Abstract
Root colonization of beneficial rhizobacteria is critical for their beneficial effects. Quorum sensing (QS) has been reported to affect the colonization of many plant pathogens. However, how QS signals regulate root colonization of beneficial rhizobacteria is unclear. In this study, the QS signal AI-2 synthetase-encoding gene luxS was completely deleted from the genome of the plant beneficial rhizobacterium Bacillus velezensis SQR9, and bioluminescence experiments showed that AI-2 production was blocked. Deletion of luxS reduced biofilm formation, motility, and root colonization of B. velezensis SQR9, while addition of exogenous AI-2 to the mutant restored this phenomenon. These results indicated that AI-2 positively affects the root colonization of B. velezensis SQR9. This study provided new insights for enhancing the colonization of beneficial rhizobacteria.
Key points
• LuxS participated in the synthesis of the quorum sensing signal AI-2 in B. velezensis.
• AI-2 enhanced motility, biofilm formation, and root colonization of B. velezensis.
• AI-2 stimulated the production of γ-polyglutamic acid by B. velezensis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plant growth–promoting rhizobacteria (PGPRs) have been widely used in agriculture for their environmentally safe and plant growth–promoting properties (Lugtenberg and Kamilova 2009). Efficient root colonization of PGPRs is necessary for the bacteria to exert their beneficial effects on plants. Chemotactic motility towards roots (chemotaxis) and the formation of biofilms on the rhizoplane are the major processes of root colonization by PGPRs (Cao et al. 2011; Li et al. 2013). In the biofilm formation process, extracellular matrix components, such as extracellular polysaccharides (EPSs) and γ-polyglutamic acid (γ-PGA), play an important role in clustering the cells together. Motility and biofilm formation are regulated by a variety of factors, including both exogenous and endogenous signals. Quorum sensing (QS), which is associated with bacterial density and generally determined by endogenous signals, has been widely studied for its effects on the motility and biofilm formation of bacteria (Huber et al. 2001), especially pathogenic bacteria. Therefore, we speculated that QS would be an important factor that influences the root colonization of PGPRs.
QS is a process for regulating specific behaviors in a cell density–dependent manner (Bassler 1999). QS is accomplished by the release, detection, and transport of small signaling molecules called autoinducers that accumulate in proportion to cell density (Wai-Leung and Bassler 2015). With some exceptions, QS signals mainly include N-acyl-homoserine lactones (AHLs), cholera autoinducer 1 (CAI-1), autoinducer-2 (AI-2), and oligopeptides (Vendeville et al. 2005). The QS systems in gram-negative and gram-positive bacteria are different. While AHLs are the most common and well-studied class of autoinducers in gram-negative bacteria, these molecules have not been detected or are not functional in gram-positive bacteria. In gram-positive bacteria, QS is usually mediated by AI-2, and this molecule could be shared by both gram-negative and gram-positive bacteria (Xavier and Bassler 2003).
AI-2 is a 2-methyl-2,3,3,4-tetrahydroxytetrahydrofuran (THMF) formed through spontaneous rearrangements of the precursor 4,5-dihydroxy 2,3-pentanedione (DPD), which is synthesized by S-ribosylhomocysteinase LuxS (Schauder et al. 2001). AI-2 was first identified in the marine bacterium Vibrio harveyi and is part of a complex multilayered QS system responsible for regulating bioluminescence and other virulence-associated traits (Bassler et al. 1994). In addition, AI-2 has been proposed as a chemokine in Escherichia coli K12 (Hegde et al. 2011). The effect of AI-2 on biofilm formation varies in different bacterial species, and the Staphylococcus aureus luxS mutant strain formed stronger biofilms than the wild-type strain under static, flowing, and anaerobic conditions (Yu et al. 2012). Bacillus species are the most important group of PGPRs; however, how AI-2 and the QS system affect the root colonization of Bacillus species is not clear.
Bacillus velezensis SQR9 is a beneficial plant bacterium isolated from the rhizosphere of cucumber plants (Cao et al. 2011). This PGPR strain has shown several beneficial effects on plants, including promoting growth (Liu et al. 2016), inducing plant resistance (Wu et al. 2018), and enhancing plant tolerance to abiotic stress (Chen et al. 2017). In this study, the potential AI-2 synthetase-encoding gene luxS was deleted, and the mutant was assessed for AI-2 production, biofilm formation, motility, and root colonization. We demonstrated that AI-2 plays positive roles in biofilm formation, γ-PGA production, motility, and root colonization of B. velezensis SQR9.
Materials and methods
Strains and culture conditions
The strains and plasmids used in this study are shown in Table 1. B. velezensis strain SQR9 (CGMCC accession no. 5808, China General Microbiology Culture Collection Center, NCBI accession NO. CP006890) was cultured at 37 °C in Luria-Bertani medium (1% (w/v) tryptone, 0.5% (w/v) yeast extract, and 0.5% (w/v) NaCl). V. harveyi (from the Han lab of Shanghai Veterinary Research Institute, CAAS) was cultured at 30 °C in AB medium (0.05 M MgSO4, 0.3 M NaCl, 1% (w/v) glycerol, 10 mM K2HPO4, 10 mM L-arginine, and 0.02% (w/v) casamino acids). To measure biofilm formation and extracellular polymeric substances, B. velezensis SQR9 and its derived mutants were cultured in MSgg medium (100 mM 3-(N-morpholino) propane sulfonic acid (MOPS), 5 mM potassium phosphate, 2 mM MgCl2, 700 μM CaCl2, 50 μM MnCl2, 50 μM FeCl3, 1 μM ZnCl2, 2 μM thiamine, 0.5% (w/v) glycerol, 0.5% (w/v) glutamate, 50 μg/mL tryptophan, 50 μg/mL phenylalanine, and 50 μg/mL threonine, pH 7.0) (Branda et al. 2001). When necessary, antibiotics were added to the growth media at the following concentrations: erythromycin, 1 μg/mL; spectinomycin, 100 μg/mL; and zeocin, 20 μg/mL for B. velezensis strains.
luxS deletion and complementation
To disrupt the target gene in the SQR9 genome, an erythromycin resistance gene was used to replace the luxS gene to obtain the luxS mutant. The upstream and downstream regions that flanked the luxS gene were amplified from the SQR9 genome. The primers used to knock out the luxS gene in SQR9 are shown in Table S1. The upstream homologous fragment, the erythromycin gene fragment with the promoter, and the downstream homologous fragment were fused by using the two-step overlapping PCR method described by Shao et al. (2015), and the resulting product was transformed into B. velezensis SQR9 to generate the luxS mutant strain of Bacillus velezensis SQR9, ∆luxS. Complementation of the luxS gene was performed by inserting the luxS operon (including its promoter) into the ΔluxS mutant at the amylase gene locus (amyE) and inserting spectinomycin (Spc, amplified from plasmid p7S6) as the screening marker. All these mutants were verified by PCR amplification and sequencing.
Detection of AI-2
Detection of AI-2 was performed following the method reported by Shao et al. (2007). AI-2 detection was carried out using Vibrio harveyi BB 170 (sensor 1−, sensor 2+) as a reporter strain. V. harveyi BB170 was cultured in AB medium overnight at 30 °C and 170 rpm. The cell suspension with an OD600 of 0.2–0.3 was diluted with AB medium at 1:5000 as the reporter solution. At the same time, supernatants of wild-type SQR9, ΔluxS, and ΔluxS/luxS cultures were collected at exponential phase, filtered through a 0.22-μm membrane and filtered through a 3-kDa exclusion filter (Millipore). Subsequently, the filtrate was lyophilized, and the powder was dissolved in PBS solution (pH 7.0). In addition, 4 μM pure AI-2 (4,5-dihydroxy 2,3-pentanedione (DPD)) (Omm Scientific Inc, TX, USA) was included as a positive control. Both the culture medium (AB medium) for the reporter strain and PBS solution were included as negative controls. Then, 20 μL of each sample was added to 180 μL of reporter solution and incubated at 30 °C and 200 rpm. Bioluminescence was measured and recorded every hour.
Plants and growth conditions
The seeds of the “jingtian” maize cultivar were soaked in 75% ethanol for 40 s, washed with sterile water, and soaked in 0.2% (w/v) NaClO solution for 5 min. The seeds were placed on sterile wet filter paper for germination at 25 °C. The germinated seeds were transferred to sterile vermiculite and cultured at a suitable temperature for 3 days. Then, maize seedlings were transplanted to 100 mL flasks containing 75 mL of sterile liquid 1/4 Hoagland medium. The plants were grown with a 16-h light/8-h dark photoperiod. The seedlings were ready for the colonization experiment after culturing for an additional 15 days.
Biofilm formation assay
The biofilm assay was carried out in MSgg medium, as described by Hsueh et al. (2006). B. velezensis SQR9 and its derived strains were cultured in LB medium to an OD600 of 1.0. The cells were collected by centrifugation at 8000 rpm. The pellets were washed with sterile water three times and suspended in an equal volume of MSgg medium. The concentrations were adjusted to an OD600 of 1.0. Then, for qualitative analysis, 10 μL of each cell suspension was inoculated into a 48-well plate containing 1 mL of MSgg medium in each well. Exogenous AI-2 was added to the corresponding wells to a final concentration of 4 μM. The 48-well plate was cultured at 37 °C without shaking. Biofilm formation was evaluated at 16 h postinoculation. For quantitative analysis, 100 μL of each cell suspension was inoculated into 6-well plates containing 10 mL of MSgg medium in each well. Sterile biofilm filters 15-1040 (Biologixb, Changzhou, China) were placed in each well to collect the biofilm for weighing. After 16 h, the biofilm was weighed. For each treatment, four replicates were included.
Motility assay
The swimming and swarming assay was performed following the method of Sperandio and Inoue (Sperandio et al. 2002; Inoue et al. 2007). B. velezensis SQR9 and its derived strains were grown in LB medium until the OD600 reached 0.8. The cultures were inoculated on petri dishes containing semisolid LB medium (0.5% (w/v) glucose and 0.6% (w/v) Eiken agar (Eiken, Nogi-machi, Japan) for the swarming assay, and 0.3% (w/v) Eiken agar for the swimming assay) by sterilized toothpicks. The petri dishes were incubated at 37 °C for 4–6 h to allow the bacteria to swim. Then, the water content of the medium in the petri dishes was reduced with dry air to terminate the swimming process. Subsequently, the petri dish was incubated overnight at room temperature.
Detection of γ-PGA
The viscosity assay was performed as described by Feng et al. (2014) to determine the production of γ-PGA. For each strain, 1 mL of cell suspension with an OD600 of 1.0 was inoculated into a 250-mL flask containing 100 mL of MSgg medium. Exogenous AI-2 was added to the corresponding wells to a final concentration of 4 μM. The bacteria were cultured with shaking at 170 rpm at 37 °C. At 24 h postinoculation, the viscosity of 20 mL of fermentation broth was measured by an NDJ-8S rotatory viscometer (Ping Xuan, Shanghai, China) with an adapted No. 0 rotator. The detection speed was set at 60 rpm for samples with a viscosity under 10 mPa s and 30 rpm for samples with a viscosity over 10 mPa s. For each treatment, three replicates were included.
The cetyltrimethylammonium bromide (CTAB) detection assay was performed as described by Ashiuchi (2011). In solutions with high ionic strength, CTAB forms complexes with proteins and non-acid polymers, so the concentration of γ-PGA can be calculated by the turbidity of the complexes. For each strain, 1 mL of cell suspension with an OD600 of 1.0 was inoculated into a 250-mL flask containing 100 mL of MSgg medium. Exogenous AI-2 was added to the corresponding wells to a final concentration of 4 μM. The bacteria were cultured with shaking at 170 rpm at 37 °C. At 24 h postinoculation, γ-PGA was purified using a previously described method (Goto and Kunioka 1992), and pure γ-PGA was dissolved in an equal volume of ddH2O for further detection. At the same time, standard γ-PGA (Yuan Ye, Shanghai, China) solutions were prepared with concentrations of 0.0391, 0.0781, 0.1563, 0.3125, 0.625, 1.25, 2.5, and 5 g/L. Then, 2 mL of each sample solution or standard γ-PGA solution was mixed with 2 mL of CTAB solution (2% NaOH, 5 g/L CTAB); subsequently, the OD250 was measured by Infinitr M200PRO (TECAN, Männedorf, Switzerland). The production of γ-PGA was calculated based on the standard curve. For each treatment, three replicates were included.
Colonization assay
The maize seedlings were grown in a hydroponic system as described above and divided into three groups for treatment. B. velezensis SQR9 and its derivative strains were cultured in LB medium to an OD600 of 1.0. The cells were collected by centrifugation at 8000 rpm. The pellets were washed with sterile water three times and resuspended in the same volume. The bacterial cells were inoculated into the maize seedling rhizosphere to a final concentration with an OD600 of 0.1. Exogenous AI-2 was added to the corresponding treatment at a final concentration of 4 μM. After incubation for 3 days, the population of the cells colonizing the rhizoplane was determined by plate counting as described previously (Liu et al. 2017). For each treatment, six replicates were included.
Result
LuxS is necessary for AI-2 production in B. velezensis SQR9
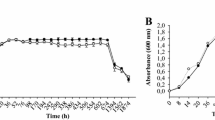
The luxS gene encoding S-ribosylhomocysteinase, which catalyzes the production of AI-2 from S-ribosylhomocysteine (SRH), was identified in the B. velezensis SQR9 genome. Deletion of luxS did not affect the growth of B. velezensis SQR9 (Figure S1). To verify its function in AI-2 production, the SQR9 wild-type strain and the luxS mutant were tested for AI-2 production by bioluminescence assay. For this experiment, both the culture medium (AB medium) for the reporter strain and the PBS solution were included as negative controls. Pure AI-2 was applied as a positive control. The results showed that deletion of luxS significantly decreased AI-2 production while reintroducing luxS into the mutant restored AI-2 production (Fig. 1). These results indicated that luxS is responsible for AI-2 signal production in SQR9.
Measurement of AI-2 production by bioluminescence assay. AI-2 in the culture medium of the SQR9 wild-type, ΔluxS and ΔluxS/luxS strains was tested using a reporter strain V. harveyi BB170, whose luminescence was enhanced with increasing concentrations of AI-2 in the environment. Pure AI-2 was applied with the final concentration of 4 μM as a positive control, and AB medium was included as a negative control. Error bars indicate the standard deviations based on six independent replicates
AI-2 signal promotes biofilm formation of B. velezensis SQR9
To test the effect of AI-2 on biofilm formation of B. velezensis SQR9, both qualitative and quantitative analysis of biofilm formation by the wild-type strain and the luxS deletion mutant in MSgg medium were performed. At 16 h postinoculation, wild-type SQR9 formed more wrinkles than ΔluxS, and the wet weight of the pellicle biofilm of the wild-type strain was 1.6-fold that of the ΔluxS strain. Reintroducing luxS into the mutant with its original promoter or adding 4 μM AI-2 restored biofilm formation. Moreover, the addition of exogenously AI-2 further increased the biofilm of the wild-type strain (Fig. 2). These results showed that AI-2 could enhance biofilm formation of B. velezensis SQR9.
Biofilm formation of B. velezensis SQR9. WT, wild-type; WT+AI-2, wild-type strain supplied with exogenous AI-2; ΔluxS, luxS mutant; ΔluxS/AI-2, ΔluxS supplied with exogenous AI-2; ΔluxS/luxS, ΔluxS complemented with luxS gene at the amylase gene locus (amyE). The final concentration of exogenous AI-2 was 4 μM. a Biofilm formation in 48-well plates. b The weight of biofilms at 16 h. Error bars indicate the standard deviations based on four independently replicated experimental values
AI-2 signal increases the synthesis of γ-PGA by B. velezensis SQR9
The extracellular matrix is important for biofilm formation. To investigate how AI-2 stimulates the biofilm formation of strain SQR9, the effect of AI-2 on the Bacillus biofilm matrix, including exopolysaccharides (EPSs), the amyloid-like protein TasA, and γ-polyglutamic acid (γ-PGA), were tested. As the extracellular matrix can be analyzed by the viscosity of the supernatant, we first measured the viscosity of the wild-type strain and ΔluxS mutant. The results showed that the SOR9 wild-type strain had a higher culture supernatant viscosity than the ΔluxS mutant. The viscosity of the culture supernatant of the wild-type strain was 7.48 mPa s, while that of ΔluxS was decreased to 2.32 mPa s. Complementation with luxS and adding 4 μM AI-2 restored the viscosity of the cultures to 5.32 mPa s and 6.33 mPa s, respectively. In addition, exogenously adding AI-2 to wild-type SQR9 cultures resulted in a viscosity of 11.64 mPa s (Fig. 3a).
The γ-PGA yield of B. velezensis SQR9. WT, wild-type; WT+AI-2, wild-type strain supplied with exogenous AI-2; ΔluxS, luxS mutant; ΔluxS/AI-2, ΔluxS supplied with exogenous AI-2; ΔluxS/luxS, ΔluxS complemented with luxS gene at the amylase gene locus (amyE). The final concentration of exogenous AI-2 was 4 μM. a The viscosities of fermentations. b The yield of γ-PGA. Error bars indicate the standard deviations based on three independently replicated experimental values
The synthesis of EPS and TasA by the SOR9 wild-type and ΔluxS mutant strains was similar (Figure S2 and Figure S3), and we speculate that biofilm matrix γ-PGA synthesis may be different between the SQR9 wild-type and ΔluxS mutant strains. CTAB can bind γ-PGA specifically to form a water-insoluble, highly dispersed micelle-like complex, which can be measured on the basis of the turbidity to determine the concentration of γ-PGA. Therefore, we used CTAB solution to react with the fermentation supernatant of each treatment. The results showed that the concentration of γ-PGA in the supernatant of the SQR9 wild-type strain was 0.92 g/L, and adding exogenous AI-2 to wild-type SQR9 increased the concentration of γ-PGA in the supernatant to 2.50 g/L. When luxS was deleted from the wild-type strain, the concentration of γ-PGA in the supernatant was decreased to 0.08 g/L, and complementation of the mutant with luxS and adding 4 μM AI-2 restored the concentration of γ-PGA to 0.40 g/L and 0.62 g/L, respectively (Fig. 3b). SDS-PAGE of the γ-PGA samples from each treatment had the same tendency (Figure S4). The results showed that AI-2 affected the synthesis of γ-PGA in B. velezensis SQR9.
AI-2 signal promotes motility of B. velezensis SQR9
Bacterial motility is critical for PGPR to colonize plant roots. To investigate the effect of AI-2 signaling on the motility of SQR9, swimming analysis of the SQR9 wild-type strain, ΔluxS mutant strain, and luxS gene complementation strain was performed; the results were compared and showed that the diameter of the motility circle of ΔluxS was significantly decreased compared with that of SQR9. Complementation with luxS or adding 4 μM AI-2 to the medium restored the motility circle of ΔluxS mutant strain to the wild-type level (Fig. 4). This result indicates that AI-2 has a positive effect on the motility ability of SQR9.
Motility analysis of B. velezensis SQR9. WT, wild-type; WT+AI-2, wild-type strain supplied with exogenous AI-2; ΔluxS, luxS mutant; ΔluxS/AI-2, ΔluxS supplied with exogenous AI-2; ΔluxS/luxS, ΔluxS complemented with luxS gene at the amylase gene locus (amyE). The final concentration of exogenous AI-2 was 4 μM. a Swimming of cells on LB swim medium solidified with 0.3% agar. b Swarming of cells on LB swarming medium solidified with 0.6% agar and 0.5% glucose. c The diameters of cell spots inoculated on swarming plates that incubated at 37 °C for 4 h. Error bars indicate the standard deviations based on six independently replicated experimental values
AI-2 signal enhances the root colonization of B. velezensis SQR9
Biofilm formation and motility are important factors in determining the root colonization of PGPRs. As shown in this study, the effects of LuxS and AI-2 on biofilm formation and motility are both positive. We proposed that LuxS/AI-2 should have a positive effect on root colonization. Quantitative analysis of colonization of maize roots was performed at 3 days postinoculation using the plate counting method. The results showed that the root colonization of ΔluxS decreased significantly by 35%. When luxS was introduced into the mutant, the colonization of the resulting strain on maize roots was restored to the same level as that of the wild-type strain. When exogenous AI-2 was added to the rhizosphere of maize, colonization of SQR9 on roots significantly increased by 16% (Fig. 5). Exogenous AI-2 also rescued the colonization decreasing caused by luxS mutation. These results indicated that AI-2 could promote root colonization of SQR9.
Root colonization of B. velezensis SQR9. WT, wild-type; WT+AI-2, wild-type strain supplied with exogenous AI-2; ΔluxS, luxS mutant; ΔluxS/AI-2, ΔluxS supplied with exogenous AI-2; ΔluxS/luxS, ΔluxS complemented with luxS gene at the amylase gene locus (amyE). The final concentration of exogenous AI-2 was 4 μM. Error bars indicate the standard deviations based on six independently replicated experimental values
Discussion
Quorum sensing is an important system that regulates specific functions in bacteria. Bacillus is the most widely used plant beneficial bacteria in agriculture, but knowledge on how QS signals regulate the rhizospheric behavior of Bacillus is lacking. In this study, the QS signal AI-2 synthetase-encoding gene luxS was deleted from the B. velezensis SQR9 genome. AI-2 production was completely blocked in the luxS mutant, and biofilm formation, motility, and root colonization of the luxS mutant were significantly reduced. These results indicated that AI-2 could act as a quorum sensing signal and promote root colonization of B. velezensis. This finding would provide new insights into the colonization of PGPR.
Quorum sensing of gram-positive bacteria is regulated by few kinds of oligopeptides and AI-2. The signaling oligopeptides are generally produced by specific cleavage of proteins. For example, bacterial pheromones, which play roles in sporulation and the production of certain secondary metabolites (Rosenberg et al. 2012), are cleaved from the ComX protein. Nevertheless, knowledge of the QS signal in gram-positive bacteria is still very poor. In comparison with other QS signals, AI-2 more widely exists in both gram-negative and gram-positive bacteria, and the structure of AI-2 is highly conserved among species, giving it the capability to act as an interspecies QS signal. However, pathways regulated by AI-2 among different bacteria are varied. Merritt et al. found that deletion of the AI-2 synthetase LuxS could affect the biofilm morphology of Streptococcus mutans (Merritt et al. 2003). Similarly, the addition of AI-2 promoted the biofilm formation of Staphylococcus epidermidis (Xue et al. 2015) and Bacillus cereus (Auger et al. 2006). This study also showed similar results that AI-2 could positively affect biofilm formation of B. velezensis SQR9. It has been reported that deletion of luxS from E. coli K-12 led to enhanced motility (Ling et al. 2010). However, in Bacillus subtilis, deletion of luxS led to reduced motility (Lombardía et al. 2006). Similarly, in this study, AI-2 also showed a positive effect on the motility of B. velezensis SQR9.
Bacillus biofilm formation is a complex process regulated by a range of regulation systems, such as Spo0A, SinI-SinR (Xu et al. 2017), and the DegSU system (Verhamme et al. 2009). In this study, AI-2 increased the production of γ-PGA. The synthesis of γ-PGA is regulated by phosphorylated DegU (Stanley and Lazazzera 2005). Interestingly, the pheromone synthesized from ComX is also involved in the regulation of the DegSU system. The pheromone could promote the transcription of degQ, which facilitates the transfer of an activating phosphate from DegS to DegU. It is proposed that AI-2 might have a similar pathway as the ComX pheromone and regulate biofilm formation of B. velezensis. Further study on how AI-2 participates in biofilm formation regulation would provide insight into the AI-2 and ComX pheromone signaling pathways.
This study revealed that AI-2 had the same effect on the synthesis of γ-PGA and biofilm formation. Therefore, it is presumed that AI-2 could affect biofilm formation in Bacillus spp. by influencing the synthesis of γ-PGA. γ-PGA is a natural polymer with many good properties, such as being nontoxic, edible, degradable, water soluble, and modifiable. γ-PGA can be used in fertilizers to enhance their effectiveness (Hoppensack et al. 2003), as well as biological flocculants for sewage treatment (Ritchie et al. 1999). γ-PGA is also used in food and cosmetics (Shyu et al. 2008), and it is widely used in medicine; it can be used as a drug carrier to improve drug solubility (Liang et al. 2006). γ-PGA is mainly produced by microbial fermentation; therefore, it is very important to improve the efficiency of microbial synthesis of γ-PGA. Our study found that the addition of AI-2 can significantly increase γ-PGA production in SQR9, which is important for the production of γ-PGA in industry.
References
Ashiuchi M (2011) Analytical approaches to poly-γ-glutamate: quantification, molecular size determination, and stereochemistry investigation. J Chromatogr B Anal Technol Biomed Life Sci 879:3096–3101. https://doi.org/10.1016/j.jchromb.2011.03.029
Auger S, Krin E, Aymerich S, Gohar M (2006) Autoinducer 2 affects biofilm formation by Bacillus cereus. Appl Environ Microbiol 72:937–941. https://doi.org/10.1128/AEM.72.1.937
Bassler BL (1999) How bacteria talk to each other: regulation of gene expression by quorum sensing. Curr Opin Microbiol 2:582–587. https://doi.org/10.1016/S1369-5274(99)00025-9
Bassler BL, Wright M, Silverman MR (1994) Multiple signalling systems controlling expression of luminescence in Vibrio harveyi: sequence and function of genes encoding a second sensory pathway. Mol Microbiol 13:273–286. https://doi.org/10.1111/j.1365-2958.1994.tb00422.x
Branda SS, González-Pastor JE, Ben-Yehuda S, Losick R, Kolter R (2001) Fruiting body formation by Bacillus. Proc Natl Acad Sci U S A 98:11621–11626. https://doi.org/10.1073/pnas.191384198
Cao Y, Zhang Z, Ling N, Yuan Y, Zheng X, Shen B, Shen Q (2011) Bacillus subtilis SQR9 can control Fusarium wilt in cucumber by colonizing plant roots. Biol Fertil Soils 47:495–506. https://doi.org/10.1007/s00374-011-0556-2
Chen L, Liu Y, Wu G, Zhang N, Shen Q, Zhang R (2017) Beneficial rhizobacterium Bacillus amyloliquefaciens SQR9 induces plant salt tolerance through spermidine production. Mol Plant-Microbe Interact 30:423–432. https://doi.org/10.1094/MPMI-02-17-0027-R
Feng J, Gao W, Gu Y, Zhang W, Cao M, Song C, Zhang P, Sun M, Yang C, Wang S (2014) Functions of poly-gamma-glutamic acid (γ-PGA) degradation genes in γ-PGA synthesis and cell morphology maintenance. Appl Microbiol Biotechnol 98:6397–6407. https://doi.org/10.1007/s00253-014-5729-0
Goto A, Kunioka M (1992) Biosynthesis and hydrolysis of poly(γ-glutamic acid) from Bacillus subtilis IF03335. Biosci Biotechnol Biochem 56:1031–1035. https://doi.org/10.1271/bbb.56.1031
Hegde M, Englert DL, Schrock S, Cohn WB, Vogt C, Wood TK, Manson MD, Jayaraman A (2011) Chemotaxis to the quorum-sensing signal AI-2 requires the Tsr chemoreceptor and the periplasmic LsrB AI-2-binding protein. J Bacteriol 193:768–773. https://doi.org/10.1128/JB.01196-10
Hoppensack A, Oppermann-Sanio FB, Steinbüchel A (2003) Conversion of the nitrogen content in liquid manure into biomass and polyglutamic acid by a newly isolated strain of Bacillus licheniformis. FEMS Microbiol Lett 218:39–45. https://doi.org/10.1016/S0378-1097(02)01132-1
Hsueh YH, Somers EB, Lereclus D, Wong ACL (2006) Biofilm formation by Bacillus cereus is influenced by PlcR, a pleiotropic regulator. Appl Environ Microbiol 72:5089–5092. https://doi.org/10.1128/AEM.00573-06
Huber B, Riedel K, Hentzer M, Heydorn A, Gotschlich A, Givskov M, Molin S, Eberl L (2001) The cep quorum-sensing system of Burkholderia cepacia H111 controls biofilm formation and swarming motility. Microbiology 147:2517–2528. https://doi.org/10.1099/00221287-147-9-2517
Inoue T, Shingaki R, Hirose S, Waki K, Mori H, Fukui K (2007) Genome-wide screening of genes required for swarming motility in Escherichia coli K-12. J Bacteriol 189:950–957. https://doi.org/10.1128/JB.01294-06
Li S, Zhang N, Zhang Z, Luo J, Shen B, Zhang R, Shen Q (2013) Antagonist Bacillus subtilis HJ5 controls Verticillium wilt of cotton by root colonization and biofilm formation. Biol Fertil Soils 49:295–303. https://doi.org/10.1007/s00374-012-0718-x
Li Q, Li Z, Li X, Xia L, Zhou X, Xu Z, Shao J, Shen Q, Zhang R (2018) FtsEX-CwlO regulates biofilm formation by a plant-beneficial rhizobacterium Bacillus velezensis SQR9. Res Microbiol 169:166–176. https://doi.org/10.1016/j.resmic.2018.01.004
Liang HF, Chen SC, Chen MC, Lee PW, Chen CT, Sung HW (2006) Paclitaxel-loaded poly(γ-glutamic acid)-poly(lactide) nanoparticles as a targeted drug delivery system against cultured HepG2 cells. Bioconjug Chem 17:291–299. https://doi.org/10.1021/bc0502107
Ling H, Kang A, Tan MH, Qi X, Chang MW (2010) The absence of the luxS gene increases swimming motility and flagella synthesis in Escherichia coli K12. Biochem Biophys Res Commun 401:521–526. https://doi.org/10.1016/j.bbrc.2010.09.080
Liu Y, Chen L, Zhang N, Li Z, Zhang G, Xu Y, Shen Q, Zhang R (2016) Plant-microbe communication enhances auxin biosynthesis by a root-associated bacterium, Bacillus amyloliquefaciens SQR9. Mol Plant-Microbe Interact 29:324–330. https://doi.org/10.1094/MPMI-10-15-0239-R
Liu Y, Chen L, Wu G, Feng H, Zhang G, Shen Q, Zhang R (2017) Identification of root-secreted compounds involved in the communication between cucumber, the beneficial Bacillus amyloliquefaciens, and the soil-borne pathogen fusarium oxysporum. Mol Plant-Microbe Interact 30:53–62. https://doi.org/10.1094/MPMI-07-16-0131-R
Lombardía E, Rovetto AJ, Arabolaza AL, Grau RR (2006) A LuxS-dependent cell-to-cell language regulates social behavior and development in Bacillus subtilis. J Bacteriol 188:4442–4452. https://doi.org/10.1128/JB.00165-06
Lugtenberg B, Kamilova F (2009) Plant-growth-promoting rhizobacteria. Annu Rev Microbiol 63:541–556. https://doi.org/10.1146/annurev.micro.62.081307.162918
Merritt J, Qi F, Goodman SD, Anderson MH, Shi W (2003) Mutation of luxS affects biofilm formation in Streptococcus mutans. Infect Immun 71:1972–1979. https://doi.org/10.1128/IAI.71.4.1972
Ritchie SMC, Bachas LG, Olin T, Sikdar SK, Bhattacharyya D (1999) Surface modification of silica- and cellulose-based microfiltration membranes with functional polyamino acids for heavy metal sorption. Langmuir 15:6346–6357. https://doi.org/10.1021/la9814438
Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F (2012) The prokaryotes: prokaryotic communities and ecophysiology. Prokaryotes Prokaryotic Communities Ecophysiol 2:1–528. https://doi.org/10.1007/978-3-642-30123-0
Schauder S, Shokat K, Surette MG, Bassler BL (2001) The LuxS family of bacterial autoinducers: biosynthesis of a novel quorum-sensing signal molecule. Mol Microbiol 41:463–476. https://doi.org/10.1046/j.1365-2958.2001.02532.x
Shao H, Lamont RJ, Demuth DR (2007) Autoinducer 2 is required for biofilm growth of Aggregatibacter (Actinobacillus) actinomycetemcomitans. Infect Immun 75:4211–4218. https://doi.org/10.1128/IAI.00402-07
Shao J, Li S, Zhang N, Cui X, Zhou X, Zhang G, Shen Q, Zhang R (2015) Analysis and cloning of the synthetic pathway of the phytohormone indole-3-acetic acid in the plant-beneficial Bacillus amyloliquefaciens SQR9. Microb Cell Factories 14:1–13. https://doi.org/10.1186/s12934-015-0323-4
Shyu YS, Hwang JY, Hsu CK (2008) Improving the rheological and thermal properties of wheat dough by the addition of γ-polyglutamic acid. LWT Food Sci Technol 41:982–987. https://doi.org/10.1016/j.lwt.2007.06.015
Sperandio V, Torres AG, Kaper JB (2002) Quorum sensing Escherichia coli regulators B and C (QseBC): a novel two-component regulatory system involved in the regulation of flagella and motility by quorum sensing in E. coli. Mol Microbiol 43:809–821. https://doi.org/10.1046/j.1365-2958.2002.02803.x
Stanley NR, Lazazzera BA (2005) Defining the genetic differences between wild and domestic strains of Bacillus subtilis that affect poly-γ-DL-glutamic acid production and biofilm formation. Mol Microbiol 57:1143–1158. https://doi.org/10.1111/j.1365-2958.2005.04746.x
Vendeville A, Winzer K, Heurlier K, Tang CM, Hardie KR (2005) Making “sense” of metabolism: autoinducer-2, LuxS and pathogenic bacteria. Nat Rev Microbiol 3:383–396. https://doi.org/10.1038/nrmicro1146
Verhamme DT, Murray EJ, Stanley-Wall NR (2009) DegU and Spo0A jointly control transcription of two loci required for complex colony development by Bacillus subtilis. J Bacteriol 91:100–108. https://doi.org/10.1128/JB.01236-08
Wai-Leung NG, Bassler B (2015) Bacterial quorum-sensing network architectures. Annu Rev Genet:197–222. https://doi.org/10.1146/annurev-genet-102108-134304.Bacterial
Wu G, Liu Y, Xu Y, Zhang G, Shen Q, Zhang R (2018) Exploring elicitors of the beneficial rhizobacterium Bacillus amyloliquefaciens SQR9 to induce plant systemic resistance and their interactions with plant signaling pathways. Mol Plant-Microbe Interact 31:560–567. https://doi.org/10.1094/MPMI-11-17-0273-R
Xavier KB, Bassler BL (2003) LuxS quorum sensing: more than just a numbers game. Curr Opin Microbiol 6:191–197. https://doi.org/10.1016/S1369-5274(03)00028-6
Xu S, Yang N, Zheng S, Yan F, Jiang C, Yu Y, Guo J, Chai Y, Chen Y (2017) The spo0A-sinI-sinR regulatory circuit plays an essential role in biofilm formation, nematicidal activities, and plant protection in Bacillus cereus AR156. Mol Plant-Microbe Interact 30:603–619. https://doi.org/10.1094/MPMI-02-17-0042-R
Xue T, Ni J, Shang F, Chen X, Zhang M (2015) Autoinducer-2 increases biofilm formation via an ica- and bhp-dependent manner in Staphylococcus epidermidis RP62A. Microbes Infect 17:345–352. https://doi.org/10.1016/j.micinf.2015.01.003
Yan X, Yu HJ, Hong Q, Li SP (2008) Cre/lox system and PCR-based genome engineering in Bacillus subtilis. Appl Environ Microbiol 74:5556–5562. https://doi.org/10.1128/AEM.01156-08
Yu D, Zhao L, Xue T, Sun B (2012) Staphylococcus aureus autoinducer-2 quorum sensing decreases biofilm formation in an icaR-dependent manner. BMC Microbiol 12:1. https://doi.org/10.1186/1471-2180-12-288
Acknowledgments
We thank Prof. Xiangan Han of Shanghai Veterinary Research Institute, Chinese Academy of Agricultural Sciences for providing us the Vibrio harveyi BB 170 reporter strain.
Funding
This work was funded by the National Natural Science Foundation of China grants 31670113 and 31870096, the National Key R & D Program of China grant 2018YFD0500201.
Author information
Authors and Affiliations
Contributions
RZ and GZ conceived and designed research. QX, DL, and HZ conducted experiments. XD and YL analyzed data. QX and YL wrote the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 446 kb).
Rights and permissions
About this article
Cite this article
Xiong, Q., Liu, D., Zhang, H. et al. Quorum sensing signal autoinducer-2 promotes root colonization of Bacillus velezensis SQR9 by affecting biofilm formation and motility. Appl Microbiol Biotechnol 104, 7177–7185 (2020). https://doi.org/10.1007/s00253-020-10713-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-10713-w