unrelated to the study. Citalopram (Cipramil) is a highly selective SSRI that is in wide clinical use for the treatment of depressive illness and panic disorder. The recommended dose is 20–60 mg daily. Citalopram has well-defined linear and dose proportional pharmacokinetics over the range 10–60 mg, has no metabolites with significant activity at SERT and has no or very low affinity for other CNS receptors [22, 23].
Seven dosing regimens of citalopram were used: (a) 10 mg daily for 2 days, (b) 20 mg daily for 2 days, (c) 40 mg daily for 3 days, (d) 40 mg daily for 2 days and 60 mg on the third day, (e) 60 mg daily for 3 days, (f) 10 mg once and (g) 40 mg daily for 7 days. These regimens were chosen to provide a suitable range of steady-state citalopram plasma concentrations. The subjects were randomised to receive one of the seven dose regimens of citalopram, with the first five subjects spanning the first five regimens.
Subjects received potassium iodate tablets (170 mg daily) for 2 days before each scan and on the scan day to saturate the thyroid gland.
Tracer
[123I]ADAM injections were manufactured under GMP standards, labelled and packed by MAP Medical Technologies Oy, Finland, and shipped to England on the day before each scan (half-life of 123I: 13.2 h). Permission to administer radioactive tracers was obtained from the United Kingdom Administration of Radioactive Substances Advisory Committee.
SPECT scanning
Dynamic SPECT data were acquired using a Prism 3000XP triple-headed scanner equipped with a 153Gd transmission source (Philips Medical Systems, Cleveland OH, USA). The three detectors were fitted with ultra-high-resolution fan-beam collimators. Primary emission data were collected in a 15% wide energy window, centred at 159 keV. Two 6% wide windows were placed either side of the peak to detect scattered primary and high-energy photons. For transmission data a 20% energy window, centred at 100 keV, was used.
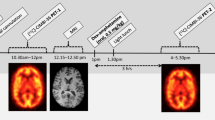
The average (±SD) amount of injected activity of [123I]ADAM was 152±31 MBq. The scanning protocol consisted of a series of five or six imaging sessions of 30 or 45 min with breaks in between, starting at the time of injection and continuing for ∼5.5 h. In the last session, transmission data were acquired simultaneously with the emission data. On most of the scanning days (except for the first and the last day), two subjects were scanned on the same day, with interleaving of the imaging sessions. Each imaging session consisted of several time frames, and the whole scanning protocol can be described as follows for the first and the last subjects: [em 18×2.5 min, br 15 min, 4×(em 6×5 min, br 30 min), em+tr 2×15 min], where em stands for emission data, tr for transmission data and br for break. For the rest it was: [em 12×2.5 min, br 40 min, 3×(em 6×5min, br 40 min), em+tr 2×15 min]. In one case (subject 6, baseline scan) the data from the first acquisition session (0–30 min p.i.) were lost owing to a technical problem.
To allow intra-subject realignment of the images from each session, four fiducial markers (containing ∼0.1 MBq of 123I) were placed on each subject’s head along the orbito-meatal line. The markers also allowed correction for head motion between different time frames within each scanning session.
Venous samples (5 ml) for the measurement of citalopram concentration were taken immediately before the start and after the end of the second scan. Plasma was separated by centrifugation.
Data processing
All image reconstruction and processing was performed using in-house software implemented in IDL 5.2 (Interactive Data Language, Research Systems Incorporated, Boulder, CO, USA). Transmission images were reconstructed using an ordered subsets implementation of the convex maximum likelihood algorithm [24] with four iterations and eight subsets. Emission data were corrected for scatter using the triple energy window method [25]. Tomographic images were reconstructed using filtered back-projection with a ramp filter [26] into a 128×128×60 matrix with 2.03×2.03×3.56 mm voxels. A 3D Butterworth filter [B(f)=(1+(f/q)2p)−1] was applied to the reconstructed images, with parameters p=4 and q=0.58 cm −1. Attenuation correction was performed using two iterations of the method proposed by Chang [27]. The final resolution of the images was ∼11 mm (full-width at half-maximum). The centroid positions of the fiducial markers were used to realign images from different sessions using a six-parameter (three translation and three rotation) rigid body transformation, by minimising the mean-square difference in the marker positions. We have estimated the accuracy of this realignment procedure to be ∼1.5 mm, with a maximum error of <3 mm.
Scanner calibration
A calibration scan was performed with a 20-cm-diameter cylindrical phantom filled with a uniform solution of 123I. The activity concentration in the phantom was determined from the known volume and the total activity (∼50 MBq), as measured with the same well counter (Capintec 120R) as was used for measuring the [123I]ADAM doses. The phantom data were reconstructed as described above and a calibration factor was determined for the purpose of estimating absolute activity concentration in the human studies.
Data analysis
An [123I]ADAM template was generated using data from the baseline scans of the first five subjects. The attenuation map from each subject was used to determine nine realignment parameters (three rotation, three shift and three scaling parameters) relative to one of the subjects (#5). The realignment parameters were applied to the integrated emission images, and an average of the five images was obtained to be used as a template. This template was originally created for an initial evaluation of the five first data sets, but was later used for all subsequently acquired data as well.
Three-dimensional volumes of interest (VOIs) were drawn manually [as a series of two-dimensional regions of interest (ROIs) in consecutive planes] on the template for the following brain regions: cerebellum (Cer), striatum (Str, including head of caudate and putamen), thalamus (Thal) and midbrain (MB).
For the purposes of analysis, the VOIs were overlaid on each subject’s SPECT data by an inverse transform procedure using attenuation map realignment parameters derived as described above. Bilateral hemispheric regions were averaged to yield a single estimate for each region per subject.
In addition, a second set of VOIs were generated by manually drawing them on the baseline scan of each subject. These VOIs were transferred to the post-citalopram scan using a six-parameter rigid body transformation, employing parameters determined by realignment of the two transmission scans on each subject. The second method of generating VOIs is less susceptible to realignment errors, while the first is less operator dependent. We also created whole-brain VOIs for estimation of whole-brain uptake of the tracer.
Tracer binding
The total brain uptake was estimated by calculating the ratio of the maximum value of the total brain activity divided by the injected activity.
Tracer binding was quantified by using cerebellum as a reference region, assumed to be devoid of SERT. Four different methods were investigated for quantification of [123I]ADAM binding: simplified reference tissue modelling (SRTM) [28], graphical analysis (GA) [29, 30], multi-linear regression (MLR) [31] and the ratio method (RM). The first three methods are kinetic analysis methods, based on dynamic data acquisition and an indirect input function obtained from cerebellum. SRTM was regarded as the “gold standard”. GA and MLR can be useful for voxel-based analysis and production of parametric images. RM requires only a static scan, and was evaluated with the aim of developing a more patient-friendly scanning protocol.
The outcome measure from the kinetic methods was binding potential, defined as BP2 = f 2·B max/K D, where f 2 is the free fraction of tracer in tissue, B max is the concentration of available binding sites and K D is the equilibrium dissociation constant [32]. BP2 is also known in the literature as V 3″.
SRTM has three free parameters: the relative plasma-to-tissue tracer delivery rate constant (R 1), the tissue-to-plasma transfer rate constant (k 2) and BP2. The number of free parameters in the other methods are 3, 2 and 1 for MLR, GA and RM, respectively. MLR was applied both on VOI data and on a voxel-by-voxel basis. To allow comparison of these two approaches, we calculated the average of the BP2 values (BP2,vox) of all voxels within each VOI used (BP2,VOI).
With RM, binding potential was calculated as follows:
where A T(t 1:t 2) and A R(t 1:t 2) are the mean activity concentrations in the time interval [t 1,t 2] in the target and reference region, respectively. BPRM was calculated for a series of time points, t, ranging from 90 to 240 min p.i. Δt was in all cases 60 min (which was the scanning time of choice for a single-scan protocol to be used in subsequent studies with this tracer). Since these time intervals would not exactly match the actual data acquisition, linear interpolation was used in the measured time-activity curves (TACs).
The different quantification methods were compared using correlation analysis.
Drug occupancy
Occupancy (E) of SERT by citalopram was calculated as follows:
where BBP2 and DBP2 are the BP values obtained from the baseline and post-citalopram scan, respectively. The estimated occupancy values were fitted to an E max model, described as follows:
where E max is the maximum occupancy, C p is plasma drug concentration and EC50 is the plasma drug concentration needed to reach 50% of E max. The goodness-of-fit was estimated as the standard deviation (SD) of the residuals.
We also investigated a method for improved estimation of drug occupancy by simultaneous modelling of the baseline and post-drug data, under the assumption that R 1 and k 2 were constant between the two scans, thereby reducing the total number of free parameters from six to four.
Handling of dropouts
The data from the subject who had only a baseline scan were used in the comparison of different methods for estimating the binding potential of [123I]ADAM. For the one scan in which partial data loss occurred, kinetic analysis was not possible. However, binding potential was estimated by the ratio method, and data from this subject were included in the drug occupancy versus plasma concentration analysis.
Results
The results obtained with the two different sets of VOIs were very similar. All results presented here were obtained with the second set of VOIs.
Tracer uptake
The average total brain uptake (±SD) of [123I]ADAM in the baseline scans was 4.1±0.8% of the injected activity. The highest tracer uptake was in midbrain. There was conspicuous uptake in thalamus and striatum, but lower than in midbrain. The average baseline binding potentials in the different regions were as follows (BP2±SD): 1.31±0.29 (MB), 0.79±0.16 (Thal), 0.66±0.13 (Str).
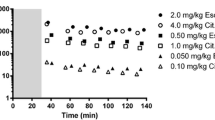
Figure 1 shows an example of TACs obtained in the baseline scan of one subject. TACs for cerebellum and MB are shown, as well as an SRTM fitted curve. (In general the measured TACs were a bit noisier than this.)
Tracer binding
There was a good agreement and correlation between BP2 values obtained with SRTM and with MLR. For the MB region the following relation was found:
There was also good correlation between SRTM and GA:
However, in this case the agreement is not as good as for MLR, since the slope is not as close to 1.There was a good agreement between voxel-based and VOI-based MLR analysis (Fig. 2). The relation between the two approaches was:
Figure 3 shows parametric BP2 images generated by MLR, averaged over 14 subjects after realignment based on parameters determined using the transmission maps. Both the baseline and the post-citalopram scans are shown.
Figure 4 shows the results of the correlation analysis comparing SRTM and RM in the MB region, including intercept of the regression line as well as the correlation coefficient (R 2), as a function of the starting point, t, of the RM time interval (Eq. 1). The intercept curve reaches a minimum at t=200 min, at which point R 2 is still close to its highest value. This means that the optimum time interval for the static scan protocol is [200,260] min p.i. Figure 5 shows BP values obtained by RM with data from this interval versus those obtained by SRTM for the MB region. The relation was:
Occupancy
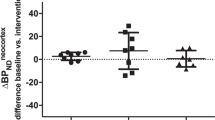
Figure 6 shows SERT occupancy as a function of plasma concentration of citalopram for MB. The occupancy in both regions shows an initial rapid increase and then plateaus at ∼80%. The fitted curves do not match the data very well, and the variability is large. The estimated curve-fit parameters were: E max=84.0%, EC50=2.47 ng/ml, SD=13.3%. Using the simultaneous fitting approach, the variability was reduced slightly (20% relative).
Discussion
Methodological considerations
Scatter correction amplifies noise and is often omitted from SPECT data processing. However, it has been shown that scatter correction can be important for the quantitation of dynamic SPECT studies using 123I-labelled tracers [33, 34], and it was thus included in the present study.
The main outcome measure in this study was occupancy of SERT by citalopram, defined as the relative change in the concentration of available binding sites (B max) after administration of the drug. For the purpose of estimating occupancy it is, however, not necessary to obtain absolute quantification of B max. In this study, the calculations were based on measured values of the binding potential BP2=f 2·B max/K D, where f 2 and K D were assumed to be constant and would therefore cancel out in Eq. 2, resulting in unbiased occupancy values.
In the comparison of different methods for estimating binding potential, we assumed that SRTM was the most robust and reliable of the methods used, although it may not be a true “gold standard”. However, any multiplicative factors influencing the BP2 values would cancel out in the occupancy calculation, in the same way as f 2 and K D. An additive term would, on the other hand, not cancel out but result in biased occupancy estimates. That is why, in the determination of the optimum scanning period for the single-scan protocol, it was important to minimise the intercept in the correlation of RM- and SRTM-determined BP2 values.
The occupancy determination would be sensitive to differences in VOI between the two scans for each subject. Nowadays it is common practice to generate ROI or VOI based on co-registered MRI images of each subject. In the present study, MRI data were not available, so the VOIs were drawn directly on the SPECT images. Since that procedure may be susceptible to errors, we used two different procedures as described above. The results from these two methods were very close, which makes it less likely that our estimated occupancy values would have been affected by VOI errors to any great degree.
We investigated the graphical methods GA and MLR, which are less time consuming than SRTM and therefore useful for voxel-by-voxel analysis. It has been reported that MLR suffers from a noise-dependent bias [35], which would make it unsuitable for voxel-based analysis. Our results showed good agreement between MLR and SRTM and also between voxel-based and VOI-based MLR analysis, indicating that this method is suitable for production of parametric images of [123I]ADAM.
Tracer binding
The average total brain uptake obtained in this study (4.1±0.8%) is in close agreement with the value of 4% reported by Kauppinen et al. [10].
We determined the optimum scanning time period for a single-scan protocol to be 200–260 min p.i. This is similar to what has been reported by other groups, i.e. 210 min p.i. [36] and 180–360 min p.i. [15], although one group chose an earlier time period, 150–210 min p.i. [14], which, according to our data, might lead to biased results.
The maximum occupancy of SERT by citalopram estimated here (E max=84%) is close to the value of 80% reported for SERT occupancy by clinical doses of citalopram in an [11C]DASB PET study [37].
Our main concern regarding the present results is the large variability in the calculated occupancy values and the poor agreement with the E max model. Several clinical studies using [123I]ADAM have reported negative findings (no difference between patient and control groups), which have been attributed to large inter-subject variability in tracer binding [38–41]. Inter-subject variability could be related to age [42] or genetic factors [43]. In this study each subject was used as his own control; therefore the calculated occupancy values should not be influenced by inter-subject variability in tracer binding, but would be affected by intra-subject (test–retest) variability. Catafau et al. [15] reported a test–retest variability of 25–37% for [123I]ADAM binding in midbrain in humans. This could explain the variability in our results, but it does not agree with the test–retest value of ∼5% obtained in non-human primate studies with [123I]ADAM [17]. A possible explanation for the discrepancy, and also for the above-mentioned large inter-subject variability in [123I]ADAM binding, could be the presence of variable amounts of labelled lipophilic metabolites. Estimation of tracer binding can be seriously compromised by lipophilic metabolites crossing the blood–brain barrier (BBB) [44, 45]. Highly variable metabolism was reported in a non-human primate study, although the labelled metabolites were not lipophilic and should therefore not cross the BBB [17]. However, in a study with [123I]ADAM in humans, a labelled lipophilic metabolite was detected in two out of six subjects [18]. Clarification of this issue will be important for future studies with this tracer.
Conclusion
Quantitative binding estimates of [123I]ADAM can be obtained using the ratio method with static data from a single scan at 200–260 min p.i. Parametric images of [123I]ADAM can be obtained by voxel-based multi-linear regression analysis of dynamic data. The estimated SSRI occupancy of SERT versus plasma drug concentration curves showed a high degree of variability, which could make this tracer less useful for studies where a detailed dose-response assessment is critical, although it might still be used to assess whether putative SSRIs achieve maximal occupancy at therapeutic doses.
References
Owens MJ, Nemeroff CB. The serotonin transporter and depression. Depress Anxiety 1998;8 Suppl 1:5–12
Mann JJ. Role of the serotonergic system in the pathogenesis of major depression and suicidal behavior. Neuropsychopharmacology 1999;21 Suppl 2:99S–105S
Dolberg OT, Iancu I, Sasson Y, Zohar J. The pathogenesis and treatment of obsessive-compulsive disorder. Clin Neuropharmacology 1996;19:129–47
Heinz A, Ragan P, Jones DW, Hommer D, Williams W, Knable MB, et al. Reduced central serotonin transporters in alcoholism. Am J Psychiatry 1998;155:1544–9
Raisman R, Cash R, Agid Y. Parkinson’s disease: decreased density of 3H-imipramine and 3H-paroxetine binding sites in putamen. Neurology 1986;36:556–60
Palmer AM, Francis PT, Benton JS, Sims NR, Mann DM, Neary D, et al. Presynaptic serotonergic dysfunction in patients with Alzheimer’s disease. J Neurochemistry 1987;48:8–15
Tejani-Butt SM, Yang J, Pawlyk AC. Altered serotonin transporter sites in Alzheimer’s disease raphe and hippocampus. Neuroreport 1995;6:1207–10
Mann JJ. The role of in vivo neurotransmitter system imaging studies in understanding major depression. Biol Psychiatry 1998;44:1077–8
Oya S, Choi SR, Hou C, Mu M, Kung MP, Acton PD, et al. 2-((2-((dimethylamino)methyl)phenyl)thio)-5-iodophenylamine (ADAM): an improved serotonin transporter ligand. Nucl Med Biol 2000;27:249–54
Kauppinen TA, Bergström KA, Heikman P, Hiltunen J, Ahonen AK. Biodistribution and radiation dosimetry of [123I]ADAM in healthy human subjects: preliminary results. Eur J Nucl Med Mol Imaging 2003;30:132–6
Newberg AB, Plossl K, Mozley PD, Stubbs JB, Wintering N, Udeshi M, et al. Biodistribution and imaging with 123I-ADAM: a serotonin transporter imaging agent. J Nucl Med 2004;45:834–41
Huang WS, Ma KH, Cheng CY, Chen CY, Fu YK, Chou YH, et al. Imaging serotonin transporters with 123I-ADAM brain SPECT in healthy non-human primates. Nucl Med Commun 2004;25:515–9
Lin KJ, Yen TC, Wey SP, Hwang JJ, Ye XX, Tzen KY, et al. Characterization of the binding sites for 123I-ADAM and the relationship to the serotonin transporter in rat and mouse brains using quantitative autoradiography. J Nucl Med 2004;45:673–81
Asenbaum S, Füger B, Zettinig G, Dudczak R. 123-I ADAM and SPECT for investigating serotonin transporter in human. Nuklearmedizin 2003;6:A158
Catafau AM, Perez V, Penengo MM, Bullich S, De-Juan R, Puigdemont D, et al. Characterization and long-term test-retest reliability of the serotonin transporter SPECT ligand 123I-ADAM in healthy volunteers. J Nucl Med 2004;45 Suppl:260P
Ahonen AK, Kauppinen TA, Heikman P, Bergström KA, Launes J, Nikkinen P, et al. 123I labeled ADAM - A selective novel radioligand for imaging of serotonin transporters in the human brain. J Nucl Med 2002;43:232P
Acton PD, Choi SR, Hou C, Plossl K, Kung HF. Quantification of serotonin transporters in nonhuman primates using [123I]ADAM and SPECT. J Nucl Med. 2001;42:1556–62
Frokjaer VG, Pinborg LH, Madsen J, Knudsen GM. Evaluation of the serotonin transporter ligand [123I]ADAM for SPECT studies in humans. Neuroimage 2004;22 Suppl 2:T176
Erlandsson K, Sivananthan T, Lui D, Townsend CE, Lucas R, Ell PJ. Development of a simplified scanning protocol for the serotonin transporter SPET tracer [123I]ADAM. Nucl Med Commun 2004;25:405
Erlandsson K, Warrington S, Sivananthan T, Lui D, Spezzi A, Townsend CE, et al. Measuring SSRI occupancy of SERT using the novel SPET tracer [123I]ADAM. J Nucl Med 2004;45:397P
Erlandsson K, Sivananthan T, Lui D, Spezzi A, Townsend CE, Mu S, et al. Estimation of SSRI occupancy of SERT using the novel SPET tracer [123I]ADAM and simultaneous modeling. Neuroimage 2004;22 Suppl 2:T144–5
Tatsumi M, Groshan K, Blakely RD, Richelson E. Pharmacological profile of antidepressants and related compounds at human monoamine transporters. Eur J Pharmacol 1997;340:249–58
Owens MJ, Morgan WN, Plott SJ, Nemeroff CB. Neurotransmitter receptor and transporter binding profile of antidepressants and their metabolites. J Pharmacol Exp Ther 1997;283:1305–22
Lange K, Fessler JA. Globally convergent algorithms for maximum a posteri transmission tomography. IEEE Trans Im Proc 1995;4:1430–8
Ogawa K, Harat Y, Ichihara T, Kubo A, Hashimoto S. A practical method for position-dependent Compton-scatter correction in single emission CT. IEEE Trans Med Im 1991;10:408–12
Kak AC, Slaney M. Principles of computerized tomographic imaging. New York: IEEE Press; 1988
Chang LT. A method for attenuation correction in radionuclide computed tomography. IEEE Trans Nucl Sci 1978;25:638–43
Lammertsma AA, Hume SP. Simplified reference tissue model for PET receptor studies. Neuroimage 1996;4:153–8
Logan J, Fowler J, Volkow ND, Wolf AP, Dewey SL, Schlyer DJ, et al. Graphical analysis of reversible radioligand binding from time-activity measurements applied to [N-11C-methyl]-(−)-cocaine PET studies in human subjects. J Cereb Blood Flow Metab 1990;10:740–7
Varga J, Szabo Z. Modified regression model for the Logan plot. J Cereb Blood Flow Metab 2002;22:240–4
Ichise M, Ballinger JR, Golan H, Vines D, Luong A, Tsai S, et al. Noninvasive quantification of dopamine D2 receptors with iodine-123-IBF SPECT. J Nucl Med 1996;37:513–20
Mintun MA, Raichle ME, Kilbourn MR, Wooten GF, Welch MJ. A quantitative model for the in vivo assessment of drug binding sites with positron emission tomography. Ann Neurol 1984;15:217–27
Kim KM, Watabe H, Onishi Y, Yonekura Y, Iida H. Effects of scatter correction on kinetic analysis in I-123 IBF SPECT study. Neuroimage 2002;16:S83
Fujita M, Varrone A, Kim KM, Watabe H, Zoghbi SS, Seneca N, et al. Effect of scatter correction on the compartmental measurement of striatal and extrastriatal dopamine D2 receptors using [123I]epidepride SPET. Eur J Nucl Med Mol Imaging 2004;31:644–54
Buchert R, Varga J, Mester J. Limitations of bi-linear regression analysis for the determination of receptor occupancy with positron emission tomography. Nucl Med Commun 2004;25:451–9
Dresel S, laFougere C, Makowski M, Meisenzahl E, Frodl T, Gildehaus F, et al. Imaging of the serotonin transporter in healthy controls: pharmacodynamics of [I-123]ADAM. J Nucl Med 2004;45 Suppl:261P
Meyer JH, Wilson AA, Ginovart N, Goulding V, Hussey D, Hood K, et al. Occupancy of serotonin transporters by paroxetine and citalopram during treatment of depression: a [11C]DASB PET imaging study. Am J Psychiatry 2001;158:1843–9
Asenbaum S, Fueger B, Zettinig G, Dudczak R. Serotonin transporter in human brain—application of 123I-ADAM, A new SPECT ligand. J Nucl Med 2003;44 Suppl:222P
Ahonen A, Koskela A, Kauppinen T, Puronto O, Nieminen-Wendt T, von Went L. Serotonin transporter availability in patients with Asperger syndrome; early results. Nuklearmedizin 2003;6:A158–9
Ahonen A, Heikman P, Kauppinen T, Koskela A, Bergström K. Serotonin transporter availability in drug free depression patients using a novel SERT ligand. Eur J Nucl Med Mol Imaging 2004;31 Suppl 2:S227–8
Koskela A, Kauppinen T, Keski-Rahkonen A, Sihvola E, Kaprio J, Bergström K, et al. Serotonin transporter availability in patients with bulimia nervosa using 123I-ADAM. Eur J Nucl Med 2004;31 Suppl 2:S228
Newberg A, Ploessl K, Wintering N, Alavi A, Udeshi M, Kung H. Age related changes in serotonin transporter binding measured by I-123 ADAM. J Nucl Med 2004;45 Suppl:279P
Mann JJ, Huang Y-Y, Arango V, Oquendo MA, Hastings R, Van Heertum RL, et al. A polymorphism of the serotonin transporter gene affects in vivo expression of transporter in human raphe nuclei as measured by positron emission tomography. Neuroimage 2004;22 Suppl 2:T47
Erlandsson K, Bressan RA, Mulligan RS, Ell PJ, Cunningham VJ, Pilowsky LS. Analysis of D2 dopamine receptor occupancy with quantitative SPET using the high affinity ligand [123I]epidepride: resolving conflicting findings. Neuroimage 2003;19:1205–14
Erlandsson K, Fujita M, Innis RB, Ell PJ, Pilowsky LS. The effect of lipophilic metabolites on reference tissue modelling of [123I]epidepride SPET data. J Nucl Med 2003;44:254P–5P
Acknowledgements
We want to thank the following for their invaluable help: At the INM, J. Bomanji, S. Gacinovic, S. Hughes, G. McNamara, N. Nagabhushan and R. Syed, and at HMR, S. Amin, S. Bakare, R. Dhadda, S. Eyers, A. Morgan, T. Nielsen, R. Ochiel, L. Stocking and P. Tsabedze. This study was conducted in accordance with applicable laws and regulations, good clinical practices, and the ethical principles that have their origin in the Declaration of Helsinki. The study was sponsored by Eli Lilly and Co. Ltd.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Erlandsson, K., Sivananthan, T., Lui, D. et al. Measuring SSRI occupancy of SERT using the novel tracer [123I]ADAM: a SPECT validation study. Eur J Nucl Med Mol Imaging 32, 1329–1336 (2005). https://doi.org/10.1007/s00259-005-1912-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00259-005-1912-y