Abstract
Vitamin B1 (VB1) is an essential coenzyme for carbohydrate metabolism and involved in energy generation in most organisms. In this study, we found that insufficient biosynthesis of VB1 in Clostridium acetobutylicum ATCC 824 is a major limiting factor for efficient acetone-butanol-ethanol (ABE) fermentation. In order to improve the fermentation performance of C. acetobutylicum ATCC 824, the VB1 biosynthesis pathway was strengthened by overexpressing the thiC, thiG, and thiE genes. The engineered strain 824(thiCGE) showed enhanced VB1 and energy synthesis, resulting in better growth, faster sugar consumption, higher solvents production, and lower acids formation than the wild-type strain in both VB1 free and normal P2 medium (1 mg/L). Compared with the wild-type strain, 824(thiCGE) produced 13.0 ± 0.1% or 12.7 ± 1.2% more butanol in VB1 free P2 medium when glucose or xylose was used as the substrate, respectively. When mixed sugar (glucose:xylose = 2:1) was used as the substrate in VB1 free P2 medium, the xylose consumption rate and butanol titer of 824(thiCGE) were 45.8 ± 1.9% and 20.4 ± 0.3% higher than those of the wild-type strain. All these results demonstrated that this metabolic engineering strategy could provide a new and effective way to improve the cellular performance of solventogenic clostridia. In addition, it may have some potential application value in ABE fermentation using simple medium and/or lignocellulosic biomass.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Clostridium acetobutylicum is an attractive anaerobe that produces the solvents acetone, butanol, and ethanol (ABE) in a ratio of 3:6:1 (Lütke-Eversloh and Bahl 2011). Among them, butanol is the most important product due to its higher production and economic values. In addition to be an important chemical precursor for paints, polymers and plastics, butanol is also considered as an ideal biofuel for replacing fossil fuels (Lee et al. 2008). Although it has broad application prospects, low product titer and productivity due to the inefficient fermentation process led to poor economic benefit, which has seriously hindered the industrialization of biobutanol. In order to overcome these obstacles, various strategies have been developed, such as strain screening (Formanek et al. 1997; Guo et al. 2011), adaptive engineering (Isar and Rangaswamy 2012; Yang and Zhao 2013), genetic modification (Harris et al. 2001; Harris et al. 2000; Jang et al. 2012; Nair et al. 1999; Zhu et al. 2011), and process optimization (Ezeji et al. 2004a; Ezeji et al. 2004b; Groot' et al. 1984; Xue et al. 2012). Recently, some efforts have been undertaken to improve the fermentation performance of C. acetobutylicum, which focused on cell growth factor, such as flavonoids (Wang et al. 2014) and biotin (Yang et al. 2016).
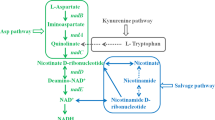
Vitamin B1 (VB1), namely thiamine, is an essential cell growth factor required by most organisms. It is very crucial for a number of central metabolic processes, such as glycolysis, pentose phosphate pathway, and tricarboxylic acid cycle (Bazurto et al. 2015). In these biochemical reactions, thiamin is served as a cofactor in thiamin pyrophosphate-dependent enzymes, including pyruvate dehydrogenase, transketolase, α-ketoglutarate dehydrogenase, pyruvate decarboxylase, branched-chain α-ketoacid dehydrogenase, and so on (Frank et al. 2007), most of which are relevant to ATP or NAD(P)H. Putative genes responsible for VB1 biosynthesis in C. acetobutylicum ATCC 824 were identified (Fig. 1), indicating that VB1 might be produced during the fermentation. However, it still needs to add a dose of VB1 to the fermentation medium, such as P2 medium (Baer et al. 1987), to maintain normal cell physiological and efficient ABE production.
VB1 is composed of a pyrimidine ring and a thiazole ring, and these two rings were linked by a methylene bridge (Manzetti et al. 2014). In prokaryotes (e.g., Escherichia coli, Bacillus subtilis, and Salmonella typhimurium), there are many enzymes involved in VB1 biosynthesis (Begley et al. 1999). For example, ThiF, ThiS, ThiG, ThiH, and ThiI are participated in the thiazole ring biosynthesis, ThiC is required for the pyrimidine ring biosynthesis, and ThiE is necessary for the catalytic reaction that linking of the thiazole ring and pyrimidine ring. In addition, some kinase also involved in VB1 biosynthesis, such as ThiL, PdxK, ThiD, and ThiM (Begley et al. 1999). In C. acetobutylicum ATCC 824, Dxs, NifS, ThiI, ThiH, ThiG, ThiC, ThiK, and ThiE may be responsible for VB1 biosynthesis based on the annotated in the genome of C. acetobutylicum ATCC 824 (Nolling et al. 2001) (Fig. 1).
In this study, in order to improve the fermentation performance of C. acetobutylicum ATCC 824, the VB1 biosynthesis pathway was strengthened by overexpressing three essential genes, including thiC. (encoding phosphomethylpyrimidine synthase), thiG (encoding thiazole synthase), and thiE (encoding thiamine monophosphate synthase) (Yu et al. 2015; Settembre et al. 2003). The results showed that when using glucose or xylose as the substrate, the fermentation performance of the engineered strain 824(thiCGE) was significantly improved. What is more, 824(thiCGE) also exhibited stronger mixed sugars utilizing ability. Therefore, this work revealed that VB1 is a pivotal cell growth factor for C. acetobutylicum ATCC 824, and the engineering strategy employed here could provide a new and effective way to improve the cellular performance of solventogenic clostridia.
Materials and methods
Bacterial strains and plasmids
The strains and plasmids used in this study are showed in Table 1. The plasmids pMTL82151 (Heap et al. 2009) and pAN2 (Heap et al. 2007) and strain C. acetobutylicum ATCC 824 were supplied by Prof. Shang-Tian Yang from the Ohio State University. E. coli DH5α was purchased from Tiangen (Beijing, China) and used for amplification and recombinant plasmids construction. E. coli TOP10 (bearing the plasmid pAN2) was used for amplification and methylation of the recombinant plasmids.
Growth conditions
E. coli DH5α and E. coli TOP10 were grown aerobically at 37 °C in liquid Luria-Bertani (LB) medium or on LB agar, and 25 μg/mL chloramphenicol or 20 μg/mL tetracycline was added as needed. C. acetobutylicum ATCC 824 or its derived strains were grown anaerobically at 37 °C in liquid reinforced clostridial medium (RCM) (Ventura et al. 2013) or on RCM agar (supplemented with 30 μg/mL thiamphenicol as needed). P2 medium (pH 6.5) (Liao et al. 2017) (supplemented with 30 μg/mL thiamphenicol as needed) was used for batch fermentation.
DNA manipulation and transformation
Axygen®AxyPrep™ Genomic DNA Kit (Corning, Wujiang, China) was used for isolating the genomic DNA from C. acetobutylicum ATCC 824, and Axygen® AxyPrep™ Plasmid Kit (Corning, Wujiang, China) was used for isolating the plasmids DNA from E. coli strains. Restriction enzymes used in this study were purchased from Thermo Scientific (Shanghai, China). PCR primers were synthesized by Sangon Biotech (Shanghai, China) and DNA ploymerase were purchased from Takara Biomedical Technology (Beijing, China). Table 1 also lists the PCR primers used in this study. Firstly, the genes thiC (CAC3014), thiG (CAC2922), and thiE (CAC0495) involved in VB1 biosynthesis were PCR amplified from the genomic of C. acetobutylicum ATCC 824 using the corresponding primers. Subsequently, the thiG and thiE were ligated through overlap PCR using thiG and thiE fragments as the templates and thiGE-F/thiGE-R as the primers, yielding gene fragment of thiGE. Finally, the gene fragment of thiC and thiGE were ligated through overlap PCR using thiC and thiGE fragments as the templates and thiCGE-F/thiCGE-R as the primers, yielding gene fragment of thiCGE. Then, the thiCGE fragment was cloned into the plasmid pMTL-Pthl, which was digested with BamHI and HindIII using ClonExpress II One Step Cloning Kit (Vazyme Biotech Co.,Ltd., Nanjing, China), yielding the recombinant plasmid pMTL-Pthl thiCGE. The recombinant plasmid was first transferred into E. coli TOP10 bearing the plasmid pAN2 for methylation. Then, the methylated recombinant plasmid was isolated and transferred into C. acetobutylicum ATCC 824 by electrotransformation as described previously (Liao et al. 2017). The primers pMTL-F/pMTL-R (Table 1) were used to identify the recombinant strains.
RNA isolation and reverse transcription-PCR analysis
C. acetobutylicum strains were inoculated in serum bottles with VB1 free P2 medium containing 80 g/L glucose and grown anaerobically at 37 °C. The samples were taken at 12, 24, and 36 h, and the cell was collected by centrifugation. The total RNA was isolated using RNAprep pure Cell/Bacteria Kit (Tiangen Biotech, Beijing, China). The PrimeScript™ RT reagent Kit with gDNA Eraser (Takara Biomedical Technology, Dalian, China) was used for cDNA Synthesis, and SYBR Premix Ex Taq II (2×) (Tli RNaseH Plus), Bulk (Takara Biomedical Technology, Beijing, China) was used for real-time PCR. PCR reactions were carried out in a LightCycler®96 instrument (Roche, Switzerland) with the protocol: 30 s at 95 °C, then 40 cycles of 5 s at 95 °C, and 30 s at 60 °C. Each transcript was performed with three PCR reactions. The pullulanase gene (CAC2679) was used as housekeeping genes (Tomas et al. 2003; Tseng et al. 2001). Primers for housekeeping genes and VB1 biosynthesis gene (thiC, thiG, and thiE) are listed in Table 1.
VB1 and ATP assays
The wild-type and 824(thiCGE) strains were inoculated in serum bottles with VB1 free P2 medium containing 80 g/L glucose and grown anaerobically at 37 °C. The samples were taken at various time points. The supernatant was collected by centrifugation and used for VB1 assay, while the cell pellets were resuspended in lysate. After the cell pellets were completely lysed, the supernatant was collected by centrifugation and used for ATP assay. The Vitamin B1 Assay Kit (Cablebridge Biotechnology, Shanghai, China) and ATP Assay Kit (Beyotime, Shanghai, China) were used for VB1 and ATP assay, respectively, according to the recommendation of the manufacturer.
Fermentation studies
C. acetobutylicum ATCC 824 or the engineered strain 824(thiCGE) was precultured to the early stationary phase in liquid RCM (supplemented with 30 μg/mL thiamphenicol as needed) as the seed culture. P2 medium with various concentrations of VB1 (0, 1, 3, or 5 mg/L) was used for ABE fermentation, and 1 mg/L represents the dose commonly added in normal P2 medium. Batch fermentations were performed in 50 mL P2 medium in serum bottles with glucose (80 g/L), xylose (80 g/L), or mixed sugars (40 g/L glucose and 20 g/L xylose). The serum bottles were filled with nitrogen to establish anaerobic conditions. The samples were taken at regular intervals to monitor the cell growth, sugar consumption, and products formation.
Analytical methods
High-performance liquid chromatography (HPLC; Waters 2695, Milford, MA) was used for measuring the concentration of sugar (Liao et al. 2017). And the concentration of products in the fermentation broth were determined by gas chromatograph (Agilent 7890A GC, Agilent Technologies) as described previously (Liao et al. 2017).
Results
Effect of exogenous VB1 addition on the fermentation performance of C. acetobutylicum
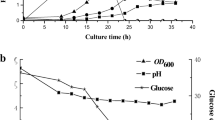
The P2 mediums with different concentrations (0, 1, 3, or 5 mg/L) of VB1 were used to study the effects of VB1 on the fermentation performance of C. acetobutylicum ATCC 824. The cell growth, glucose consumption and products formation were examined, and the results are shown in Fig. 2. Fermentation without VB1 addition was used as the control. As the VB1 concentration increased from 0 to 3 mg/L, the cellular performance of C. acetobutylicum ATCC 824 was enhanced. The best results were obtained with 3 mg/L VB1, and the max OD600, glucose consumption and total solvents titer were 7.5 ± 0.3, 79.01 ± 0.14 g/L, and 20.53 ± 0.27 g/L, respectively, which were significantly higher than those of the control (max OD600 = 6.0 ± 0.4, glucose consumption = 65.10 ± 0.57 g/L, total solvents titer = 15.90 ± 0.82 g/L). In addition from Fig. 2b, c, we can see that the glucose consumption rate and solvents formation rate were faster than the control, resulting in an increase solvents productivity. With 3 mg/L VB1, the residual glucose was only about 4.72 ± 1.35 g/L at 48 h, while it still had 20.43 ± 2.83 g/L for the control; the total solvents titer reached 20.18 ± 0.52 g/L at 48 h, which was much higher than that of the control (12.18 ± 0.49 g/L). These results indicated that VB1 plays an important role in cellular performance of C. acetobutylicum ATCC 824.
Real-time PCR and relative gene expression analysis
In general, transcriptions of thiC, thiG, and thiE in C. acetobutylicum ATCC 824 and 824(thiCGE) were increased from 12 to 24 h, then decreased at 36 h (Fig. 3). After the recombinant plasmid pMTL-Pthl thiCGE was transferred into C. acetobutylicum ATCC 824, all three measured genes were highly overexpressed. For example, the relative expression levels of thiC, thiG, and thiE of 824(thiCGE) were increased by 38.9 ± 4.6, 16.1 ± 2.7, and 41.4 ± 3.4-fold compared to those in the wild-type strain at 12 h (Fig. 3). The enhancement of the transcription level in 824(thiCGE) indicates that the thiC, thiG, and thiE were effectively overexpressed.
Effect of thiC, thiG, and thiE overexpressing on VB1 and ATP synthesis
As shown in Fig. 4a, the extracellular VB1 concentration was significantly increased during the logarithmic and stationary phases (0–36 h), then it was decreased at the end of the fermentation. During the whole fermentation process, the VB1 concentration of the engineered strain 824(thiCGE) was significantly higher than that of the wild-type strain. For example, VB1 produced by C. acetobutylicum was increased by 1.88 ± 0.09 times at 36 h, from 0.73 ± 0.07 to 2.10 ± 0.14 μM (~ 0.6 mg/L). It means that the metabolic engineering strategy to improve VB1 biosynthesis was succeeded by overexpressing thiC, thiG, and thiE.
Since VB1 is served as a coenzyme participating in some enzymatic catalytic processes of energy generation, such as glycolysis and tricarboxylic acid cycle (Manzetti et al. 2014), the ATP concentration was also measured during the fermentation process. As shown in Fig. 4b, the ATP concentrations of the engineered strain 824(thiCGE) and the wild-type strain were decreased gradually after 24 h; however, the ATP concentration of 824(thiCGE) was always higher than that of the wild-type strain. This result indicated that the energy generation was also enhanced by reinforcing the VB1 biosynthesis.
ABE fermentation using glucose as the substrate
In order to confirm the effect of VB1 biosynthesis on the fermentation performance of C. acetobutylicum ATCC 824, batch fermentation of glucose was carried out. The results showed that 824(thiCGE) exhibited better fermentation performance in both VB1 free and normal P2 medium than the wild-type strain (Table 2). When engineered strain 824(thiCGE) was cultured in the VB1 free P2 medium, the max OD600 reached up to 6.8 ± 0.2, which was significantly higher than that of the wild-type strain (max OD600 = 5.4 ± 0.3) (Fig. 5a). The engineered strain 824(thiCGE) also showed much faster glucose consumption rate and solvents production rate (Fig. 5a, b). As a result, butanol productivity was increased by 12.6 ± 0.1% compared with the wild-type strain. The total solvents produced by 824(thiCGE) reached 18.61 ± 0.22 g/L, 16.8 ± 0.9% higher than that of the wild-type strain (Fig. 5b). In addition, the organic acids concentrations of 824(thiCGE) were always lower than those of the wild-type strain throughout the fermentation process (Fig. 5c). Similar phenomenon was observed when C. acetobutylicum was cultured in the normal P2 medium (Fig. 5d–f). It should be noted that the fermentation performance of 824(thiCGE) in VB1 free P2 medium was comparable to the wild-type strain in normal P2 medium, indicating that the enhanced VB1 biosynthesis in 824(thiCGE) could replace exogenous VB1 addition in normal P2 medium (Table 2).
Batch fermentation profiles of C. acetobutylicum ATCC 824 and 824(thiCGE) cultivated in VB1 free P2 medium and in normal P2 medium with 80 g/L glucose. a The profiles of growth and glucose consumption in VB1 free P2 medium. b The profiles of solvents production in VB1 free P2 medium. c The profiles of organic acids production in VB1 free P2 medium. d The profiles of growth and glucose consumption in normal P2 medium. e The profiles of solvents production in normal P2 medium. f The profiles of organic acids production in normal P2 medium. The data are the means and standard deviations of three replicates
ABE fermentation using xylose or mixed sugars as the substrate
Since VB1 also plays an important role in the pentose phosphate pathway (Bazurto et al. 2015; Manzetti et al. 2014; Tittmann 2009), then we assumed that the enhanced VB1 biosynthesis could also promote the xylose metabolism. In order to confirm this hypothesis, the ABE fermentation by C. acetobutylicum ATCC 824 using 80 g/L xylose as the substrate was carried out. From Fig. 6, we can see that the xylose consumption rate and solvents production rate of C. acetobutylicum in normal P2 medium (Fig. 6d–f) were significantly faster than that in VB1 free P2 medium (Fig. 6a–c). The final butanol titer and xylose consumption of C. acetobutylicum ATCC 824 in normal P2 medium were 10.40 ± 0.11 g/L and 66.32 ± 0.18 g/L, respectively, 16.1 ± 2.3% and 14.1 ± 0.1% higher than those in VB1 free P2 medium (Table 3). As expected, compared with the wild-type strain, the engineered strain 824(thiCGE) exhibited better fermentation performances in both VB1 free and normal P2 medium (Fig. 6 and Table 3), although xylose utilization was much slower than glucose. When xylose fermentation was carried out in VB1 free and normal P2 medium, the butanol titer of 824(thiCGE) were 10.10 ± 0.21 and 11.95 ± 0.21 g/L, respectively, which were 12.7 ± 1.2% and 14.9 ± 0.8% higher than those of the wild-type strain (Table 3).
Batch fermentation profiles of C. acetobutylicum ATCC 824 and 824(thiCGE) cultivated in VB1 free P2 medium and in normal P2 medium with 80 g/L xylose. a The profiles of growth and xylose consumption in VB1 free P2 medium. b The profiles of solvents production in VB1 free P2 medium. c The profiles of organic acids production in VB1 free P2 medium. d The profiles of growth and xylose consumption in normal P2 medium. e The profiles of solvents production in normal P2 medium. f The profiles of organic acids production in normal P2 medium. The data are the means and standard deviations of three replicates
In addition, the fermentation performance was also studied on mixed sugars containing 40 g/L glucose and 20 g/L xylose, based on the general ratio of glucose and xylose in the lignocellulosic hydrolysate (Aristidou and Penttilä 2000). The results showed that the engineered strain 824(thiCGE) exhibited better growth, faster sugars consumption, higher solvents production, and lower acids formation than those of the wild-type strain (Fig. 7 and Table 4), as observed in glucose or xylose fermentation.
Batch fermentation profiles of C. acetobutylicum ATCC 824 and 824(thiCGE) cultivated in VB1 free P2 medium with mixed sugars (40 g/L glucose and 20 g/L xylose). a The profiles of growth. b The profiles of sugars consumption. c The profiles of solvents production. d The profiles of organic acids production. The data are the means and standard deviations of three replicates
Discussion
Improving the fermentation performance is very important for ABE fermentation. Previous studies have shown that VB1 is a crucial molecule in some enzymatic catalytic processes of carbohydrate metabolism, which participate in energy generation from different carbohydrate sources. These enzymatic catalytic processes involves in glycolysis, pentose phosphate pathway, and tricarboxylic acid cycle (Bazurto et al. 2015; Manzetti et al. 2014). In addition, VB1 could improve the cellular resistance to oxidative stress, heat stress, high light stress, and biotic stresses (Dong et al. 2015; Medina-Silva et al. 2006; Rapala-Kozik et al. 2008; Tunc-Ozdemir et al. 2009; van der Graaff et al. 2004; Wolak et al. 2015). In this study, we found that the cellular performance of C. acetobutylicum ATCC 824 was greatly affected by VB1. Although the putative genes responsible for VB1 biosynthesis were found in C. acetobutylicum ATCC 824 (Fig. 1), the VB1 biosynthesis was insufficient to maintain normal cell growth and efficient solvent production (Fig. 2). With the VB1 concentration increasing, C. acetobutylicum ATCC 824 exhibited better growth, faster sugar utilization, and solvent production, indicating that VB1 play a key role for efficient ABE fermentation.
In order to improve the fermentation performance of C. acetobutylicum ATCC 824, the VB1 biosynthesis pathway was strengthened by overexpressing three essential genes, including thiC, thiG, and thiE (Fig. 1). Although there are many enzymes participating in VB1 biosynthesis, and most of them have been validated in some prokaryotes, such as E. coli, B. subtilis, and S. typhimurium (Begley et al. 1999). We choose these three enzymes based on the structure of VB1 and previous reports. VB1 is formed from a pyrimidine ring and a thiazole ring, which are linked by a methylene bridge; thus, there are separate biosynthetic pathways for the pyrimidine moiety and thiazole moiety. As shown in Fig. 1, pyrimidine is formed from 5-aminoimidazole ribotide, and ThiC (encoded by thiC) is the key enzyme participated in pyrimidine ring synthesis; thiazole is formed from pyruvate, glyceraldehyde 3-phosphate, tyrosine and cysteine, and the oxidative condensation of these metabolic products (1-deoxy-D-xylulose-5-phosphate, thiocarboxy-[sulfur-carrier protein], and iminoglycine) into thiazole is catalyzed by ThiG (encoded by thiG); ThiE (encoded by thiE) is necessary for linking of the pyrimidine moiety and thiazole moiety to form thiamin monophosphate (Chakravortty et al. 2015; Settembre et al. 2003). The function of these genes have been validated in other bacteria, pyrimidine was needed when thiC was knocked out, deletion mutations of thiG resulted in a thiazole auxotrophy, and mutants of thiE resulted in thiamin insufficient (Backstrom et al. 1995; Dong et al. 2015; Kelleher et al. 1998; Begley et al. 1999; Webb et al. 1996; Zhang et al. 1997). A previous study has also shown that the increase in VB1 concentration was not obvious in Arabidopsis thaliana when overexpressing thiC or thi1 (equivalent to thiG in C. acetobutylicum) singly, while it was dramatically improved by overexpressing thiC and thi1 simultaneously (Dong et al. 2015). Therefore, we attempted to reinforce the VB1 biosynthesis by co-expressing these three essential genes (thiC, thiG, and thiE). The results showed that both the transcript levels of the thiC, thiG, and thiE (Fig. 3) and VB1 concentration (Fig. 4a) of the engineered strain 824(thiCGE) were significantly higher than those of the wild-type strain, demonstrating that this metabolic engineering strategy was successful in C. acetobutylicum.
The fermentation results in VB1 free P2 medium showed that both the total glucose consumption and the glucose metabolic rate of 824(thiCGE) were higher than those of the wild type strain (Fig. 5a). This may be attributed to the improved enzymes activities of glucose metabolism by reinforcing the VB1 biosynthesis, which was consistent with the previous research that VB1 supplement enhanced the metabolic flux and enzyme activity (phosphofructokinase) of glycolysis pathway (Xu et al. 2008). In addition, since 824(thiCGE) can produce sufficient energy from glucose metabolism due to the enhanced VB1 biosynthesis (Fig. 4b), the acids concentrations were decreased significantly (Fig. 5c). With the improved efficiency of glucose metabolism and ATP production, the butanol production of 824(thiCGE) was also enhanced when compared with the wild-type strain (Fig. 5b). For example, the butanol titer in the VB1 free and normal P2 medium were 10.94 ± 0.08 g/L and 13.18 ± 0.23 g/L for 824(thiCGE), respectively, 13.0 ± 0.1% and 19.8 ± 0.6% higher than that of the wild-type strain (Table 2). It should be noted that the fermentation performances of 824(thiCGE) in VB1 free P2 medium were comparable to the wild-type strain in normal P2 medium, which means that the engineered strain 824(thiCGE) could produce adequate amounts of VB1 to replace exogenous VB1 addition in normal P2 medium (Table 2 and Table 3). Therefore, this strategy could reduce the investment of ABE fermentation and improve the economic benefit.
Pentose phosphate pathway was essential for xylose metabolism, and xylose utilization could be improved by overexpressing the genes involved in pentose phosphate pathway in C. acetobutylicum (Gu et al. 2009; Jin et al. 2014). As a coenzyme, VB1 also participates in the pentose phosphate pathway (Bazurto et al. 2015; Manzetti et al. 2014; Tittmann 2009). Therefore, it is expected that VB1 will have a positive effect on xylose metabolism. When fermentation was carried out in normal P2 medium using xylose as the substrate, the butanol productivity of C. acetobutylicum ATCC 824 reached 0.108 ± 0.004 g/L·h, exceeding that of in VB1 free P2 medium by more than 16.1 ± 3.7% (Table 3), demonstrating that VB1 also plays an important role in xylose metabolism. Furthermore, 824(thiCGE) produced 12.7 ± 1.2% and 14.9 ± 0.8% more butanol than the wide-type in VB1 free and normal P2 medium (Table 3), respectively. It is worth noting that when mixed sugars (glucose:xylose = 2:1) was used as the substrate in VB1 free P2 medium, the engineered strain 824(thiCGE) also showed better fermentation performance than the wild-type strain (Fig. 7). Both glucose and xylose utilization were improved by reinforcing the intracellular VB1 biosynthesis, although xylose consumption was still lagged behind glucose. As a result, the xylose metabolic rate and butanol titer of 824(thiCGE) were increased by 45.8 ± 1.9% and 20.4 ± 0.3% compared with the wild-type strain (Table 4). Similarly, the comparative transcriptomes analysis among the mutant strains with improved xylose utilization and the native strain of Saccharomyces cerevisiae showed that the most prominently changed (upregulated) genes were involved in VB1 biosynthesis (Zeng et al. 2017). Improving xylose utilization of C. acetobutylicum is of great significance to ABE fermentation with lignocellulosic biomass. Therefore, this metabolic engineering strategy may provide a new way for enhanced use of lignocellulosic biomass.
In this study, we have successfully constructed a robust engineering bacterium by reinforcing the VB1 biosynthetic pathway in C. acetobutylicum for the first time. The engineered strain 824(thiCGE) showed greatly improved performance grown on glucose in both VB1 free and normal P2 medium. What is more, improved xylose utilization was also observed when xylose or mixed sugars were used as the substrate. All these results demonstrated that this strategy will have some potential application value in ABE fermentation using simple medium and/or lignocellulosic biomass. On the other hand, the VB1 biosynthesis in 824(thiCGE) is still insufficient to reach the optimal condition (3 mg/L). Therefore, further improve the VB1 biosynthesis in C. acetobutylicum ATCC 824 is necessary in future research. To achieve this objective, the methods of modifying promoter and regulatory parts or overexpressing the genes in chromosome instead of in plasmid can be used, which have been proved to be effective in improving gene expression (Shi et al. 2013; Yang et al. 2016).
References
Aristidou A, Penttilä M (2000) Metabolic engineering applications to renewable resource utilization. Curr Opin Biotechnol 11(2):187–198. https://doi.org/10.1016/S0958-1669(00)00085-9
Backstrom AD, McMordie RAS, Begley TP (1995) Biosynthesis of thiamin I: the function of the thiE gene product. J Am Chem Soc 117(8):2351–2352. https://doi.org/10.1021/ja00113a025
Baer SH, Blaschek HP, Smith TL (1987) Effect of butanol challenge and temperature on lipid composition and membrane fluidity of butanol-tolerant Clostridium acetobutylicum. Appl Environ Microbiol 53(12):2854–2861. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC204212/
Bazurto JV, Heitman NJ, Downs DM (2015) Aminoimidazole carboxamide ribotide exerts opposing effects on thiamine synthesis in Salmonella enterica. J Bacteriol 197(17):2821–2830. http://jb.asm.org/content/197/17/2821.abstract
Begley TP, Downs D, Ealick SE, McLafferty FW, Van Loon AP, Taylor S, Campobasso N, Chiu H-J, Kinsland C, Reddick JJ, Xi J (1999) Thiamin biosynthesis in prokaryotes. Arch Microbiol 171:293–300. https://doi.org/10.1007/s002030050713
Dong W, Stockwell VO, Goyer A (2015) Enhancement of thiamin content in Arabidopsis thaliana by metabolic engineering. Plant Cell Physiol 56(12):2285–2296. https://doi.org/10.1093/pcp/pcv148
Ezeji TC, Qureshi N, Blaschek HP (2004a) Acetone butanol ethanol (ABE) production from concentrated substrate: reduction in substrate inhibition by fed-batch technique and product inhibition by gas stripping. Appl Microbiol Biotechnol 63(6):653–658. https://doi.org/10.1007/s00253-003-1400-x
Ezeji TC, Qureshi N, Blaschek HP (2004b) Butanol fermentation research: upstream and downstream manipulations. Chem Rec 4(5):305–314. https://onlinelibrary.wiley.com/doi/abs/10.1002/tcr.20023
Formanek J, Mackie R, Blaschek HP (1997) Enhanced butanol production by Clostridium beijerinckii BA101 grown in semidefined P2 medium containing 6 percent maltodextrin or glucose. Appl Environ Microbiol 63(6):2306–2310. http://aem.asm.org/content/63/6/2306.abstract
Frank RA, Leeper FJ, Luisi BF (2007) Structure, mechanism and catalytic duality of thiamine-dependent enzymes. Cell Mol Life Sci 64(7–8):892–905. https://doi.org/10.1007/s00018-007-6423-5
Groot' WJ, van den Qever CE, Kossen NWF (1984) Pervaporation for simultaneous product recovery in the butanol/isopropanol batch fermentation. Biotechnol Lett 6(11):709–714. https://doi.org/10.1007/BF00133061
Gu Y, Li J, Zhang L, Chen J, Niu L, Yang Y, Yang S, Jiang W (2009) Improvement of xylose utilization in Clostridium acetobutylicum via expression of the talA gene encoding transaldolase from Escherichia coli. J Biotechnol 143(4):284–287. https://doi.org/10.1016/j.jbiotec.2009.08.009
Guo T, Tang Y, Xi YL, He AY, Sun BJ, Wu H, Liang DF, Jiang M, Ouyang PK (2011) Clostridium beijerinckii mutant obtained by atmospheric pressure glow discharge producing high proportions of butanol and solvent yields. Biotechnol Lett 33(12):2379–2383. https://doi.org/10.1007/s10529-011-0702-9
Harris LM, DR P, WN E, PE T (2000) Characterization of recombinant strains of the Clostridium acetobutylicum butyrate kinase inactivation mutant: need for new phenomenological models for solventogenesis and butanol inhibition? Biotechnol Bioeng 67(1):1–11. https://onlinelibrary.wiley.com/doi/abs/10.1002/%28SICI%291097-0290%2820000105%2967%3A1%3C1%3A%3AAID-BIT1%3E3.0.CO%3B2-G
Harris L, Blank L, Desai R, Welker N, Papoutsakis E (2001) Fermentation characterization and flux analysis of recombinant strains of Clostridium acetobutylicum with an inactivated solR gene. J Ind Microbiol Biotechnol 27(5):322–328. https://doi.org/10.1038/sj.jim.7000191
Heap JT, Pennington OJ, Cartman ST, Carter GP, Minton NP (2007) The ClosTron: a universal gene knock-out system for the genus Clostridium. J Microbiol Methods 70(3):452–464. https://doi.org/10.1016/j.mimet.2007.05.021
Heap JT, Pennington OJ, Cartman ST, Minton NP (2009) A modular system for Clostridium shuttle plasmids. J Microbiol Methods 78(1):79–85. https://doi.org/10.1016/j.mimet.2009.05.004
Isar J, Rangaswamy V (2012) Improved n-butanol production by solvent tolerant Clostridium beijerinckii. Biomass Bioenergy 37:9–15. https://doi.org/10.1016/j.biombioe.2011.12.046
Jang Y-S, Lee JY, Lee J, Park JH, Im JA, Eom M-H, Lee J, Lee S-H, Song H, Cho J-H, Seung DY, Lee SY (2012) Enhanced butanol production obtained by reinforcing the direct butanol-forming route in Clostridium acetobutylicum. mBio 3(5):e00314–e00312. http://mbio.asm.org/content/3/5/e00314-12.abstract
Jin L, Zhang H, Chen L, Yang C, Yang S, Jiang W, Gu Y (2014) Combined overexpression of genes involved in pentose phosphate pathway enables enhanced D-xylose utilization by Clostridium acetobutylicum. J Biotechnol 173:7–9. https://doi.org/10.1016/j.jbiotec.2014.01.002
Kelleher NL, Taylor SV, Grannis D, Kinsland C, Chiu HJ, Begley TP, McLafferty FW (1998) Efficient sequence analysis of the six gene products (7-74 kDa) from the Escherichia coli thiamin biosynthetic operon by tandem high-resolution mass spectrometry. Protein Sci 7(8):1796–1801. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC2144080/
Lee SY, Park JH, Jang SH, Nielsen LK, Kim J, Jung KS (2008) Fermentative butanol production by clostridia. Biotechnol Bioeng 101(2):209–228. https://doi.org/10.1002/bit.22003
Liao Z, Zhang Y, Luo S, Suo Y, Zhang S, Wang J (2017) Improving cellular robustness and butanol titers of Clostridium acetobutylicum ATCC824 by introducing heat shock proteins from an extremophilic bacterium. J Biotechnol 252:1–10. http://www.sciencedirect.com/science/article/pii/S0168165617301943
Lütke-Eversloh T, Bahl H (2011) Metabolic engineering of Clostridium acetobutylicum: recent advances to improve butanol production. Curr Opin Biotechnol 22(5):634–647. https://doi.org/10.1016/j.copbio.2011.01.011
Manzetti S, Zhang J, van der Spoel D (2014) Thiamin function, metabolism, uptake, and transport. Biochemistry 53(5):821–835. https://doi.org/10.1021/bi401618y
Medina-Silva R, Barros MP, Galhardo RS, Netto LE, Colepicolo P, Menck CF (2006) Heat stress promotes mitochondrial instability and oxidative responses in yeast deficient in thiazole biosynthesis. Res Microbiol 157(3):275–281. https://doi.org/10.1016/j.resmic.2005.07.004
Nair RV, Green EM, Watson DE, Bennett GN, Papoutsakis ET (1999) Regulation of the sol locus genes for butanol and acetone formation in Clostridium acetobutylicum ATCC 824 by a putative transcriptional repressor. J Bacteriol 181(1):319–330.http://jb.asm.org/content/181/1/319.abstract
Nolling J, Breton G, Omelchenko MV, Makarova KS, Zeng Q, Gibson R, Lee HM, Dubois J, Qiu D, Hitti J, Production GTCSC, Finishing BT, Wolf YI, Tatusov RL, Sabathe F, Doucette-Stamm L, Soucaille P, Daly MJ, Bennett GN, Koonin EV, Smith DR (2001) Genome sequence and comparative analysis of the solvent-producing bacterium Clostridium acetobutylicum. J Bacteriol 183(16):4823–4838.http://jb.asm.org/content/183/16/4823.abstract
Rapala-Kozik M, Kowalska E, Ostrowska K (2008) Modulation of thiamine metabolism in Zea mays seedlings under conditions of abiotic stress. J Exp Bot 59(15):4133–4143. https://doi.org/10.1093/jxb/ern253
Settembre E, Begley TP, Ealick SE (2003) Structural biology of enzymes of the thiamin biosynthesis pathway. Curr Opin Struct Biol 13(6):739–747. https://doi.org/10.1016/j.sbi.2003.10.006
Shi A, Zhu X, Lu J, Zhang X, Ma Y (2013) Activating transhydrogenase and NAD kinase in combination for improving isobutanol production. Metab Eng 16:1–10. https://doi.org/10.1016/j.ymben.2012.11.008
Tittmann K (2009) Reaction mechanisms of thiamin diphosphate enzymes: redox reactions. FEBS J 276(9):2454–2468. https://doi.org/10.1111/j.1742-4658.2009.06966.x
Tomas CA, Welker NE, Papoutsakis ET (2003) Overexpression of groESL in Clostridium acetobutylicum results in increased solvent production and tolerance, prolonged metabolism, and changes in the cell’s transcriptional program. Appl Environ Microbiol 69(8):4951–4965. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC169105/
Tseng GC, Oh M-K, Rohlin L, Liao JC, Wong WH (2001) Issues in cDNA microarray analysis: quality filtering, channel normalization, models of variations and assessment of gene effects. Nucleic Acids Res 29(12):2549–2557. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC55725/
Tunc-Ozdemir M, Miller G, Song L, Kim J, Sodek A, Koussevitzky S, Misra AN, Mittler R, Shintani D (2009) Thiamin confers enhanced tolerance to oxidative stress in Arabidopsis. Plant Physiol 151(1):421–432. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC2735988/
van der Graaff E, Hooykaas P, Lein W, Lerchl J, Kunze G, Sonnewald U, Boldt R (2004) Molecular analysis of “de novo” purine biosynthesis in solanaceous species and in Arabidopsis thaliana. Front Biosci 9:1803–1816. https://doi.org/10.2741/1344
Ventura J-RS, Hu H, Jahng D (2013) Enhanced butanol production in Clostridium acetobutylicum ATCC 824 by double overexpression of 6-phosphofructokinase and pyruvate kinase genes. Appl Microbiol Biotechnol 97(16):7505–7516. https://doi.org/10.1007/s00253-013-5075-7
Wang L, Xia M, Zhang L, Chen H (2014) Promotion of the Clostridium acetobutylicum ATCC 824 growth and acetone–butanol–ethanol fermentation by flavonoids. World J Microbiol Biotechnol 30(7):1969–1976. https://doi.org/10.1007/s11274-014-1619-y
Webb E, Febres F, Downs DM (1996) Thiamine pyrophosphate (TPP) negatively regulates transcription of some thi genes of Salmonella typhimurium. J Bacteriol 178(9):2533–2538.http://jb.asm.org/content/178/9/2533.abstract
Wolak N, Tomasi M, Kozik A, Rapala-Kozik M (2015) Characterization of thiamine uptake and utilization in Candida spp. subjected to oxidative stress. Acta Biochim Pol 62(3):445–455. https://doi.org/10.18388/abp.2015_1044
Xu G-Q, Chu J, Zhuang Y-P, Wang Y-H, Zhang S-L (2008) Effects of vitamins on the lactic acid biosynthesis of Lactobacillus paracasei NERCB 0401. Biochem Eng J 38(2):189–197. https://doi.org/10.1016/j.bej.2007.07.003
Xue C, Zhao J, Lu C, Yang S-T, Bai F, Tang IC (2012) High-titer n-butanol production by Clostridium acetobutylicum JB200 in fed-batch fermentation with intermittent gas stripping. Biotechnol Bioeng 109:2746–2756. https://onlinelibrary.wiley.com/doi/abs/10.1002/bit.24563
Yang ST, Zhao J (2013) Adaptive engineering of Clostridium for increased butanol production. US Patent 8450093
Yang Y, Lang N, Yang G, Yang S, Jiang W, Gu Y (2016) Improving the performance of solventogenic clostridia by reinforcing the biotin synthetic pathway. Metab Eng 35:121–128. https://doi.org/10.1016/j.ymben.2016.02.006
Yu X, Liang X, Liu K, Dong W, Wang J, Zhou M-g (2015) The thiG Gene Is Required for Full Virulence of Xanthomonas oryzae pv. oryzae by Preventing Cell Aggregation. PLoS ONE 10(7):e0134237. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC4519133/
Zeng WY, Tang YQ, Gou M, Sun ZY, Xia ZY, Kida K (2017) Comparative transcriptomes reveal novel evolutionary strategies adopted by Saccharomyces cerevisiae with improved xylose utilization capability. Appl Microbiol Biotechnol 101(4):1753–1767. https://doi.org/10.1007/s00253-016-8046-y
Zhang YI, Taylor S, Chiu H-J, P Begley T (1997) Characterization of the Bacillus subtilis thiC operon involved in thiamine biosynthesis. J Bacteriol 179:3030–3035. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC179069/
Zhu L, Dong H, Zhang Y, Li Y (2011) Engineering the robustness of Clostridium acetobutylicum by introducing glutathione biosynthetic capability. Metab Eng 13(4):426–434. https://doi.org/10.1016/j.ymben.2011.01.009
Funding
This work was funded by the National Natural Science Foundation of China (21676098), the Fundamental Research Funds for the Central Universities (2017PY013, 2017BQ084), and the China Postdoctoral Science Foundation Funded Project (2017M612667).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Liao, Z., Suo, Y., Xue, C. et al. Improving the fermentation performance of Clostridium acetobutylicum ATCC 824 by strengthening the VB1 biosynthesis pathway. Appl Microbiol Biotechnol 102, 8107–8119 (2018). https://doi.org/10.1007/s00253-018-9208-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-9208-x