Abstract
2-Phenylethanol (2-PE) is an important flavor compound but also impairs cell growth severely, which in turn blocks its bioproduction. However, the molecular mechanism of 2-PE tolerance is unclear. In this study, a superb 2-PE stress-tolerant and producing yeast, Candida glycerinogenes, was selected to uncover the underlying mechanism of 2-PE tolerance. We discovered that Hap5 is an essential regulator to 2-PE resistance, and its induction by 2-PE stress occurs at the post-transcriptional level, rather than at the transcriptional level. Under 2-PE stress, Hap5 is activated and imported into the nucleus rapidly. Then, the nuclear Hap5 binds to the glutathione synthetase (gsh2) promoter via CCAAT box, to induce the expression of gsh2 gene. The increased gsh2 expression contributes to enhanced cellular glutathione content, and consequently alleviates ROS accumulation, lipid peroxidation, and cell membrane damage caused by 2-PE toxicity. Specifically, increasing the expression of gsh2 is effective in improving not just 2-PE tolerance (33.7% higher biomass under 29 mM 2-PE), but also 2-PE production (16.2% higher). This study extends our knowledge of 2-PE tolerance mechanism and also provides a promising strategy to improve 2-PE production.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Recently, microbial biosynthesis of 2-phenylethanol (2-PE) has gained great attention, since chemically synthesized 2-PE contains carcinogenic byproducts and extracted one from flowers is costly (Etschmann et al. 2002; Hua and Xu 2011; Mei et al. 2009). However, due to the aromatic structure, 2-PE exhibits much higher cell toxicity than ethanol. Most bacteria and yeasts completely lost growth at the concentration between 16 and 25 mM 2-PE (Stark et al. 2003), resulting in very low 2-PE yield (almost below 33 mM) (Chen et al. 2016; Liu et al. 2018a; Wang et al. 2019). High levels of 2-PE damages the cell membrane integrity and nutrient transport, disrupts mitochondrial respiration, and inhibits DNA, RNA, and protein synthesis (Corre et al. 1990; Lester 1965; Terenzi and Storck 1969).
Mechanisms of ethanol tolerance have been extensively studied including gene expression responses, signal-transduction pathways, and regulatory networks (Ma and Liu 2010a). For example, numerous genes involved in fatty acid, ergosterol, and glycolipid metabolism were up- or downregulated to remodel cell membrane components to resist ethanol stress (Alexandre et al. 2001; Chandler et al. 2004; Diniz et al. 2017; Ma and Liu 2010b). Improving microbial tolerance to ethanol through engineering specific gene targets, e.g., relating to efflux pumps, cell membrane, and energy generation, can lead to higher production (Tan et al. 2017; Teixeira et al. 2012; Zhang et al. 2018, 2019). However, unlike ethanol, 2-PE tolerance mechanism is poorly understood, which limits the improvement of 2-PE tolerance through the knowledge-based rational engineering strategy. To overcome 2-PE stress, several strategies including in situ product recovery (ISPR) techniques, screening more resistant strains, and cells immobilization have been studied (Martínez-Avila et al. 2018). However, these strategies are either costly or time-consuming. Importantly, they do not shed light on the underlying genetic information of 2-PE tolerance. Genome-scale engineering, by contrast, is a powerful tool to uncover genetic mechanisms and engineer microorganisms (Liu et al. 2018b; Si et al. 2017).
Candida glycerinogenes is a GRAS diploid industrial yeast with an excellent multi-stress tolerant property (Hou et al. 2019; Yang et al. 2018; Zhuge et al. 2001). Except for the application in commercial scale production of glycerol, C. glycerinogenes has also emerged as one of the best 2-PE producers (Lu et al. 2016; Zhuge et al. 2001), and exhibits superb 2-PE tolerance, which can even grow at 33 mM 2-PE, surpassing most commonly used 2-PE producers, such as Saccharomyces cerevisiae, Kluyveromyces marxianus, and Escherichia coli (Kang et al. 2014; Lu et al. 2016; Seward et al. 2013). This merit suggests that C. glycerinogenes has evolved efficient defense mechanisms against 2-PE toxicity, which makes it a promising candidate to explore molecular mechanism of 2-PE tolerance and provide efficient gene targets for constructing other robust industrial strains.
To improve 2-PE production, it is necessary to explore the mechanism of 2-PE tolerance. In this study, we identified a 2-PE-resistant gene (gsh2) in C. glycerinogenes by screening a plasmid-based genomic library. We further demonstrated that increasing gsh2 expression was effective in improving 2-PE tolerance and production. Additionally, the upstream regulation mechanism of gsh2 and its exact role in resisting 2-PE stress were then investigated.
Materials and methods
Strains, media, and culture conditions
All strains used for this study are described in Table S1. Escherichia coli DH5α used for plasmid construction was cultured in Luriae-Bertani medium. C. glycerinogenes WL2002-5 (CCTCC M93018) and its derived strains were grown in YPD medium (20 g/L glucose, 20 g/L peptone, and 10 g/L yeast extract, pH 6.0). These derived strains were screened on YNB medium (20 g/L glucose and 6.7 g/L YNB without amino acids) or YPD medium supplemented with 2 g/L 5-fluoroanthranilic acid (FAA) agar plates. For 2-PE production, C. glycerinogenes strains were grown in fermentation medium (90 g/L glucose, 7 g/L L-phenylalanine, 5 g/L KH2PO4, 0.5 g/L MgSO4·7H2O, 6.7 g/L YNB without amino acids and ammonium sulfate, pH 4.5). S. cerevisiae W303-1A used for isolation of 2-PE-resistant genes was cultured in YPD medium, and its derived strains were selected on synthetic complete leucine drop-out medium (SC-leu, 20 g/L glucose, 6.7 g/L YNB without amino acids, 40 mg/L each of histidine, tryptophan, uracil and adenine) agar plates. Agar at 2% was supplemented for plates. All yeast strains were incubated at 30 °C with shaking at 200 rpm.
Screening for genes to improve 2-PE tolerance
The C. glycerinogenes genomic DNA library, which was constructed using yeast episomal plasmid YEp51 with a 2 μ origin of replication and a leu2 marker as previously described (Li et al. 2005), was used to screen for genes conferring 2-PE resistance when overexpressed. Briefly, YEp51 genomic library of C. glycerinogenes was transformed into S. cerevisiae W303-1A. S. cerevisiae transformants were firstly screened on SC-Leu medium agar plates, then washed into SC-Leu liquid medium containing 25 mM 2-PE to enrich transformants with higher resistance to 2-PE. Finally, 2-PE tolerant transformants were obtained from selecting the large colonies from agar plate containing 25 mM 2-PE, while control strain with empty vector YEp51 did not grow. The isolated plasmids from 2-PE tolerant transformants were sequenced and compared with C. glycerinogenes genome sequence to identify the insert genes. To validate whether these insert genes are also involved in enhanced 2-PE tolerance in C. glycerinogenes, they were single overexpressed in C. glycerinogenes as described in the following section.
DNA manipulation
For construction of gsh2 gene (GenBank accession number: MN159189) overexpression plasmid, gsh2 was amplified from the genomic DNA of C. glycerinogenes using the primers (Table S2). Then the PCR products were ligated into the integrated expression plasmid pGAPa (driven by GAP promoter) with restriction sites BamH I and Sac II, generating plasmid pGAPa-gsh2. For construction of the gsh2 gene silencing plasmids, two gsh2 positions, − 300 to − 1 bp (A) and − 100 to + 100 bp (B), were amplified and reverse inserted into pGAPa between BamH I and Sac II sites, generating plasmids pGAPa-asgsh2-A and pGAPa-asgsh2-B. pGAPa and its derived plasmids were linearized with Hind III, then integrated into the chromosome of C. glycerinogenes UA5 at 18S rDNA gene site using LiAC/SS carrier DNA/PEG method (Ji et al. 2017), and screened from YNB medium agar plates, obtaining target genes overexpressing or silencing C. glycerinogenes strains. The general procedure of deleting genes in C. glycerinogenes used transient CRISPR-Cas9 method according to Zhu et al. (2019). Take hap5 gene for example, the 20-bp guide sequences of hap5 was analyzed by bioinformatics tool sgRNACas9, then sgRNA gene was synthesized and ligated into pMY plasmid by Synbio Technology (Suzhou, China) to generate single-guide RNA targeting plasmid pMY-sghap5. The sgRNA and CgCas9 cassettes were respectively amplified from pMY-sghap5 and pMYC9 both using the primer pair CgGAPp_F and PpAOX1t_R. The donor DNA containing auxotroph marker gene ura5 gene was amplified with primers, which had 50-bp sequence homologous to upstream and downstream of hap5 gene. The donor DNA, sgRNA, and CgCas9 cassettes were co-transformed into C. glycerinogenes UA5 using LiAC/SS carrier DNA/PEG method (Ji et al. 2017), and screened from YNB medium agar plates, yielding hap5 knockout strain. Repair template primers and synthetic sgRNA sequences targeting hap5, yap1, and rgt1 genes (GenBank accession numbers: MN159190-92) were shown in Table S2 and Table S3, respectively.
Spot, growth, and viability assays and 2-PE fermentation analysis
These procedures of spot and growth assay were performed as previously described (Lu et al. 2016). For viability measure, yeast cells grown at YPD liquid medium with or without 41 mM 2-PE, 0.1 mM glutathione, and 0.5 mM DEM were appropriately diluted and plated onto YPD plates at certain time intervals, respectively. Cell viability was determined by counting colony numbers with respect to ones at initial time. 2-PE fermentation analysis was performed as previously described (Lu et al. 2016).
qRT-PCR analysis
Exponentially growing yeast cells were treated with or without 25 mM 2-PE for 2 h and then collected by centrifugation. The total RNA was extracted using Trizol as described by Ji et al. (2016). Genomic DNA elimination and cDNA synthesis were performed with HiScript II 1st Strand cDNA Synthesis Kit (Vazyme, China). Gene expression levels were detected by UltraSYBR mixture kit (CWBIO, China) using a Bio-Rad CFX96 Real-Time PCR system, and calculated via the 2-∆∆Ct method (Livak and Schmittgen 2001). 18S rRNA was used as internal control. All qRT-PCR primers used were listed in Table S2.
Yeast one-hybrid assays
The yeast one-hybrid assay was performed using Matchmaker Gold Yeast One-Hybrid Library Screening System (Clontech, USA). The DNA bait, 1.0 kb DNA fragment of gsh2 promoter, was amplified by PCR and ligated into the pAbAi plasmid at Xho I and Hind III sites to obtain plasmid pAbAi-Pgsh2. Then linearized bait plasmid was integrated into the chromosome of Y1HGold yeast and screened from SD/-Ura agar plates (Clontech, USA), which created the bait strain Y1HGold (Pgsh2-AbAi). A YIH cDNA library was constructed by inserting cDNA fragments of C. glycerinogenes into pGATD7-Rec according to the manufacturer’s manual and transformed into the bait strain Y1HGold (Pgsh2-AbAi). The positive colonies were screened from SD/-Leu/AbA (0.2 mM) agar plates (Clontech, USA) and further used to retest and sequence.
Intracellular ROS detection
The intracellular ROS levels were measured using 2′, 7′-dichlorofluorescin diacetate (Sigma-Aldrich, USA) according to a previous study (Du and Takagi 2007) with minor mortification. Briefly, exponentially growing yeast cells were treated with or without 25 mM 2-PE for 2 h and then collected by centrifugation. The cells were washed twice by distilled water and resuspended in 0.5 mL PBS buffer (pH = 7.4). Then 2′, 7′-dichlorofluorescin diacetate was added to the cell suspensions at the final concentration of 50 μM and incubated at 30 °C for 30 min. The fluorescence of cells was detected by using a spectrofluorometer (Hitachi, Japan) with excitation wavelength at 488 nm and emission wavelength at 525 nm. Finally, ROS levels were represented as the relative units of fluorescence per OD600.
Glutathione assay
Glutathione levels were measured using glutathione assay kit (Jiancheng, China). Yeast cells were cultured in YPD medium with or without 25 mM 2-PE for 12 h then collected by centrifugation. The cells were washed twice by distilled water and resuspended in 250 μL of cold 1% 5-sulfosalicylic acid. Cells were disrupted by sonication for 30 min in an ice-bath. The supernatant containing intracellular glutathione was obtained by centrifugation (12,000g, 4 °C, 20 min). Glutathione levels were measured following the manufacturer’s instructions. The protein content was determined by the Bradford method (Kruger 1988).
Cell membrane integrity assay
Cell membrane integrity was measured using propidium iodide (PI; Sigma-Aldrich, USA) as previously described (Fang et al. 2017). Briefly, exponentially growing yeast cells were treated with or without 25 mM 2-PE for 4 h then collected by centrifugation. The cells were washed twice by distilled water and resuspended in 500 μL PBS buffer (pH = 7.4). Then PI was added to the cell suspensions at the final concentration of 10 μM and kept in the dark for 15 min at 30 °C. The fluorescence of cells was detected by using a Leica TCS SP8 Confocal System microscope (Wetzlar, Germany).
Lipid peroxidation assay
Lipid peroxidation was measured using the previous study (Yagi 1976). Briefly, exponentially growing yeast cells were treated with or without 25 mM 2-PE for 4 h. Then, the cells were washed twice by distilled water and harvested by centrifugation. One milliliter of thiobarbituric acid (TBA) reagent (0.25 M HCl, 15% trichloroacetic acid, and 0.375% TBA) was added and incubated at 100 °C for 15 min, and then centrifuged at 12,000g, 4 °C for 20 min. Lipid peroxidation levels were monitored the absorbance of the supernatants at 532 nm using a UV spectrophotometer (Shimadzu, Japan). An equal volume of TBA reagent with distilled water was used as a blank. Lipid peroxidation levels were represented as moles of TBARS per gram protein as described by Mejia-Barajas et al. (2017).
Binding site identification
The promotor fragment (− 1000 to − 1 bp) of gsh2 gene was cloned and inserted into the BamH I and Stu I sites of plasmid pUGA (Hou et al. 2019), generating the element reporter Pgsh2-GFP. The mutant allele Pgsh2m-GFP was generated from Pgsh2-GFP after altering each position of NYF by a transversion mutation. These element reporters, Pgsh2-GFP and Pgsh2m-GFP, were individually knocked in C. glycerinogenes chromosome at trp1 gene site according to previous method (Zhu et al. 2019). These engineered strains were respectively cultured in YPD medium with or without 25 mM 2-PE for 12 h then collected and washed twice with PBS buffer. The fluorescence of cells was detected by Microplate Reader (BioTek, USA) with excitation wavelength at 488 nm and emission wavelength at 520 nm. Fluorescence intensity was represented as the relative units of fluorescence per OD600.
Subcellular localization
The GFP gene fragment was PCR amplified and inserted into the Stu I and Sac II sites of plasmid pGAPa, generating the recombinant plasmid pGAPa-GFP. Then, the gsh2 gene fragment without stop codon was cloned into pGAPa-GFP by using a one-step cloning kit (Vazyme, China); the construct was named pGAPa-gsh2-GFP. The pGAPa-gsh2-GFP was linearized and integrated into the chromosome of C. glycerinogenes UA5 at 18S rDNA gene site. The transformant carrying gsh2-GFP was used for fluorescence localization analysis. Firstly, these yeast cells were cultured in YPD medium for 12 h, subsequently treated with 25 mM 2-PE for 30 min. Then yeast cells were pelleted, stained with DAPI (5 mg/L; Sangon Biotech, China), and mounted on a glass slide. Images were obtained using a Leica TCS SP8 Confocal System microscope (Wetzlar, Germany).
Results
Identification of target genes to improve 2-PE tolerance in C. glycerinogenes
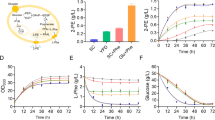
As one major barrier to 2-PE maximum titer is its high cytotoxicity, improving microbial tolerance to 2-PE may lead to higher production. Plasmid-based genomic library screen is a promising strategy to improve microbial tolerance. However, C. glycerinogenes does not have episomal plasmid, leading to an inefficient acquisition of positive transformants. Since many orthologues genes from C. glycerinogenes exhibited functional complementation in S. cerevisiae mutants (Ji et al. 2014; Li et al. 2005; Liang et al. 2018; Zhao et al. 2019), we transformed previously constructed genomic library of C. glycerinogenes (Li 2005; Li et al. 2005) into S. cerevisiae W303-1A to identify potential 2-PE-resistant genes (Fig. 1a). One of the 2-PE tolerant S. cerevisiae transformations was determined to contain a complete sequence of gsh2 gene. To examine whether gsh2 is also involved in improved 2-PE tolerance in C. glycerinogenes, it was overexpressed in C. glycerinogenes, generating Cggsh2 strain. Spot dilution assay (Fig. 1b) and growth curves (Fig. S1) showed that the cell growth of wild-type and Cggsh2 strains were similar in the absence of 2-PE, whereas Cggsh2 strain exhibited improved cell growth by 25.1% and 33.7% at 25 and 29 mM 2-PE, respectively, under 2-PE stress. Besides, the final biomass (OD600) and 2-PE production of Cggsh2 strain reached 26.5 and 33.6 mM under fermentation condition, which increased by 14.7% and 16.2% relative to wild-type strain, respectively (Fig. 1c). These results suggest that gsh2 is effective in improving 2-PE tolerance and production in C. glycerinogenes.
Screen for 2-PE-resistant genes and confirmatory tests in C. glycerinogenes. a Overall scheme of 2-PE-resistant gene screening in C. glycerinogenes. This screening process is performed as described in the “Materials and methods” section and identifies a 2-PE-resistant gene (gsh2). b Comparing growth of wild-type (wt) and Cggsh2 strains on YPD agar containing different concentrations of 2-PE shows that overexpression of gsh2 contributes to enhanced tolerance to 2-PE stress. c Comparing 2-PE yield of wild-type (wt) and Cggsh2 strains shows that overexpression of gsh2 contributes to enhanced 2-PE production. Values are means and standard deviations (n = 3)
Downregulation of gsh2 causes a 2-PE sensitive phenotype
To further investigate the function of gsh2 on 2-PE tolerance, we tried to knockout gsh2 gene in C. glycerinogenes by CRISPR-Cas9 method but without success. No colonies were grown on the selection medium even though many strategies, e.g., changes of SgRNA and length of homology arms of repair templates, have been studied, suggesting that gsh2 may be essential for cell viability of C. glycerinogenes. Therefore, we knocked down the expression of gsh2 by a previously established antisense RNA technique in C. glycerinogenes (Yang et al. 2018). Two antisense RNA fragments were chosen as pictured in Fig. 2a. The gsh2 gene silencing strains (named as Cgasgsh2-A and Cgasgsh2-B) showed no impact on cell growth relative to wild-type strain in the absence of 2-PE. Whereas at 25 mM 2-PE, their growth were inhibited severely and the biomass of Cgasgsh2-A and Cgasgsh2-B strains decreased by 27.4% and 30.4% compared with wild-type strain, respectively (Fig. 2b, c). These data further indicate that gsh2 plays an essential role in resisting 2-PE stress in C. glycerinogenes.
Downregulated expression of gsh2 effects 2-PE tolerance in C. glycerinogenes. a Two RNA-interference fragments (− 300 to − 1 bp and − 100 to + 100 bp) of gsh2 gene are used to reduce gsh2 expression. b, c Comparing growth of wild-type (wt), Cgasgsh2-A, and Cgasgsh2-B strains on YPD agar (b) or in liquid YPD medium (c) containing different concentrations of 2-PE shows that downregulation of gsh2 causes a 2-PE sensitive phenotype. Values are means and standard deviations (n = 3)
Glutathione plays a resistant role against 2-PE
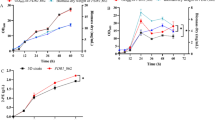
Gsh2 encodes glutathione synthetase converting glycine and L-γ-glutamyl-L-cysteine to glutathione. The intracellular glutathione levels of Cggsh2 and Cgasgsh2 strains were 23.1% higher and 30.5% lower than that of wild-type strain, respectively, at 25 mM 2-PE (Fig. 3a), suggesting that the positive role of gsh2 in response to 2-PE stress is largely dependent on the glutathione content. To validate this hypothesis, we examined the effects of exogenously added glutathione or DEM (depleting intracellular glutathione) on 2-PE tolerance. As shown in Fig. 3b, addition of glutathione increased intracellular glutathione content by 18.0% and that of added DEM decreased by 40.8%. Cell viability assay showed that 27.8% of the control cells survived under 2-PE stress, while the glutathione-enriched cells (added glutathione) and depleted cells (added DEM) exhibited 64.8% and 12.3% cell survival, representing a 37.0% increase and 15.5% decrease compared with that of the control cells, respectively (Fig. 3c). Spot dilution assay also showed similar results (Fig. 3d). These data indicate that the contribution of gsh2 to 2-PE tolerance is largely dependent on intracellular glutathione levels.
Glutathione effects 2-PE tolerance in C. glycerinogenes. a Comparing glutathione content in wild-type (wt), Cggsh2, Cgasgsh2-A, and Cgasgsh2-B strains shows that influence of glutathione synthesis by genetic engineering of gsh2 effects 2-PE tolerance. b–d Glutathione content (b), Cell viability (c), and cell growth (d) of C. glycerinogenes cultured with YPD exogenously added 2-PE, glutathione (GSH), or DEM. These data indicate that glutathione plays a resistant role against 2-PE stress. Values are means and standard deviations (n = 3). *P < 0.05, **P < 0.01, ***P < 0.001, Student’s t test
The mechanism of glutathione to improve 2-PE tolerance
Since glutathione plays a vital role in resisting oxidative stress, ROS levels in wild-type and Cggsh2 strains treated with 2-PE were detected. As shown in Fig. 4a, ROS levels significantly increased in both strains as subjected to 2-PE stress, whereas Cggsh2 strain showed a 14.2% less compared with the wild-type strain under 2-PE stress. Besides, lipids are highly susceptible to free radicals attack; thus, lipid peroxidation and cell membrane integrity assays were also performed in wild-type and Cggsh2 strains treated with or without 2-PE stress. The data showed that no significant difference of TBARS levels and PI-stained cells were found between wild-type and Cggsh2 strains in the absence of 2-PE, whereas obvious decreases of TBARS levels (35.1%) and PI-stained cells (43.2%) were observed in Cggsh2 strain compared with wild-type strain under 2-PE stress (Fig. 4b, c, d). These results indicate that glutathione protects cells against 2-PE stress by reducing ROS accumulation, lipid peroxidation, and cell membrane damage.
The mechanisms of glutathione to improve 2-PE tolerance. a–c The physiological changes, including ROS accumulation (a), lipid peroxidation (b), and cell membrane integrity (c) were detected in wild-type (wt) and Cggsh2 strains treated with or without 25 mM 2-PE, which shows that gsh2 overexpression made a significant contribution to reduce ROS accumulation, lipid peroxidation, and membrane damage caused by 2-PE stress. d Representative fluorescence microscopy photographs of wild-type (wt) and Cggsh2 strains treated with or without 3 g/L 2-PE. The exponentially growing yeast cells were treated with or without 25 mM 2-PE for 4 h and then were stained with PI and visualized by fluorescence microscopy. The less number of red-stained Cggsh2 strain compared with wild-type strain upon 2-PE stress shows that overexpression of gsh2 enhances cell membrane integrity. Values are means and standard deviations (n = 3). *P < 0.05, **P < 0.01, ***P < 0.001, Student’s t test
Identification of transcription factors regulating gsh2 gene expression
Since the expression level of gsh2 was induced by 2-PE stress (Fig. S2), we next examine whether there are potential transcription factors positively regulate gsh2 expression to resist 2-PE stress. To identify these potential transcription factors, the yeast one-hybrid assay was performed by using 1.0-kb promoter region from the gsh2 gene as bait. Three proteins (Hap5, Yap1, and Rgt1) that bind to gsh2 promoter were identified in yeast. As shown in Fig. 5a, compared with the background growth of yeast cells containing either pAbAi-Pgsh2 plus pGADT7 vectors or pAbAi plus pGADT7-hap5 vectors, yeast cells harboring pAbAi-Pgsh2 and pGADT7-hap5 vectors exhibit apparent growth in the presence of AbA, similar results to Yap1 and Rgt1, indicating that these transcription factors can bind to the promoter of gsh2.
Identification of transcription factors regulating gsh2 gene expression. a Yeast one-hybrid assay shows that Hap5, Yap1, and Rgt1 can bind to gsh2 promoter. The concentration of AbA is 0.2 mM. b Comparing mRNA levels of gsh2 in wild-type (wt), Cghap5∆, Cgyap1∆, and Cgrgt1∆ strains shows that only Hap5 and Yap1 can upregulate gsh2 expression under 2-PE stress, but not Rgt1. Values are means and standard deviations (n = 3). *P < 0.05, **P < 0.01, ***P < 0.001, Student’s t test
To further verify whether the upregulation of gsh2 expression under 2-PE stress is dependent on the above transcription factors, gsh2 mRNA levels were detected in wild-type and transcription factor single deleted strains. As shown in Fig. 5b, mRNA levels of gsh2 in all strain were similar in the absence of 2-PE. However, gsh2 mRNA levels in Cghap5∆ and Cgyap1∆ strains significantly decreased but in Cgrgt1∆ strain remained unchanged compared with wild-type strain under 2-PE stress. These data indicate that Hap5 and Yap1 serve as transcriptional activators to regulate gsh2 expression under 2-PE stress, but not Rgt1. Since gsh2 regulation in the Yap1-dependent manner has already been reported in detail by Sugiyama et al. (2000), Hap5 protein was therefore further investigated in this study.
Hap5 binds to gsh2 promoter via CCAAT box
Hap5 is a NF-Y-binding transcription factor, which recognizes the CCAAT box motif presented in the promoters and regulates the expression of its target genes. Two CCAAT boxes, designated NFY1 and NFY2, were identified in the gsh2 promoter (Fig. 6a). To determine whether Hap5 binds to gsh2 promoter by these potential NFYs, NFY1 and NFY2 were both mutated in the context of the full-length gsh2 promoter fused to the GFP gene and integrated into C. glycerinogenes chromosome. GFP mRNA level of Pgsh2-GFP reporter was respectively assayed in yeast cells with or without 2-PE treatment. As shown in Fig. 6b, GFP mRNA level of Pgsh2-GFP reporter was significantly increased with 2-PE treatment in wild-type strain, which confirms again that the expression of gsh2 is induced by 2-PE stress. However, this 2-PE stress-induced expression of GFP gene was obviously eliminated since NFY sites were mutated in gsh2 promoter and/or disruption of hap5. Similar results were also obtained by measuring fluorescence intensity (Fig. 6c). These results indicate that Hap5 plays an important role for the upregulation of gsh2 through CCAAT-binding sites under 2-PE stress.
Hap5 binds to gsh2 promoter by CCAAT box. a Schematic of putative binding sites in the gsh2 promoter. The full-length gsh2 promoter has two CCAAT motifs (NFY1 and NFY2, in square box). The CCAAT motifs were altered by a transversion mutation. b, c Comparing GFP mRNA level (b) and fluorescence intensity (c) of wild-type or mutated Pgsh2-GFP reporters in wild-type (wt) and Cghap5∆ strains treated with or without 25 mM 2-PE shows that Hap5 upregulates gsh2 through CCAAT-binding sites under 2-PE stress. Values are means and standard deviations (n = 3). *P < 0.05, **P < 0.01, ***P < 0.001, Student’s t test
2-PE stress leads to nuclear localization of Hap5
The expression levels of gsh2 remarkably decreased in hap5∆ strain under 2-PE stress, whereas no significant difference was observed between wild-type and hap5∆ strains in the absence of 2-PE, (Figs. 5b and 6b), suggesting that Hap5 activity is also triggered by 2-PE stress. To investigate this hypothesis, the mRNA levels of hap5 under 2-PE stress were analyzed, whereas the similar results were observed in 2-PE-treated and untreated cells (Fig. S3), indicating that the activation of Hap5 by 2-PE stress does not occur at transcriptional level. Then, the localization of Hap5 tagged with GFP was examined. As shown in Fig. 7, Hap5-GFP was distributed in the cytosol and nucleus in the absence of 2-PE, whereas at 25 mM 2-PE for 30 min, Hap5-GFP was concentrated in the nucleus. These results indicate that 2-PE stress response action of Hap5 occurs at post-translational level.
Hap5 is essential for cell growth under 2-PE stress
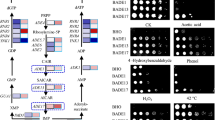
As mentioned above, Hap5 activity is triggered by 2-PE stress, suggesting that Hap5 may be essential for 2-PE resistance. To address this possibility, we compared the cell growth between wild-type and Cghap5∆ strains. Spot dilution assay showed a similar phenotype among these strains at 0 mM 2-PE, whereas Cghap5∆ strain was much more sensitive than wild-type strain under 2-PE stress (Fig. 8a). Similar results were also obtained by using a liquid medium, which Cghap5∆ strain showed 32.7% and 66.9% lower biomass than that of the wild-type strain under 25 mM or 29 mM 2-PE stress, respectively (Fig. 8b). In addition, we also examined the cell growth of Cghap5∆ under other stressors. As shown in Fig. 8c, except furfural and heat stress, Cghap5∆ exhibited severe defective growth under H2O2, NaCl, acetic acid, isobutanol, ethanol, and cold stress. These data indicate that Hap5 has a cross-protection against various types of stress.
Effects of hap5 knockout on stress tolerance. a, c Cell growth of wild-type (wt) and Cghap5∆ strains were performed by spotting onto the solid YPD supplemented with one of the following reagents: 2-PE (a), H2O2, NaCl, acetic acid, isobutanol, ethanol, cold, heat, and furfural stress (c). b Growth curves of wild-type (wt) and Cghap5∆ strains under 2-PE stress. These data show that Hap5 is essential for cell growth under 2-PE, H2O2, NaCl, acetic acid, isobutanol, ethanol, and cold, stress, but not under heat and furfural stress. Values are means and standard deviations (n = 3)
Discussion
Microbial biosynthesis of 2-PE is a promising alternative to satisfy the demands of natural aromas. However, the high cell toxicity of 2-PE limits its bioproduction (Stark et al. 2003). Due to the unclear molecular mechanism of 2-PE tolerance, engineering of relevant gene targets to breed a 2-PE tolerant strain remains a major challenge. In this study, by using a plasmid-based genomic library screen, a candidate gene target, gsh2, was identified to be effective to improve 2-PE tolerance and production, and its tolerant mechanism was also investigated.
Isolating relevant gene targets is an essential step in engineering strains for desired traits (Woodruff et al. 2013b). Although adaptive laboratory evolution (ALE) has gained great success in this field, the entire process is time-consuming (Barrick and Lenski 2013; Fletcher et al. 2017; Pontrelli et al. 2018). In comparison with ALE, genome-scale screen enables a rapid and efficient mapping of the gain- and loss-of-function mutations (Woodruff et al. 2013a, b). Existing approaches such as plasmid-based genomic library screen (Fang et al. 2017; Hong et al. 2010), transposon-based mutagenesis (Horton and Kumar 2015; Suzuki et al. 2003), and CRISPR-based library screen (Liu et al. 2018b; Schwartz et al. 2019) have mainly been applied in yeast. While the latter two methods cannot be easily implemented in C. glycerinogenes. Since this strain is a diploid yeast with inefficiency of homologous recombination, transposon-based mutagenesis would lead to generate an exceptional high proportion of single-gene deletion mutants (Suzuki et al. 2003; Zhu et al. 2019). These heterozygous diploid null mutants are hard to isolate due to the functional complementation of homologous gene. In addition, the expensive synthesis of large numbers of guide RNAs (Lane et al. 2015) and lacking of efficient nonhomologous end-joining (Zhu et al. 2019) also limit CRISPR-based library screen applied in C. glycerinogenes. In the previous study (Li 2005; Li et al. 2005), a plasmid-based genomic library of C. glycerinogenes was constructed by using yeast episomal plasmid YEp51. This library consists of about 6100 insert-containing clones with an average insert size of 7.2 kb, providing a 4-fold coverage of C. glycerinogenes genome (10.9 mb). Thus, with this powerful tool, plasmid-based genomic library screen was performed and enabled a more rapid selection of 2-PE-resistant genes in C. glycerinogenes.
During microbial production process, cells are exposed to multiple stressors that often cause oxidative stress and thus inhibit cell growth (Berterame et al. 2018). In this study, we found that 2-PE also induces oxidative stress. One possible reason is that mitochondria are a target for 2-PE damage, the compromised mitochondria result in respiratory chain deficiency and lead to more ROS production (Auesukaree et al. 2009; Wilkie and Maroudas 1969). In response to oxidative stress, microorganisms have evolved multiple mechanisms to sense and induce production of various antioxidant metabolisms, such as glutathione (Huang and Kao 2018; Sugiyama et al. 2000; Wang et al. 2018). Similarly, glutathione synthesis gene gsh2 expression and intracellular glutathione content were also increased with 2-PE stress in C. glycerinogenes. Since glutathione can reduce hydroxyl radicals (Mei et al. 2009), which are highly susceptible to attack lipids (Xu et al. 2017), alleviation of lipid peroxidation and cell membrane damage were observed in Cggsh2 strain under 2-PE stress. In addition to the role of antioxidant molecule, glutathione also possesses many other physiological functions, including maintaining redox potential, protein folding, preventing protein aggregation, and transportation of organic sulfur (Cotgreave and Gerdes 1998; Inoue et al. 1999; Sies 1999). These roles may be other reasons for glutathione protecting cells against 2-PE stress. Although enhanced tolerance does not always lead to improved production (Fu et al. 2016), in this study, tolerance engineering contributed to 2-PE production directly. Compared with ISPR, this strategy decreases the cost of extraction agent and simplifies downstream purification.
Several transcription factors are involved in antioxidant genes response (Rodrigues-Pousada et al. 2004). In C. glycerinogenes, transcription factor Hap5 plays a vital role for the upregulation of gsh2. Hap5 belongs to a heterotrimeric transcription factor HAP (also maned as NFY or CBF), which consist of other two subunits Hap2 and Hap3 (Matuoka and Chen 1999). Hap2/3/5 trimer binds to the promoters of its target genes via CCAAT box (Sinha et al. 1995), which is in agreement with our results. However, unlike Hap2/3/5 trimer of mammalian which has the ability to induce target gene expression, Hap2/3/5 trimer in yeast is only responsible for DNA binding, while gene activation requires another subunit Hap4, which recruited to Hap2/3/5 trimer via a small domain of Hap5 (Olesen and Guarente 1990). The general role of the HAP complex in yeast is implicated in upregulation of cytochrome gene expression to utilization of non-fermentable carbon sources such as lactate (Bolotin-Fukuhara 2017). In addition, HAP complex is also involved in iron homeostasis and oxidative stress response. For example, deletion of Hap4 in Hansenula polymorpha led to hypersensitivity to H2O2 (Petryk et al. 2014). Hap5 in Candida glabrata and Candida albicans induced several antioxidant genes (e.g., cat1, sod2, and oye2) expression to resist excess iron and H2O2 stress (Thiebaut et al. 2017; Wong et al. 2003). In S. cerevisiae, transcriptomic studies showed that deletion of Schap5 did not alter Scgsh2 mRNA level under normal condition (grown in YPD medium) (Hu et al. 2007), which is in agreement with our results detected by qRT-PCR in C. glycerinogenes. However, the regulation of Scgsh2 by ScHap5 protein under stress condition is no further explored. Since five CCAAT boxes are located at − 854 to − 207 in the Scgsh2 promoter, and ScHap complex is essential for oxidative stress resistance (Pinkham et al. 1997), we speculate that ScHap5 may also bind to CCAAT box triggered by stress to upregulate gsh2 for protecting cell against stress, similar to Hap5-gsh2 interaction in C. glycerinogenes. Moreover, as 2-PE can induce oxidative stress, this may be one reason that deletion of hap5 led to a significant lower biomass of C. glycerinogenes under 2-PE stress. Importantly, except for 2-PE and H2O2 stress, we first verified that hap5 was also essential in yeast to resist other stress (e.g., NaCl, acetic acid, isobutanol, ethanol, and cold stress), which suggested that hap5 is an important factor for stress tolerance.
The regulation of transcription factors involved in stress response is mainly mediated at transcriptional and/or post-transcriptional level (Huang et al. 2013; Zheng et al. 2019). The action of Hap5 in plants occurs at transcription level dealing with stress such as photooxidation and drought (Hackenberg et al. 2012; Nelson et al. 2007). However, this action of C. glycerinogenes Hap5 (CgHap5) occurs at the post-translational level though regulated of protein import into the nucleus, which enables a rapid, location-specific, and transient reaction to regulate the expression of the downstream genes compared with transcription (Deribe et al. 2010; Reinders and Sickmann 2007). Similar to other yeast and higher eukaryotes (Bolotin-Fukuhara 2017), CgHap5 also lacks independent nuclear localization signal (NLS), which indicates that Hap5p alone cannot transport into the nucleus. In mammalian cells, Hap3 and Hap5 enter the nucleus as a dimer via the importin 13 nuclear import system. Then Hap3/5 dimer interacts with Hap2, which is imported into the nucleus by an importin β mechanism, to generate the mature heterotrimeric complex (Kahle et al. 2005; Mattia et al. 2004). In fungi, Hap3/5 dimer is mediated by the non-conserved NLS Hap2 to the nucleus through a “piggy-back” mechanism (Steidl et al. 2004). In contrast to Hap2/3/5 trimer of mammalian and fungi that assembled in a stepwise manner, the assembly of heterotrimeric HAP complex in S. cerevisiae uses a one-step pathway requiring Hap2/Hap3/Hap5 subunits simultaneously (McNabb and Pinto 2005). However, how the Hap2/Hap3/Hap5 subunits transport into the nucleus in yeast is unknown (Bolotin-Fukuhara 2017). It is worth noting that CgHap5 is concentrated in the nucleus rapidly (within 30 min), suggesting that some regulator may involve in inducing CgHap5 under stress. In addition, in contrast to cellular localization of Hap5 in C. glycerinogenes, Hap5 in A. nidulans was located in the nucleus, but its nuclear localization was abolished upon oxidative stress, because Hap5 cannot interact with the oxidized Hap3 caused by the thioredoxin system, and consequently impairs Hap2-mediated piggy back transport into the nucleus. However, the upstream regulation and nuclear translocation mechanism of Hap5 in yeast is not yet clear (Bolotin-Fukuhara 2017; Mao and Chen 2019), and further studies are needed.
Based on the results in this study, we propose a hypothetical working model of 2-PE tolerance mechanism in C. glycerinogenes (Fig. 9). Under 2-PE stress, transcription factor Hap5 enters the nucleus rapidly, where it activates the transcription of gsh2, resulting in an increase of intracellular glutathione pool. The increased glutathione alleviates ROS accumulation, lipid peroxidation, and cell membrane damage, which in turn improves 2-PE tolerance and production. These findings highlight new insights into 2-PE tolerance mechanisms and also provide a good strategy to improve 2-PE production by tolerance engineering.
A hypothetical working model of 2-PE tolerance mechanism in C. glycerinogenes. Under 2-PE stress, Hap5 protein rapidly enters the nucleus independently (a) or mediated by Hap2 and Hap3 (b). In the nucleus, Hap2, Hap3, and Hap5 subunits form into a functional trimeric HAP and bind to gsh2 promoter by CCAAT box, which in turn induce gsh2 expression. The increased gsh2 expression contributed to improve cellular glutathione levels, and consequently alleviates ROS accumulation, lipid peroxidation, and cell membrane damage caused by 2-PE toxicity
References
Alexandre H, Ansanay-Galeote V, Dequin S, Blondin S (2001) Global gene expression during short-term ethanol stress in Saccharomyces cerevisiae. FEBS Lett 498:98–103. https://doi.org/10.1016/s0014-5793(01)02503-0
Auesukaree C, Damnernsawad A, Kruatrachue M, Pokethitiyook P, Boonchird C, Kaneko Y, Harashima S (2009) Genome-wide identification of genes involved in tolerance to various environmental stresses in Saccharomyces cerevisiae. J Appl Genet 50:301–310. https://doi.org/10.1007/BF03195688
Barrick JE, Lenski RE (2013) Genome dynamics during experimental evolution. Nat Rev Genet 14:827–839. https://doi.org/10.1038/nrg3564
Berterame NM, Martani F, Porro D, Branduardi P (2018) Copper homeostasis as a target to improve Saccharomyces cerevisiae tolerance to oxidative stress. Metab Eng 46:43–50. https://doi.org/10.1016/j.ymben.2018.02.010
Bolotin-Fukuhara M (2017) Thirty years of the HAP2/3/4/5 complex. Biochim Biophys Acta Gene Regul Mech 1860:543–559. https://doi.org/10.1016/j.bbagrm.2016.10.011
Chandler M, Stanley GA, Rogers P, Chambers P (2004) A genomic approach to defining the ethanol stress response in the yeast Saccharomyces cerevisiae. Ann Microbiol 54:427–454. https://doi.org/10.1081/labt-200039529
Chen X, Wang Z, Guo X, Liu S, He X (2016) Regulation of general amino acid permeases Gap1p, GATA transcription factors Gln3p and Gat1p on 2-phenylethanol biosynthesis via Ehrlich pathway. J Biotechnol 242:83–91. https://doi.org/10.1016/j.jbiotec.2016.11.028
Corre J, Lucchini JJ, Mercier GM, Cremieux A (1990) Antibacterial activity of phenethyl alcohol and resulting membrane alterations. Res Microbiol 141:483–497. https://doi.org/10.1016/0923-2508(90)90074-z
Cotgreave IA, Gerdes RG (1998) Recent trends in glutathione biochemistry glutathione-protein interactions: a molecular link between oxidative stress and cell proliferation? Biochem Biophys Res Commun 242:1–9. https://doi.org/10.1006/bbrc.1997.7812
Deribe YL, Pawson T, Dikic I (2010) Post-translational modifications in signal integration. Nat Struct Mol Biol 17:666–672. https://doi.org/10.1038/nsmb.1842
Diniz RHS, Villada JC, Alvim MCT, Vidigal PMP, Vieira NM, Lamas-Maceiras M, Cerdan ME, Gonzalez-Siso MI, Lahtvee PJ, da Silveira WB (2017) Transcriptome analysis of the thermotolerant yeast Kluyveromyces marxianus CCT 7735 under ethanol stress. Appl Microbiol Biotechnol 101:6969–6980. https://doi.org/10.1007/s00253-017-8432-0
Du X, Takagi H (2007) N-Acetyltransferase Mpr1 confers ethanol tolerance on Saccharomyces cerevisiae by reducing reactive oxygen species. Appl Microbiol Biotechnol 75:1343–1351. https://doi.org/10.1007/s00253-007-0940-x
Etschmann MM, Bluemke W, Sell D, Schrader J (2002) Biotechnological production of 2-phenylethanol. Appl Microbiol Biotechnol 59:1–8. https://doi.org/10.1007/s00253-002-0992-x
Fang Z, Chen Z, Wang S, Shi P, Shen Y, Zhang Y, Xiao J, Huang Z (2017) Overexpression of OLE1 enhances cytoplasmic membrane stability and confers resistance to cadmium in Saccharomyces cerevisiae. Appl Environ Microbiol 83. https://doi.org/10.1128/AEM.02319-16
Fletcher E, Feizi A, Bisschops MMM, Hallstrom BM, Khoomrung S, Siewers V, Nielsen J (2017) Evolutionary engineering reveals divergent paths when yeast is adapted to different acidic environments. Metab Eng 39:19–28. https://doi.org/10.1016/j.ymben.2016.10.010
Fu Y, Chen L, Zhang W (2016) Regulatory mechanisms related to biofuel tolerance in producing microbes. J Appl Microbiol 121:320–332. https://doi.org/10.1111/jam.13162
Hackenberg D, Keetman U, Grimm B (2012) Homologous NF-YC2 subunit from Arabidopsis and tobacco is activated by photooxidative stress and induces flowering. Int J Mol Sci 13:3458–3477. https://doi.org/10.3390/ijms13033458
Hong ME, Lee KS, Yu BJ, Sung YJ, Park SM, Koo HM, Kweon DH, Park JC, Jin YS (2010) Identification of gene targets eliciting improved alcohol tolerance in Saccharomyces cerevisiae through inverse metabolic engineering. J Biotechnol 149:52–59. https://doi.org/10.1016/j.jbiotec.2010.06.006
Horton BN, Kumar A (2015) Genome-wide synthetic genetic screening by transposon mutagenesis in Candida albicans. Methods Mol Biol 1279:125–135. https://doi.org/10.1007/978-1-4939-2398-4_8
Hou Q, He Q, Liu G, Lu X, Zong H, Chen W, Zhuge B (2019) Identification and application of novel low pH-inducible promoters for lactic acid production in the tolerant yeast Candida glycerinogenes. J Biosci Bioeng 128:8–12. https://doi.org/10.1016/j.jbiosc.2019.01.005
Hu Z, Killion PJ, Iyer VR (2007) Genetic reconstruction of a functional transcriptional regulatory network. Nat Genet 39:683–687. https://doi.org/10.1038/ng2012
Hua D, Xu P (2011) Recent advances in biotechnological production of 2-phenylethanol. Biotechnol Adv 29:654–660. https://doi.org/10.1016/j.biotechadv.2011.05.001
Huang M, Kao KC (2018) Identifying novel genetic determinants for oxidative stress tolerance in Candida glabrata via adaptive laboratory evolution. Yeast. https://doi.org/10.1002/yea.3352
Huang W, Miao M, Kud J, Niu X, Ouyang B, Zhang J, Ye Z, Kuhl JC, Liu Y, Xiao F (2013) SlNAC1, a stress-related transcription factor, is fine-tuned on both the transcriptional and the post-translational level. New Phytol 197:1214–1224. https://doi.org/10.1111/nph.12096
Inoue Y, Matsuda T, Sugiyama K, Izawa S, Kimura A (1999) Genetic analysis of glutathione peroxidase in oxidative stress response of Saccharomyces cerevisiae. J Biol Chem 274:27002–27009. https://doi.org/10.1074/jbc.274.38.27002
Ji H, Lu X, Wang C, Zong H, Fang H, Sun J, Zhuge J, Zhuge B (2014) Identification of a novel HOG1 homologue from an industrial glycerol producer Candida glycerinogenes. Curr Microbiol 69:909–914. https://doi.org/10.1007/s00284-014-0670-0
Ji H, Zhuge B, Zong H, Lu X, Fang H, Zhuge J (2016) Role of CgHOG1 in stress responses and glycerol overproduction of Candida glycerinogenes. Curr Microbiol 73:827–833. https://doi.org/10.1007/s00284-016-1132-7
Ji H, Lu X, Zong H, Zhuge B (2017) A synthetic hybrid promoter for D-xylonate production at low pH in the tolerant yeast Candida glycerinogenes. Bioengineered 8:700–706. https://doi.org/10.1080/21655979.2017.1312229
Kahle J, Baake M, Doenecke D, Albig W (2005) Subunits of the heterotrimeric transcription factor NF-Y are imported into the nucleus by distinct pathways involving importin beta and importin 13. Mol Cell Biol 25:5339–5354. https://doi.org/10.1128/MCB.25.13.5339-5354.2005
Kang Z, Zhang C, Du G, Chen J (2014) Metabolic engineering of Escherichia coli for production of 2-phenylethanol from renewable glucose. Appl Biochem Biotechnol 172:2012–2021. https://doi.org/10.1007/s12010-013-0659-3
Kruger N (1988) The Bradford method for protein quantitation. Methods Mol Biol 32:9–15. https://doi.org/10.1385/0-89603-268-X:9
Lane AB, Strzelecka M, Ettinger A, Grenfell AW, Wittmann T, Heald R (2015) Enzymatically generated CRISPR libraries for genome labeling and screening. Dev Cell 34:373–378. https://doi.org/10.1016/j.devcel.2015.06.003
Lester G (1965) Inhibition of growth, synthesis, and permeability in Neurospora crassa by phenethyl alcohol. J Bacteriol 90:29–37. https://doi.org/10.1111/j.1365-2672.1965.tb02178.x
Li Y (2005) Cloning of marker gene and integrated site gene from Candida glycerinogenes. Dissertation, Jiangnan University
Li Y, Shen W, Wang Z, Liu JQ, Rao Z, Tang X, Fang H, Zhuge J (2005) Isolation and sequence analysis of the gene URA3 encoding the orotidine-5′-phosphate decarboxylase from Candida glycerinogenes WL2002-5, an industrial glycerol producer. Yeast 22:423–430. https://doi.org/10.1002/yea.1211
Liang Z, Liu D, Lu X, Zong H, Song J, Zhuge B (2018) Identification and characterization from Candida glycerinogenes of hexose transporters having high efficiency at high glucose concentrations. Appl Microbiol Biotechnol 102:5557–5567. https://doi.org/10.1007/s00253-018-9027-0
Liu J, Jiang J, Bai Y, Fan TP, Zhao Y, Zheng X, Cai Y (2018a) Mimicking a new 2-phenylethanol production pathway from Proteus mirabilis JN458 in Escherichia coli. J Agric Food Chem 66:3498–3504. https://doi.org/10.1021/acs.jafc.8b00627
Liu R, Liang L, Choudhury A, Garst AD, Eckert CA, Oh EJ, Winkler J, Gill RT (2018b) Multiplex navigation of global regulatory networks (MINR) in yeast for improved ethanol tolerance and production. Metab Eng 51:50–58. https://doi.org/10.1016/j.ymben.2018.07.007
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Lu X, Wang Y, Zong H, Ji H, Zhuge B, Dong Z (2016) Bioconversion of L-phenylalanine to 2-phenylethanol by the novel stress-tolerant yeast Candida glycerinogenes WL2002-5. Bioengineered 7:418–423. https://doi.org/10.1080/21655979.2016.1171437
Ma M, Liu ZL (2010a) Mechanisms of ethanol tolerance in Saccharomyces cerevisiae. Appl Microbiol Biotechnol 87:829–845. https://doi.org/10.1007/s00253-010-2594-3
Ma M, Liu ZL (2010b) Quantitative transcription dynamic analysis reveals candidate genes and key regulators for ethanol tolerance in Saccharomyces cerevisiae. BMC Microbiol 10:169. https://doi.org/10.1186/1471-2180-10-169
Mao Y, Chen C (2019) The hap complex in yeasts: structure, assembly mode, and gene regulation. Front Microbiol 10:1645. https://doi.org/10.3389/fmicb.2019.01645
Martínez-Avila O, Sánchez A, Font X, Barrena R (2018) Bioprocesses for 2-phenylethanol and 2-phenylethyl acetate production: current state and perspectives. Appl Microbiol Biotechnol 102:9991–10004. https://doi.org/10.1007/s00253-018-9384-8
Mattia F, Carol I, Isabella M, Roberto M (2004) Cell cycle regulation of NF-YC nuclear localization. Cell Cycle 3:205–210
Matuoka K, Chen KY (1999) Nuclear factor Y (NF-Y) and cellular senescence. Exp Cell Res 253:365–371. https://doi.org/10.1006/excr.1999.4605
McNabb DS, Pinto I (2005) Assembly of the Hap2p/Hap3p/Hap4p/Hap5p-DNA complex in Saccharomyces cerevisiae. Eukaryot Cell 4:1829–1839. https://doi.org/10.1128/EC.4.11.1829-1839.2005
Mei J, Min H, Lü Z (2009) Enhanced biotransformation of L-phenylalanine to 2-phenylethanol using an in situ product adsorption technique. Process Biochem 44:886–890. https://doi.org/10.1016/j.procbio.2009.04.012
Mejia-Barajas JA, Montoya-Perez R, Salgado-Garciglia R, Aguilera-Aguirre L, Cortes-Rojo C, Mejia-Zepeda R, Arellano-Plaza M, Saavedra-Molina A (2017) Oxidative stress and antioxidant response in a thermotolerant yeast. Braz J Microbiol 48:326–332. https://doi.org/10.1016/j.bjm.2016.11.005
Nelson DE, Repetti PP, Adams TR, Creelman RA, Wu J, Warner DC, Anstrom DC, Bensen RJ, Castiglioni PP, Donnarummo MG, Hinchey BS, Kumimoto RW, Maszle DR, Canales RD, Krolikowski KA, Dotson SB, Gutterson N, Ratcliffe OJ, Heard JE (2007) Plant nuclear factor Y (NF-Y) B subunits confer drought tolerance and lead to improved corn yields on water-limited acres. Proc Natl Acad Sci U S A 104:16450–16455. https://doi.org/10.1073/pnas.0707193104
Olesen JT, Guarente L (1990) The HAP2 subunit of yeast CCAAT transcriptional activator contains adjacent domains for subunit association and DNA recognition: model for the HAP2/3/4 complex. Genes Dev 4:1714–1729. https://doi.org/10.1101/gad.4.10.1714
Petryk N, Zhou YF, Sybirna K, Mucchielli MH, Guiard B, Bao WG, Stasyk OV, Stasyk OG, Krasovska OS, Budin K, Reymond N, Imbeaud S, Coudouel S, Delacroix H, Sibirny A, Bolotin-Fukuhara M (2014) Functional study of the Hap4-like genes suggests that the key regulators of carbon metabolism HAP4 and oxidative stress response YAP1 in yeast diverged from a common ancestor. PLoS One 9:e112263. https://doi.org/10.1371/journal.pone.0112263
Pinkham JL, Wang Z, Alsina J (1997) Heme regulates SOD2 transcription by activation and repression in Saccharomyces cerevisiae. Curr Genet 31:281–291. https://doi.org/10.1007/s002940050207
Pontrelli S, Fricke RCB, Sakurai SSM, Putri SP, Fitz-Gibbon S, Chung M, Wu HY, Chen YJ, Pellegrini M, Fukusaki E, Liao JC (2018) Directed strain evolution restructures metabolism for 1-butanol production in minimal media. Metab Eng 49:153–163. https://doi.org/10.1016/j.ymben.2018.08.004
Reinders J, Sickmann A (2007) Modificomics: posttranslational modifications beyond protein phosphorylation and glycosylation. Biomol Eng 24:169–177. https://doi.org/10.1016/j.bioeng.2007.03.002
Rodrigues-Pousada CA, Nevitt T, Menezes R, Azevedo D, Pereira J, Amaral C (2004) Yeast activator proteins and stress response: an overview. FEBS Lett 567:80–85. https://doi.org/10.1016/j.febslet.2004.03.119
Schwartz C, Cheng JF, Evans R, Schwartz CA, Wagner JM, Anglin S, Beitz A, Pan W, Lonardi S, Blenner M, Alper HS, Yoshikuni Y, Wheeldon I (2019) Validating genome-wide CRISPR-Cas9 function improves screening in the oleaginous yeast Yarrowia lipolytica. Metab Eng 55:102–110. https://doi.org/10.1016/j.ymben.2019.06.007
Seward R, Willetts JC, Dinsdale MG, Lloyd D (2013) The effects of ethanol, hexan-1-ol, and 2-phenylethanol on cider yeast growth, viability, and energy status; synergistic inhibition. J Inst Brew 102:439–443. https://doi.org/10.1002/j.2050-0416.1996.tb00928.x
Si T, Chao R, Min Y, Wu Y, Ren W, Zhao H (2017) Automated multiplex genome-scale engineering in yeast. Nat Commun 8:15187. https://doi.org/10.1038/ncomms15187
Sies H (1999) Glutathione and its role in cellular functions. Free Radic Biol Med 27:916–921. https://doi.org/10.1016/s0891-5849(99)00177-x
Sinha S, Maity SN, Lu J, Crombrugghe B (1995) Recombinant rat CBF-C, the third subunit of CBF/NFY, allows formation of a protein-DNA complex with CBF-A and CBF-B and with yeast HAP2 and HAP3. Proc Natl Acad Sci U S A 92:1624–1628. https://doi.org/10.1073/pnas.92.5.1624
Stark D, Zala D, Münch T, Sonnleitner B, Marison IW, Stockar UV (2003) Inhibition aspects of the bioconversion of L-phenylalanine to 2-phenylethanol by Saccharomyces cerevisiae. Enzym Microb Technol 32:212–223. https://doi.org/10.1016/s0141-0229(02)00237-5
Steidl S, Tüncher A, Goda H, Guder C, Papadopoulou N, Kobayashi T, Tsukagoshi N, Kato M, Brakhage AA (2004) A single subunit of a heterotrimeric CCAAT-binding complex carries a nuclear localization signal: piggy back transport of the pre-assembled complex to the nucleus. J Mol Biol 342:515–524. https://doi.org/10.1016/j.jmb.2004.07.011
Sugiyama K, Izawa S, Inoue Y (2000) The Yap1p-dependent induction of glutathione synthesis in heat shock response of Saccharomyces cerevisiae. J Biol Chem 275:15535–15540. https://doi.org/10.1074/jbc.275.20.15535
Suzuki C, Hori Y, Kashiwagi Y (2003) Screening and characterization of transposon-insertion mutants in a pseudohyphal strain of Saccharomyces cerevisiae. Yeast 20:407–415. https://doi.org/10.1002/yea.970
Tan Z, Khakbaz P, Chen Y, Lombardo J, Yoon JM, Shanks JV, Klauda JB, Jarboe LR (2017) Engineering Escherichia coli membrane phospholipid head distribution improves tolerance and production of biorenewables. Metab Eng 44:1–12. https://doi.org/10.1016/j.ymben.2017.08.006
Teixeira MC, Godinho CP, Cabrito TR, Mira NP, Sá-Correia I (2012) Increased expression of the yeast multidrug resistance ABC transporter Pdr18 leads to increased ethanol tolerance and ethanol production in high gravity alcoholic fermentation. Microb Cell Factories 11:98. https://doi.org/10.1186/1475-2859-11-98
Terenzi HF, Storck R (1969) Stimulation of fermentation and yeast-like morphogenesis in Mucor rouxii by phenethyl alcohol. J Bacteriol 97:1248–1261. https://doi.org/10.1016/S0901-5027(05)81368-2
Thiebaut A, Delaveau T, Benchouaia M, Boeri J, Garcia M, Lelandais G, Devaux F (2017) The CCAAT-binding complex controls respiratory gene expression and iron homeostasis in Candida glabrata. Sci Rep 7:3531. https://doi.org/10.1038/s41598-017-03750-5
Wang D, Wu D, Yang X, Hong J (2018) Transcriptomic analysis of thermotolerant yeast Kluyveromyces marxianus in multiple inhibitors tolerance. RSC Adv 8:14177–14192. https://doi.org/10.1039/c8ra00335a
Wang Y, Zhang H, Lu X, Zong H, Zhuge B (2019) Advances in 2-phenylethanol production from engineered microorganisms. Biotechnol Adv 37:403–409. https://doi.org/10.1016/j.biotechadv.2019.02.005
Wilkie D, Maroudas NG (1969) Induction of cytoplasmic respiratory deficiency in yeast by phenethyl alcohol. Genet Res 13:107–111. https://doi.org/10.1017/s0016672300002792
Wong CM, Ching YP, Zhou Y, Kung HF, Jin DY (2003) Transcriptional regulation of yeast peroxiredoxin gene TSA2 through haplp, Rox1p, and Hap2/3/5p. Free Radic Biol Med 34:585–597. https://doi.org/10.1016/s0891-5849(02)01354-0
Woodruff LB, Boyle NR, Gill RT (2013a) Engineering improved ethanol production in Escherichia coli with a genome-wide approach. Metab Eng 17:1–11. https://doi.org/10.1016/j.ymben.2013.01.006
Woodruff LB, Pandhal J, Ow SY, Karimpour-Fard A, Weiss SJ, Wright PC, Gill RT (2013b) Genome-scale identification and characterization of ethanol tolerance genes in Escherichia coli. Metab Eng 15:124–133. https://doi.org/10.1016/j.ymben.2012.10.007
Xu P, Qiao K, Stephanopoulos G (2017) Engineering oxidative stress defense pathways to build a robust lipid production platform in Yarrowia lipolytica. Biotechnol Bioeng 114:1521–1530. https://doi.org/10.1002/bit.26285
Yagi K (1976) A simple fluorometric assay for lipoperoxide in blood plasma. Biochem Med 15:212–216. https://doi.org/10.1016/0006-2944(76)90049-1
Yang F, Lu X, Zong H, Ji H, Zhuge B (2018) Gene expression profiles of Candida glycerinogenes under combined heat and high-glucose stresses. J Biosci Bioeng 126:464–469. https://doi.org/10.1016/j.jbiosc.2018.04.006
Zhang J, Astorga MA, Gardner JM, Walker ME, Grbin PR, Jiranek V (2018) Disruption of the cell wall integrity gene ECM33 results in improved fermentation by wine yeast. Metab Eng 45:255–264. https://doi.org/10.1016/j.ymben.2017.12.012
Zhang MM, Xiong L, Tang YJ, Mehmood MA, Zhao ZK, Bai FW, Zhao XQ (2019) Enhanced acetic acid stress tolerance and ethanol production in Saccharomyces cerevisiae by modulating expression of the de novo purine biosynthesis genes. Biotechnol Biofuels 12:116. https://doi.org/10.1186/s13068-019-1456-1
Zhao M, Shi D, Lu X, Zong H, Zhuge B, Ji H (2019) Ethanol fermentation from non-detoxified lignocellulose hydrolysate by a multi-stress tolerant yeast Candida glycerinogenes mutant. Bioresour Technol 273:634–640. https://doi.org/10.1016/j.biortech.2018.11.053
Zheng M, Li D, Zhao Z, Shytikov D, Xu Q, Jin X, Liang J, Lou J, Wu S, Wang L, Hu H, Zhou Y, Gao X, Lu L (2019) Protein phosphatase 2A has an essential role in promoting thymocyte survival during selection. Proc Natl Acad Sci U S A 116:12422–12427. https://doi.org/10.1073/pnas.1821116116
Zhu M, Sun L, Lu X, Zong H, Zhuge B (2019) Establishment of a transient CRISPR-Cas9 genome editing system in Candida glycerinogenes for co-production of ethanol and xylonic acid. J Biosci Bioeng 128:283–289. https://doi.org/10.1016/j.jbiosc.2019.03.009
Zhuge J, Fang HY, Wang ZX, Chen DZ, Jin HR, Gu HL (2001) Glycerol production by a novel osmotolerant yeast Candida glycerinogenes. Appl Microbiol Biotechnol 55:686–692. https://doi.org/10.1007/s002530100596
Funding
This work was supported by the National Natural Science Foundation of China (31601456, 31570052, 21708016).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 340 kb).
Rights and permissions
About this article
Cite this article
Wang, Y., Zhang, Z., Lu, X. et al. Transcription factor Hap5 induces gsh2 expression to enhance 2-phenylethanol tolerance and production in an industrial yeast Candida glycerinogenes. Appl Microbiol Biotechnol 104, 4093–4107 (2020). https://doi.org/10.1007/s00253-020-10509-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-10509-y