Abstract
Alkyl hydroperoxide reductase (AhP), catalase G (KatG), and catalase E (KatE) are the main enzymes to scavenge the excessive hydrogen peroxide in E. coli. It was found the concentration of endogenous H2O2 was submicromolar in a mutant strain E. coli MG1655/ΔAhpΔKatEΔKatG, which was enough to cause damage to DNA and proteins as well as concomitant cell growth and metabolism. However, few studies explored how submicromolar intracellular hydrogen peroxide alters protein function and regulates the signaling pathways at the proteome level. In order to study the effect of endogenous oxidative stress caused by submicromolar hydrogen peroxide, this study first constructed a mutant strain E. coli MG1655/ΔAhpΔKatEΔKatG. Then, label-free quantitative proteomic analysis was used to quantify the differentially expressed proteins between the wild-type strain and the mutant strain. A total of 265 proteins were observed as differentially expressed proteins including 108 upregulated proteins and 157 downregulated proteins. Among them, three differentially expressed proteins were also validated by parallel reaction monitoring (PRM) methodology. The 265 differentially expressed proteins are not only involved with many metabolism pathways including the TCA cycle, the pentose phosphate pathway, and the glyoxylic acid cycle, but also activated the DNA repair and cellular antioxidant signaling pathway. These findings not only demonstrated that ahp, katE, and katG played the critical role in aerobic growth but also delineated proteins network and pathway regulated by submicromolar intracellular hydrogen peroxide, which allowed a deeper understanding of oxidative signaling in E. coli. The findings of this study also demonstrate that the mutant E. coli may serve as a cell model to investigate the effect of endogenous oxidative stress and downstream signaling pathways.
Key points
• The mutant strain E. coli MG1655/ΔAhpΔKatEΔKatG was constructed to study the effect of endogenous oxidative stress in E. coli.
• A total of 265 differentially expressed proteins were quantified and enriched in metabolic pathways and antioxidant systems by using label-free proteomics analysis.
• The findings of this study demonstrate that the mutant E. coli may serve as an effective tool to investigate the endogenous oxidative stress.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Reactive oxygen species (ROS) are continuously produced in prokaryotic cells during the aerobic respiration (Dixon and Stockwell 2014). They are a group of oxidative molecules that includes hydrogen peroxide (H2O2), superoxide anion, hydroxyl radicals (OH), and singlet oxygen (Imlay 2013). The excessive ROS not only is able to attack nucleic acid and alter protein function in cells (Temple et al. 2005) but also might even cause cell damage or cell death (Ezraty et al. 2017). Therefore, many living organisms have evolved the antioxidant defense systems that can scavenge the excessive ROS and maintain the redox balance in cells.
The OxyR system is a well-known antioxidant defense system in E. coli. Oxidized OxyR can promote the transcription and induce the expression of various enzymes, such as alkyl hydroperoxide reductase (AhP) encoded by ahp (Seaver and Imlay 2001a) and catalase G (KatG) encoded by katG (Panek and O'Brian 2004), both of which can scavenge H2O2. AhP, KatG, and RpoS-induced catalase E (KatE) encoded by katE are the three main enzymes to scavenge H2O2 in E. coli. When the intracellular H2O2 concentration is relatively low, AhP plays a critical role to remove the excess H2O2. However, catalases (KatG, KatE) occupy the dominant position as H2O2 level rises to 20 μM (Imlay 2013). Taking together, the three enzymes AhP, KatG, and KatE control the level of H2O2 synergistically in E. coli (Greenberg and Demple 1989) and prevent oxidative damage to the cell (Zheng et al. 2001).
In order to study the biological effect of hydrogen peroxide, E. coli is typically stimulated with a large dose of H2O2. For example, after addition of millimolar H2O2 into the bacterial culture medium, reactive oxygen species were elevated and many oxidatively damaged proteins were identified (Tamarit et al. 1998), and it was also observed that the OxyR antioxidant defense system has been activated in E. coli (Seaver and Imlay 2001b). A recent study also conducted time-lapse quantitative proteomic analysis on E. coli cells treated with 0.1 mM and 1 mM H2O2. The findings indicated non-lethal H2O2 boosted bacterial survival and AhP and KatG played an important role against oxidative stress (Rodriguez-Rojas et al. 2020).
However, most of aforementioned studies were typically used the millimolar hydrogen peroxide. It is a big challenge to study the oxidative damage with a small dose of H2O2, especially at close to physiological concentration. One of difficulties is the high scavenging activity within the cells, which causes the low dose of H2O2 to disappear rapidly. Park and his colleagues solved this problem by construction of mutants of E. coli, which lack AhP, KatG, and KatE enzymes to scavenge H2O2 (Park et al. 2005). It was found the concentration of endogenous H2O2 was submicromolar in E. coli, which was enough to cause damage to DNA and proteins as well as concomitant cell growth and metabolism (Varghese et al. 2007). So far, few studies explored how submicromolar endogenous hydrogen peroxide alters protein function and regulates the signaling pathways at the proteome level. Therefore, in the present work, we constructed the mutant strain E. coli MG1655/ΔAhpΔKatEΔKatG and applied label-free quantitative proteomic method to quantify the differentially expressed proteins between the wild-type strain and the mutant strain. It was found that the knockout mutant strain disrupted the redox balance and caused a series of adaptive responses. These findings not only demonstrated that ahp, katE, and katG played the critical role in growth but also delineated proteins network and pathway regulated by submicromolar intracellular hydrogen peroxide, which allowed a deeper understanding of oxidative signaling in E. coli.
Materials and methods
Bacterial strains and chemical reagents
The wild-type strain E. coli MG1655 is maintained by our laboratory. The mutant strain (E. coli MG1655/ΔAhpΔKatEΔKatG) was constructed by homologous recombination with plasmids. The two strains were transferred to sterile LB liquid medium and cultured at 37 °C with shaking to reach log phase. Each bacterial strain was mixed with an equal volume of sterile 50% glycerol, and then stored at −80°C for further use. Urea, dithiothreitol (DTT), and ammonium bicarbonate were purchased from Sigma-Aldrich (Steinheim, Germany). Trypsin was purchased from Promega (Madison, WI). Formic acid and acetonitrile (ACN) were purchased from Fisher Scientific Canada (Edmonton, Canada). The protease inhibitor was supplied from Roche (Mannheim, Germany). Water was obtained from a Milli-Q Plus purification system (Millipore, Bedford, MA). BCA protein assay kit and BCA peptide assay kit were purchased from Thermo Fisher Scientific.
Construction of the mutant strain of E. coli MG1655
The 50 bp sequences on both sides of the selected genes were selected as the upstream and downstream homologous recombination arms. Then, the gene target fragments were obtained by PCR using plasmids pKD3, pUCmT-Gm, and pKD4 as templates, respectively. Gene targeting fragments were used to electroporate the E. coli strain carrying helper plasmid pKD46 and positive clones were selected by the associated antibiotic resistance. Finally, the plasmid pCP20 was used to remove the resistance genes, and the mutant strain was obtained. The process of ahp, katE, and katG gene knockout sequentially followed the steps above. The primers are listed in Table S1. The mutation was confirmed by PCR and agarose gel electrophoresis of the PCR products.
Growth and cell morphology of wild-type and mutant strains
Both wild-type and mutant strains were cultured in sterilized fresh LB liquid medium at 37 °C with shaking at 150 rpm. The absorbance at OD600 for each strain was measured every hour by three biological replicates. In light of the values at OD600, the growth curves of both E. coli MG1655 and E. coli MG1655/ΔAhpΔKatEΔKatG were plotted. The two strains were grown to the exponential phase (OD600 ~0.6) and the cell morphology was visualized on a Leica TCS-SP5 microscope (Leica Microsystems, Mannheim, Germany). For proteomic analysis, each strain was prepared in four biological replicates. Every sample was centrifuged at 10000 g for 10 min at 4 °C and the pellet was collected and washed twice with 50 mM PBS.
Assay of hydrogen peroxide by horseradish peroxidase-amplex red
In the presence of H2O2, horseradish peroxidase (HRP) can oxidize amplex red (AR) to the fluorescent product resorufin (Messner and Imlay 2002). One milligram of AR was dissolved in 0.78 ml of DMSO, and 0.25 ml of this solution was then diluted into 24 ml of 50 mM potassium phosphate (KPi, pH 7.8) to generate a 50 μM stock solution. HRP was dissolved in 50 mM KPi (pH 7.8) to 0.01 mg/ml (Seaver and Imlay 2001a). The mixture contains 50 μM AR reagent and 0.01 mg/ml HRP in potassium phosphate and 50 μL samples. The fluorescence was measured with a fluorescence-based microplate reader using excitation at 530 nm and detection at 585 nm. The concentration of hydrogen peroxide of samples was calculated using a standard curve prepared with known concentrations of H2O2.
Protein extraction, digestion, and desalting
The bacterial pellet was resuspended in the lysis buffer (8 M Urea, 2 mM EDTA, 1 mM PMSF, 50 mM NH4HCO3). Then, the cells were disrupted on ice by sonication for 8 min. After that, the sample was centrifuged at 15000 g for 45 min at 4 °C, and supernatant was transferred into a new tube. The protein concentration was measured by the BCA protein method. Each sample was reduced with 10 mM DTT for 30 min at 56 °C, alkylated with 50 mM iodoacetamide in the dark for 30 min, and then diluted and digested for 16 h at 37°C by trypsin at an enzyme/protein ratio of 1:50. The residual trypsin activity was quenched by the addition of 3% formic acid (v/v). The peptides were desalted using a C18 solid-phase extraction (SPE) column (Supelco, Bellefonte, PA) and dried using a vacuum centrifuge. Peptide concentration was determined by the BCA peptide assay.
LC-MS/MS analysis
The peptide mixture was dissolved in water containing 0.1% FA and analyzed using an on-line U3000-nano coupled with an Orbitrap Q-Exactive HFX mass spectrometer (Thermo Fisher Scientific, Massachusetts, USA). Peptides were separated through using a 15 cm house-made C18 reversed-phase column (100-μm inner diameter, 1.9 μm resin) and a 90 min elution gradient. Mobile phase A consisted of 0.1% FA and H2O and mobile phase B consisted of 20% H2O and 80% ACN. A 90 min gradient (mobile phase B: 5% at 0 min, 10% at 16 min, 22% at 60 min, 35% at 78 min, 99% at 83 min, 99% at 85 min, 5% at 86 min, 0% at 90 min) was used at a flow rate of 300 nl/min. The data were acquired in a data-dependent mode. For mass spectrometry parameters, the m/z range was set to 350–1500 for the MS scan, and the accumulation time was 0.25 s. The top 20 most intense ions in MS1 were selected for MS/MS analysis and the dynamic exclusion time was 20 s.
Data analysis and bioinformatics
The RAW mass spectrometry files were processed using MaxQuant with an integrated Andromeda search engine using false discovery rate (FDR) <0.01 at protein and peptide level. Tandem mass spectra were searched against the Uniprot E.coli database (2019/10/29, taxonomy ID: 83333, 4,391 sequences) concatenated with a reverse decoy database. The following parameters were used: 20 ppm first search peptide tolerance, 4.5 ppm main search peptide tolerance; trypsin enzyme specificity, a maximum of two missed cleavages; fixed modification: Carbamidomethyl (C), variable modification: oxidation (M); the option of match between runs was enabled with a matching time window of 0.7 min and alignment window of 20 min. The other parameters in MaxQuant were set with default values. The built-in label-free quantification algorithm (LFQ) in Maxquant was applied to quantification (Cox et al. 2014). Then, the Maxquant results were imported into Perseus software for statistics analysis. The protein identifications classified as reverse, only identified by site and contaminants was excluded from the data. In the protein expression matrix, we filtered out the rows with missing values across the wild-type and mutant biological replicates, and then, expression values were log-transformed with base 2. In order to obtain the significantly differentially expressed proteins, two-tailed t tests were performed in Perseus using cutoffs set an FDR of 0.05 and S0 value of 2. The DAVID 6.8 bioinformatics tools (https://david.ncifcrf.gov/) were used for the analysis of functional categorization of differentially expressed proteins. GO (gene ontology) cellular components, GO molecular function, and KEGG (Kyoto encyclopedia of gene and genomes) pathways were used for further analysis (Huang et al. 2009). The results were visualized by the R package “ggplot2.” In addition, the Cytoscape (version 3.6.1) plugin ClueGO (version 2.5.4) and Cluepedia (version 1.5.4) were used to show the KEGG pathways and PPI (protein-protein interaction) of related proteins (Bindea et al. 2009). Only pathways with p values <0.05 were shown. The kappa score was set to 0.7, which was calculated based on the number of proteins shared between pathways, as an indicator of grouping pathways. The PPI information was downloaded from the STRING database. To reduce data redundancy, the kappa score of PPI was also set to 0.7.
Parallel reaction monitoring analysis
The differentially expressed proteins between the wild-type strain and the mutant strain were selected for further targeted quantification by parallel reaction monitoring (PRM). The Skyline software (MacCross Laboratory, University of Washington) was used to analyze the PRM data and calculate the normalized peak area generated from precursor ions in the wild-type strain and the mutant strain (MacLean et al. 2010). Relative protein quantification was calculated by the ratio of the peptide peak area of mutant strain to wild strain, and then, the ratios were log-transformed with base 2. The MS proteomics data for PRM were deposited in the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD021467.
Results
Effect of the knockout of the ahp, katE, katG gene on the growth and morphology and H2O2 concentration
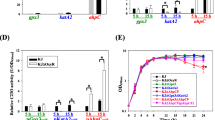
As shown in Fig. S1, the mutant strain has been successfully constructed. To determine if the lack of ahp, katE, and katG genes would affect the growth of E. coli, we monitored the growth of both the wild-type and mutant strains in LB medium at 37 °C under aerobic conditions. As shown in Fig. 1a, the mutant strain grew as well as the wild-type strain for the initial 2 h, but it grew slower than the wild strain for next three hours and spent much more time to reach the logarithmic phase, which is probably due to the accumulation of H2O2 in the mutant strain. As shown in Fig. 1b, the level of intracellular hydrogen peroxide was both submicromolar in two strains. The concentration of hydrogen peroxide in the mutant strain is significantly higher than that of the wild-type. The observed phenotype of mutant strain is consistent with previous studies (Park et al. 2005). Furthermore, the cell morphology of the two strains was observed by the microscope in the Fig. 1c and Fig. 1d. Compared with the wild-type, the mutant strain showed filamentation and aggregation, which is consistent with the Imlay’s research (Seaver and Imlay 2001a).
Comparison of wild-type and mutant strains. a Effect of the knockout of the ahp, katE, katG genes on the growth of the mutant strain of E.coli in LB medium. The mutant strain (blue) exhibits growth lag for 2–3 h compared with the wild-type strain (red) in the logarithmic phase in aerobic media. b The concentration of hydrogen peroxide in the two strains. The hydrogen peroxide level of the mutant strain (blue) is significantly higher than that of the wild-type strain (red). The cell morphology of the wild-type (C) and mutant strains (D) observed under a light microscope
Comparative proteomics analysis of total proteins between the wild-type and mutant strains
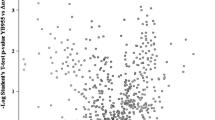
To investigate the effect of knockout genes on protein expression in E.coli, total proteins were extracted and quantified from the wild-type (Parent) and mutant strains (ΔAhpΔKatEΔKatG) using label-free quantitative proteomic strategy. Each group has four biological replicates. A total of 2335 proteins were identified by Maxquant. To obtain a general overview of the data quality, we performed principal components analysis (PCA), which revealed that the proteomes of each group cluster tightly and are distinct from other group (Fig. 2a). In addition, the Pearson correlation coefficient of intra-group and inter-group was ~0.99 and ~0.97, respectively, which indicated that reproducibility between biological replicates was robust (Fig. 2b). The heatmap showed the expression levels of identified proteins of the wild-type strain (Parent) and the mutant strain (ΔAhpΔKatEΔKatG), which also suggested a reliable quantification analysis (Fig. 2c). In order to obtain the significantly differentially expressed proteins, the volcano plot was made using cutoffs set an FDR of 0.05 and S0 value of 2. As shown in Fig. 3, 265 proteins were quantified as differentially expressed proteins between the wild-type strain and the mutant strain. Compared to the wild-type strain, 108 proteins were upregulated and 157 proteins were downregulated in mutant strain (Table S2, Table S3). It can be seen that the proteins with a larger fold-change and smaller p values appear in the upper left and right areas of the volcano plot (Fig. 3). As listed in Table 1, the upregulated proteins with large fold-change are associated with the antioxidant defense systems and DNA damage repair to prevent the cells from oxidative damage, while the downregulated proteins are associated with the metabolism of glutamate and glutathione.
Quality assessment of proteomic data between wild-type strain and mutant strain. Each strain has four biological replicates. a Principal components analysis of wild-type and mutant strains. The first and second components are shown. b Correlation of the protein intensity among four biological replicates of the wild-type strain and the mutant strain, as analyzed with R language. The Pearson color scale bar is shown below. c The heat map showing the 2335 proteins abundance identified in the wild-type strain and the mutant strain
a Volcano plot showing the proteins in the proteomic analysis of parent and mutant strains. The significantly differentially expressed proteins in blue dots and orange dots represent downregulated and upregulated proteins, respectively (S0 = 2; FDR = 5%). The non-significant proteins are shown in grey. b One hundred eight proteins were upregulated and 157 proteins were downregulated in the mutant strain, compared to the wild-type strain
Gene ontology analysis
In order to obtain functional information about differentially expressed proteins, GO and KEGG analysis were performed on the upregulated and downregulated proteins with p value less than 0.05. The total number of annotations of upregulated and downregulated protein enrichment is shown in Fig. 4a and Fig. 4b, respectively, which included four categories: biological processes, cellular components, molecular functions, and KEGG pathway. The top 10 terms for upregulated and downregulated proteins are shown in Fig. 4c and Fig. 4d, respectively. The top five biological processes enriched by the upregulated proteins were small molecule metabolism, redox, organic acid metabolism, keto acid metabolism, and carboxylic acid metabolism, while the downregulated proteins were enriched in cellular homeostasis. In the cellular components category, the upregulated proteins were located in the cytoplasm, cytosol, and cytoplasm, while the downregulated proteins were located in the cell and cell part. As for the molecular functions, the upregulated proteins were enriched in ion binding, metal ion binding, and cation binding, while the downregulated proteins were highly enriched in oxidoreductase activity. To further analyze the key enrichment pathways of differentially expressed proteins, we performed KEGG analysis. As shown in Fig. 4e, it can be observed that the differentially expressed proteins are involved in the TCA cycle, glyoxylic acid and dicarboxylic acid metabolism, and pyruvate metabolism. The partial list of differentially expressed proteins involved in energy metabolism was shown in Table 2. We also found that the carbon metabolism, amino acid biosynthesis, and secondary metabolite biosynthesis were enriched, indicating that genes knockout in the mutant may affect cell growth.
Gene ontology and KEGG pathway enrichment analysis of differentially expressed proteins. Charts showing the total number of enriched annotations for upregulated (a) and downregulated proteins (b). Four categories were considered including biological process, cell composition, molecular function, and KEGG pathway. The top 10 significantly enriched GO annotations associated with upregulated (c) and downregulated (d) proteins belonging to the term of biological process, cell composition, and molecular function. e The top eight enriched pathways are shown along with p value and gene number. The color represents the significance of enrichment
Protein-protein interaction and biological pathways networks
Proteins always assemble together and interact with each other to play a vital role in the cell. Therefore, we constructed a protein-protein interaction and biological pathways networks for the differentially expressed proteins to obtain a comprehensive understanding of how they work. As shown in Fig. 5, differentially expressed proteins between the wild-type strain and the mutant strain are enriched in ten pathways, including TCA cycle, glyoxylic acid and dicarboxylic acid metabolism, pentose phosphate metabolism pathway, pyruvate metabolism, alanine and aspartic acid and glutamic acid metabolism, glutathione metabolism, selenium-containing compound metabolism, nitrogen metabolism, and nitrotoluene degradation. Among these pathways, nearly half of the pathways are associated with energy metabolism, which are discussed in details as follows.
Protein-protein interaction and biological pathways networks for differentially expressed proteins of the wild-type strain and the mutant strain. The node size represented the percent of associated genes of proteins per node and the different pathways were displayed in different colors. Furthermore, the protein-protein interaction networks were constructed using the Cluepedia plugin and STRING database with STRING score threshold of 0.7. The protein-protein interactions were marked as gray line. It can be found that the protein-protein interaction maps were connected with ten different pathways, which provided new insights into protein functions, signaling pathways, and functional protein modules
Validation of differentially expressed proteins by PRM
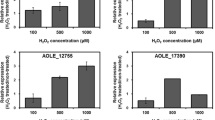
The label-free quantitative proteomic analysis revealed the differentially expressed proteins of the wild-type strain and the mutant strains. In order to validate the accuracy of label-free method, the upregulated proteins Antigen 43 (AG43), the downregulated protein 6-phosphogluconolactonase (6PgL) and one knockout protein Catalase G (KatG) were selected and quantified by PRM analysis. As expected, the amount of the Antigen 43 in the mutant strain was increased compared with the wild-type strain, while the abundance of 6-phosphogluconolactonase in the mutant strain was decreased. Moreover, the catalase G was also downregulated in the mutant strain according to the PRM data. Therefore, the quantification results of three proteins were in complete agreement with the label-free quantitative proteomic analysis data. The fold changes of three proteins by PRM were listed in Table 3.
Discussion
Alkyl hydroperoxide reductase, catalase G, and catalase E play a very important role in the process of scavenging H2O2 in E. coli. When the concentration of H2O2 in the cell reaches at 100 nM, the OxyR is oxidized and oxidized OxyR can induce the expression of ahp and katG (Varghese et al. 2007). Therefore, H2O2 can be quickly scavenged by enzymes to ensure that the cells are not damaged. In this study, we constructed a mutant strain E. coli MG1655/ΔAhpΔKatEΔKatG to explore how submicromolar endogenous hydrogen peroxide alters protein function and regulates the signaling pathways at the proteome level. A total of 265 differentially expressed proteins were identified in the mutant strain based on label-free proteomic strategy and then analyzed by the bioinformatic tools. Based on the differentially expressed proteins, as shown in Fig. 6, we proposed a new model to depict the responses of antioxidant defense system, DNA repair, and energy metabolic pathways in the mutant strain, indicating the implication for the roles of knockout genes in oxidative stress and metabolism.
The proposed model depicting the responses of antioxidant defense system, DNA repair, and energy metabolic pathways in the mutant strain, indicating the implication for the roles of knockout genes in oxidative stress and metabolism. The upregulated and downregulated proteins are shown in red and green circles, respectively. ROS including hydrogen peroxide (H2O2), superoxide anion(O2-), and hydroxyl radicals(·OH) were shown in yellow circle
The activation of antioxidant defense system
The mutant strain failed to scavenge excessive H2O2 rapidly due to the knockout of ahp, katE, and katG genes. In order to minimize the damage caused by H2O2, the antioxidant mechanism in the mutant strain was activated and the antioxidant enzymes were upregulated, such as thioredoxin Thio2, cytochrome C peroxidase homolog YhjA, and disulfide conjugate isomerase DsbG. Besides the OxyR and SoxR system, the antioxidant defense systems in E. coli also include the glutathione system, the thioredoxin system, and the cytochrome C peroxidase system (Staerck et al. 2017). These antioxidant defense systems were all activated when mutant strains were under oxidative stress and minimized the damage caused by H2O2 to cells.
It is well known that the OxyR system is a critical antioxidant defense system in E. coli. The OxyR is an important global regulator under oxidative stress. Based on the label-free quantitative proteomic method, we found AG43 outer membrane protein, glutathione reductase GshR, disulfide conjugate isomerase DsbG, thioredoxin 2 Thio2, ferric uptake regulation protein Fur, ferro chelatase HemH were upregulated. Previous studies have also found a number of proteins were induced by OxyR, including alkyl peroxidase, catalase G, AG43 outer membrane protein, and ferric uptake regulation protein, and glutathione reductase (Zheng et al. 2001). In this study, the level of hydrogen peroxide was increased in the mutant E.coli, and the cysteine residues in OxyR were oxidized to disulfide bonds (Imlay 2013). Although alkyl peroxidase and catalase G were knocked out, we still found other OxyR-induced proteins, such as AG43 outer membrane protein, were upregulated 39-fold compared with the control group. Interestingly, AG 43 protein is the most upregulated protein among the differentially expressed proteins and controls the morphological variation and self-aggregation of colonies. Ag43-mediated cell aggregation has a significant protective effect on hydrogen peroxide (Schembri et al. 2003). As shown in Fig. 1c and d, we observed the cell morphology of the two strains and found that the mutant strains showed filamentation and self-aggregation, and a previous study also found the mutant strain showed filaments when cultured under aerobic conditions (Varghese et al. 2007). When the OxyR system was activated, the DsbG encoded by the dsbG gene, one of the disulfide conjugate isomerases, was also upregulated in the mutant E. coli. The periplasmic protein DsbG contains a special catalytic motif to protect cysteine residues from oxidation (Ezraty et al. 2017). The oxidation of these cysteine residues will affect the function of protein. Thus, the upregulation of DsbG is possible to ensure the proteins perform function properly.
The glutathione system is one of the antioxidant defense systems in E. coli. It is composed of glutathione reductase, glutathione peroxidase, and glutaredoxin (Dickinson and Forman 2002). The glutathione reductase GshR was upregulated in mutant strain. It can catalyze the reduction of glutathione disulfide (GSSG) to glutathione GSH. Glutathione is capable of preventing damage caused by excessive hydrogen peroxide and maintains the reducing environment of the cell. The glutathione peroxidase is a cytosolic enzyme that catalyzes the reduction of hydrogen peroxide H2O2 to H2O via oxidation of reduced glutathione GSH into its disulfide form (GSSG). It is surprising that the amount of glutathione peroxidase BtuE decreased in the mutant strain, indicating there is an alternative way to remove the excessive H2O2. The glutaredoxin 2 Glrx2 and glutaredoxin 3 Glrx3 were both downregulated in mutant strain. The glutaredoxins appear to play a critical role in oxidative stress responses, which are able to catalyze the reduction of disulfides and convert oxidized proteins to reduced proteins. However, this reaction needs to consume glutathione. The downregulation of Glrx2 and Glrx3 in the mutant strain might decrease the consumption of glutathione to counteract oxidative stress.
The thioredoxin system is composed of several proteins, such as thioredoxin peroxidase, thioredoxin reductase (Hall et al. 2011), and OxyR-induced thioredoxin Thio2 (Lu and Holmgren 2014), which play a cytoprotective role in the cell. The expression of thioredoxin (Thio2) and the thioredoxin reductase (TrxB) was both upregulated in the mutant strain. Thio2 participates in various redox reaction (Carmel-Harel and Storz 2000) and reverse oxidized protein by initiating the reduction reaction (Collet and Messens 2010). TrxB can reduce oxidized thioredoxin to thioredoxin, ensuring that there is enough reduced thioredoxin to repair the oxidized protein. The upregulation of these two proteins may maintain cellular redox homeostasis and protect cells against various oxidative stresses.
As a cytochrome C peroxidase homolog, the YhjA protein was upregulated in the mutant E.coli, indicating the cytochrome C peroxidase system was probably activated. It has been reported that the YhjA protein can degrade a small dose of H2O2 in the E. coli periplasm (Khademian and Imlay 2017), which enable E. coli to use H2O2 as an anaerobic electron acceptor to respire through its linkage to the quinone pool.
DNA damage and DNA repair
Fur protein plays a key role in iron balance (Nunoshiba et al. 1999). It is a global negative control factor that can directly sense the level of iron in the cytoplasm. It has been reported that the fur gene is induced by OxyR. With the elevation of the hydrogen peroxide, the OxyR system is activated to induce the overexpression of fur gene. In mutant strains, the upregulated Fur protein can bind ferrous ions to decrease the Fenton reaction and reduce the cell damage. However, the overexpressed hydroxyl free radicals (·OH) still would attack the DNA and cause DNA damage, which is one of the possible reasons to slow down the mutant strain’s growth. Besides the changes in the expression of OxyR regulated genes, the expression of OxyR-independent genes also altered for oxidative defense (Zheng et al. 2001) such as RecA protein, which can specifically bind to the single-stranded DNA and activate DNA repair. When DNA is damaged, cell repair mechanisms are initiated. The upregulated RecA in the mutant strain acts as an activator of sos genes, which is required for homologous recombination (Mendoza-Chamizo et al. 2018). The expressions of OxyR regulated genes and OxyR-independent genes were summarized in Table S4 and Table S5.
Effects on metabolic pathways
As shown Fig. 4e and Fig. 5, the KEGG pathways and protein-protein interaction network were mapped in order to provide new insights into protein functions, signaling pathways, and functional protein modules. Among these pathways in the protein-protein interaction network, it is obviously found that the differentially expressed proteins are mainly involved in glucose metabolism pathways, especially in the TCA cycle, glyoxylate cycle, and pentose phosphate pathway. As shown in Fig. 6, all eight proteins involved in the TCA cycle were upregulated in the mutant strain, including citrate synthase CisY, succinate dehydrogenase SucC, anaerobic fumarate hydratase FumA, and malate enzyme Mdh. The glyoxylate cycle is a variation of TCA cycle. The expression of isocitrate lyase AceA and malate enzyme Mdh was both upregulated in glyoxylate cycle. It indicated that the TCA and glyoxylate cycle was both promoted, which allowed the cells to generate more energy against oxidative stress and maintain cell growth. From the Fig. 1a, it can be also found the mutant strain (red) grew slower than wild-type strain due to the accumulated ROS in the logarithmic phase. However, after a while, the mutant strain boosted the growth and reaches the stationary phase like the wild-type, which is probably due to the more energy production in vivo.
In the pentose phosphate pathway, six proteins were downregulated such as glucose dehydrogenase Dhg, transketolase 2 Tkt2, and Transaldolase A TalA. The phosphate pathway is regulated by intracellular NADPH concentration. In most cases, the concentration of NADPH is elevated under oxidative stress, while the concentration of NADH is reduced (Singh et al. 2008). NADPH is formed in the pentose phosphate pathway during reactions catalyzed by two enzymes encoded by zwf and gnd genes, expression of which was not affected in the mutant strain. In the pentose phosphate pathway, downregulated proteins including quinoprotein glucose dehydrogenase are not directly involved in the synthesis of NADPH. NADP-dependent malic enzyme Mao2 can catalyze the formation of NADPH and pyruvate from NADP and malic acid, which is upregulated in the mutant strain. NADPH might be overproduced due to the enhanced level of Mao2. Thus, the pentose phosphate pathway was inhibited to some extent.
In this study, the ahpC/F and katE and katG knockout E. coli is successfully constructed and validated how the cell compensates for the lack of the Ahp, KatG, and CatE enzymes, which constitute the first line of defense against hydrogen peroxide at the proteome level. By using the label-free proteomics strategy, a total of 265 proteins were observed as differentially expressed proteins between the wild-type and mutant strains including 108 upregulated proteins and 157 downregulated proteins. The differentially expressed proteins were involved in the activation of antioxidant defense system, DNA repair, and metabolism pathways, which indicated the redox balance was disrupted and caused a series of adaptive responses within the mutant strain. The results of this study not only improve our understanding of the mechanisms underlying the response of mutant E. coli to endogenous hydrogen peroxide but also suggest that the mutant E. coli become an effective model to investigate the oxidative stress effect, especially to the small dose of oxidant, and it may also serve as a potential tool to screen the antioxidant medicine in vitro.
Data availability
The label-free proteomics data have been deposited to the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org) via the PRIDE partner repository with the dataset identifier PXD020291. The MS proteomics data for PRM were deposited in the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD021467.
References
Bindea G, Mlecnik B, Hackl H, Charoentong P, Tosolini M, Kirilovsky A, Galon J (2009) ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25(8):1091–1093. https://doi.org/10.1093/bioinformatics/btp101
Carmel-Harel O, Storz G (2000) Roles of the glutathione- and thioredoxin-dependent reduction systems in the Escherichia coli and Saccharomyces cerevisiae responses to oxidative stress. Annu Rev Microbiol 54:439–461. https://doi.org/10.1146/annurev.micro.54.1.439
Collet JF, Messens J (2010) Structure, function, and mechanism of thioredoxin proteins. Antioxid Redox Signal 13(8):1205–1216. https://doi.org/10.1089/ars.2010.3114
Cox J, Hein MY, Luber CA, Paron I, Nagaraj N, Mann M (2014) Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, Termed MaxLFQ. Mol Cell Proteomics 13(9):2513–2526. https://doi.org/10.1074/mcp.M113.031591
Dickinson DA, Forman HJ (2002) Glutathione in defense and signaling - lessons from a small thiol. Ann N Y Acad Sci 973:488–504. https://doi.org/10.1111/j.1749-6632.2002.tb04690.x
Dixon SJ, Stockwell BR (2014) The role of iron and reactive oxygen species in cell death. Nat Chem Biol 10(1):9–17. https://doi.org/10.1038/nchembio.1416
Ezraty B, Gennaris A, Barras F, Collet JF (2017) Oxidative stress, protein damage and repair in bacteria. Nat Rev Microbiol 15(7):385–396. https://doi.org/10.1038/nrmicro.2017.26
Greenberg JT, Demple B (1989) A global response induced in Escherichia-Coli by redox-cycling agents overlaps with that induced by peroxide stress. J Bacteriol 171(7):3933–3939. https://doi.org/10.1128/jb.171.7.3933-3939.1989
Hall A, Nelson K, Poole LB, Karplus PA (2011) Structure-based insights into the catalytic power and conformational dexterity of peroxiredoxins. Antioxid Redox Signal 15(3):795–815. https://doi.org/10.1089/ars.2010.3624
Huang DW, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4(1):44–57. https://doi.org/10.1038/nprot.2008.211
Imlay JA (2013) The molecular mechanisms and physiological consequences of oxidative stress: lessons from a model bacterium. Nat Rev Microbiol 11(7):443–454. https://doi.org/10.1038/nrmicro3032
Khademian M, Imlay JA (2017) Escherichia coli cytochrome c peroxidase is a respiratory oxidase that enables the use of hydrogen peroxide as a terminal electron acceptor. Proc Natl Acad Sci U S A 114(33):E6922–E6931. https://doi.org/10.1073/pnas.1701587114
Lu J, Holmgren A (2014) The thioredoxin antioxidant system. Free Radic Biol Med 66:75–87. https://doi.org/10.1016/j.freeradbiomed.2013.07.036
MacLean B, Tomazela DM, Shulman N, Chambers M, Finney GL, Frewen B, MacCoss MJ (2010) Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26(7):966–968. https://doi.org/10.1093/bioinformatics/btq054
Mendoza-Chamizo B, ner-Olesen A, Charbon G (2018) Coping with reactive oxygen species to ensure genome stability in Escherichia coli. Genes-basel 9(11). https://doi.org/10.3390/genes9110565
Messner KR, Imlay JA (2002) In vitro quantitation of biological superoxide and hydrogen peroxide generation. Methods Enzymol 349:354–361. https://doi.org/10.1016/S0076-6879(02)49351-2
Nunoshiba T, Obata F, Boss AC, Oikawa S, Mori T, Kawanishi S, Yamamoto E (1999) Role of iron and superoxide for generation of hydroxyl radical, oxidative DNA lesions, and mutagenesis in Escherichia coli. J Biol Chem 274(49):34832–34837. https://doi.org/10.1074/jbc.274.49.34832
Panek HR, O'Brian MR (2004) KatG is the primary detoxifier of hydrogen peroxide produced by aerobic metabolism in Bradyrhizobium japonicum. J Bacteriol 186(23):7874–7880. https://doi.org/10.1128/Jb.186.23.7874-7880.2004
Park S, You XJ, Imlay JA (2005) Substantial DNA damage from submicromolar intracellular hydrogen peroxide detected in Hpx- mutants of Escherichia coli. Proc Natl Acad Sci U S A 102(26):9317–9322. https://doi.org/10.1073/pnas.0502051102
Rodriguez-Rojas A, Kim JJ, Johnston PR, Makarova O, Eravci M, Weise C, Hengge R, Rolff J (2020) Non-lethal exposure to H2O2 boosts bacterial survival and evolvability against oxidative stress. PLoS Genet 16(3):1553–7404. https://doi.org/10.1371/journal.pgen.1008649
Schembri MA, Hjerrild L, Gjermansen M, Klemm P (2003) Differential expression of the Escherichia coli autoaggregation factor antigen 43. J Bacteriol 185(7):2236–2242. https://doi.org/10.1128/Jb.185.7.2236-2242.2003
Seaver LC, Imlay JA (2001a) Alkyl hydroperoxide reductase is the primary scavenger of endogenous hydrogen peroxide in Escherichia coli. J Bacteriol 183(24):7173–7181. https://doi.org/10.1128/Jb.183.24.7173-7181.2001
Seaver LC, Imlay JA (2001b) Hydrogen peroxide fluxes and compartmentalization inside growing Escherichia coli. J Bacteriol 183(24):7182–7189. https://doi.org/10.1128/Jb.183.24.7182-7189.2001
Singh R, Lemire J, Mailloux RJ, Appanna VD (2008) A novel strategy involved anti-oxidative defense: the conversion of NADH into NADPH by a metabolic network. PLoS One 3(7):e2682. https://doi.org/10.1371/journal.pone.0002682
Staerck C, Gastebois A, Vandeputte P, Calenda A, Larcher G, Gillmann L, Papon N, Bouchara JP, Fleury MJJ (2017) Microbial antioxidant defense enzymes. Microb Pathog 110:56–65. https://doi.org/10.1016/j.micpath.2017.06.015
Tamarit J, Cabiscol E, Ros J (1998) Identification of the major oxidatively damaged proteins in Escherichia coli cells exposed to oxidative stress. J Biol Chem 273(5):3027–3032. https://doi.org/10.1074/jbc.273.5.3027
Temple MD, Perrone GG, Dawes IW (2005) Complex cellular responses to reactive oxygen species. Trends Cell Biol 15(6):319–326. https://doi.org/10.1016/j.tcb.2005.04.003
Varghese S, Wu A, Park S, Imlay KRC, Imlay JA (2007) Submicromolar hydrogen peroxide disrupts the ability of Fur protein to control free-iron levels in Escherichia coli. Mol Microbiol 64(3):822–830. https://doi.org/10.1111/j.1365-2958.2007.05701.x
Zheng M, Wang X, Templeton LJ, Smulski DR, LaRossa RA, Storz G (2001) DNA microarray-mediated transcriptional profiling of the Escherichia coli response to hydrogen peroxide. J Bacteriol 183(15):4562–4570. https://doi.org/10.1128/Jb.183.15.4562-4570.2001
Acknowledgments
We also gratefully acknowledge the technical support for mass spectrometer from the Analysis & Testing Center of Beijing Institute of Technology.
Funding
This work was supported by the National Key Research and Development Program of China (Grant NO. 2017YFC0108504).
Author information
Authors and Affiliations
Contributions
Y.Z. and Y.D. conceived and designed research. F.L., J.H., and G.C. conducted experiments. F.L. and R.M. analyzed data. F.L. wrote the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(PDF 498 kb)
Rights and permissions
About this article
Cite this article
Liu, F., Min, R., Hong, J. et al. Quantitative proteomic analysis of ahpC/F and katE and katG knockout Escherichia coli—a useful model to study endogenous oxidative stress. Appl Microbiol Biotechnol 105, 2399–2410 (2021). https://doi.org/10.1007/s00253-021-11169-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-021-11169-2