Abstract
Fucosyl-N-acetylglucosamine disaccharides are present in many biologically important oligosaccharides, such as human milk oligosaccharides, Lewis carbohydrate antigens, and glycans on cell-surface glycoconjugate receptors, and thus have vast potential for infant formulas, prebiotics, and pharmaceutical applications. In this work, in order to screen biocatalysts for enzymatic synthesis of fucosyl-N-acetylglucosamine disaccharides, we performed sequence analysis of 12 putative and one known α-L-fucosidases of Bacteroides fragilis NCTC9343 and constructed a phylogenetic tree of the nine GH29 α-L-fucosidases. After that, five GH29A α-L-fucosidases were cloned, and four of them were successfully heterogeneous expressed and screened for transglycosylation activity, and a GH29A α-L-fucosidase (BF3242) that synthesized a mix of Fuc-α-1,3/1,6-GlcNAc disaccharides using pNPαFuc as donor and GlcNAc as acceptor was characterized. The effects of initial substrate concentration, pH, temperature, and reaction time on its transglycosylation activity were studied in detail. Under the optimum conditions of 0.05 U/mL enzyme, 20 mM pNPαFuc, and 500 mM GlcNAc in sodium buffer (pH 7.5) at 37 °C for 45 min, BF3242 efficiently synthesized Fuc-α-1,3/1,6-GlcNAc at a maximum yield of 79.0% with the ratio of 0.48 for 1,3/1,6. The molecular dynamics simulation analysis revealed that Loop-4 (His220-Ser245) in the putative 3D model of BF3242 displayed significant changes throughout the thermal simulations, might being responsible for the changes in the ratio of two regioisomeric products at different temperatures. This work provided not only a potential synthetic tool for enzymatic synthesis of fucosyl-N-acetylglucosamine disaccharides but also a possibility for the formation of regioisomeric products in glycosidase-catalyzed transglycosylation.
Key points
• Sequence analysis of α-L-fucosidases of Bacteroides fragilis NCTC9343
• Obtainment of an α-L-fucosidase with high transglycosylation activity
• Explanation why temperature affected the ratio of two regioisomeric products
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fucosyl-N-acetylglucosamine disaccharides have attracted great interest because of their presence in many biologically important oligosaccharides. Fucosyl-α-1,3-N-acetylglucosamine (Fuc-α-1,3-GlcNAc) and fucosyl-α-1,4-N-acetylglucosamine (Fuc-α-1,4-GlcNAc) are found in the key structures of human milk oligosaccharides (HMOs) which have been shown to exhibit prebiotic effects (Bode 2012; Kobata 2010), and they are also present in type I and type II Lewis carbohydrate antigens that may be used as the tumor biomarkers (Cordon-Cardo et al. 1986; Mariano et al. 2000; Tübel et al. 2012; Yu et al. 2012). Additionally, a recent work has further demonstrated that Fuc-α-1,3-GlcNAc, naturally present in HMOs, can be used as an effective prebiotic for enrichment of specific Lactobacillus and Bifidobacterium bacterial species, while fucosyl-α-1,6-N-acetylglucosamine (Fuc-α-1,6-GlcNAc), usually present in the core structure of N-Glycan of glycoproteins in mammal, can be used as anti-adhesin that provides decoy binding sites for enteropathogens to inhibit adhesion of different extents of certain enteropathogenic Escherichia coli strains to human colon adenocarcinoma epithelial (HT29) cells (Becerra et al. 2015).
Currently, chemical method for synthesizing fucosidic bonds is challenging because of its acid lability (Yu et al. 2012). As an alternative, enzymatic synthesis of fucosidic bonds can be accomplished through a simple step under mild, environmental-friendly conditions using fucosyltransferase or α-L-fucosidase enzymes (Chen et al. 2015; Li et al. 2017a, b; Rodríguez-Díaz et al. 2013; Yang et al. 2017; Yu et al. 2017). Fucosyltransferases effectively catalyze stereo- and regioselective reactions both in vivo and in vitro, but their applications are somewhat hampered by the high cost of guanosine 5′-diphosphate-L-fucose (GDP-Fuc) (Tseng et al. 2017; Weijers et al. 2008). Usually, α-L-fucosidases are responsible for hydrolyzing fucosidic bonds in vivo (Fan et al. 2016; Li et al. 2017a, b). Recently, some works describe the synthesis of fucosidic bonds by α-L-fucosidases through transglycosylation in vitro (Ajisaka and Shirakabe 1992; Ajisaka et al. 1998; Becerra et al. 2020; Eneyskaya et al. 2001; Liu et al. 2016; Murata et al. 1999; Rodríguez-Díaz et al. 2013; Vetere et al. 1997). By using of comparatively inexpensive glycosyl donor pNPαFuc, the synthesis of fucosylated oligosaccharides by α-L-fucosidases is more economical (Crout and Vic 1998; Guzmán-Rodríguez et al. 2018).
Based on the amino acid sequence similarities and the mechanisms of hydrolysis, the α-L-fucosidases can be classified into the glycoside hydrolase (GH) families 29 and 95 in the CAZy database (http://www.cazy.org). GH29 α-L-fucosidases are configuration-retaining enzymes and catalyze the hydrolysis reactions via a double-displacement mechanism, resulting in retention of anomeric stereochemistry (Zechel and Withers 2000). During the double-displacement reaction, firstly, the general acid catalyst protonates the glycosidic oxygen with bond cleavage and the catalytic nucleophile directly attacks at the anomeric center of the sugar residue to form a covalent glycosyl-enzyme intermediate. Subsequently, the carbohydrate moiety was released with the general base catalyst deprotonating the water (Nakai et al. 2010; Okuyama et al. 2017). GH29 family has been further divided into two subfamilies GH29A and GH29B according to phylogenetic relationships and substrate specificities (Ashida et al. 2009; Sakurama et al. 2012a). GH29A α-L-fucosidases (EC 3.2.1.51) have more relaxed substrate specificities than GH29B enzymes, and can hydrolyze α-1,2, α-1,3, α-1,4, and/or α-1,6 fucosidic linkages as well as artificial pNPαFuc (Dawson and Tsay 1977; DiCioccio et al. 1982; Eneyskaya et al. 2001; Lezyk et al. 2016; Li et al. 2017a; Liu et al. 2016; Megson et al. 2015; Rodríguez-Díaz et al. 2011). GH29B α-L-fucosidases, referred to as 1,3/1,4-α-L-fucosidases (EC 3.2.1.111), can specifically hydrolyze α-1,3 or α-1,4 fucosidic linkages, but not pNPαFuc (Sakurama et al. 2012b). In contrast, GH95 α-L-fucosidases (1,2-α-L-fucosidases, EC 3.2.1.63) mediate an inversion mechanism to specifically hydrolyze terminal α-1,2 linkages and also do not hydrolyze pNPαFuc (Katayama et al. 2004). Although α-L-fucosidases are widely distributed in microorganisms, plants, and animals, only GH29A α-L-fucosidases have been reportedly capable of hydrolyzing the artificial substrate pNPαFuc, and therefore have the potential to synthesize fucosylated compounds. Using pNPαFuc as fucosyl donor, GH29A α-L-fucosidases have been successfully used to synthesize fucosyl-N-acetylglucosamine disaccharides at 20 to 58% yields (Ajisaka and Shirakabe 1992; Ajisaka et al. 1998; Eneyskaya et al. 2001; Rodríguez-Díaz et al. 2013; Vetere et al. 1997), and other fucosylated oligosaccharides or fucosylated compounds, such as fucosyl-N-acetyllactosamine and its methyl derivative (Murata et al. 1999), fucosyllactose and its methyl derivative (Lezyk et al. 2016; Murata et al. 1999), fucosylgalactoside and its methyl derivative (Ajisaka and Shirakabe 1992; Becerra et al. 2020), fucosylglucoside (Ajisaka and Shirakabe 1992; Becerra et al. 2020), fucosyl-diacetylchitobiose (Becerra et al. 2020), fucosyl-N-acetylglucosamine-asparagine (Becerra et al. 2020), and 4-methylumbelliferone-α-L-fucoside (Liu et al. 2016). Therefore, the exploitation of α-L-fucosidases with high transfucosylation activity would be of importance for biosynthetic purposes.
Bacteroides fragilis is a gram-negative anaerobic bacterium and primarily colonizes in the human lower gastrointestinal tract. A large portion of its genome is devoted to encoding carbohydrate metabolism, such as the degradation of dietary polysaccharides (Troy and Kasper 2010). B. fragilis has been proved as an excellent source for glycosidases. The hydrolysis activities of several glycosidases have been reported. A fructanase (FruA1) from B. fragilis BF-1 was able to hydrolyze sucrose, raffinose, inulin, and levan but not melezitose (Blatch and Woods 1993). Two sialidases purified from B. fragilis SBT3182 preferentially hydrolyzed sialyl α-2,8 linkage rather than α-2,3 and α-2,6 bonds (Tanaka et al. 1994). A recombinant sialidase (rNanH1) from B. fragilis YHC46 also preferentially hydrolyzed sialyl α-2,8 linkage to cleave sialic acids from mucin and serum proteins (Yamamoto et al. 2018). Two α-galactosidases (BfGal110A and BfGal110B) from B. fragilis NCTC9343 were identified to be general α-1,3-linkage-specific galactosidases for hydrolyzing blood group B antigens (Liu et al. 2008). A mannanase (BfMan26) from B. fragilis NCTC9343 was found to produce mannobiose exclusively from mannans (Kawaguchi et al. 2014). An α-L-fucosidase (BF3242) from B. fragilis NCTC9343 was capable of efficient removal of the core fucose and hydrolyzed other fucosidic linkages in various glycans, glycocopeptides, and glycoproteins (Tsai et al. 2017). Recently, we have reported several works on glycosidases with transglycosylation activity from B. fragilis NCTC9343, including the identification of transglycosylation activity of four β-N-acetylhexosaminidases (BF0669, BF0953, BF1181, and BF4033) (Chen et al. 2016), and the discovery of an α-galactosidase (AgaBf3S) and an exo-α-sialidase (BfGH33C) for the production of globotriose and 6′-sialyllactose, respectively (Gong et al. 2016; Guo et al. 2018). However, attempts to study the transglycosylation activity of α-L-fucosidases of B. fragilis have not been made so far.
In this work, in order to screen GH29A α-L-fucosidases for enzymatic synthesis of fucosyl-N-acetylglucosamine disaccharides using pNPαFuc as donor, through sequence analysis and phylogenetic tree construction, five GH29A α-L-fucosidases were selected as the candidates from 12 putative and one known α-L-fucosidases of B. fragilis NCTC9343 for gene cloning and heterogeneous expression in E. coli, four of which were expressed successfully as soluble proteins and the recombinant enzymes were purified and determined for hydrolysis activity and transglycosylation activity. As a result, an α-L-fucosidase (BF3242) with high transfucosylation activity was obtained. BF3242 synthesized Fuc-α-1,3/1,6-GlcNAc at a maximum yield of 79.0% with pNPαFuc as donor and GlcNAc as acceptor, and the ratio of two regioisomeric products could be remarkably affected by the reaction temperature. Furthermore, the molecular dynamics (MD) simulation analysis provided a possibility for regioisomeric products formation in glycosidase-catalyzed transglycosylation.
Materials and methods
Materials
Antigen H disaccharide, Lea and Lex trisaccharides, Fuc-α-1,3-GlcNAc, Fuc-α-1,4-GlcNAc, Fuc-α-1,6-GlcNAc, and N-acetyl-D-glucosamine (GlcNAc) were purchased from Carbosynth (Berkshire, UK). p-Nitrophenyl-α-L-fucopyranoside (pNPαFuc) and other nitrophenyl glycosides were purchased from Sigma (St. Louis, USA). Glucose, fructose, galactose, cellobiose, maltose, and lactose were purchased from Sangon Biotech (Shanghai China). Restriction endonucleases (BamH I, Nde I, and Xho I) were obtained from NEB (Ipswich, USA). T4 DNA ligase was from TaKaRa (Dalian, China). FastPfu Fly DNA polymerase was obtained from Transgen (Beijing, China). L-Fucose Assay Kit was purchased from Megazyme (Wicklow, Ireland). Other chemicals used were of analytical grade.
Microorganisms
B. fragilis NCTC9343 was cultured anaerobically using Forma anaerobic system (Thermo, USA) under a mixture of nitrogen, hydrogen, and carbon dioxide (84.9:10:5.1, v/v/v) at 37 °C. The cultured medium contained 0.5 mg vitamin K1, 5 mg hemin, 5 g NaCl, 5 g yeast extract, 20 g proteose peptone, and 60 g glucose in 1 L water at pH 7.2.
Escherichia coli strains DH5α and BL21 (DE3) harboring pGEX-4T-1 plasmid were grown at 37 °C in Luria–Bertani (LB) medium that was supplemented with 50 μg/mL ampicillin.
Sequence analysis
Through cross-referencing the genomic data of B. fragilis NCTC9343 (GenBank accession no. CR626927.1) and CAZy database (http://www.cazy.org) by using BLAST method (https://blast.ncbi.nlm.nih.gov/Blast.cgi), 12 putative and one known α-L-fucosidases, including nine GH29 α-L-fucosidases (BF0028, BF0664, BF0804, BF0810, BF1796, BF3083, BF3201, BF3242, and BF3591) and four GH95 α-L-fucosidases (BF0474, BF0570, BF0855, and BF4255), were obtained. To further identify the GH29A, a phylogenetic tree was constructed using the predicted protein sequences of enzymes. The following 18 GH29 α-L-fucosidases from other species were selected for the analysis: AlfA (Uniprot B3W8U6), AlfB (Uniprot B3WB08), and AlfC (Uniprot B3WBB5) from Lactobacillus casei BL23, BT1625 (Uniprot Q8A7A0), BT2192 (Uniprot Q8A5P6), and BT2970 (Uniprot Q8A3I4) from Bacteroides thetaiotaomicron VPI-5482, TM0306 (Uniprot Q9WYE2) from Thermotoga maritima MSB8, AfcB (Uniprot C5NS94) from Bifidobacterium bifidum JCM1254, Blon2336 (Uniprot B7GNN8), Blon0248 (Uniprot B7GTT5), and Blon0426 (Uniprot B7GN40) from Bifidobacterium longum ATCC15697, ALfuk1 (Uniprot E3PQQ9) from Paenibacillus thiaminolyticus, BFO2737 (Uniprot G8UMQ6) from Tannerella forsythia ATCC43037, FucA1 (Uniprot Q8P6S7) from Xanthomonas campestris ATCC33913, FUCA1 (Uniprot P48300) from Canis lupus familiaris, AtFuc1 (Uniprot Q8GW72) from Arabidopsis thaliana, FucA1 (Uniprot P04066) from Homo sapiens, and FucA1 (Uniprot P17164) from Rattus norvegicus. The sequences of these 27 GH29 α-L-fucosidases were firstly subjected to the multiple alignments by the ClustalX and then converted to a phylogenetic tree by the MEGA5. All positions containing gaps and missing data were eliminated. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1000 replicates) was shown next to the branches. Then, the multiple alignments of the partial deduced amino acid sequences of the catalytic domains of nine GH29 α-L-fucosidases from B. fragilis NCTC9343 and other four characterized GH29 enzymes (FucA1 from H. sapiens, TM0306 from T. maritima MSB8, AfcB from B. bifidum JCM1254, and BT1625 from B. thetaiotaomicron VPI-5482) were analyzed by ClustalX (http://www.clustal.org/).
Gene cloning and heterogeneous expression
The genomic DNA of B. fragilis NCTC9343 was extracted by TIANamp Bacteria DNA Kit from Tiangen Biotech (Beijing, China) and used as template for PCR. The genes encoding five GH29A α-L-fucosidases (BF0028, BF0810, BF1796, BF3242, and BF3591) from B. fragilis NCTC9343 were amplified with the primers (Table S1) that were designed based on the genome sequence of B. fragilis NCTC9343 (GenBank accession no. CR626927.1). PCR cycling conditions consisted of an initial step of 5 min at 94 °C, followed by 30 cycles of 30 s at 94 °C, 30 s at 60 °C to 65 °C, and 1 min 30 s at 72 °C, and a final step of 10 min at 72 °C. The PCR products were ligated into pGEX-4 T-1 fusion expression vector and transformed into E. coli BL21. The proper E. coli transformants were grown at 37 °C in LB medium containing 50 μg/mL ampicillin. When the cell density reached 0.6 to 0.8 at OD600, the culture temperature was decreased to 16 °C and 0.2 mM isopropyl-1-thio-β-D-galactoside (IPTG) was added for induction for 12 to 14 h. Then the cells were harvested and lysed by sonication. The lysates were centrifuged and the recombinant enzymes in the supernatant were purified through GST affinity chromatography (GE Healthcare, Sweden). The purified protein samples were analyzed by the 10% (w/v) of sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE). Proteins in the gel were visualized by Coomassie brilliant blue R-250 staining.
Enzyme assays
The α-L-fucosidase activity was measured by adding 20 μL enzyme solution to 60 μL 2 mM pNPαFuc in 50 mM sodium phosphate buffer (pH 7.0) with 20% (v/v) dimethyl sulfoxide (DMSO). The reaction was performed at 37 °C for 10 min, and then stopped by adding 120 μL of 1 M Na2CO3 solution. The release of p-nitrophenol was measured at 405 nm. One unit of enzyme activity is defined as the amount of enzyme required to release 1 μmol p-nitrophenol per min under the assay conditions. Assays for the other nitrophenyl glycosides were performed under the same conditions. Protein concentrations were determined by measuring absorbance at 280 nm using Nanodrop 2000 calibrated with the extinction coefficient values predicted by online analytical tool: ExPASy (http://web.expasy.org/protparam/).
Substrate specificity of the enzymes for hydrolysis
To evaluate the fucosidic linkage specificity of the enzyme, reaction mixture (10 μL) containing enzyme and 2 mM various substrates in 50 mM NaPB buffer (pH 7.0) was incubated at 37 °C for 10 min. The tested substrates included pNPαFuc, antigen H disaccharide, Lea and Lex trisaccharides, Fuc-α-1,3-GlcNAc, Fuc-α-1,4-GlcNAc, and Fuc-α-1,6-GlcNAc. The released fucose in the reaction was measured by the L-Fucose Assay Kit using L-fucose as a standard.
Screening of the enzymes for transfucosylation activity
Transfucosylation activity of four recombinant α-L-fucosidases was detected by incubating the enzyme (0.05 U/mL) with 20 mM pNPαFuc as donor and 200 mM GlcNAc, glucose, fructose, galactose, cellobiose, maltose, or lactose as acceptor in 20% (v/v) DMSO at 37 °C for 1 h. The reaction was stopped by adding three times ethanol, and the product was analyzed by thin layer chromatography (TLC).
Effects of pH and temperature on enzyme activity of BF3242
The optimum pH was determined by assaying the enzyme activity at 37 °C for 10 min with pNPαFuc at pH values ranging from 2.5 to 12.0 in 50 mM buffer containing citric acid, KH2PO4, boric acid, and barbitone and using NaOH to adjust the pH. The pH stability was determined by measuring the residual activity under the conditions as described in enzyme essay after incubating the enzyme in the presence of the above various pH buffers at 4 °C for 12 h. The optimal temperature for enzyme reaction was determined by assaying the enzyme activity with pNPαFuc in 50 mM sodium phosphate buffer (pH 7.0) for 10 min at 20 to 65 °C. Thermo-stability was determined by assaying residual enzymatic activity in 50 mM sodium phosphate buffer (pH 7.0) after incubating the enzyme for 1 h at 20 to 65 °C.
Synthesis of fucosyl-N-acetylglucosamine disaccharides by BF3242
The reaction for the synthesis of fucosyl-N-acetylglucosamine disaccharides was performed by incubating BF3242 with pNPαFuc as the glycosyl donor and GlcNAc as the acceptor in 20% (v/v) DMSO. To achieve the maximum yield, the reaction conditions including initial substrate concentrations, pH, temperature, and reaction time were carefully evaluated. The enzyme was used at a concentration of 0.05 U/mL. The effects of GlcNAc concentrations (100, 200, 300, 400, 500, and 600 mM) were determined using 20 mM pNPαFuc at pH 7.0 and 37 °C for 1 h. The effects of pNPαFuc concentrations (10, 20, 30, 40, 50, and 60 mM) were measured using 500 mM GlcNAc at pH 7.0 and 37 °C for 1 h. The effects of pH were examined at 14 different pH values (CKBB buffer at 4.0, 4.5, 5.5, 6.0, 6.5, 7.0, 7.5, 8.0, 8.5, 9.0, 9.5, 10.0, 10.5, and 11.0) at 37 °C by using 500 mM GlcNAc in the presence of 20 mM pNPαFuc for 1 h. The effects of temperature were investigated by using 500 mM GlcNAc and 20 mM pNPαFuc at pH 7.5 at 10 different temperatures (20 °C, 25 °C, 30 °C, 37 °C, 40 °C, 45 °C, 50 °C, 55 °C, 60 °C, and 65 °C) for 1 h. The effects of reaction time were evaluated by using 500 mM GlcNAc and 20 mM pNPαFuc at pH 7.5 and 37 °C with interval sampling within 3 h. All the reactions were stopped by adding three times ethanol and the resulting products were analyzed by HPLC. The yield of the product was defined as the ratio of the concentration of the synthesized glycoside product (mM) to the initial concentration of donor (mM). The ratio of the transglycosylation to hydrolysis activity (RT/H) was calculated by dividing synthesized product concentration (mM) by hydrolyzed donor concentration (mM).
Isolation of the fucosyl-N-acetylglucosamine disaccharides
The oligosaccharide product was separated by a Bio-Gel P2 (Bio-Rad, USA) column (1.6 × 90 cm) with distilled water as the eluent. The eluted fractions were collected and subjected to sugar determination by TLC. The identified fractions were concentrated by evaporation. The regioisomeric products were separated by HPLC and the corresponding fucoside-containing fractions were combined and lyophilized to dry powder.
HPLC and TLC analyses
HPLC was performed by an Agilent 1200 series coupled with a UV detector (G1314B) using an Acchrom XAmide analysis column (4.6 × 250 mm) at 30 °C. Samples were eluted with 91% (v/v) acetonitrile as the mobile phase at a flow rate of 1.0 mL/min and detected at 210 nm.
TLC was performed by loading samples on silica gel 60 F254 plates (Merck, Germany). The loaded samples were developed by a mixture of n-butanol/ethanol/water (15:1:1, v/v/v) and subsequently visualized by spraying with diphenylamine-aniline-phosphoric acid reagent and heating at 86 °C for 30 min.
MS and NMR analysis
Mass spectra were detected and recorded on a Shimadzu LCMS-IT-TOF instrument (Kyoto, Japan) equipped with an ESI source and operated in positive ion mode. 1H and 13C NMR data were recorded at room temperature with an Agilent DD2 600 MHz instrument operating at 600 and 150 MHz, respectively. Chemical shifts were given in ppm downfield from internal TMS of D2O. Chemical shifts and coupling constants were calculated from a first-order analysis of the spectra. Assignments of proton and carbon atoms were achieved through 1H, 13C, and homonuclear (COSY) and heteronuclear (HMQC, HMBC) following the standard Agilent pulse programs.
Molecular modeling and MD simulations
Homology modeling of BF3242 was performed using SWISS-MODEL (https://swissmodel.expasy.org/interactive), and the 3D structure of α-L-fucosidase BT2970 (PDB entry 4WSJ) from Bacteroides thetaiotaomicron ATCC29148 served as the template. BF3242 shared 32% sequence identity with the template.
The MD simulations of BF3242 were studied by principal component analysis (PCA) (Liu et al. 2014). The covariance matrix was created using the atomic coordinates of protein backbone atoms. The eigenvectors were generated by diagonalization of the covariance matrix. Each of them had a respective eigenvalue (g_covar). The trajectory was projected onto a particular eigenvector to reveal the concerted motions of enzyme (g_anaeig). The mathematical formulas used for PCA have been made in previous study (Liu et al. 2014; Jiang et al. 2017). Root-mean-square fluctuation (RMSF) analysis was carried out by using gmx rmsf tool. In order to ensure the accuracy and reliability of the measuring data, the last 20 ns simulation trajectories were used to calculate all the time-averaged properties. PyMOL 2.1 (http://www.pymol.org) (Robert and Gouet 2014) and MATLAB 2016b (https://ww2.mathworks.cn/products/matlab.html) were used to analyze and visualize enzyme structure and simulation data.
Results
Sequence analysis of α-L-fucosidases from B. fragilis NCTC9343
By cross-referencing the genome sequence of B. fragilis and CAZy database, we found there were 142 putative glycosidases (as of November 2019) including 13 α-L-fucosidases in the genome of B. fragilis NCTC9343. Among these 13 α-L-fucosidases, nine α-L-fucosidases were predicted to belong to GH29 and four α-L-fucosidases were predicted to belong to GH95 (Table 1). All of these 13 α-L-fucosidases did not contain transmembrane region, and 11 of them had signal peptides in their deduced protein sequences except two enzymes BF0570 and BF0664. They had theoretical molecular weight between 47 and 93 kDa. Since only the α-L-fucosidases in GH29A had the potential to synthesize the fucosylated compounds using pNPαFuc as the donor, we constructed a phylogram to identify which of the nine GH29 α-L-fucosidases belonged to GH29A. The results (Fig. 1) showed that five α-L-fucosidases (BF0028, BF0810, BF1796, BF3242, and BF3591) belonged to GH29A and four α-L-fucosidases (BF0664, BF0804, BF3083, and BF3201) belonged to GH29B. Furthermore, the result (Fig. 2) of multiple alignments of the partial deduced amino acid sequences of the catalytic domains of these nine GH29 α-L-fucosidases and other four characterized GH29 enzymes showed that a conserved aspartate residue as the catalytic nucleophile in GH29 enzymes was identified in all the nine αL-fucosidases of B. fragilis and other four characterized GH29 α-L-fucosidases. However, the acid/base residue (glutamate) was conserved in GH29B but not in GH29A. We then chose the five GH29A α-L-fucosidases for further study.
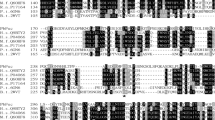
Phylogenetic analysis of 26 GH29 α-L-fucosidases. The sequences of these α-L-fucosidases were firstly subjected to the multiple alignments by the ClustalX and then converted to a phylogenetic tree by the MEGA5. The sequences of the nine GH29 α-L-fucosidases, BF0028, BF0664, BF0804, BF0810, BF1796, BF3083, BF3201, BF3242, and BF3591, from B. fragilis NCTC9343 were highlighted in red
Multiple alignment of the partial amino acid sequences of GH29A and GH29B α-L-fucosidases. The sequences for alignment included the nine GH29 α-L-fucosidases from B. fragilis NCTC9343, FucA1 from H. sapiens, TM0306 from T. maritima, AfcB from B. bifidum JCM 1254, and BT1625 from B. thetaiotaomicron. The alignment was generated with ClustalX. The regions of sequence identity were shaded in black, and the regions of sequence similarity were in gray. The blue triangle denotes the conserved nucleophile of GH29A and GH29B. The red triangle denotes the conserved acid/base residues of GH29B. Red boxes indicate experimentally confirmed acid/base residues
Cloning, and heterogeneous expression of GH29A α-L-fucosidases from B. fragilis NCTC9343
The genes of five GH29A α-L-fucosidases (BF0028, BF0810, BF1796, BF3242, and BF3591) were cloned from genome DNA of B. fragilis NCTC9343 and expressed in E. coli BL21. Four of them (BF0028, BF0810, BF3242, and BF3591) were expressed successfully as soluble proteins. As shown in SDS-PAGE (Fig. S1), these four recombinant GH29 α-L-fucosidases, BF0028, BF0810, BF3242, and BF3591, were purified by GST affinity purification and they migrated as nearly single protein bands with molecular masses in agreement with the calculated masses fused with the GST tag (about 26 kDa) (Table 1).
The hydrolysis activity of these four recombinant α-L-fucosidases was determined using 12 different nitrophenyl glycosides. All the four recombinant enzymes hydrolyzed pNPαFuc, but showed no activities towards other 11 artificial substrates with β-fucosidic linkage (pNP-β-L-fucopyranoside) and without fucose in the glycon moieties including pNP-α/β-D-galactopyranoside, pNP-α/β-D-glucopyranoside, pNP-α/β-D-mannopyranoside, pNP-N-acetyl-α/β-D-galactosaminide, and pNP-N-acetyl-α/β-D-glucosaminide. The specific activities for pNPαFuc of BF0028, BF0810, BF3242, and BF3591 were 0.5, 8.7, 11.7, and 0.2 U/mg, respectively.
Fucosidic linkage specificity of the enzymes for hydrolysis
The fucosidic linkage specificities of the four recombinant α-L-fucosidases against different substrates including pNPαFuc, antigen H disaccharide, Lea and Lex trisaccharides, Fuc-α-1,3-GlcNAc, Fuc-α-1,4-GlcNAc, and Fuc-α-1,6-GlcNAc disaccharides are shown in Table 2. The results indicated that these four enzymes had different fucosidic linkage specificities for various natural oligosaccharides and pNPαFuc. Three of them (BF0028, BF0810, and BF3242) showed higher hydrolytic activities towards pNPαFuc than the natural oligosaccharides, and BF3591 showed higher cleavage rate for α-1,3 linkages of Lex trisaccharide and Fuc-α-1,3-GlcNAc. BF3242 and BF3591 were capable of hydrolyzing α-1,2-, α-1,3-, α-1,4-, and α-1,6-linked fucosides. Particularly, BF3242 had higher hydrolytic activity towards disaccharides than trisaccharides. BF3591 showed higher activity towards α-1,3-linked fucosides than α-1,2-, α-1,4-, and α-1,6-linked fucosides and displayed higher activity to nature substrate Lex trisaccharides and Fuc-α-1,3-GlcNAc. BF0028 showed weak hydrolytic activity towards the α-1,2- and α-1,3-linked fucosides of antigen H disaccharide, Lea trisaccharides, and Fuc-α-1,3-GlcNAc, and it preferred Lea trisaccharide to Fuc-α-1,3-GlcNAc. BF0810 only hydrolyzed artificial substrate pNPαFuc.
Screening of α-L-fucosidases for transglycosylation
The transglycosylation activities of the four recombinant GH29A α-L-fucosidases (BF0028, BF0810, BF3242, and BF3591) were examined using pNPαFuc as donor and GlcNAc, glucose, fructose, galactose, cellobiose, maltose, or lactose as acceptor. The results (Figs. 3a and S2) showed that only BF3242 showed transglycosylation activity towards GlcNAc, glucose, and maltose, and the other three enzymes (BF0028, BF0810, and BF3591) did not show transglycosylation activity with all the tested acceptors. As shown in Fig. 3a, using GlcNAc as acceptor, two novel spots of products (named P1 and P2) appeared below the spot of GlcNAc on TLC plate. P2 was formed firstly within 30 min reaction, and then P1 accumulated during the next 30 min. P1 and P2 had the same migration rates as the standard samples of Fuc-α-1,3-GlcNAc and Fuc-α-1,6-GlcNAc on the TLC plate, respectively. The results of HPLC analysis also revealed that the peak of P1 and P2 shared the identical retention times to those of standard samples of Fuc-α-1,3-GlcNAc and Fuc-α-1,6-GlcNAc, respectively (Fig. 3b). Therefore, BF3242 was selected for further study.
Analysis of transglycosylation reaction mixture of four α-L-fucosidases by TLC (a) and analysis of transglycosylation products of BF3242 by HPLC (b). a Transglycosylation activities of the four purified recombinant enzymes (BF0028, BF0810, BF3242, BF3596) were tested by incubation of 0.05 U/mL enzyme with 20 mM pNPαFuc as donor and 200 mM GlcNAc as acceptor in 20% (v/v) DMSO at 37 °C for 0.5 h and 1 h. STD standard saccharides. b Control and BF3242 reaction, inactivated BF3242 and BF3242 were used in the reactions as (a) for 1 h, respectively. The retention times of standard Fuc-α-1,3-GlcNAc and Fuc-α-1,6-GlcNAc were 27 and 38 min, respectively
BF3242 showed an optimal temperature at 45 °C, and it was stable below 45 °C (Fig. S3a). The enzyme was highly active at pH 5.5 to 7.5 and stable at pH 5.0 to 11.0 (Fig. S3b).
Synthesis of fucosyl-N-acetylglucosamine disaccharides by BF3242
Transglycosylation reactions were performed by incubating BF3242 with pNPαFuc and GlcNAc under various conditions. Figure 4a shows the effects of the acceptor concentration on disaccharide synthesis. When the GlcNAc concentrations were increased from 100 to 600 mM, the overall yield of two products increased from a minimum of 19.4% (7.5% of P1 and 11.9% of P2) at 100 mM to a maximum of 66.8% (20.1% of P1 and 46.7% of P2) at 500 mM, and then slightly decreased to 53.4% (14.3% of P1 and 39.1% of P2) at 600 mM. Therefore, the subsequent reactions were performed with 500 mM GlcNAc. Figure 4b shows the influence of donor concentrations on the yield of products. When the pNPαFuc concentrations were increased from 10 to 20 mM, the overall yield of two products was 50.9% (8.9% of P1 and 42.0% of P2) at 10 mM and reached a maximum of 67.3% (20.3% of P1 and 47.0% of P2) at 20 mM. Continuous increase in donor concentration from 20 to 60 mM reduced product yields. Thus, the subsequent reactions were performed using 500 mM GlcNAc and 20 mM pNPαFuc.
The effects of substrate concentration (a, b), pH (c), temperature (d), and reaction time (e) on product yields. a The acceptor concentrations from 100 to 600 mM were tested at pH 7.0 and 37 °C in the presence of 20 mM pNPαFuc for 1 h. b The donor concentrations were tested by incubation with 500 mM GlcNAc at pH 7.0 and 37 °C in the presence of pNPαFuc from 10 to 60 mM for 1 h. c pH tests were carried out at 37 °C by incubating the enzyme with 500 mM GlcNAc in the presence of 20 mM pNPαFuc for 1 h in buffers ranging from pH 4.0 to 11.0. d Temperature was tested at pH 7.5 by incubation of the enzyme with 500 mM GlcNAc and 20 mM pNPαFuc at 20 to 65 °C for 1 h. e Reaction time was determined at pH 7.5 by incubation of the enzyme with 500 mM GlcNAc and 20 mM pNPαFuc at 37 °C within 180 min. Data points represent the means ± S.D. of three replicates
The pH values also strongly affected the yield of products. As showed in Fig. 4c, the overall yield of two products increased from pH 4.0 to 7.5 and then tended to stabilize at the pH range of 7.5 to 9.0. The maximum overall yield of two products of 73.4% (30.3% of P1 and 43.1% of P2) was obtained at pH 7.5. Once the pH values exceeded 9.0, the overall yield of two products dramatically dropped. Thus, the subsequent reactions were performed at pH 7.5.
The reaction temperature remarkably affected the product formation. It was worth noting that the two products reached their maximum yields at different temperatures. As showed in Fig. 4d, the maximum yield of P2 at 48.9% was obtained at 37 °C while the yield of P1 was 25.4%, which presented a maximum overall yield of two products of 74.3%. When the temperatures increased above 37 °C, the yield of P2 decreased markedly, but the yield of P1 increased. When the temperatures increased to 50 °C, the yield of P1 reached the maximal value at 46.9% while the yield of P2 was 21.2%. Continuous increase in the temperature increased from 50 to 60 °C, the overall yield of two products sharply decreased. Thus, the next reactions were carried out in the conditions of 500 mM GlcNAc, 20 mM pNPαFuc, pH 7.5, and 37 °C. The time curves (Fig. 4e) showed that P1 reached a maximum yield of 27.9% at 30 min, and P2 reached a maximum yield of 53.4% at 45 min. The overall yield of two products reached peak value of 79.0% (25.5% of P1 and 53.5% of P2) at 45 min; in this case, 15.8 mM fucosyl-N-acetylglucosamine disaccharides were synthesized and 4.2 mM pNPαFuc was hydrolyzed, indicating that the RT/H was 3.8.
Isolation and identification of fucosyl-N-acetylglucosamine disaccharides
The transglycosylation reactions were performed under the optimal conditions of 500 mM GlcNAc, 20 mM pNPαFuc, pH 7.5, and 37 °C for 45 min. The products were purified by Bio-Gel P2 column chromatography, and the two regioisomeric products were further separated by HPLC, and then analyzed by MS and NMR spectroscopy (Fig. S4–15).
The positive ion ESI mass (Fig S4 and S10) showed the peaks of [M + H]+ and [M + Na]+ at m/z 368.1628 and m/z 390.1457 for P1 and m/z 368.1617 and m/z 390.1471 for P2, respectively, consistent with the theoretic molecular mass of fucosyl-N-acetylglucosamine disaccharides (367). The chemical shifts and configurations of the sugar residues are summarized in Table 3. As for P1, H-5 of the Fuc residue appeared at δ 4.17 ppm, and C-3 of the α/β-GlcNAc residues appeared at δ 77.9 ppm and δ 80.4 ppm, respectively. The lower-field shifts of the C-3 signals revealed that the Fuc residue is bound to O-3 of the GlcNAc residue; therefore, the chemical structure of P1 was Fuc-α-1,3-GlcNAc. Similarly, as for P2, H-5 of the Fuc residue appeared at δ 3.96 ppm, and C-6 of the α/β-GlcNAc residues appeared at δ 67.7 ppm and δ 67.2 ppm, respectively. These lower-field shifts revealed a binding of the Fuc residue to O-6 of the GlcNAc residue. Thus, the P2 structure was completely characterized as Fuc-α-1,6-GlcNAc.
The MD simulation analysis of BF3242 at different temperatures
In order to further understand why the temperature affected the ratio of two regioisomeric products, we constructed the putative 3D model of BF3242 (Fig. 5a) and used it to study the effects of temperature on protein structure and dynamics. Clearly, BF3242 shared the common characteristic of all known structures of GH29 enzymes (Fig. 5a). We then analyzed structural flexibility according to the RMSF of Cα atoms with respect to the initial 3D structure. The RMSF reflect the mobility of residues from their time-averaged position over the course of a simulations and higher RMSF coincide with more protein flexibilities and potentially less thermal stabilities. As shown in Fig. 5b, c, Loop-4 (His220-Ser245) displayed significant changes throughout the thermal simulations, suggesting that Loop-4 was the most temperature sensitive regions of BF3242. Moreover, Fig. 5d shows the RMSF value for Loop-4 at 300 K was about 0.43 nm and it increased to 0.67 nm at 350 K which indicated the ΔRMSF of Loop-4 region fluctuates more flexible than other regions with increasing temperature (300 to 350 K).
Molecular modeling and molecular dynamics simulation analysis of BF3242. a Putative 3D model of BF3242. b, c Visualization of PCA displacement on BF3242 at 300 K and 350 K. The most thermal-sensitive loops are indicated with the red circles. d Root-mean-square fluctuations (RMSF) for Cα atoms in BF3242 based on the last 20 ns simulations at 300 K and 350 K. ΔRMSF: the absolute values of difference of RMSF between 300 K and 350 K
Discussion
Although the fucosyl-N-acetylglucosamine disaccharides are present in many biologically important oligosaccharides, so far, the source of glycosidases for synthesis of fucosyl-N-acetylglucosamine disaccharides is still limited. A few microbial GH29A α-L-fucosidases have been reported to be capable of synthesizing fucosyl-N-acetylglucosamine disaccharides with pNPαFuc as glycosyl donor and GlcNAc as acceptor. Fuc-α-1,3-GlcNAc has been synthesized at various yields by two α-L-fucosidases from Aspergillus niger (24% and 58%) (Ajisaka and Shirakabe 1992; Vetere et al. 1997), an α-L-fucosidase from Penicillium multicolor (49%) (Ajisaka et al. 1998), and an α-L-fucosidase from L. casei (23%) (Rodríguez-Díaz et al. 2013). Fuc-α-1,6-GlcNAc has been synthesized at the yield of 56% by an α-L-fucosidase from L. casei (Rodríguez-Díaz et al. 2013). Two Fuc-α-GlcNAc regioisomeric products (linkage not determined) have been synthesized with a yield of 20% by an α-L-fucosidase from Thermus sp. Y5 (Eneyskaya et al. 2001). Discovering α-L-fucosidases and their biochemical characterization is not only of academic interest but also can lead to the exploitation of useful synthetic tools for important oligosaccharide production.
It was reported that the large proportion (out of 4300 genes) of the genome of B. fragilis is devoted to encoding carbohydrate metabolism (Coyne and Comstock 2008; Xu et al. 2003; Zhao et al. 2019). As we mentioned in the introduction of this paper, the hydrolysis activity and transglycosylation activity of several glycosidases of B. fragilis have been reported (Blatch and Woods 1993; Chen et al. 2016; Gong et al. 2016; Guo et al. 2018; Kawaguchi et al. 2014; Liu et al. 2008; Tanaka et al. 1994; Tsai et al. 2017; Yamamoto et al. 2018), but there has been no report on the transglycosylation activity of its α-L-fucosidases. In this work, we found there were 12 putative and one known α-L-fucosidases in the genome of B. fragilis NCTC9343 by cross-referencing the genome data of B. fragilis NCTC9343 (GenBank accession no. CR626927.1) and CAZy database. Among them, nine GH29 α-L-fucosidases were further divided into two subfamilies, five α-L-fucosidases belonging to GH29A and four α-L-fucosidases belonging to GH29B, through phylogenetic tree analysis. The five GH29A α-L-fucosidases were then chosen as the candidates for gene cloning and heterogeneous expression in E. coli BL21 to screen the biocatalyst for synthesis of fucosyl-N-acetylglucosamine disaccharides. Four GH29A α-L-fucosidases (BF0028, BF0810, BF3242, and BF3591) were successfully expressed as soluble proteins and characterized for substrate specificity and transglycosylation activity. However, BF1796 was expressed in the form of inclusion body, which might be due to lacking the C-terminal domain (Fig. S16). Similar result has been reported that the C-terminal domain of a lichenase from Clostridium thermocellum was necessary to maintain enzymatic activity (Niu et al. 2010). BF3242 was found to be the only one that possessed transglycosylation activity. After optimization of the conditions for synthetic reaction, BF3242 exhibited excellent transglycosylation activity for the synthesis fucosyl-N-acetylglucosamine disaccharides with pNPαFuc as donor and GlcNAc as acceptor at the overall yield of 79.0%, presenting the highest transglycosylation efficiency in all the reported α-L-fucosidases so far.
It was very essential to study the reaction conditions including initial substrate concentrations, pH, temperature, and reaction time to obtain the optimized synthetic conditions for BF3242. The substrate concentrations significantly affected the yield of products. It has been reported that the increase of acceptor concentrations could reduce water activity within the enzyme catalytic center, which might prefer the transglycosylation reaction to the hydrolysis reaction (Zeuner et al. 2014). In this work, the yield of fucosyl-N-acetylglucosamine disaccharides was improved with the concentrations of acceptor substrate GlcNAc increased from 100 to 500 mM. Nevertheless, when the concentration of GlcNAc was raised to 600 mM, the yield of fucosyl-N-acetylglucosamine disaccharides observably dropped. The phenomenon was similar to the result that the transglycosylation yields of an α-galactosidase (AgaBf3S) from B. fragilis were decreased when the lactose concentrations increased from 500 to 600 mM (Gong et al. 2016). The excessive amount of acceptor substrate could cause molecular crowding effect which might partly inhibit the transglycosylation reaction (Kim and Yethiraj 2009).
BF3242 could synthesize both Fuc-α-1,3-GlcNAc and Fuc-α-1,6-GlcNAc in one reaction. It was worth mentioning that the reaction temperature distinctly influenced the ratio of the two regioisomeric products (Fig. 4d). At the lower temperature range from 25 to 37 °C, BF3242 preferred synthesizing Fuc-α-1,6-GlcNAc with the proportion of products from 88 to 68%, while at the higher temperature range from 37 to 50 °C, the proportion of Fuc-α-1,3-GlcNAc increased from 32 to 68%. The similar results were also reported in the synthesis of galactosyl-N-acetylglucosamine disaccharides by a GH42 β-galactosidase from Bacillus circulans with lactose as donor and GlcNAc as acceptor when temperature was below 50 °C, the enzyme tended to synthesize Gal-β-1,4-GlcNAc other than Gal-β-1,6-GlcNAc, and when temperature increased to 60 °C, Gal-β-1,6-GlcNAc accounted for the vast majority (Sakai et al. 1992). In order to understand why temperature affects the regioselectivity in transglycosylation, we further analyzed the structural fluctuations of BF3242 using the MD simulations. Fast atomic thermal fluctuations are considered lubricants for conformational changes of large proteins. These changes allow sufficient flexibility for biological functions (Dong et al. 2018; Sigtryggsdóttir et al. 2014). The functions of the flexible regions identified through the MD simulations have been verified by protein mutagenesis technique. It was reported that the mutations (L435A/G and F432I/L) in a flexible region of monooxygenases produced a series of substituted lactones with inversed configuration (Hu et al. 2019). Similarly, mutation V7A in a flexible region in N-terminal domain of a caseinolytic protease from Staphylococcus aureus inactivated the enzyme (Vahidi et al. 2018). Such reports suggested that the flexible regions of enzyme could be of great importance to the catalytic activities, and the rationally designed mutations in the regions could influence the product configuration or enzymatic activity. In this work, the loop-4 region (His220-Ser245) located obliquely above the catalytic chamber in the putative 3D model of BF3242 (Fig. 5a) and the results of the MD simulation analysis showed that the fluctuations of loop-4 region were more sensitive to temperature, which seems to be consistent with that the reaction temperature could influence the ratio of two regioisomeric products. Therefore, the flexible regions loop-4 of BF3242 might be responsible for the changes of the ratio of two regioisomers in the products at different reaction temperatures.
In conclusion, five GH29A α-L-fucosidases from 12 putative and one known α-L-fucosidases of B. fragilis NCTC9343 were cloned, heterogeneously expressed, and screened for transglycosylation activity, and a GH29A α-L-fucosidases (BF3242) with the outstanding ability for synthesis of Fuc-α-1,3/1,6-GlcNAc at a maximum yield of 79.0% with the ratio of 0.48 for 1,3/1,6 was obtained. BF3242 could be an attractive candidate for enzymatic synthesis of fucosyl-N-acetylglucosamine disaccharides.
References
Ajisaka K, Shirakabe M (1992) Regioselective synthesis of α-L-fucosylcontaining disaccharides by use of α-L-fucosidases of various origins. Carbohydr Res 224:291–299. https://doi.org/10.1016/0008-6215(92)84115-9
Ajisaka K, Fujimoto H, Miyasato M (1998) An α-L-fucosidase from Penicillium multicolor as a candidate enzyme for the synthesis of α(1→3)-linked fucosyl oligosaccharides by transglycosylation. Carbohydr Res 309:125–129. https://doi.org/10.1016/s0008-6215(98)00112-8
Ashida H, Miyake A, Kiyohara M, Wada J (2009) Two distinct alpha-L-fucosidases from Bifidobacterium bifidum are essential for the utilization of fucosylated milk oligosaccharides and glycoconjugates. Glycobiology 19:1010–1017. https://doi.org/10.1093/glycob/cwp082
Becerra JE, Coll-Marqués JM, Rodríguez-Díaz J, Monedero V, Yebra MJ (2015) Preparative scale purification of fucosyl-N-acetylglucosamine disaccharides and their evaluation as potential prebiotics and antiadhesins. Appl Microbiol Biotechnol 99:7165–7176. https://doi.org/10.1007/s00253-015-6666-2
Becerra JE, Rodríguez-Díaz J, Gozalbo-Rovira R, Palomino-Schätzlein M, Zúñiga M, Monedero V, Yebra MJ (2020) Unique microbial catabolic pathway for the human core N-glycan constituent fucosyl-α-1,6-N-acetylglucosamine-asparagine. mBio 11:e02804–e02819. https://doi.org/10.1128/mBio.02804-19
Blatch GL, Woods DR (1993) Molecular characterization of a fructanase produced by Bacteroides fragilis BF-1. J Bacteriol 175:3058–3066. https://doi.org/10.1128/jb.175.10.3058-3066.1993
Bode L (2012) Human milk oligosaccharides: every baby needs a sugar mama. Glycobiology 22:1147–1162. https://doi.org/10.1093/glycob/cws074
Chen C, Zhang Y, Xue M, Liu XW, Li Y, Chen X, Wang PG, Wang F, Cao H (2015) Sequential one-pot multienzyme (OPME) synthesis of lacto-N-neotetraose and its sialyl and fucosyl derivatives. Chem Commun 51:7689–7692. https://doi.org/10.1039/c5cc01330e
Chen X, Xu L, Jin L, Sun B, Gu G, Lu L, Xiao M (2016) Efficient and regioselective synthesis of β-GalNAc/GlcNAc-Lactose by a bifunctional transglycosylating β-N-acetylhexosaminidase from Bifidobacterium bifidum. Appl Environ Microbiol 82:5642–5652. https://doi.org/10.1128/AEM.01325-16
Cordon-Cardo C, Lloyd KO, Sakamoto J, McGroarty ME, Old LJ, Melamed MR (1986) Immunohistologic expression of blood-group antigens in normal human gastrointestinal tractand colonic carcinoma. Int J Cancer 37:667–676. https://doi.org/10.1002/ijc.2910370505
Coyne MJ, Comstock LE (2008) Niche-specific features of the intestinal bacteroidales. J Bacteriol 190:736–742. https://doi.org/10.1128/JB.01559-07
Crout DH, Vic G (1998) Glycosidases and glycosyl transferases in glycoside and oligosaccharide synthesis. Curr Opin Chem Biol 2:98–111. https://doi.org/10.1002/chin.199832334
Dawson G, Tsay G (1977) Substrate specificity of human α-L-fucosidase. Arch Biochem Biophys 184:12–23. https://doi.org/10.1016/0003-9861(77)90321-6
DiCioccio RA, Barlow JJ, Matta KL (1982) Substrate specificity and other properties of α-L-fucosidase from human serum. J Biol Chem 257:714–718. https://doi.org/10.1016/0165-022X(82)90005-7
Dong YW, Liao ML, Meng XL, Somero GN (2018) Structural flexibility and protein adaptation to temperature: molecular dynamics analysis of malate dehydrogenases of marine molluscs. PNAS 115:1274–1279. https://doi.org/10.1073/pnas.1718910115
Eneyskaya EV, Kulminskaya AA, Kalkkinen N, Nifantiev NE, Arbatskii NP, Saenko AI, Chepurnaya OV, Arutyunyan AV, Shabalin KA, Neustroev KN (2001) An α-L-fucosidase from Thermus sp. with unusually broad specificity. Glycoconj J 18:827–834. https://doi.org/10.1023/A:1021163720282
Fan S, Zhang H, Chen X, Lu L, Xu L, Xiao M (2016) Cloning, characterization, and production of three α-L-fucosidases from Clostridium perfringens ATCC 13124. J Basic Microbiol 56:347–357. https://doi.org/10.1002/jobm.201500582
Gong W, Xu L, Gu G, Lu L, Xiao M (2016) Efficient and regioselective synthesis of globotriose by a novel α-galactosidase from Bacteroides fragilis. Appl Microbiol Biotechnol 100:6693–6702. https://doi.org/10.1007/s00253-016-7464-1
Guo L, Chen X, Xu L, Xiao M, Lu L (2018) Enzymatic synthesis of 6′-Sialyllactose, a dominant sialylated human milk oligosaccharide, by a novel exo-α-sialidase from Bacteroides fragilis NCTC9343. Appl Environ Microbiol 84:e00071–e00018. https://doi.org/10.1128/AEM.00071-18
Guzmán-Rodríguez F, Alatorre-Santamaría S, Gómez-Ruiz L, Rodríguez-Serrano G, García-Garibay M, Cruz-Guerrero A (2018) Synthesis of a fucosylated trisaccharide via transglycosylation by α-L-fucosidase from Thermotoga maritima. Appl Biochem Biotechnol 186:681–691. https://doi.org/10.1007/s12010-018-2771-x
Hu Y, Wang J, Cen Y, Zheng H, Huang M, Lin X, Wu Q (2019) “Top” or “bottom” switches of a cyclohexanone monooxygenase controlling the enantioselectivity of the sandwiched substrate. Chem Commun (Camb) 55:2198–2201. https://doi.org/10.1039/c8cc09951k
Jiang X, Li W, Chen G, Wang L (2017) Dynamic perturbation of the active site determines reversible thermal inactivation in glycoside hydrolase family 12. J Chem Inf Model 57:288–297. https://doi.org/10.1021/acs.jcim.6b00692
Katayama T, Sakuma A, Kimura T, Makimura Y (2004) Molecular cloning and characterization of Bifidobacterium bifidum 1,2-alpha-L-fucosidase (AfcA), a novel inverting glycosidase (glycoside hydrolase family 95). J Bacteriol 186:4885–4893. https://doi.org/10.1128/JB.186.15.4885-4893.2004
Kawaguchi K, Senoura T, Ito S, Taira T, Ito H, Wasaki J, Ito S (2014) The mannobiose-forming exo-mannanase involved in a new mannan catabolic pathway in Bacteroides fragilis. Arch Microbiol 196:17–23. https://doi.org/10.1007/s00203-013-0938-y
Kim JS, Yethiraj A (2009) Effect of macromolecular crowding on reaction rates: a computational and theoretical study. Biophys J 96:1333–1340. https://doi.org/10.1016/j.bpj.2008.11.030
Kobata A (2010) Structures and application of oligosaccharides in human milk. Proc Jpn Acad Ser B Phys Biol Sci 86:731–747. https://doi.org/10.2183/pjab.86.731
Lezyk M, Jers C, Kjaerulff L, Gotfredsen CH, Mikkelsen MD, Mikkelsen JD (2016) Novel α-L-Fucosidases from a soil metagenome for production of fucosylated human milk oligosaccharides. PLoS One 11:e0147438. https://doi.org/10.1371/journal.pone.0147438
Li T, Li M, Hou L, Guo Y, Wang L, Sun G, Chen L (2017a) Identification and characterization of a core fucosidase from the bacterium Elizabethkingia meningoseptica. J Biol Chem 293:1243–1258. https://doi.org/10.1074/jbc.M117.804252
Li C, Zhu S, Ma C, Wang LX (2017b) Designer α1,6-fucosidase mutants enable direct core fucosylation of intact N-glycopeptides and N-glycoproteins. J Am Chem Soc 139:15074–15087. https://doi.org/10.1021/jacs.7b07906
Liu QP, Yuan H, Bennett EP, Levery SB, Nudelman E, Spence J, Pietz G, Saunders K, White T, Olsson ML, Henrissat B, Sulzenbacher G, Clausen H (2008) Identification of a GH110 subfamily of alpha 1,3-galactosidases: novel enzymes for removal of the alpha 3Gal xenotransplantation antigen. J Biol Chem 283:8545–8554. https://doi.org/10.1074/jbc.M709020200
Liu M, Wang L, Sun X, Zhao X (2014) Investigating the impact of Asp181 point mutations on interactions between PTP1B and phosphotyrosine substrate. Sci Rep 4:5095. https://doi.org/10.1038/srep05095
Liu S, Kulinich A, Cai ZP, Ma HY, Du YM, Lv YM, Liu L, Voglmeir J (2016) The fucosidase-pool of Emticicia oligotrophica: biochemical characterization and transfucosylation potential. Glycobiology 26:871–879. https://doi.org/10.1093/glycob/cww030
Mariano A, Di Carlo A, Santonastaso C, Oliva A, D’ Armiento M, Macchia V (2000) Expression of Lewis carbohydrate antigens and chromogranin A in human prostatic cancer. Int J Oncol 17:167–171. https://doi.org/10.3892/ijo.17.1.167
Megson ZA, Koerdt A, Schuster H (2015) Characterization of an α-L-fucosidase from the periodontal pathogen Tannerella forsythia. Virulenc 6:282–292. https://doi.org/10.1080/21505594.2015.1010982
Murata T, Morimoto S, Zeng X, Watannabe S, Usui T (1999) Enzymatic synthesis of α-L-fucosyl-N-acetyllactosamines and 3-O-α-L-fucosyllactose utilizing α-L-fucosidases. Carbohydr Res 320:192–199. https://doi.org/10.1002/chin.200011204
Nakai H, Baumann MJ, Petersen BO, Westphal Y, Hachem MA, Dilokpimol A, Duus JØ, Schols HA, Svensson B (2010) Aspergillus nidulans alpha-galactosidase of glycoside hydrolase family 36 catalyses the formation of alpha-galacto-oligosaccharides by transglycosylation. FEBS J 277:3538–3551. https://doi.org/10.1111/j.1742-4658.2010.07763.x
Niu D, Zhou XX, Yuan TY, Lin ZW, Ruan H, Li WF (2010) Effect of the C-terminal domains and terminal residues of catalytic domain on enzymatic activity and thermostability of lichenase from Clostridium thermocellum. Biotechnol Lett 32:963–967. https://doi.org/10.1007/s10529-010-0241-9
Okuyama M, Matsunaga K, Watanabe KI, Yamashita K, Tagami T, Kikuchi A, Ma M, Klahan P, Mori H, Yao M, Kimura A (2017) Efficient synthesis of α-galactosyl oligosaccharides using a mutant Bacteroides thetaiotaomicron retaining α-galactosidase (BtGH97b). FEBS J 284:766–783. https://doi.org/10.1111/febs.14018
Robert X, Gouet P (2014) Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res 42:W320–W324. https://doi.org/10.1093/nar/gku316
Rodríguez-Díaz J, Monedero V, Yebra MJ (2011) Utilization of natural fucosylated oligosaccharides by three novel alpha-L-fucosidases from a probiotic Lactobacillus casei strain. Appl Environ Microbiol 77:703–705. https://doi.org/10.1128/AEM.01906-10
Rodríguez-Díaz J, Carbajo RJ, Pineda-Lucena A, Monedero V, Yebra MJ (2013) Synthesis of fucosyl-N-acetylglucosamine disaccharides by transfucosylation using α-L-fucosidases from Lactobacillus casei. Appl Environ Microbiol 79:3847–3850. https://doi.org/10.1128/AEM.00229-13
Sakai K, Katsumi R, Ohi H, Usui T, Ishido Y (1992) Enzymatic syntheses of N-acetyllactosamine and N-acetylallolactosamine by the use of β-D-galactosidases. J Carbohydr Chem 11:553–565. https://doi.org/10.1080/07328309208016148
Sakurama H, Tsutsumi E, Ashida H, Katayama T, Yamamoto K, Kumagai H (2012a) Differences in the substrate specificities and active-site structures of two α-L-fucosidases (glycoside hydrolase family 29) from Bacteroides thetaiotaomicron. Biosci Biotechnol Biochem 76:1022–1024. https://doi.org/10.1271/bbb.111004
Sakurama H, Fushinobu S, Hidaka M, Yoshida E (2012b) 1,3-1,4-alpha-L-fucosynthase that specifically introduces Lewis a/x antigens into type-1/2 chains. J Biol Chem 287:16709–16719. https://doi.org/10.1074/jbc.M111.333781
Sigtryggsdóttir AR, Papaleo E, Thorbjarnardóttir SH, Kristjánsson MM (2014) Flexibility of cold- and heat-adapted subtilisin-like serine proteinases evaluated with fluorescence quenching and molecular dynamics. Biochim Biophys Acta 1844:705–712. https://doi.org/10.1016/j.bbapap.2014.02.009
Tanaka H, Ito F, Iwasaki T (1994) Two sialidases which preferentially hydrolyze sialyl alpha 2-8 linkage from Bacteroides fragilis SBT3182. J Biochem 115:318–321. https://doi.org/10.1093/oxfordjournals.jbchem.a124335
Troy EB, Kasper DL (2010) Beneficial effects of Bacteroides fragilis polysaccharides on the immune system. Front Biosci (Landmark Ed) 15:25–34. https://doi.org/10.2741/3603
Tsai TI, Li ST, Liu CP, Chen KY, Shivatare SS, Lin CW, Liao SF, Lin CW, Hsu TL, Wu YT, Tsai MH, Lai MY, Lin NH, Wu CY, Wong CH (2017) An effective bacterial fucosidase for glycoprotein remodeling. ACS Chem Biol 12:63–72. https://doi.org/10.1021/acschembio.6b00821
Tseng TH, Lin TW, Chen CY, Chen CH, Lin JL, Hsu TL, Wong CH (2017) Substrate preference and interplay of fucosyltransferase 8 and N-acetylglucosaminyltransferases. J Am Chem Soc 139:9431–9434. https://doi.org/10.1021/jacs.7b03729
Tübel J, Saldamli B, Wiest I, Jeschke U, Burgkart R (2012) Expression of the tumor markers sialyl Lewis A, sialyl Lewis X, Lewis Y, Thomsen-Friedenreich antigen, galectin-1 and galectin-3 in human osteoblasts in vitro. Anticancer Res 32:2159–2164. https://doi.org/10.1038/onc.2010.612
Vahidi S, Ripstein ZA, Bonomi M, Yuwen T, Mabanglo MF, Juravsky JB, Rizzolo K, Velyvis A, Houry WA, Vendruscolo M, Rubinstein JL, Kay LE (2018) Reversible inhibition of the ClpP protease via an N-terminal conformational switch. PNAS 115:E6447–E6456. https://doi.org/10.1073/pnas.1805125115
Vetere A, Galateo C, Paoletti S (1997) All-aqueous, regiospecific tranglycosylation synthesis of 3-O-α-L-fucopyranosyl-2-acetamido-2-deoxy-D-glucopyranose, a building block for the synthesis of branched oligosaccharides. Biochem Biophys Res Commun 234:358–361. https://doi.org/10.1006/bbrc.1997.6630
Weijers CAGM, Franssen MCR, Visser GM (2008) Glycosyltransferase-catalyzed synthesis of bioactive oligosaccharides. Biotechnol Adv 26:436–456. https://doi.org/10.1016/j.biotechadv.2008.05.001
Xu J, Bjursell MK, Himrod J, Deng S, Carmichael LK, Chiang HC, Hooper LV, Gordon JI (2003) A genomic view of the human-Bacteroides thetaiotaomicron symbiosis. Science 299:2074–2076. https://doi.org/10.1126/science.1080029
Yamamoto T, Ugai H, Nakayama-Imaohji H, Tada A, Elahi M, Houchi H, Kuwahara T (2018) Characterization of a recombinant Bacteroides fragilis sialidase expressed in Escherichia coli. Anaerobe 50:69–75. https://doi.org/10.1016/j.anaerobe.2018.02.003
Yang Q, Zhang R, Cai H, Wang LX (2017) Revisiting the substrate specificity of mammalian α1,6-fucosyltransferase reveals that it catalyzes core fucosylation of N-glycans lacking α1,3-arm GlcNAc. J Biol Chem 292:14796–14803. https://doi.org/10.1074/jbc.M117.804070
Yu H, Lau K, Li Y, Sugiarto G, Chen X (2012) One-pot multienzyme synthesis of Lewis x and sialyl Lewis x antigens. Curr Protoc Chem Biol 4:233–247. https://doi.org/10.1002/9780470559277.ch110277
Yu H, Li Y, Wu Z, Li L, Zeng J, Zhao C, Wu Y, Tasnima N, Wang J, Liu H, Gadi MR, Guan W, Wang PG, Chen X (2017) H. pylori α1-3/4-fucosyltransferase (Hp3/4FT)-catalyzed one-pot multienzyme (OPME) synthesis of Lewis antigens and human milk fucosides. Chem Commun (Camb) 53:11012–11015. https://doi.org/10.1039/c7cc05403c
Zechel DL, Withers SG (2000) Glycosidase mechanisms: anatomy of a finely tuned catalyst. Acc Chem Res 33:11–18. https://doi.org/10.1021/acs.accounts.9b00052
Zeuner B, Jers C, Mikkelsen JD, Meyer AS (2014) Methods for improving enzymatic trans-glycosylation for synthesis of human milk oligosaccharide biomimetics. J Agric Food Chem 62:9615–9631. https://doi.org/10.1021/jf502619p
Zhao S, Lieberman TD, Poyet M, Kauffman KM, Gibbons SM, Groussin M, Xavier RJ, Alm EJ (2019) Adaptive evolution within gut microbiomes of healthy people. Cell Host Microbe 25:656–667. https://doi.org/10.1016/j.chom.2019.03.007
Funding
This work was partly supported by National Key Research and Development Program of China (2018YFA0902000), the National Natural Science Foundation of China (31872626 and 31670062), and the Science and Technology Development Project of Shandong Province (2016GGH4502).
Author information
Authors and Affiliations
Contributions
PL and MX conceived and designed research. PL and HZ conducted experiments. YW contributed analytical tools. PL, XC, and LJ analyzed data. PL, LX, and MX wrote the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 830 kb).
Rights and permissions
About this article
Cite this article
Liu, P., Zhang, H., Wang, Y. et al. Screening and characterization of an α-L-fucosidase from Bacteroides fragilis NCTC9343 for synthesis of fucosyl-N-acetylglucosamine disaccharides. Appl Microbiol Biotechnol 104, 7827–7840 (2020). https://doi.org/10.1007/s00253-020-10759-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-10759-w