Abstract
Porcine epidemic diarrhea virus (PEDV), an enteric coronavirus, induces severe vomiting and acute watery diarrhea in unweaned piglets. The pig industry has suffered tremendous financial losses due to the high mortality rate of piglets caused by PEDV. Consequently, a simple and rapid on-site diagnostic technology is crucial for preventing and controlling PEDV. This study established a detection method for PEDV using recombinase-aided amplification (RAA) and Pyrococcus furiosus Argonaute (PfAgo), which can detect 100 copies of PEDV without cross-reactivity with other pathogens. The entire reaction of RAA and PfAgo to detect PEDV does not require sophisticated instruments, and the reaction results can be observed with the naked eye. Overall, this integrated RAA-PfAgo cleavage assay is a practical tool for accurately and quickly detecting PEDV.
Key points
• PfAgo has the potential to serve as a viable molecular diagnostic tool for the detection and diagnosis of viral genomes
• The RAA-PfAgo detection technique has a remarkable level of sensitivity and specificity
• The RAA-PfAgo detection system can identify PEDV without needing advanced equipment
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Porcine diarrhea in the global swine industry is most commonly caused by PEDV, and PEDV is an enveloped virus that belongs to the family Coronaviridae and the genus Alphacoronavirus (Li et al. 2021a). PEDV infects pigs of all ages, causing vomiting, acute watery diarrhea, dehydration, and weight loss. The mortality rate of piglets reaches 100% (Tian et al. 2021). Since the discovery of PED in the UK in 1971 (Chasey and Cartwright 1978), it has spread to many countries, including the USA (Huang et al. 2013), Europe (Jung and Saif 2015), and China (Sun et al. 2012), where it has resulted in significant economic losses for the pig industry. However, PEDV is frequently co-infected with other coronaviruses in pig farms, including porcine delta coronavirus (PDCoV) and transmissible gastroenteritis virus (TGEV) (Ding et al. 2020a), which can cause similar clinical symptoms and make differential diagnosis challenging. Consequently, developing novel, simple, and effective approaches for detecting PEDV is urgently needed.
Currently, numerous techniques for detecting PEDV have been reported, including conventional virus isolation (Valkó et al. 2019), enzyme-linked immune sorbent (ELISA) (Gerber et al. 2014), and molecular biological techniques (Kim et al. 2007). Virus isolation and identification is a laborious process that takes a lot of time and necessitates specialized equipment and personnel; ELISA has been widely used to detect the level of antibodies in serum, but it was not a good choice for early diagnosis of viral infection as antibody generation required a few days, it was also laborious and time-consuming (Pewlaoo et al. 2022; Wang et al. 2020). Several molecular biological techniques commonly employed for PEDV diagnostics are quantitative polymerase chain reaction (qPCR) (Pan et al. 2020), recombinase polymerase amplification (RPA) (Li et al. 2021b), and loop-mediated isothermal amplification (LAMP) (Yu et al. 2015). Nevertheless, these technologies possess many drawbacks, including requiring skilled workers or costly and intricate equipment for qPCR (Yu et al. 2015), alongside elevated rates of false positives when solely employing RPA (Qian et al. 2023; Qin et al. 2022; Xu et al. 2022), complex primer design for LAMP (Kim et al. 2021; Li et al. 2022). The introduction and development of CRISPR diagnostic technology have provided us with an improved option for addressing the issues mentioned above. Bacteria and archaea possess adaptive immune systems known as clustered, regularly interspaced short palindromic repeat (CRISPR)-Cas systems. Currently, the CRISPR system consists primarily of Cas9, Cas12, Cas13, and Cas14. Cas9 and Cas12a are used primarily to cut double-stranded DNA, and Cas14a and Cas13a are primarily used to cut single-stranded DNA and single-stranded RNA, respectively. Currently, these nucleases are utilized in various virus detection techniques (Ding et al. 2020b; Kellner et al. 2019; Mali et al. 2013; Yang et al. 2021a). Current PEDV detection methods based on Cas12a are fast, specific, sensitive, and reliable; these attributes will be essential in the future development of point-of-care testing (POCT) molecular diagnostic technologies for viral infectious diseases (Liu et al. 2022; Qian et al. 2022; Yang et al. 2021b). However, the CRISPR system also has some limitations. Firstly, Cas proteins can function only at room temperature and become inactive at high temperatures, thereby influencing cleavage activity. Cas proteins require a section of gRNA guidance to cleave the target nucleic acid; however, RNA is susceptible to degradation, which makes storage and transport difficult. Additionally, gRNA synthesis is quite expensive.
A typical DNA-guided endonuclease in pAgos is the archaea Pyrococcus furiosus Argonaute (PfAgo), which cleaves the phosphodiester bond between the 10th and 11th bases of the target DNA from the 5′ end (Swarts et al. 2015). Compared to the CRISPR system, PfAgo only requires gDNA guidance to cut single-stranded DNA. PfAgo can withstand high temperatures, making it more suitable for transportation and preservation. PfAgo also provides greater gDNA selection versatility because it does not necessitate a PAM (protospacer adjacent motif) site. Applications of the PfAgo to molecular detections have emerged in recent years (He et al. 2019; Wang et al. 2021; Yang et al. 2023; Zhao et al. 2022).
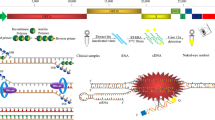
Recombinase-aided amplification (Wang et al. 2023) is an isothermal amplification technique that combines recombinant enzymes with other enzymes to achieve DNA or RNA amplification in vitro. In this study, the authors employed a combination of recombinase-aided amplification (RAA) and PfAgo to develop a novel detection technique for PEDV. The principle of the RAA-PfAgo process is presented in Fig. 1. In short, the PEDV RNA is isolated and then reverse-transcribed into cDNA. Subsequently, target fragments were amplified on cDNA using RAA at 39 °C for 30 min. PfAgo can specifically cleave the RAA amplified fragment strand that is complementary to the sequence of gDNA (the first cleavage) and release 16 bp of new gDNA by using the gDNAs guide, which is a set of gDNAs targeting the conserved region of the PEDV N gene (gDNA1 and gDNA2 are closely adjacent and complement to the N gene’s sense chain, while gDNA3 is complementary to the antisense chain). The newly released genomic DNA fragment possessing a 5′-phosphorylation can interact with unoccupied PfAgo proteins, resulting in the cleavage of molecular probe. The molecular probe that has been cleaved can emit fluorescence when exposed to blue light. This method can detect 100 copies of PEDV nucleic acid and has no cross-reactivity with other pathogens. The process can be completed using only two specific temperatures and does not necessitate costly equipment. This study offers significant evidence to support the effective management and containment of the PEDV outbreak.
Methods
The nucleic acid of viruses
TIANamp Virus DNA/RNA Kit (Tiangen, Beijing, China) was used to extract RNA or DNA from porcine epidemic diarrhea virus (PEDV), porcine deltacoronavirus (PDCoV), porcine transmissible gastroenteritis virus (TGEV), pseudorabies virus (PRV), porcine circovirus 2 (PCV2), and porcine circovirus 3 (PCV3). Then, cDNAs were synthesized with the PrimeScript RT Master Mix (Takara, Beijing, China) and stored at − 80 °C until use.
The utilization of oligonucleotide primers for amplification, gDNAs, and probe synthesis
A set of PCR and RAA primers was developed targeting the conserved region of the PEDV N gene. PCR-F/R was used to amplify the entire N gene of PEDV, while RAA-F/R was used for RAA amplification. Additionally, a conservative group of gDNAs was also designed to cleave the double-stranded DNA of RAA products, resulting in a 16 bp single-stranded DNA for probe reporter cleavage. The primers, 5′-phosphorylated guide DNAs (gDNAs), fluorescent single-stranded DNA (ssDNA) reporter (5′6-FAM-ssDNA-BHQ1-3′), and lateral flow dipstick test reporter (5′6-FAM-ssDNA-Biotin-3′) were manufactured by Sangon Biotech, located in Shanghai, China. The primers, gDNAs, and probe reporter sequences are provided in Table 1.
Construction of standard plasmid
In a reaction mixture of 20 μL, the components included 1 μL of template, 0.5 μL PCR-F and PCR-R (10 μM), 10 μL of 2 × Taq Plus Master Mix (Vazyme biotech co. Ltd, 137 Nanjing, China), and 8 μL of ddH2O. The PCR reaction was conducted with the following parameters: an initial denaturation step at 95 °C for 5 min, followed by 35 cycles of denaturation at 95 °C for 15 s, annealing at 60 °C for 15 s, and extension at 72 °C for 45 s. The entire N gene of PEDV was isolated and inserted into the pMD19-T plasmid. The concentration of the DNA was determined using a Nanodrop 2000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA) after being stored at − 20 °C.
Expression and purification of PfAgo
The pET28a-His-PfAgo was chemically synthesized by Sangon Biotech (Shanghai, China) based on the nucleic acid sequence of PfAgo (GenBank accession number: OR402834). Subsequently, the manufactured construct was introduced into E. coli Rosetta (DE3) pLysS Singles Competent Cells (Millipore, Burlington, MA, USA) by a transformation process. Overnight cultures (10 mL) were added to 400 mL LB broth (with kanamycin, 50 μg/mL) and grown at 37 °C to an OD600 of 0.6. Isopropyl-β-D-1-thiogalactopyranoside (IPTG, 1 mM) was incorporated into the culture medium, and induction was carried out at 16 °C and 180 rpm for 20 h. The supernatant protein was purified using a His-tag Protein Purification Kit (Sangon Biotech, Shanghai, China) after the cells were centrifuged at 15,000 g for 20 min at 4 °C. This study employed a 50 KD ultrafiltration column (Millipore, Darmstadt, Germany) to concentrate the pure target protein. Subsequently, the identification of the purified protein was conducted using SDS-PAGE.
The RAA reaction
RAA reaction mixture (ZC Bio-Sci &Tech Co. Ltd, Hangzhou, Zhejiang, China) containing primer RAA-F (10 μM, 2 μL), primer RAA-R (10 μM, 2 μL), buffer A (25 μL), ultrapure water (13.5 μL), buffer B (2.5 μL), and template DNA or cDNA (5 μL) was used to carry out the RAA reaction. Furthermore, to achieve the most favorable reaction conditions for the RAA process, a series of experiments were conducted at various temperatures (37, 38, 39, 40, 41, and 42 °C) and reaction durations (0, 20, 30, 40, and 50 min). The temperature and duration of the RAA reaction were controlled using PTC-200 thermal cyclers manufactured by MJ Research in Waltham, MA, USA. The resulting compounds were subsequently identified through the utilization of agarose gel electrophoresis.
The PfAgo reaction
The PfAgo reaction system (20 μL) which includes 2 μM PfAgo, 0.25 μM each gDNA (total three gDNAs), 0.5 μM probe, 0.5 mM Mncl2, and 10 μL RAA production. The tube was placed in PTC-200 Thermal cyclers (MJ Research, Waltham, MA, USA) at 95 °C for 30 min, and the fluorescence signal following the PfAgo-mediated cleavage reaction was observed under blue light.
RAA-PfAgo detection system evaluation
Determination of analytic sensitivities of RAA-PfAgo detection systems was carried out by using pMD19-PEDV-N DNA ranging from 1 × 104 to 1× 101 copies/μL and the RAA-PfAgo sensitivity reaction was conducted through PTC-200 Thermal cyclers (MJ Research, Waltham, MA, USA) or real-time fluorescence quantitative PCR instrument (CFX96TM, Bio-Rad, Hercules, CA, USA) at 95 °C for 30 min. The genomic cDNA or DNA of PRV, PCV2, PCV3, PEDV, TGEV, and PDCoV were utilized to test the detection system’s specificity, and the result was detected under blue light after 30 min of the RAA-PfAgo reaction at 95 °C.
The RAA-PfAgo-LFD reaction
The RAA-PfAgo-LFD reaction system (20 μL) including 2 μM PfAgo, 0.25 μM each gDNA (total three gDNAs), 125 nM probe, 0.5 mM Mncl2, and 10 μL RAA production. For 30 min, the tube was placed in PTC-200 Thermal cyclers (MJ Research, Waltham, MA, USA) at a temperature of 95 °C. After the RAA-PfAgo reaction, 30 μL of ultrapure water was added to the reaction tube, mixed well, and a test strip (JY030, Tiosbio, Beijing, China.) was inserted into the tube, followed by observing results after 5 min.
Identification of PEDV in clinical samples
Newborn piglets with diarrhea were collected from seven farms in Sichuan, China, yielding 53 clinical specimens (including feces and intestinal samples). Total RNAs were extracted from these samples and then reverse-transcribed to cDNA, serving as templates in PCR (Tian et al. 2021) or RAA-PfAgo/RAA-PfAgo-LFD assay.
Results
PfAgo expression and purification outcomes
The PfAgo protein expression was induced by 1 mM IPTG for 20 h at 16 °C. The pellets were resuspended after harvesting, then lysed, and finally concentrated. The expressed PfAgo (about 87 KDa) was dissolved primarily in the supernatant and eluted by elution buffer (Fig. 2).
The results of optimization of the RAA reaction
ddH2O and 1 × 105 copies/μL of the pMD19-PEDV-N DNA were used as the RAA template. After 30 min of RAA reaction at 39 °C, there were specifically sized bands in PEDV-positive DNA but none in the negative control (Fig. 3A), demonstrating the efficacy of the RAA primers (RAA-F/R). To enhance the efficiency of the RAA reaction, the reaction was conducted at various temperatures and durations to explore optimal conditions. According to the data, the optimum conditions for the RAA reaction were achieved at a temperature of 39 °C (Fig. 3B) and a reaction period of 30 min (Fig. 3C).
The results of the RAA-PfAgo detection system
PfAgo and RAA were combined to increase the sensitivity of PfAgo detection. The procedure is as follows: reverse transcription (RT) of the extracted viral RNA, amplification of the obtained cDNA by RAA at 39 °C for 30 min, mixing of RAA product with PfAgo, reacted at 95 °C for 30 min, and results were observed under the blue light (Fig 4A). For the RAA-PfAgo detection system, 1 × 105 copies/μL of the pMD19-PEDV-N DNA and ddH2O was used as the RAA template, and the amplification products were added to the PfAgo system to measure fluorescence value. This study’s findings demonstrate a significant increase in positive fluorescence values compared to negative fluorescence values when exposed to blue light (Fig. 4B). Additionally, the results depicted in Fig. 4C indicate that PfAgo could cleave the probe and generate fluorescence signals only in the presence of both gDNAs and templates. Based on these observations, it can be concluded that the RAA-PfAgo system can be utilized to detect PEDV. The sensitivity of the RAA-PfAgo reaction was evaluated using a 10-fold serially diluted template ranging from 1 × 104 to 1× 101 copies/μL of the pMD19-PEDV-N DNA. The obtained findings demonstrated the potential of the developed detection system to detect as low as 1 × 102 copies/μL of the dsDNA template (Fig. 4D) under blue light. This observed limit of detection (LOD) was consistent with the value detected by real-time fluorescence detection (Fig. 4E). Figure 4F demonstrates the capability of the RAA-PfAgo detection system to detect PEDV and no cross-reaction with other viruses. The results suggest that the established RAA-PfAgo detection method for PEDV has high sensitivity, specificity, and reproducibility.
Establishing the RAA-PfAgo assay. A Schematic representation of RAA-PfAgo assay to detect PEDV. B The outcome of using blue light to identify PEDV via the RAA-PfAgo reaction. C RAA-PfAgo reaction outcomes when gDNAs and templates are present at the same time. D The sensitivity of the RAA-PfAgo to detect PEDV under blue light. The detection limit of the RAA-PfAgo assay was determined by testing 10-fold subsequent dilutions of the DNA standard over a range of 104 to 10 copies/μL. E The sensitivity of the RAA-PfAgo to detect PEDV by real-time fluorescence detection, the detection limit of the RAA-PfAgo assay was determined by testing 10-fold successive dilutions of the DNA standard over a range of 104 to 10 copies/μL (n = 3 replicates; error bars represent SD. ***p < 0.001, two-sample t-test). F The RAA-PfAgo assay’s specificity for identifying PEDV in blue light. NC, negative control
The results of the RAA-PfAgo-LFD reaction
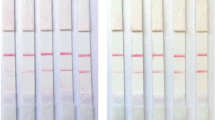
In the RAA-PfAgo-LFD method, the RNA of the virus was isolated and subsequently converted into cDNA using reverse transcription. The resulting cDNA was then used for amplification using the RAA technique, which was carried out at 39 °C for 30 min. The RAA product was added to the PfAgo system and allowed to react at 95 °C for 30 min, followed by an LFD reaction. If the 5′-6-FAM-ssDNA-Biotin-3′ reporter gene was utilized, the FAM modification group binds to the gold NP anti-FAM antibody, forming a complex caught by the antibody detection line. A chain avidin (control band) line can capture biotin-modified groups. Bands are seen exclusively at the control line when PfAgo did not destroy ssDNA reporter molecules and are captured by the streptavidin system. After the PfAgo-mediated degradation of the ssDNA reporter, although the streptavidin line (control band) captured the biotin-modified group owing to the cutoff of the ssDNA reporter molecule, the broken FAM-modified group can also be captured by the antibody detection line, resulting in the appearance of bands on both the control and the detection lines (Fig. 5A). As demonstrated in Fig. 5B, it can be observed that both the detection and control lines of the positive sample exhibited bands. However, only the control line of the negative ddH2O or no gDNAs control presented a band. This observation suggests that the developed RAA-PfAgo-LFD technique is feasible. The LOD of the RAA-PfAgo-LFD for PEDV was 1 × 102 copies/μL (Fig. 5C), and specific tests suggested that the method cannot detect other pathogens (Fig. 5D).
Establishing the RAA-PfAgo-LFD. A Schematic illustration of RAA-PfAgo-LFD assay to detect PEDV. B The result of the RAA-PfAgo-LFD reaction to detect PEDV. C The assay’s sensitivity in detecting PEDV using the RAA-PfAgo-LFD method. D The assay’s specificity in detecting PEDV using the RAA-PfAgo-LFD method. NC, negative control
The results of RAA-PfAgo/RAA-PfAgo-LFD to detect PEDV in clinical samples
A total of 53 typical clinical samples were successfully evaluated (nine were identified as PEDV positive by the PCR) to establish the validity of RAA-PfAgo tests. The RAA-PfAgo and RAA-PfAgo-LFD PEDV detection rates were consistent with the PCR (Table 2).
Discussion
PEDV is a highly contagious swine enteric infection that results in significant economic losses for the pig industry (Li et al. 2021a). Despite inactivated or attenuated vaccinations being used to combat PEDV (Li et al. 2017), PEDV is still prevalent worldwide, resulting in enormous economic losses to farmers. China has a large pig farming industry. Although agriculture is increasingly mechanized, not all farms have professional virus detection equipment and personnel. Virus detection requires the shipment of collected samples to professional laboratories, which frequently wastes significant human and material resources. Currently, the majority of available methods for PEDV surveillance have numerous drawbacks, such as insensitivity and unsuitability for field detection. Therefore, a simple, convenient, and fast detection method is urgently required for PEDV detection.
Isothermal amplification technology is a recent detection method that includes loop-mediated isothermal amplification (LAMP) (Yu et al. 2015), cross primer amplification (CPA) (Qiao et al. 2015), rolling circle amplification (RCA) (Zhang et al. 2021), and recombinase polymerase amplification (RPA). The limitations of LAMP (Kim et al. 2021; Li et al. 2022) include time-consuming primer design and vulnerability to nonspecific amplification. CPA (Frączyk et al. 2016) requires multiple primers, increasing reaction cost and complexity. One limitation of the RCA (Zhu et al. 2023) technique is its requirement for circular DNA as a template. In contrast, it should be noted that RPA (Wang et al. 2017) exhibits a notable degree and effectiveness in amplification. Furthermore, it is worth mentioning that RPA does not necessitate intricate primer design and can extract a greater quantity of target nucleic acids within a comparatively brief duration. However, because RPA is susceptible to false positives, it is frequently necessary to combine RPA and CRISPR systems to increase the specificity of the detection (Kellner et al. 2019; Wu et al. 2022). Currently, there are numerous detection methods for the combination of RPA and CRISPR (Chen et al. 2022a; Lei et al. 2022), but CRISPR has its limitations, such as the necessity for crRNA guidance to cut target nucleic acids or its sensitivity to high temperatures, which limits its application in testing.
When comparing CRISPR to PfAgo, it is evident that PfAgo exhibits the advantage of requiring a smaller amount of gDNA to cleave the target sequence. Furthermore, PfAgo and gDNA resist elevated temperatures, making PfAgo a more appropriate choice for regions with limited and distant resources (He et al. 2019; Swarts et al. 2015; Wang et al. 2021). In recent years, PfAgo-based detection methods such as PCR combined with PfAgo (He et al. 2019), LAMP combined with PfAgo (Xun et al. 2021), and RPA combined with PfAgo have emerged progressively (Yang et al. 2023; Zhao et al. 2022). Previous studies have described using RPA and PfAgo to detect enterocytozoon hepatopenaei (Yang et al. 2023). Although the technique exhibits exceptional specificity and sensitivity, including the PfAgo reaction after the purification of the RPA amplification product inevitably introduces a higher level of intricacy to the overall procedure. In our research, we focused on RAA because PCR requires complex instruments and a long time, while LAMP is quite challenging in constructing primers. RAA, similar to RPA, has the advantages of rapid nucleic acid amplification at room temperature and simple operation (Wu et al. 2022). However, due to the need for purification, nucleic acid electrophoresis, and the tendency for false positives to occur when using RAA alone (Chen et al. 2022b; Wang et al. 2023), we combine PfAgo with RAA to increase the convenience and specificity of detection based on the high sensitivity and specificity of PfAgo cleavage activity. In using RAA and PfAgo, a visual detection approach was developed for PEDV in this article. To minimize the possibility of false negative, we developed primers and gDNA based on the conservative PEDV N gene. Furthermore, the enhancement of RAA amplification efficiency and specificity was achieved by optimizing amplification settings. To streamline operations and minimize environmental pollution, the RAA product obtained from the reaction at 39 °C for 30 min was directly transferred to the PfAgo system without any purification steps. Subsequently, the transferred product was heated at 95 °C for an additional 30 min. The analysis of the reaction outcomes involved subjecting the substance to blue light. The detection strategy significantly benefits from its simplicity and versatility since it requires heating and blue light equipment. In order to enhance the examination of the outcomes, the RAA-PfAgo reaction was integrated with LFD (lateral flow dipstick) and assessed for its specificity, sensitivity, and practicality in the implemented approach. According to the results, RAA-PfAgo and RAA-PfAgo-LFD could detect 100 copies of viral nucleic acid without cross-reactivity with other organisms. Clinical samples were then utilized to assess the method’s dependability, which was compatible with the PCR results.
In conclusion, a simple, portable, economical, and highly sensitive detection platform was developed for PEDV. This technique provides new technical support for detecting and preventing PEDV without requiring precise instruments or complex experimental procedures.
Data availability
The data presented in this study are available on request from the corresponding author.
References
Chasey D, Cartwright SF (1978) Virus-like particles associated with porcine epidemic diarrhoea. Res Vet Sci 25:255–256. https://doi.org/10.1016/S0034-5288(18)32994-1
Chen J, Huang Y, Xiao B, Deng H, Gong K, Li K, Li L, Hao W (2022a) Development of a RPA-CRISPR-Cas12a assay for rapid, simple, and sensitive detection of Mycoplasma hominis. Front Microbiol 13:842415. https://doi.org/10.3389/fmicb.2022.842415
Chen Y, Zong N, Ye F, Mei Y, Qu J, Jiang X (2022b) Dual-CRISPR/Cas12a-assisted RT-RAA for ultrasensitive SARS-CoV-2 detection on automated centrifugal microfluidics. Anal Chem 94:9603–9609. https://doi.org/10.1021/acs.analchem.2c00638
Ding G, Fu Y, Li B, Chen J, Wang J, Yin B, Sha W, Liu G (2020a) Development of a multiplex RT-PCR for the detection of major diarrhoeal viruses in pig herds in China. Transbound Emerg Dis 67:678–685. https://doi.org/10.1111/tbed.13385
Ding X, Yin K, Li Z, Lalla RV, Ballesteros E, Sfeir MM, Liu C (2020b) Ultrasensitive and visual detection of SARS-CoV-2 using all-in-one dual CRISPR-Cas12a assay. Nat Ccmmun 11:4711. https://doi.org/10.1038/s41467-020-18575-6
Frączyk M, Woźniakowski G, Kowalczyk A, Niemczuk K, Pejsak Z (2016) Development of cross-priming amplification for direct detection of the African Swine Fever Virus, in pig and wild boar blood and sera samples. Lett Appl Microbiol 62:386–391. https://doi.org/10.1111/lam.12569
Gerber PF, Gong Q, Huang YW, Wang C, Holtkamp D, Opriessnig T (2014) Detection of antibodies against porcine epidemic diarrhea virus in serum and colostrum by indirect ELISA. Vet J 202:33–36. https://doi.org/10.1016/j.tvjl.2014.07.018
He R, Wang L, Wang F, Li W, Liu Y, Li A, Wang Y, Mao W, Zhai C, Ma L (2019) Pyrococcus furiosus Argonaute-mediated nucleic acid detection. Chem Commun 55:13219–13222. https://doi.org/10.1039/c9cc07339f
Huang YW, Dickerman AW, Piñeyro P, Li L, Fang L, Kiehne R, Opriessnig T, Meng XJ (2013) Origin, evolution, and genotyping of emergent porcine epidemic diarrhea virus strains in the United States. Mbio 4:e00737–e00713. https://doi.org/10.1128/mBio.00737-13
Jung K, Saif LJ (2015) Porcine epidemic diarrhea virus infection: etiology, epidemiology, pathogenesis and immunoprophylaxis. Vet J 204:134–143. https://doi.org/10.1016/j.tvjl.2015.02.017
Kellner MJ, Koob JG, Gootenberg JS, Abudayyeh OO, Zhang F (2019) SHERLOCK: nucleic acid detection with CRISPR nucleases. Nat Protoc 14:2986–3012. https://doi.org/10.1038/s41596-019-0210-2
Kim JK, Kim HR, Kim DY, Kim JM, Kwon NY, Park JH, Park JY, Kim SH, Lee KK, Lee C, Joo HD, Lyoo YS, Park CK (2021) A simple colorimetric detection of porcine epidemic diarrhea virus by reverse transcription loop-mediated isothermal amplification assay using hydroxynaphthol blue metal indicator. J Virol Methods 298:114289. https://doi.org/10.1016/j.jviromet.2021.114289
Kim SH, Kim IJ, Pyo HM, Tark DS, Song JY, Hyun BH (2007) Multiplex real-time RT-PCR for the simultaneous detection and quantification of transmissible gastroenteritis virus and porcine epidemic diarrhea virus. J Virol Methods 146:172–177. https://doi.org/10.1016/j.jviromet.2007.06.021
Lei R, Li L, Wu P, Fei X, Zhang Y, Wang J, Zhang D, Zhang Q, Yang N, Wang X (2022) RPA/CRISPR/Cas12a-based on-site and rapid nucleic acid detection of toxoplasma gondii in the environment. ACS Synth Biol 11:1772–1781. https://doi.org/10.1021/acssynbio.1c00620
Li C, Liang J, Yang D, Zhang Q, Miao D, He X, Du Y, Zhang W, Ni J, Zhao K (2022) Visual and rapid detection of porcine epidemic diarrhea virus (PEDV) using reverse transcription loop-mediated isothermal amplification method. Animals (Basel) 12:2712. https://doi.org/10.3390/ani12192712
Li D, Li Y, Liu Y, Chen Y, Jiao W, Feng H, Wei Q, Wang J, Zhang Y, Zhang G (2021a) Isolation and identification of a recombinant porcine epidemic diarrhea virus with a novel insertion in S1 domain. Front Microbiol 12:667084. https://doi.org/10.3389/fmicb.2021.667084
Li G, Wu M, Li J, Cai W, Xie Y, Si G, Xiao L, Cong F, He D (2021b) Rapid detection of porcine deltacoronavirus and porcine epidemic diarrhea virus using the duplex recombinase polymerase amplification method. J Virol Methods 292:114096. https://doi.org/10.1016/j.jviromet.2021.114096
Li Y, Wang G, Wang J, Man K, Yang Q (2017) Cell attenuated porcine epidemic diarrhea virus strain Zhejiang08 provides effective immune protection attributed to dendritic cell stimulation. Vaccine 35:7033–7041. https://doi.org/10.1016/j.vaccine.2017.10.052
Liu J, Tao D, Chen X, Shen L, Zhu L, Xu B, Liu H, Zhao S, Li X, Liu X, Xie S, Niu L (2022) Detection of four porcine enteric coronaviruses using CRISPR-Cas12a combined with multiplex reverse transcriptase loop-mediated isothermal amplification assay. Viruses (Basel) 14:833. https://doi.org/10.3390/v14040833
Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, Norville JE, Church GM (2013) RNA-guided human genome engineering via Cas9. Science 339:823–826. https://doi.org/10.1126/science.1232033
Pan Z, Lu J, Wang N, He WT, Zhang L, Zhao W, Su S (2020) Development of a TaqMan-probe-based multiplex real-time PCR for the simultaneous detection of emerging and reemerging swine coronaviruses. Virulence 11:707–718. https://doi.org/10.1080/21505594.2020.1771980
Pewlaoo S, Phanthong S, Kong-Ngoen T, Santajit S, Tunyong W, Buranasinsup S, Kaeoket K, Thavorasak T, Pumirat P, Sookrung N, Chaicumpa W, Indrawattana N (2022) Development of a rapid reverse transcription-recombinase polymerase amplification couple nucleic acid lateral flow method for detecting porcine epidemic diarrhoea virus. Biology (Basel) 11:1018. https://doi.org/10.3390/biology11071018
Qian B, Liao K, Zeng D, Peng W, Wu X, Li J, Bo Z, Hu Y, Nan W, Wen Y, Cao Y, Xue F, Zhang X, Dai J (2022) Clustered regularly interspaced short palindromic repeat/Cas12a mediated multiplexable and portable detection platform for GII genotype porcine epidemic diarrhoea virus rapid diagnosis. Front Microbiol 13:920801. https://doi.org/10.3389/fmicb.2022.920801
Qian W, Wang X, Huang J, Liu J, Chen S, Wang T, Zhang D, Li Y (2023) Sensitive and rapid RT-RPA-Cas12a-mediated detection method capable of human rhinovirus A and/or C species by targeting VP4. Virus Res 323:199001. https://doi.org/10.1016/j.virusres.2022.199001
Qiao B, Cui JY, Sun L, Yang S, Zhao YL (2015) Cross-priming amplification targeting the coagulase gene for rapid detection of coagulase-positive Staphylococci. J Appl Microbiol 119:188–195. https://doi.org/10.1111/jam.12836
Qin C, Liu J, Zhu W, Zeng M, Xu K, Ding J, Zhou H, Zhu J, Ke Y, Li LY, Sheng G, Li Z, Luo H, Jiang S, Chen K, Ding X, Meng H (2022) One-pot visual detection of african swine fever virus using CRISPR-Cas12a. Front Vet Sci 9:962438. https://doi.org/10.3389/fvets.2022.962438
Sun RQ, Cai RJ, Chen YQ, Liang PS, Chen DK, Song CX (2012) Outbreak of porcine epidemic diarrhea in suckling piglets, China. Emerg Infect Dis 18:161–163. https://doi.org/10.3201/eid1801.111259
Swarts DC, Hegge JW, Hinojo I, Shiimori M, Ellis MA, Dumrongkulraksa J, Terns RM, Terns MP, van der Oost J (2015) Argonaute of the archaeon Pyrococcus furiosus is a DNA-guided nuclease that targets cognate DNA. Nucleic Acids Res 43:5120–5129. https://doi.org/10.1093/nar/gkv415
Tian Y, Yang X, Li H, Ma B, Guan R, Yang J, Chen D, Han X, Zhou L, Song Z, Xie X, Wang H (2021) Molecular characterization of porcine epidemic diarrhea virus associated with outbreaks in southwest China during 2014-2018. Transbound Emerg Dis 68:3482–3497. https://doi.org/10.1111/tbed.13953
Valkó A, Albert E, Cságola A, Varga T, Kiss K, Farkas R, Rónai Z, Biksi I, Dán Á (2019) Isolation and characterisation of porcine epidemic diarrhoea virus in Hungary - short communication. Acta Vet Hung 67:307–313. https://doi.org/10.1556/004.2019.031
Wang F, Liang Q, Lv R, Ahmad S, Bano M, Weng G, Wen R (2023) Optimization and validation of reverse transcription recombinase-aided amplification (RT-RAA) for sorghum mosaic virus detection in sugarcane. Pathogens (Basel) 12:1055. https://doi.org/10.3390/pathogens12081055
Wang F, Yang J, He R, Yu X, Chen S, Liu Y, Wang L, Li A, Liu L, Zhai C, Ma L (2021) PfAgo-based detection of SARS-CoV-2. Biosens Bioelectron 177:112932. https://doi.org/10.1016/j.bios.2020.112932
Wang J, Wang J, Geng Y, Yuan W (2017) A recombinase polymerase amplification-based assay for rapid detection of African swine fever virus. Can J Vet Res 81:308–312
Wang X, He S, Zhao N, Liu X, Cao Y, Zhang G, Wang G, Guo C (2020) Development and clinical application of a novel CRISPR-Cas12a based assay for the detection of African swine fever virus. BMC Microbiol 20:282. https://doi.org/10.1186/s12866-020-01966-6
Wu X, Liu Y, Gao L, Yan Z, Zhao Q, Chen F, Xie Q, Zhang X (2022) Development and Application of a reverse-transcription recombinase-aided amplification assay for porcine epidemic diarrhea virus. Viruses (Basel) 14:591. https://doi.org/10.3390/v14030591
Xu B, Gong P, Zhang Y, Wang Y, Tao D, Fu L, Khazalwa EM, Liu H, Zhao S, Zhang X, Xie S (2022) A one-tube rapid visual CRISPR assay for the field detection of Japanese encephalitis virus. Virus Res 319:198869. https://doi.org/10.1016/j.virusres.2022.198869
Xun G, Lane ST, Petrov VA, Pepa BE, Zhao H (2021) A rapid, accurate, scalable, and portable testing system for COVID-19 diagnosis. Nat Commun 12:2905. https://doi.org/10.1038/s41467-021-23185-x
Yang H, Chen J, Yang S, Zhang T, Xia X, Zhang K, Deng S, He G, Gao H, He Q, Deng R (2021a) CRISPR/Cas14a-Based isothermal amplification for profiling plant microRNAs. Anal Chem 93:12602–12608. https://doi.org/10.1021/acs.analchem.1c02137
Yang K, Liang Y, Li Y, Liu Q, Zhang W, Yin D, Song X, Shao Y, Tu J, Qi K (2021b) Reverse transcription-enzymatic recombinase amplification coupled with CRISPR-Cas12a for rapid detection and differentiation of PEDV wild-type strains and attenuated vaccine strains. Anal Bioanal Chem 413:7521–7529. https://doi.org/10.1007/s00216-021-03716-7
Yang L, Guo B, Wang Y, Zhao C, Zhang X, Wang Y, Tang Y, Shen H, Wang P, Gao S (2023) Pyrococcus furiosus Argonaute combined with recombinase polymerase amplification for rapid and sensitive detection of Enterocytozoon hepatopenaei. J Agric Food Chem 71:944–951. https://doi.org/10.1021/acs.jafc.2c06582
Yu X, Shi L, Lv X, Yao W, Cao M, Yu H, Wang X, Zheng S (2015) Development of a real-time reverse transcription loop-mediated isothermal amplification method for the rapid detection of porcine epidemic diarrhea virus. Virol J 12:76. https://doi.org/10.1186/s12985-015-0297-1
Zhang K, Zhang H, Cao H, Jiang Y, Mao K, Yang Z (2021) Rolling circle amplification as an efficient analytical tool for rapid detection of contaminants in aqueous environments. Biosensors (Basel) 11:352. https://doi.org/10.3390/bios11100352
Zhao C, Yang L, Zhang X, Tang Y, Wang Y, Shao X, Gao S, Liu X, Wang P (2022) Rapid and sensitive genotyping of SARS-CoV-2 key mutation L452R with an RPA-PfAgo method. Anal Chem 94:17151–17159. https://doi.org/10.1021/acs.analchem.2c03563
Zhu Z, Guo Y, Wang C, Yang Z, Li R, Zeng Z, Li H, Zhang D, Yang L (2023) An ultra-sensitive one-pot RNA-templated DNA ligation rolling circle amplification-assisted CRISPR/Cas12a detector assay for rapid detection of SARS-CoV-2. Biosens Bioelectron 228:115179. https://doi.org/10.1016/j.bios.2023.115179
Acknowledgements
The authors would like to thank Tiejun Zhang, Cailiang Song, Kailu Wang, Wenjun Yan, and Chengyao Hou from College of Life Sciences, Sichuan University, for their help in data analysis.
Funding
This work was supported by the Sichuan Science and Technology Programs (2021ZDZX0010, 2020YFN0147, and 2021YFYZ0030).
Author information
Authors and Affiliations
Contributions
YZ, CZ, and BG designed research and conducted experiments. XY and HW wrote the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Ethics approval
This article contains no studies with human participants or animals performed by authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhao, Y., Zhou, C., Guo, B. et al. Pyrococcus furiosus Argonaute-mediated porcine epidemic diarrhea virus detection. Appl Microbiol Biotechnol 108, 137 (2024). https://doi.org/10.1007/s00253-023-12919-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00253-023-12919-0