Abstract
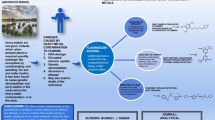
Heavy metal(loid)s such as Cd and Hg adversely affect human health and are therefore strictly regulated and monitored; however, their quantitation in the environment is usually performed by expensive and time-consuming instrumental analysis techniques, which necessitates the search for more practical alternatives. Herein, we prepare enhanced green fluorescent protein (eGFP)–based biomolecules for metal(loid) sensing by insertion of metal-binding loops (MBLs) into a loop region of eGFP to render this protein inactive and show that the binding of metal ions to MBLs induces a conformational change and restores the original activity. Specifically, eGFP with an MBL sequenced as CTTCGCG regains fluorescence upon exposure to Cd and Hg, which allows the above metals to be quantified in the concentration range of 0–5 μM. For practical applicability verification, the developed sensing platform is used to quantify Cd in artificially amended soil and water samples. Although the obtained results imply that sensor performance needs to be significantly improved, the presented design concept is believed to be of high value to researchers in the field of heavy metal sensing and facilitate the development of new biosensors.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Environmental monitoring is considered to be of high importance for human health, as we live in environmental systems and are directly/indirectly affected by these systems and the hazardous materials present therein (Järup 2003), e.g., heavy metals cause plant photosynthesis malfunctions and accumulate in diverse food chains (Küpper et al. 1998; Peralta-Videa et al. 2009; Singh and Prasad 2011). Although most heavy metals are strictly controlled, even traces of them may adversely affect human health, and the severity of environmental pollution often becomes apparent when it is too late to act, as the consequences of this pollution are not immediately obvious. Thus, the development of fast and simple methods of environmental pollution monitoring is a task of high practical significance.
Traditionally, harmful materials in environmental systems (e.g., atmosphere, water, and soil) have been quantified by time-consuming and expensive instrumental analysis techniques (Ammann 2002; Yuan et al. 2004), which determine the total amount of heavy metals even if separated extraction steps are employed and hence do not consider the complexity of environmental matters. Although total heavy metal content is an important parameter, it does not equal the content of bioavailable heavy metals, which is only a fraction of the former and depends on the environmental system (Baumann and van der Meer 2007; Turpeinen et al. 2003; Yoon et al. 2016a). For example, soil remediation increases the bioavailability of As even if the total As content is concomitantly reduced (Yoon et al. 2016b). Therefore, both total and biologically active pollutant contents should be considered for pollutant control and related risk assessment.
The past decades have witnessed the development and implementation of diverse biosensors for environmental pollutant quantitation, as these biosensors employ fast inexpensive processes and are consequently free from the inherent drawbacks of traditional techniques. Moreover, as biosensing utilizes bacterial cells, enzymes, or electrochemical methods, it allows one to quantify the amounts of bioavailable metals and not just the total amounts (Belkin 2003; Kuswandi 2003; Zhylyak et al. 1995). In particular, much attention has been directed at the development of bacterial cell–based biosensors, or whole-cell bioreporters (WCBs), which typically comprise sensing and reporter domains (Belkin 2003; Rawson et al. 1989). In these systems, stress-responsive operon components are usually employed as a sensing (target-recognizing) domain, while genes encoding enzymes or fluorescent proteins are used as reporter domains; i.e., the presence of targets is determined by monitoring the expression of reporter genes. However, the shortcomings of WCBs, e.g., low target selectivity/sensitivity and limited number of genetic systems, preclude the widespread application of these systems to environmental monitoring. In fact, WCB selectivity depends on the interaction between target and regulatory proteins on sensing domains, and low sensitivity is caused by the operation of microorganism defense systems to maintain homeostasis (Brocklehurst et al. 1999; Schalk et al. 2011). In our previous studies, we showed that the selectivity and sensitivity of WCBs can be improved by genetic engineering (Kang et al. 2018a; Kang et al. 2018b; Yoon et al. 2018). Briefly, the selectivity of WCB was increased and modified by introducing point mutations into the metal-binding loop of regulatory proteins, while sensitivity was increased by deleting genes involved in metal transportation. Nonetheless, these improvements proved to be insufficient, as the number of hazardous materials significantly exceeds that of genetic systems, which inspired us to adopt a new strategy, namely to develop biosensors for environmental pollutant monitoring as an alternative to typical WCBs.
The new strategy demonstrated herein involves the use of split-protein systems. Such systems are commonly employed to measure protein–protein interactions, since mature protein behavior is observed when split protein parts are combined (Shekhawat and Ghosh 2011; Stynen et al. 2012). Inspired by the use of split green fluorescent protein (GFP) to measure the interaction between two proteins in cells (Cabantous et al. 2013; Cabantous et al. 2005), we utilized enhanced GFP (eGFP) as a split-protein system and replaced one of its loop regions (between β-strands 9 and 10, sequenced as GDGPVL) with other amino acid sequences to afford metal-binding loops (MBLs). In the absence of metal ions, the modified proteins were inactive, whereas metal ion binding to MBLs resulted in structural changes leading to active GFP formation. Despite the multitude of factors affecting the performance of such novel metal–sensing biomolecules and the associated challenges, we concluded that the suggested strategy can be used as a platform to generate new biosensors targeting other pollutants.
Materials and methods
Materials
Escherichia coli (E. coli) DH5α was used as the host strain for gene cloning, protein expression, and biosensor fabrication. The eGFP-encoding gene (egfp) used as a template was cloned from pEGFP-N1 (Novagen), and the promoter of the Zn-responsive operon (znt-operon) was obtained from the genomic DNA of E. coli DH5α. All tested plasmids were constructed using pET-21(a) as a vector. Heavy metal(loid)–containing compounds, namely CdCl2, K2Cr2O7, CuCl2·2H2O, HgCl2, NiCl2, AsCl3, PbCl2, AuCl2, SbCl2, and ZnCl2, were purchased from Sigma-Aldrich (St. Louis, MO, USA) and used to prepare 10 mM stock solutions. Landwirtschaftliche Untersuchungs- und Forschungsanstalt (LUFA) standard soil (LUFA Speyer, Germany) was used to prepare Cd-amended soil samples. The intensity of eGFP fluorescence was measured using an FS-2 fluorescence spectrophotometer (Scinco, Seoul, Korea) equipped with a Xe lamp as a light source and bandwidth-adjustable filters for excitation and emission wavelengths.
Plasmid construction
Peptide sequences expected to bind metal ions were inserted into the loop region of eGFP. Nucleotide sequences encoding the designated peptide sequences were located in primers, and MBL insertion was performed by overlapping PCR. PCR fragments were ligated into pET-21(a) with BamHI/XhoI restriction sites. Since gene expression was controlled by the T7 promoter requiring isopropyl β-D-thiogalactoside (IPTG) and lacI genes, the above promoter was replaced by the znt-operon promoter (Pznt), which was amplified by PCR from the genomic DNA of E. coli and inserted into BglII and XbaI restriction sites. The DNA construct harboring the fusion of the znt promoter and the engineered eGFP was denoted as pZnt-eGFP-HJ1. The plasmid harboring the znt promoter with wild-type eGFP, denoted as pZnt-eGFP and constructed as described elsewhere, was used as a control for comparing eGFP expression levels (Yoon et al. 2016c). MBL introduction into eGFP was further confirmed by DNA sequencing, and the plasmids were finally transformed into E. coli DH5α for the verification of metal sensing properties.
Characterization of the znt promoter
Since the znt promoter is not tightly regulated, the genes placed downstream were constitutively expressed. To verify the properties of the znt promoter, pZnt-eGFP and pZnt-eGFP-HJ1 were compared side-by-side. Specifically, we inoculated overnight cultures in fresh lysogeny broth (LB) and then investigated the time course of emission intensity in the absence of added metal(loid)s over 12 h.
In vivo and in vitro metal(loid) sensing assays
The selectivity and specificity of metal(loid) detection were assessed by in vivo and in vitro assays, in which biosensor fluorescence intensity was measured after 0-–6-h exposure to metal(loid) solutions (0–50 μM) (Yoon et al. 2016a, b). In the former assay, the fluorescence intensity of whole E. coli cells was measured, while supernatants of E. coli cells were employed in the latter case to exclude cell-inseparable small particles. The in vivo assay could be applied only to water samples, while the in vitro assay could be applied to all environmental samples.
In vivo assay
E. coli DH5α cells harboring pZnt-eGFP-HJ1 were grown overnight at 37 °C in LB broth containing ampicillin (50 μg/mL) and then inoculated in fresh LB broth. For metal(loid) selectivity screening, cells were grown at 30 °C and exposed to diverse metal(loid) solutions (10 μM) when the optical density at 600 nm (OD600) reached 0.4. For fluorescence measurements, 1-mL aliquots of cell suspensions were collected at different times and re-suspended in 50 mM Tris-HCl (pH 7.4).
In vitro assay
The in vitro assay was employed when environmental samples contained small cell–inseparable particles inhibiting signal acquisition. In this case, the supernatants of E. coli cells were used instead of whole cells for fluorescence measurements. The sample preparation and analysis procedures were identical to those used in the in vivo assay except for the fact that cells were disrupted by sonication and centrifuged at 10,000 g to afford the abovementioned supernatants. Herein, the in vitro assay was applied to determine the concentration of Cd in amended soil samples.
Fluorescence measurements
Fluorescence intensity was measured using an FS-2 fluorescence spectrophotometer (Scinco, Seoul, Korea). Excitation/emission wavelengths of 470/510 nm were employed for eGFP, while the corresponding bandwidths equaled 5 nm. The expression level of engineered eGFP was represented by the induction coefficient, which, in turn, was defined as [fluorescence intensity of heavy metal–exposed biosensor]/[fluorescence intensity of non-exposed biosensor].
Cd sensing properties of the recombinant protein
Since eGFP-HJ1 behaved as a sensing molecule in E. coli cells, we tested the Cd sensing ability of the recombinant protein. Cells harboring pZnt-eGFP-HJ1 were grown at 30 °C for 6 h in the absence of metal(loid)s, harvested by centrifugation, and sonicated. The supernatants containing recombinant protein were partitioned between test tubes, exposed to Cd2+ concentrations of 0–1 mM, and subjected to fluorescence intensity measurements after different exposure times.
Preparation of artificially amended water and soil samples
The applicability of new biosensors to real-life systems was probed by analysis of artificially amended water and soil samples. To prepare amended water samples, distilled water was spiked with Cd and Hg stock solutions to levels of 5, 10, 20, and 25 μM. In the case of soil samples, only Cd-amended samples were prepared. Briefly, LUFA soils were amended with Cd to reach final loadings of 5, 10, 20, 50, and 100 mg/kg, and samples were stored in the dark for 7 days prior to analysis.
Quantitation of Cd in water and soil samples
The Cd contents of artificially contaminated water and soil samples were determined by linear regression analysis of standard curves. Briefly, 5-mL WCB samples (OD600 = 0.4) were treated with Cd-free water (0.5 mL) and soil (0.25 g) to account for environmental matrix–caused interferences and then exposed to Cd ions at concentrations of 0–5 μM. Induction coefficient values were measured after 2-h exposure, and plots of induction coefficient vs. Cd concentration were used as standard curves for quantitation. Water samples were analyzed using both in vivo and in vitro assays, and the results were compared, while soil samples were analyzed solely using the in vitro assay.
Results
Generation of new biosensor molecules based on split eGFP
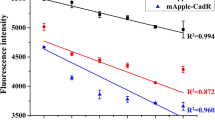
As the T7 promoter in the pET-21(a) expression vector was replaced by the znt promoter, the properties of the latter were characterized in detail. In fact, the T7 promoter could also be used for overexpression in E. coli BL21 stain; however, additional steps such as strain change and the addition of IPTG for induction were required. Although the znt promoter was driven from Zn-responsive operon in E. coli, its regulation was not very tight, which could result in constitutive expression of genes downstream. To verify this assumption, two E. coli DH5α cells harboring pZnt-eGFP and pZnt-eGFP-HJ1 were tested and compared, and fluorescence intensities of 3000 and 300 arbitrary units (AU) were observed, respectively (Fig. 1). This result confirmed that the insertion of an MBL sequenced as CTTCGCG interrupted the fluorescence of eGFP. Moreover, the expression of eGFP increased with increasing incubation time, which confirmed the constitutive nature of gene expression.
Fluorescence intensity of E. coli cells harboring pZnt-eGFP and pZnt-eGFP-HJ1 as a function of incubation time. After 6-h incubation, eGFP signal intensity increased to 3000 arbitrary units (AU) for wild-type eGFP, while an only modest increase to 300 AU was observed for eGFP with HJ1. The error bars represented as standard deviation and the experiments were replicated over 3 times
Selectivity of engineered eGFP for metal(loid)s
As mentioned above, two different methods were employed to determine the fluorescence intensity of E. coli biosensors, namely in vivo and in vitro assays. Briefly, the in vivo assay measured the fluorescence intensity of samples containing whole E. coli cells, while the in vitro assay analyzed cell supernatants. However, metal(loid) selectivity was screened only using the in vivo assay because of its simplicity and ease of operation. The in vitro assay was employed to account for diverse interferences caused by environmental matters, especially by small particles in soils, and its application is discussed in a later section. For selectivity screening, E. coli DH5α cells possessing pZnt-eGFP-HJ1 were exposed to solutions of diverse metal(loid)s (10 μM), and fluorescence intensity was measured after 2 h (Fig. 2). The sample not exposed to any metal(loid) ions was used as a control, and responses to metal(loid)s were represented by induction coefficient values. Notably, only Cd and Hg induced the appearance of fluorescence, i.e., the developed biosensor could selectively sense the above metals.
Selectivity of E. coil cells with pZnt-eGFP-HJ1 to heavy metal(loid) ions. Fluorescence intensities of cells treated with 10 μM metal(loid) ion solutions were measured after 2-h incubation, and the obtained values were converted to induction coefficients, which were defined as [fluorescence intensity of metal(loid)-exposed cells/fluorescence intensity of non-exposed cells]. The error bars represented as standard deviation and the experiments were replicated over 3 times. The asterisk (*) indicates statistical significance at the 95% level (p < 0.05), when compared to control
Sensitivity of engineered eGFP to metal(loid)s
Based on the results of selectivity screening, we further probed the sensitivity of the developed biosensor to Cd and Hg by investigating the effects of 2-h exposure to Cd and Hg solutions on fluorescence intensity. Figure 3 shows the thus obtained induction coefficient values, revealing that Cd response strength increased with increasing Cd concentration up to 5 μM (Fig. 3a). The decrease observed at higher Cd levels was ascribed to the inhibition of E. coli growth, i.e., to the cytotoxicity of Cd. In the range of 0–5 μM, the relationship between Cd concentration and induction coefficient could be well fitted by a straight line (R2 = 0.996; Fig. 3a inbox), which implied that Cd levels within this range could be quantitatively determined by our biosensor. A similar trend was observed for Hg, i.e., a strong dose–response correlation (R2 = 0.996, Fig. 3b) was observed within the concentration range of 0–5 μM, with cytotoxicity becoming obvious at higher concentrations (Fig. S1). Unlike to Cd, the correlation was non-linear and the response toward Hg was observed from 2.5 μM. It was inferred that the novel biosensors showed higher sensitivity toward Cd than Hg.
Sensitivity of E. coli harboring pZnt-eGFP-HJ1 to Cd and Hg. Responses to a Cd2+ (0–20 μM) and b Hg2+ solutions (0–7.5 μM) and regression analysis of the relationship between induction coefficient and metal ion concentration (insets). The error bars represented as standard deviation and the experiments were replicated over 3 times. The asterisk (*) indicates statistical significance at the 95% level (p < 0.05)
Sensitivity of recombinant protein
Unlike typical stress–responsive operon–based WCBs, the biosensor developed herein utilized an expressed recombinant protein, namely engineered eGFP. Therefore, we tested the response of this recombinant protein to Cd. The supernatants of sonication-produced cell lysates were exposed to Cd at concentrations of 0–1000 μM, and the fluorescence of each test set was measured during 12-h incubation at 30 °C. Fluorescence intensity increased upon Cd exposure for up to 5 h, decreasing at longer exposure times, and hence, only induction coefficients obtained after 0.5, 2, and 5 h were considered (Fig. 4). The responses toward Cd showed non-linear response and the response was saturated at 200 μM with 2-h and 5-h exposures. Notably, the response was much weaker than that observed for E. coli cells, which was attributed to protein instability in an extracellular environment. The issue of split eGFP instability was raised in a previous study, where point mutations were employed for stability improvement (Cabantous et al. 2005), and is discussed later in the “Discussion” section. Nonetheless, the obtained results supported our hypothesis that MBL-containing eGFP acted as a genuine biosensor.
Induction coefficients of recombinant eGFP-HJ1 obtained at diverse concentrations of Cd. Values obtained for 0.5-h (white bar), 2-h (gray), and 5-h (dark gray) exposure to 0–1 mM Cd2+. The error bars represented as standard deviation and the experiments were replicated over 3 times. The asterisk (*) indicates statistical significance at the 95% level (p < 0.05)
Quantitation of Cd in artificially amended water samples
To evaluate the suitability of the developed biosensor to real-life applications, it was used to quantify Cd and Hg in artificially contaminated water samples. Standard curves were constructed using induction coefficient values determined for cells exposed to diverse Cd/Hg concentrations. Since the concentration of these metals in artificially contaminated water samples equaled 5–25 μM, these samples were diluted prior to analysis. In the case of Cd, a linear correlation between induction coefficient and concentration was obtained (R2 = 0.987; Fig. S2A), whereas a non-linear correlation (R2 = 0.998) was obtained for Hg (Fig. S2B). The above correlations were used to quantitate Cd and Hg in contaminated samples. Table 1 summarizes the obtained results, demonstrating that accuracies of 81–98% were achieved and thus showing that the developed approach was well suited for Cd and Hg quantitation.
To verify the reliability of the established technique, cells used in the in vivo assay were employed to perform the in vitro assay. Samples were sonicated in 50 mM Tris-HCl (1 mL, pH 7.3) containing 160 mM KCl, and the fluorescence intensity of supernatants was recorded. The induction coefficients were found to be different from those obtained in the in vivo assay, although the trends observed in both cases were similar (Fig. S2C and D). The quantitation procedure was identical to that used in the in vivo assay and afforded similar results (Table 1), e.g., accuracies of ~ 90% were observed. Thus, the in vivo and in vitro assays complemented each other and were well suited for the quantitation of Cd and Hg.
Quantitation of Cd in artificially amended soil samples
The Cd contents of artificially amended soil samples were determined by the in vitro assay, as the presence of cell-inseparable soil particles did not allow one to obtain reliable fluorescence signals in the in vivo assay. E. coli cells were exposed to Cd-amended (5, 10, 20, 50, and 100 mg/kg) 0.5-g soil samples for 2 h. Subsequently, 1-mL aliquots were collected, and the intensity of supernatant fluorescence at 510 nm was determined by fluorescence spectrophotometry. The Cd contents of soil samples were determined using the standard curve method and linear regression analysis, with the obtained results listed in Table 2. The overall recovery was less than 27%, i.e., significantly lower than in the case of water samples. However, this finding was not unexpected, as the bioavailability of metal(loid)s in soils is known to be low and vary with species, soil physicochemical properties, organic matter content, and aging duration. In fact, compared to analytical instruments, biosensors provide information on the content of bioavailable heavy metals and not on their total content, which is considered to be both their weak point and an advantage.
Discussion
Bacterial cell–based bioreporters and protein-based biosensors have been extensively researched in the past few decades. These biosensors offer important advantages (e.g., simplicity, low cost, and convenience) over instrumental analysis but are not as powerful, especially in the case of environmental samples. Generally, regulation policies are based on the total amounts of pollutants in environmental systems. Although the total pollutant amount is a meaningful parameter, its use may result in risk exaggeration, as the bioavailable pollutant amount is usually much less than the total amount. Therefore, much effort has been directed at the development of new biosensors and at emphasizing their importance.
The living cell–based biosensors, so called whole-cell bioreporter (WCBs), were generated by employing the promoter of stress-responsive operon as sensing domains and genes encoding enzymes and bioluminescent or fluorescent proteins as reporter domains. The promoter–reporter combinations for WCBs had been identified from diverse microorganisms, and they have been employed to generate biosensors sensing specific targets (Harms et al. 2006; Robbens et al. 2010). Because of its nature to assess bioavailability, it had been widely applied to environmental systems to monitor and to quantify pollutants. Although it was traditionally assessed by instrumental and chemical analysis, WCBs were considered as a complementary analysis. However, the number of genetic systems was limited and many of them showed multi-target specificity. To overcome these shortcomings and to enhance the properties of WCBs, many research groups have put their efforts on genetic modification of sensing domains and organisms.
In the present study, we demonstrated a novel strategy of biosensor development and investigated the applicability of biosensors to the analysis of contaminated water and soil systems. Unlike WCB systems, which are based on stress-responsive operons in microorganisms, our biosensor utilized a split-protein system (Shekhawat and Ghosh 2011), i.e., eGFP that was split into N- and C-terminal parts via MBL insertion between Ile188 and Leu196. Instead of using split-eGFP, two part of eGFP was connected via potential MBLs. Since MBLs were longer than the loop region, the stability of eGFP was decreased and lost the fluorescence. In our recent study, the structures of engineered eGFP with MBLs were built and analyzed (Lee et al. 2019). Interestingly, the overall structures were similar to native eGFP. However, it was noticed that the coordination of chromophore composed of T65, Y66, and G67 was changed. We believed this was the reason eGFP with MBL was not fluorescent without metal ions. As shown in Fig. 1, E. coli cells harboring pZnt-eGFP-HJ1 showed no fluorescence, while wild-type eGFP was fluorescent, which was ascribed to the fact that the replacement of a loop region (sequenced GDGPVL) located between β-strands 9 and 10 by MBL HJ1 (sequenced CTTCGCG) destabilized the recombinant protein. In the presence of Cd ions, fluorescence was regained, as their binding to the MBL resulted in a conformational change that brought the two parts of engineered eGFP close enough for association.
As mentioned in the “Results” section, E. coli cells harboring engineered eGFP, eGFP-HJ1, responded to both Cd and Hg in a concentration-dependent manner and could therefore be used as biosensors for the quantitation of these metals (Figs. 2 and 3). The suitability of these biosensors to real-life applications was probed by analysis of artificially amended water and soil samples using procedures similar to those employed in WCB assays (Yoon et al. 2016a, c) despite the differences in the working mechanism. The quantitation of Cd and Hg in water samples was performed by in vivo and in vitro assays, and the observation of ~ 90% recoveries in both cases confirmed that the former assay was as reliable as the latter (Table 1 and Fig. S2).
Soil samples were analyzed (by the standard curve method) using solely the in vitro assay, as the presence of cell-inseparable particles in these samples disrupted fluorescence signal acquisition. The thus determined Cd concentrations (Table 2) were significantly different from the originally spiked values, in agreement with the well-known fact that the content of bioavailable metal ions in soil samples is mostly much less than their total content in view of the strong interaction of these ions with soil particles and organic matter. Unlike in the case of water samples, the fraction of bioavailable metal(loid)s in soil is known to depend on diverse factors such as metal(loid) species, soil composition, pH, organic matter content, and other physicochemical properties. In our case, the fractions of bioavailable Cd in 5-day-aged LUFA soils were as low as 18–27%.
At this point, one should mention that environmental monitoring and risk management largely deal with total pollutant concentrations rather than with those of bioavailable pollutants, which may result in risk overestimation, particularly for soil systems. Thus, accurate risk assessment should be performed taking into account pollutant bioavailability. In fact, the importance of bioavailability has long been recognized and emphasized. However, the amount of tools for assessing bioavailability has remained limited, although numerous bacterial cell–based biosensors have been reported. Herein, we suggested a novel approach for the development of biosensors based on split-protein systems, demonstrating a metal-sensing biosensor based on eGFP and showing that the concept of fusing target–binding modules and split-protein systems offers a platform for the design of numerous other sensors, even though the performance of the described sensor was far from ideal.
References
Ammann AA (2002) Speciation of heavy metals in environmental water by ion chromatography coupled to ICP–MS. Anal Bioanal Chem 372(3):448–452
Baumann B, van der Meer JR (2007) Analysis of bioavailable arsenic in rice with whole cell living bioreporter bacteria. J Agric Food Chem 55(6):2115–2120
Belkin S (2003) Microbial whole-cell sensing systems of environmental pollutants. Curr Opin Microbiol 6(3):206–212
Brocklehurst KR, Hobman JL, Lawley B, Blank L, Marshall SJ, Brown NL, Morby AP (1999) ZntR is a Zn (II)-responsive MerR-like transcriptional regulator of zntA in Escherichia coli. Mol Micribiol 31(3):893–902
Cabantous S, Terwilliger TC, Waldo GS (2005) Protein tagging and detection with engineered self-assembling fragments of green fluorescent protein. Nat Biotechnol 23(1):102–107
Cabantous S, Nguyen HB, Pedelacq J-D, Koraïchi F, Chaudhary A, Ganguly K, Lockard MA, Favre G, Terwilliger TC, Waldo GS (2013) A new protein-protein interaction sensor based on tripartite split-GFP association. Sci Rep 3:2854
Harms H, Wells MC, van der Meer JR (2006) Whole-cell living biosensors—are they ready for environmental application? Appl Microbiol Biotechnol 70(3):273–280
Järup L (2003) Hazards of heavy metal contamination. Br Med Bull 68(1):167–182
Kang Y, Lee W, Jang G, Kim B-G, Yoon Y (2018a) Modulating the sensing properties of Escherichia coli-based bioreporters for cadmium and mercury. Appl Microbiol Biotechnol 102(11):4863–4872
Kang Y, Lee W, Kim S, Jang G, Kim B-G, Yoon Y (2018b) Enhancing the copper-sensing capability of Escherichia coli-based whole-cell bioreporters by genetic engineering. Appl Microbiol Biotechnol 102(3):1513–1521
Küpper H, Küpper F, Spiller M (1998) In situ detection of heavy metal substituted chlorophylls in water plants. Photosynth Res 58(2):123–133
Kuswandi B (2003) Simple optical fibre biosensor based on immobilised enzyme for monitoring of trace heavy metal ions. Anal Bioanal Chem 376(7):1104–1110
Lee W, Kim H, Kang Y, Lee Y, Yoon Y (2019) A biosensor platform for metal detection based on enhanced green fluorescent protein. Sensors 19(8):1846
Peralta-Videa JR, Lopez ML, Narayan M, Saupe G, Gardea-Torresdey J (2009) The biochemistry of environmental heavy metal uptake by plants: implications for the food chain. Int J Biochem Cell Biol 41(8–9):1665–1677
Rawson DM, Willmer AJ, Turner AP (1989) Whole-cell biosensors for environmental monitoring. Biosensors 4(5):299–311
Robbens J, Dardenne F, Devriese L, De Coen W, Blust R (2010) Escherichia coli as a bioreporter in ecotoxicology. Appl Microbiol Biotechnol 88(5):1007–1025
Schalk IJ, Hannauer M, Braud A (2011) New roles for bacterial siderophores in metal transport and tolerance. Environ Microbiol 13(11):2844–2854
Shekhawat SS, Ghosh I (2011) Split-protein systems: beyond binary protein–protein interactions. Curr Opin Chem Biol 15(6):789–797
Singh A, Prasad SM (2011) Reduction of heavy metal load in food chain: technology assessment. Rev Environ Sci Bio 10(3):199–214
Stynen B, Tournu H, Tavernier J, Van Dijck P (2012) Diversity in genetic in vivo methods for protein-protein interaction studies: from the yeast two-hybrid system to the mammalian split-luciferase system. Microbiol Mol Biol Rev 76(2):331–382
Turpeinen R, Virta M, Häggblom MM (2003) Analysis of arsenic bioavailability in contaminated soils. Environ Toxicol Chem 22(1):1–6
Yoon Y, Kang Y, Chae Y, Kim S, Lee Y, Jeong S-W, An Y-J (2016a) Arsenic bioavailability in soils before and after soil washing: the use of Escherichia coli whole-cell bioreporters. Environ Sci Pollut Res 23(3):2353–2361
Yoon Y, Kim S, Chae Y, Jeong S-W, An Y-J (2016b) Evaluation of bioavailable arsenic and remediation performance using a whole-cell bioreporter. Sci Total Environ 547:125–131
Yoon Y, Kim S, Chae Y, Kang Y, Lee Y, Jeong S-W, An Y-J (2016c) Use of tunable whole-cell bioreporters to assess bioavailable cadmium and remediation performance in soils. PLoS One 11(5):e0154506
Yoon Y, Kang Y, Lee W, Oh K-C, Jang G, Kim B-G (2018) Modulating the properties of metal-sensing whole-cell bioreporters by interfering with Escherichia coli metal homeostasis. J Microbiol Biotechnol 28(2):323–329
Yuan C-g, Shi J-b, He B, Liu J-f, Liang L-n, Jiang G-b (2004) Speciation of heavy metals in marine sediments from the East China Sea by ICP-MS with sequential extraction. Environ Int 30(6):769–783
Zhylyak G, Dzyadevich S, Korpan Y, Soldatkin A, El’Skaya A (1995) Application of urease conductometric biosensor for heavy-metal ion determination. Sens and Actuators B Chem 24(1–3):145–148
Funding
This study was supported by the Basic Science Research Program of the National Research Foundation of Korea (NRF) funded by the Ministry of Science, ICT, and Future Planning (NRF-2017R1E1A1A01073894 to Y. Y.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 206 kb)
Rights and permissions
About this article
Cite this article
Kim, H., Lee, W. & Yoon, Y. Heavy metal(loid) biosensor based on split-enhanced green fluorescent protein: development and characterization. Appl Microbiol Biotechnol 103, 6345–6352 (2019). https://doi.org/10.1007/s00253-019-09908-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-019-09908-7