Abstract
Numerous methoxylated flavonoids exhibit pronounced bioactivities. Their biotechnological production and diversification are therefore of interest to pharmaceutical and nutraceutical industries. We used a set of enzymes from sweet basil (Ocimum basilicum) to construct five strains of Saccharomyces cerevisiae producing 8- and/or 6-substituted, methoxylated flavones from their natural precursor apigenin. After identifying several growth parameters affecting the overall yields and flux, we applied optimized conditions and explored the ability of the generated strains to utilize alternative substrates. The yeast cells produced substantial amounts of 6-hydroxylated, methylated derivatives of naringenin and luteolin while the corresponding derivatives of flavonol kaempferol were only detected in trace amounts. Analysis of the intermediates and by-products of the different bioconversions suggested that the substrate specificity of both the hydroxylases and the flavonoid O-methyltransferases is imposing barriers on yields obtained with alternative substrates and highlighted steps that appear to represent bottlenecks en route to increasing the strains’ efficiencies. Additionally, analysis of flavonoid localization during fermentation revealed unequal distribution with strong intracellular accumulation of a number of methylated flavonoids and extracellular enrichment of several pathway intermediates. This work establishes a platform for the production of complex methoxylated flavonoids and discusses strategies for its improvement.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Efforts to engineer flavonoid metabolism in microorganisms have been conducted for over a decade, and tremendous progress in process optimization and increases in yield including de novo biosynthesis from simple and cheap carbon sources have been achieved (Chouhan et al. 2017; Pandey et al. 2016; Trantas et al. 2015). Among the target modifications and molecules, the production of lipophilic (poly)methoxylated flavonoids has not been extensively addressed. These compounds occur in numerous land plant families and often possess pronounced bioactivities (Berim and Gang 2016; Koirala et al. 2016). Ready access to larger quantities of pure (poly)methoxylated flavonoids would facilitate a better assessment of their pharmacological potential as well as the generation of novel chemical entities.
In the first experiments aimed at the production of methoxylated flavonoids, six flavonoid O-methyltransferases (FOMTs) from peppermint (Mentha × piperita) were expressed in Escherichia coli and assayed in vitro with an array of potential substrates (Willits et al. 2004). Their biodiversification potential was then tested in vivo with quercetin as fed substrate using single or mixed cultures of E. coli cells expressing individual enzymes. These experiments revealed low substrate selectivity and high regiospecificity of the isolated FOMTs and enabled the production of several mono- and dimethylated and one trimethylated derivative of quercetin (Willits et al. 2004). Other studies employing one or two FOMTs from a plant or a microorganism and E. coli as a host and feeding flavonoid precursors were published more recently (Jeon et al. 2009; Kim et al. 2008; Kim et al. 2006a; Kim et al. 2005; Kim et al. 2006b; Koirala et al. 2014; Lee et al. 2017). In addition, successful production of several 7-O-methylated flavonoids from phenylpropanoid acids (Leonard et al. 2006a; Malla et al. 2012) or from tyrosine (Lee et al. 2015) in E. coli has been reported.
The structural diversity of (poly)methoxylated flavonoids is significantly expanded by the “decorative” hydroxylations that are introduced after the formation of their 5,7,4′-hydroxylated backbone (Anarat-Cappillino and Sattely 2014; Ververidis et al. 2007). Most frequently, positions 3′ and 5′ of ring B and 6 and 8 of ring A carry such decorative hydroxyl groups that can be subsequently methylated. The production of 5′- and/or 3′-hydroxylated flavonoids has been achieved using both de novo and precursor feeding strategies (Amor et al. 2010; Cress et al. 2017; Leonard et al. 2006b; Rodriguez et al. 2017; Trantas et al. 2009). In contrast, there are just exploratory reports concerning the microbial production of flavonoids with additional hydroxylations of ring A. Using cytochrome P450 enzymes from Nocardia farcinica, daidzein was converted into 6- or 8-hydroxydaidzein, albeit at low rates (Choi et al. 2009). The caffeic acid 3-hydroxylase, Sam5, from Saccharothrix espanaensis was expressed in E. coli and found to catalyze the 3′-, 6-, and 8-hydroxylation of select flavonoids, leading to production of 8-hydroxyluteolin and 3′-hydroxygenistein (Lee et al. 2014). Incubation with Rhodotorula glutinis strain KCh735 resulted in 8-hydroxylation of luteolin (LUT), apigenin (API), and 5,7-dihydroxyflavone chrysin and 6- and 8-hydroxylation of naringenin (NAR) (Sordon et al. 2016). The enzyme(s) catalyzing these conversions have not been isolated yet. Very recently, an efficient 6-hydroxylation of chrysin by human CYP1A1 expressed in yeast was reported (Williams et al. 2017).
In the past few years, we elucidated the biosynthesis of methoxylated flavonoids in trichomes of sweet basil (Ocimum basilicum). Flavones substituted at position 6 or both 6 and 8, and carrying up to four methyl moieties at positions 6, 7, 8, and 4′, occur in this tissue (Grayer et al. 1996; Grayer et al. 2001). We identified a set of regioselective FOMTs with distinctive properties and regiospecific flavonoid 6- and 8-hydroxylases (Berim and Gang 2016). To obtain commercially unavailable substrates for our biochemical investigation, we combined appropriate hydroxylase(s) and FOMTs and constructed five Saccharomyces cerevisiae strains that are designed to produce flavones found in basil when fed API as substrate. The flavones produced (and the corresponding strain designation) are the dimethylated cirsimaritin (CIRM) and ladanein (LAD), trimethylated salvigenin (SALV), and 8-substituted derivatives of SALV, 8-hydroxysalvigenin (8HS), and gardenin B (GB) (Fig. 1). Initially, different growth conditions and periods were compared using the SALV strain (designed to accumulate SALV as the major end product; the same nomenclature is used to refer to the other strains included in this investigation). We then probed the utility of these yeast strains for biodiversification by feeding the cells with three additional compounds representing different flavonoid subclasses. These experiments revealed that the constructed strains can be used to produce a number of rare compounds not reported to occur in basil and that the substrate specificity in both of the FOMTs and the flavonoid hydroxylases restricts the product yields.
a Steps of flavone biosynthesis in sweet basil used for flavonoid production by constructed yeast strains. Bold arrows indicate major pathways, thinner arrows indicate side reactions. Enzyme designations have been published previously: ObFOMT1–6 (Berim et al. 2012), flavone 6-hydroxylase CYP82D33 (Berim and Gang 2013b), ObF8OMT-1 (Berim and Gang 2013a), flavone 8-hydroxylase ObF8H-1 (Berim et al. 2014). b Structures of flavonoids offered as alternative substrates in this study. Flavonoid backbone nomenclature is shown on the structure of apigenin
Materials and methods
Chemicals
All general chemicals used were of analytical grade and obtained from common vendors (Fisher, VWR, Sigma). Flavonoids fed as precursors were from Indofine except for NAR which was from Sigma-Aldrich. Yeast nitrogen base without amino acids and appropriate amino acid mixes for the preparation of dropout media were purchased from United States Biological.
Construction of expression plasmids
pESC vector series (Agilent) were used for the expression of all plant genes, and S. cerevisiae strain INVSc-1 (Invitrogen) was used as expression host. A summary of all cloning details including all primer sequences, corresponding vectors, and strains where they are used, is shown in Supplementary Table S1. Vectors constructed previously for the functional expression of basil FOMTs in bacteria (Berim and Gang 2013a; Berim et al. 2012) were used as templates for producing yeast expression plasmids. The pESC-URA plasmid harboring basil flavonoid 6-hydroxylase has been published previously (Berim and Gang 2013b). As described earlier, all components of the basil flavonoid 8-hydroxylase system were expressed as fusions with yeast Yah1p mitochondrial transit peptide (Berim et al. 2014). All constructs were validated by direct sequencing.
Yeast growth and cultivation conditions
Seed cultures were grown for 16–22 h at 29 °C and 210 rpm (New Brunswick C25KC shaker) in 30 mL appropriate selection medium (SD minus histidine (HIS), tryptophan (TRP), URA for CIRM, LAD, and SALV strains and SD minus HIS, TRP, URA, LEU for strains 8HS and GB) with 2% glucose. The main cultures were 10 mL-aliquots of either appropriate selection or rich YP medium (10 g L−1 each yeast extract and peptone) either in 125-mL Erlenmeyer flasks or in 50 mL vertically positioned conical tubes, with 2% galactose in all cases, and were inoculated with 1 mL of seed culture (OD600 at 1:20 dilution = 0.5). All substrates were supplied at the time of main culture inoculation as 100 μL aliquots of 10 mM solutions in DMSO per culture vessel, with a final concentration of 100 μM. Main cultures were grown at 29 °C and 210 rpm. After removing the aliquots for analysis on days 2 and 4 of growth, 500 μL of 20% galactose solution was added to appropriate cultures. For the evaluation of alternative substrates, yeast cells were grown in rich medium and flasks, with a single harvest after 2 days of incubation with the fed precursor.
Cell densities were determined at 600 nm in 1:20 or 1:100 dilutions using a Lambda 35 spectrophotometer (PerkinElmer). Calculation of cell numbers was based on colony counts after plating serial dilutions of a culture with known cell density.
Extraction of fermentation products
Aliquots of 500 μL culture broth with cells were removed from the culture vessels. When necessary, cells were separated from the culture medium by centrifugation at 21,000×g for 1 min. The supernatants were then transferred to another plastic tube while the cell pellet was suspended in 500 μL water. All samples were acidified with 25 μL 6 N HCl and extracted twice with 550 μL ethyl acetate. Combined organic fractions were dried in a centrifugal vacuum concentrator. The residues were dissolved in 200 μL 50% aqueous methanol with 0.1% formic acid, containing 25 μM quercetagetin (Extrasynthese) or 7,8,3′,4′-tetrahydroxyflavone (Indofine) as internal standard.
Flavonoid analysis by LC-MS
A Synapt G2-S quadrupole-ion mobility spectrometry-time of flight mass spectrometer system (Waters) equipped with an Acquity UPLC system with a photodiode array detector was used for LC-MS analysis of extracts. Extracted metabolites were separated on an Acquity BEH C18 UPLC column (50 mm length, 2.1 mm diameter, particle size 1.7 μm) using acetonitrile with 0.1% formic acid as solvent B and water with 0.1% formic acid as solvent A at a flow rate of 400 μL min−1 and the following linear gradient extending over 14 min: 0 min, 3% B; 1.86 min, 5% B, 6.86 min: 35% B; 9.69 min, 100% B; 10.52 min, 100% B; 11.02 min, 3% B; and 14 min, 3% B. Mass spectra were collected in positive mode over a range of m/z 50–1000 with a scan time of 0.2 s. The capillary was set at 3 kV, the sampling cone at 40 V, the source at 120 °C, and the desolvation temperature at 250 °C. Cone gas and desolvation gas flows were 0 and 850 L h−1, respectively. The collision energy for MS/MS fragmentation was 30 V. Calibration accuracy cutoff was 1 ppm. Leucine enkephalin was used for post-acquisition mass (lock mass) correction. UV data were collected over a range of 210–500 nm. Quantification of products was carried out using the UV data and calibration curves for the corresponding fed substrate under the previously validated assumption that the extinction coefficient for band I does not change significantly with methylations/6-hydroxylation (Grayer et al. 1996). The wavelengths used correspond to band I UV maximum of individual substrates (335, 345, 290, and 366 nm for API, LUT, NAR, and kaempferol (KAEM), respectively). All concentrations were recalculated to milligram per liter of culture from the 10 mL batches. Quantitative analysis was conducted using the TargetLynx module of MassLynx v.4.1. Statistical analysis was performed using Statistica 13 (TIBCO) and SPSS Statistics 23 (IBM).
GenBank accession numbers
The GenBank nucleotide sequence accession numbers of the proteins used in this study are as follows: ObFOMT1, JQ653275; ObFOMT3, JQ653277; ObFOMT4, JQ653278; ObFOMT5, JQ653279; ObF8OMT-1, KC354402; basil flavonoid 6-hydroxylase ObF6H-1, JX162212; basil flavonoid 8-hydroxylase ObF8H-1: KJ765356; basil ferredoxin ObFdx, KJ765361; and basil ferredoxin NADPH-reductase ObFNR, KJ765362.
Results
Strains producing CIRM, LAD, SALV, 8HS, and GB from API
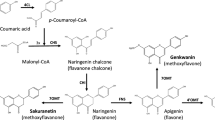
Combinations of previously characterized regioselective FOMTs with flavonoid ring A hydroxylases were used for strain construction. All strains harbored basil ObFOMT1, a flavonoid 7-O-methyltransferase (F7OMT) that very efficiently converts API into genkwanin (GENK; Fig. 1) (Berim et al. 2012). All strains also expressed the basil flavonoid 6-hydroxylase (F6H) encoded by CYP82D33 (ObF6H-1; Berim and Gang 2013b). It converts its preferred substrate GENK into scutellarein-7-methyl ether (S7Me; Fig. 1). The other components varied between strains.
The strain producing CIRM additionally harbored ObFOMT4, which catalyzes the 6-O-methylation of S7Me (Berim et al. 2012). After 2 days of growth under preferred conditions described below the titer of CIRM was 12.1 ± 1.7 mg L−1 (Supplementary Table S2). ObFOMT4 only exhibits very low flavonoid 4′-O-methyltransferase (F4′OMT) activity when incubated with appropriate substrates such as GENK or CIRM for an extended amount of time. Therefore, the formation of by-products apigenin-7,4′-dimethyl ether (AdM) and SALV is minimal in this strain (Fig. 2; Supplementary Fig. S1). One readily detectable by-product is GENK (Supplementary Fig. S1), the product of the initial API 7-O-methylation that has not been completely 6-hydroxylated.
Biofermentation products of apigenin with five constructed strains. UV335 traces of bioconversion products extracted from the whole cell suspension after 2 days of growth in shake flasks/rich medium. Strains are indicated in the upper right corner of each trace. IS internal standard quercetagetin. Other compound abbreviations are as in text. Vertical dotted lines connect the same compounds across strains
Instead of ObFOMT4, the strain-producing LAD expressed ObFOMT5, an enzyme whose preferred substrate is S7Me and which is strongly selective for the 4′-OH moiety as acceptor (Berim et al. 2012). The concentration of LAD was 6.2 ± 0.2 mg L−1 under applied growth conditions (Supplementary Table S2). ObFOMT5 is active with GENK and also shows some F6OMT activity with LAD. The accumulation of the by-product AdM (and a small signal for SALV) is therefore readily detectable at UV335 (Fig. 2; Supplementary Fig. S1).
The strain-producing trimethylated SALV expressed ObFOMT3 (Berim et al. 2012) as the only FOMT other than ObFOMT1. This enzyme shares 92% identity with ObFOMT5 and acts primarily as the F4′OMT of S7Me in planta. However, in contrast to ObFOMT5, ObFOMT3 has significant 6-O-methylating activity with LAD (17.06% relative to turnover with S7Me; Berim et al. 2012). By taking advantage of this bifunctional F6/4′OMT, we aimed to circumvent the need of using a separate F6OMT such as ObFOMT4. The concentration of SALV reached 5.0 ± 0.9 mg L−1 after 2 days of growth (Supplementary Table S2). As is the case with ObFOMT5, ObFOMT3 displays F4′OMT activity with GENK, leading to accumulation of AdM as by-product (Fig. 2; Supplementary Fig. S1).
The strains producing 8HS and GB were derived from the SALV strain. They both additionally harbored three enzymes necessary for the 8-hydroxylation of SALV. Basil flavone 8-hydroxylase (F8H) is a Rieske-type oxygenase that requires reduced ferredoxin (Fdx) as an electron donor (Berim et al. 2014). In turn, Fdx is reduced by Fdx-NADP+ reductase (FNR). Constructs encoding all three plant enzymes had to be introduced into yeast cells as the activity of ObF8H-1 in this host is extremely low in the absence of its native redox partners (Berim et al. 2014). The 8HS strain accumulated 8HS at a titer of 0.81 ± 0.04 mg L−1. In comparison to the 8HS strain, the strain producing GB additionally harbored the basil flavonoid 8-O-methyltransferase (F8OMT) designated ObF8OMT-1 (Berim and Gang 2013a). The yield of GB was 0.44 ± 0.12 mg L−1 (Supplementary Table S2). Both strains accumulated AdM and SALV as by-products (Fig. 2; Supplementary Fig. S1).
Characterization of flavone accumulation in the SALV strain
Prior to testing alternative substrates, we evaluated the effect of several basic growth parameters on product yields. These tests were conducted using the SALV strain. In an initial test, we compared four possible combinations of rich versus selection media and conical tubes (50 mL) versus unbaffled Erlenmeyer flasks (125 mL, with 10 mL medium in all conditions) in a bioconversion that lasted 48 h. Cells growing in rich medium and shake flasks (treatment designated RF) accumulated the highest amounts of the desired product SALV and moderate amounts of the major by-product AdM (Supplementary Fig. S2) as compared to the other three treatments designated SF (selection medium/flask), RT (rich medium/tube), and ST (selection medium/tube). The product specificity was estimated by the molar ratio between the desired 6-hydroxylated product SALV and the by-product AdM, and it was highest in SF cultures (Supplementary Fig. S2). For the subsequent tests, the cultivation time was extended up to 6 days, and RF growth conditions were compared to SF and RT while ST treatments were not continued based on data shown in Supplementary Fig. S2. To maintain the activity of the GAL promoters driving the expression of all heterologous genes (Hovland et al. 1989), all cells except for the cultures designated RF-G (grown in rich medium and flasks) were fed with galactose on days 2 and 4 of incubation. After 2 days of growth, RF and RF-G cultures, which were treated identically to this time point, accumulated the highest amounts of SALV after 2 days of growth (Fig. 3a). SALV abundance increased by about 30% to day 4 in RF cultures and did not increase significantly after an additional 48 h of incubation. SALV yield was significantly lower in SF and RT cultures in comparison to RF treatment. The accumulation of SALV increased significantly throughout the monitored time course in RT cultures grown in conical tubes (Fig. 3a). The main by-product, AdM, was accumulated at highest levels in RT cultures (Fig. 3b). In RF cultures, its abundance was highest on day 2 and decreased with longer incubation. The apparent consumption of AdM confirms the ability of ObF6H-1 to hydroxylate this flavone, as observed in vitro (Berim and Gang 2013b). Analysis of biosynthesis intermediates revealed that LAD and S7Me were most abundant in SF cultures (Supplementary Fig. S3a,b) while GENK was most abundant in RT cultures (Supplementary Fig. S3c).
Comparison of growth conditions using the SALV strain. The SALV strain was grown in rich medium and flask (RF), selection medium and flask (SF), or rich medium and conical tube (RT) for 6 days and supplemented with galactose on days 2 and 4 after inoculation except for treatment RF-G (grown in rich medium and shake flask). a Accumulation of SALV. b Accumulation of AdM. c Molar SALV/AdM ratios. d Cell numbers. Error bars represent standard deviation (n = 3). Different lowercase letters indicate the means differed significantly across treatments for the same time point. Different uppercase letters indicate the means differed significantly across time for the same treatment (two-way mixed ANOVA, p ≤ 0.05)
The SALV/AdM ratio was highest in SF cultures on day 2. However, it increased significantly with extended growth time in RF cultures (Fig. 3c). The lowest SALV/AdM ratios were measured in RT cultures. The periodic addition of the expression inducer galactose seems to support SALV production and improve the SALV/AdM ratio, as suggested by a comparison between RF and RF-G cultures (Fig. 3a–c).
The cell growth was not equal under the different treatments (Fig. 3d). It was significantly slower in RT cultures as compared to RF cultures, and the cell numbers increased steadily throughout the incubation. It is possible that the cell growth in tightly capped tubes is restricted by oxygen supply, and the amount of nutrients suffices to sustain the cell growth at a lower rate but over a longer period of time. Notably, overall “per cell” yields of SALV together with AdM were similar between RF and RT cultures (Supplementary Fig. S3d,e). The cell growth was also slow in SF cultures, and no increase in cell numbers was observed over the monitored time period. The selection medium might thus be depleted after 2 days of growth.
Localization of flavones during bioconversion of API
To analyze the localization of the flavones during biofermentation, yeast cells were separated from the spent medium by centrifugation, and the two fractions were extracted individually. SALV strain cells grown under RF, SF, and RT conditions were compared in this study. Approximately 90% (of the total amount extracted from whole cell suspension) of the main products SALV and AdM were found in yeast cells in RF cultures on day 2 after inoculation (Fig. 4). The fed substrate API also accumulated in yeast cells. Remarkably, the proportion of intracellularly accumulated AdM and SALV was significantly lower in cells of SF cultures. Analysis of pathway intermediates revealed that GENK was also strongly enriched in cells. By contrast, the relative abundances of LAD and S7Me in the medium were significantly higher than those of SALV, GENK, and AdM in RF cultures. LAD content in the medium amounted to 30–40% of its abundance in whole cell suspension; S7Me content ranged at 30–50%. Notably, the content of LAD and S7Me in cells was conspicuously low, ranging 22–60% of the total (and as a result, the relative abundances do not add up to 100% of the amount found in fermentation broth with cells; Fig. 4). Currently, we do not have an explanation for this observation, which has been made in a number of independent experiments. Overall, the high relative abundance of LAD and S7Me in spent medium suggests that partitioning and translocation of flavones involves certain selectivity, as both of these compounds accumulate at only low levels.
Distribution of flavones between cells and medium in the SALV strain. The abundances of flavones were calculated as percent of abundance in extracts from fermentation broth containing both cells and medium after 2 days of incubation with API. Error bars represent standard deviation (n = 3). Designations of culture conditions are RF rich, flask; SF selection, flask; and RT rich, tube. Different lowercase letters indicate the means differed significantly (comparison within the same fraction and treatment, one-way ANOVA, p < 0.05)
In a separate test, we analyzed the distribution of the main products in the SALV, CIRM, and LAD strains grown in parallel under RF conditions. The distribution of the fed substrate API was comparable across strains (overall 10–19% in the medium and 74–85% in cells). By contrast, CIRM partitioned nearly equally (40.5% medium/55.1% cells). As the main product of the LAD strain, LAD was more abundant in spent medium (60.2% medium/29.0% cells), consistently with its location when monitored as a low-abundance intermediate of SALV biosynthesis in the SALV strain (Fig. 4). As expected based on the above results, SALV was enriched in cells (16.6% medium/77.4% cells).
Biofermentation using flavanone NAR as fed substrate
NAR differs from API by having a saturated C2-C3 bond (Fig. 1). As a result, the molecule has a chiral carbon (C2). Natural flavanones typically have the S configuration at this position. The substrate that we fed to yeast cells was an unspecified mixture of the two enantiomers.
In the yeast fermentation system, all of the target dihydrogenated (abbreviated as dh) products, i.e., dhCIRM, dhLAD, dhSALV, dh8HS, and dhGB, were formed by their respective strains, but in differing amounts (Fig. 5; Supplementary Figs. S4 and S5; Supplementary Table S2). The accumulation of dhCIRM reached a final level of 1.85 ± 0.34 mg L−1. The titer of dhLAD reached 1.56 ± 0.54 mg L−1 in the LAD strain. In the SALV, 8HS and GB strains, residual dhLAD accumulated at levels ranging from 0.57 to 0.74 mg L−1 (Supplementary Fig. S5). The dihydro-derivative of SALV accumulated at 0.60 ± 0.07 mg L−1 in the SALV strain. Notably, its abundance in the 8HS and GB strains was only about 20% of the concentration in SALV strain, even though its downstream conversion to 8-hydroxylated products was quantitatively insignificant.
Biofermentation of naringenin by the constructed yeast strains. a UV290 chromatograms of extracts from whole cell suspensions. b Selected ion chromatograms of the same extracts from 8HS and GB strains. Maximum peak height indicated in the upper right corner. Ion monitored is indicated in the upper left corner. All compound abbreviations are as in text. Strains are indicated in the upper right corner. Vertical dotted lines connect the same metabolites across strains. IS internal standard quercetagetin. NdM elutes at 6.13 min, but the signal is compromised by co-eluting compounds in several traces and hence left unlabeled
No signals for the 8-hydroxylated derivatives of NAR were visible in UV290 traces. However, candidate peaks for both dh8HS and dhGB could be easily identified in selected ion chromatograms from the MS data (Fig. 5b). The accurate masses and isotope models matched the predictions for the specific compounds (Supplementary Fig. S4d,e), with only one plausible candidate signal being detected for each compound.
The first biosynthetic intermediate, 7-methylnaringenin (sakuranetin, SAK), was readily detectable in all five strains. Its level was lower in the CIRM strain and about equal in the other four strains (Supplementary Fig. S5). Remarkably, the flavanone analog of S7Me, carthamidin-7-methyl ether (C7Me), accumulated in high relative abundance (0.32–0.38 mg L−1) in all but the CIRM strain, where only traces of this intermediate were present (ca. 0.02 mg L−1). Naringenin-7,4′-dimethyl ether (NdM) also accumulated in all strains (with the CIRM strain only containing trace amounts; Supplementary Fig. S5).
We analyzed the distribution of NAR and its fermentation products in the SALV strain and found that over 80% of NAR was present in the culture media (Supplementary Table S3). By contrast, NdM and SAK were enriched in cells. Like their flavone analogs, C7Me and dhLAD were predominantly located in spent medium. Surprisingly, over 50% of dhSALV was also present in spent medium.
Biofermentation using 3′-hydroxylated flavone LUT as fed substrate
LUT differs from API by a hydroxyl residue at the 3′ position of the ring B (Fig. 1). Some methoxylated derivatives of LUT occur in O. basilicum (Grayer et al. 1996). Notably, these compounds are substituted at position 6, but not at position 8 of the backbone (Grayer et al. 2001).
The first pathway intermediate, 7-methylluteolin (L7Me), was readily detectable in UV345 traces of all five strains (Fig. 6; Supplementary Figs. S6 and S7). The 7,4′-dimethylated LUT also accumulated in considerable amounts in all but the CIRM strain. All five strains accumulated high amounts of pedalitin (PED), the 3′-hydroxylated derivative of S7Me (Fig. 6; Supplementary Figs. S6 and S7), suggesting that the downstream conversion of this intermediate is slower than its production. Its titers reached 2.76–4.08 mg L−1 in the CIRM, LAD, and SALV strains and were lower in the 8HS and GB strains. In the CIRM strain, accumulation of cirsiliol (CIRL, 3′-OH-CIRM) reached 1.74 ± 0.15 mg L−1. Nuchensin (NUCH), the 3′-hydroxylated analog of LAD, not only reached a final titer of 0.74 ± 0.04 mg L−1 in the LAD strain but was also incompletely converted in the SALV, 8HS, and GB strains, where it was present at 0.22–0.40 mg L−1. In the SALV strain, eupatorin (EUP, 3′-OH-SALV) accumulated at a final concentration of 0.40 ± 0.22 mg L−1. The concentration of EUP in the 8HS and GB strains was lower in comparison to the SALV strain even though the downstream conversion to 8-hydroxylated derivatives was negligible (Supplementary Fig. S7). Signals with accurate masses and isotope models matching those predicted for 3′-OH-8HS and 3′-OH-GB were detectable in selected ion chromatograms (Supplementary Fig. S8). While there was only one plausible candidate peak for 3′-OH-GB, two candidate 3′-OH-8HS signals with very similar retention times and accurate masses matching the expected formula were detected. As those signals were specific to this strain and substrate, it is possible that more than one product is formed. Further analysis is therefore required for unambiguous identity assignment.
Biofermentation of luteolin by the constructed yeast strains. UV345 chromatograms of extracts from whole cell suspensions. Strains are indicated in the upper right corner. Vertical lines connect the same metabolites across strains. IS internal standard quercetagetin. All other abbreviations are as in text
The distribution of LUT and its fermentation products was analyzed in the SALV strain. Approximately equal proportions of LUT were present in the cells and the culture media (Supplementary Table S3). Similar distribution was observed for L7Me and EUP. Luteolin-7,4′-dimethyl ether (LdM) was enriched in cells. Both PED and NUCH were enriched in the culture medium.
Biofermentation using flavonol KAEM as fed substrate
In comparison to flavones, flavonols carry an additional hydroxyl moiety at position 3 (Fig. 1). Flavonols have not been reported to accumulate in sweet basil’s trichomes. In our fermentation system utilizing KAEM as fed substrate, the two abundant and easily detectable products were kaempferol-7-methyl ether (K7Me), which was present in all strains and most abundant in the CIRM strain, and kaempferol-7,4′-dimethyl ether (KdM), which was present in considerable amounts in all but the CIRM strain (Supplementary Figs. S9 and S10). Evaluation using the MS data revealed that a signal with the accurate mass corresponding to that of hydroxylated CIRM was present in the CIRM strain (Fig. 7a, d, e). A signal with the same accurate mass but a different retention time (and thus likely representing 3-OH-LAD) was present in LAD strain (Fig. 7b, d, f). Its intensity was lower than that of putative 3-OH-CIRM. An even weaker signal with the accurate mass corresponding to 3-OH-SALV was present in the SALV strain (Fig. 7c, g, h). Under the applied cell growth and flavonoid analysis conditions, no promising signals with accurate masses corresponding to hydroxylated 8HS and GB were detected.
Chromatograms and mass spectra of 6-hydroxylated biofermentation products with kaempferol as fed substrate. a–c Selected ion chromatograms. The strains and height of largest peak are indicated in the upper right corner, while the ion is monitored in the left upper corner. d–h Predicted and observed isotope models and accurate masses of the peaks shown in a–c. Metabolites shown, formulae, and mass errors are indicated in the upper right corner
The localization of KAEM and its bioconversion products was analyzed in the SALV strain. Similarly to other mono- and dimethylated, 6-unsubstituted derivatives of fed substrates in our study, KdM and K7Me were enriched in cells (Supplementary Table S3). About 50% of KAEM found in extracts from whole cell suspension was recovered from yeast cells.
Discussion
Faster cell growth outweighs the benefits of selection medium
Previous work with baker’s yeast often used dropout media for the fermentation stage to avoid recombination or plasmid loss over time (Chemler et al. 2007; Eichenberger et al. 2017; Leonard et al. 2005; Wang et al. 2011; Yan et al. 2005). To the best of our knowledge, the only study directly comparing rich to selection media for flavonoid production was the analysis of NAR conversion into eriodictyol by a flavonoid 3′-hydroxylase by Amor et al. (2010). Cells grown in selection medium returned better yields in that study, with full consumption of NAR being reached within 30 h of incubation as compared to 71% conversion in rich medium. After this growth period, only 58% of cells grown without selection contained plasmid (Amor et al. 2010). Our results indicate that use of rich medium both affords higher overall flavonoid yield and is suitable for obtaining acceptable flavonoid composition, especially with extended cultivation time (Fig. 3a–c). Plasmid retention was not monitored, and it is to assume that plasmid loss occurs as reported previously (Amor et al. 2010; Jiang and Morgan 2004). However, the benefits of the higher cell growth rate seem to outweigh the reduction in biosynthesis rate due to plasmid loss. The comparison between the shake flasks and conical tubes as growth vessel showed that both the cell growth (Fig. 3d) and the performance of the oxygen-dependent ObF6H-1 are reduced in cells growing in tubes. The latter conclusion is based on high relative abundance of 6-unsubstituted by-products GENK and AdM in RT cultures of the SALV strain (Fig. 3b; Supplementary Fig. S3c). The use of conical tubes would enable an increase in throughput (replicates/conditions) in our laboratory setting which might be beneficial in future experiments. As limited aeration is likely to be the reason for the restricted growth and ObF6H-1 activity, modification of growth conditions, e.g., shaking under angle or use of air-permeable caps, could be tested as an approach to alleviating the observed drawbacks.
Remarkably, the levels of the intermediates S7Me and LAD, though overall low, were highest in selection medium (Supplementary Fig. S3a,b). In the constructed pathway, S7Me is the direct precursor of LAD, which is then converted into SALV by the same enzyme, ObFOMT3. The most obvious speculation would therefore be that this enzyme is not performing at sufficiently high levels in SF cultures. Corresponding expression data are necessary to help interpret the observations made at the metabolite level.
Analysis of flavonoid production over 6 days revealed that the yeast cells maintained biosynthetic activity during this time period (Fig. 3). This finding is in line with earlier studies where an increase of flavonoid concentration was still observed after 144–200 h (Koopman et al. 2012; Trantas et al. 2009) but overall hardly comparable because different growth conditions and inoculation densities were used.
While the results obtained here in a small-scale laboratory setting do not directly apply to engineering of flavonoid production on an industrial scale, they are encouraging for similar research endeavors and may help increase the yields and detectability of products.
Flavone production using API as fed precursor: enzymes perform well and in accordance with biochemical properties with their native substrate
The product profiles obtained with API as fed precursor largely reflected the in vitro biochemical properties of the employed enzymes. The small amounts of by-products observed with the CIRM strain are due to the regiospecificity of ObFOMT4, which is a highly specialized F6OMT and which therefore had only one substrate it could be active with (S7Me) (Berim et al. 2012). In the LAD, SALV, and derived strains, the permissive F4′OMTs, ObFOMT3, and ObFOMT5 could act upon an additional substrate (GENK) yielding the by-product AdM (Figs. 1 and 2). Notably, just as the plasticity of the F4′OMTs led to undesired side reactions, the relaxed substrate specificity of ObF6H-1 was beneficial for the product profile as it allowed for the 6-hydroxylation and thus partial redirection of the formed AdM in those strains.
The SALV strain and its derivatives (8HS and GB strains) were constructed using ObFOMT3 as a bifunctional F6/4′OMT. Based on substrate specificities (Berim et al. 2012), a better enzyme combination might comprise ObFOMT4 and ObFOMT3, where LAD formed from S7Me by ObFOMT3 would be 6-O-methylated by ObFOMT4, and CIRM formed by ObFOMT4 would be converted to SALV by ObFOMT3. At the time of strain construction, yield increase was not an objective; hence, this hypothesis has not been tested experimentally. On the one hand, the intermediate LAD does not accumulate in the SALV strain at high levels, suggesting that it is efficiently converted into SALV. On the other hand, the presence of two FOMTs consuming S7Me may create a pull effect on the ObF6H-1 and increase the overall flux into this branch of biosynthesis.
The 8-substituted flavones 8HS and GB only accumulated at very low levels. There are multiple possible reasons. The expression levels of ObF8H-1 and the redox partners FNR and Fdx may be insufficient. It is relevant to mention that the in vitro turnover rates of ObF8H-1 were rather low, yet these values may not accurately reflect its physiological properties as it is a membrane protein that had to be solubilized during purification (Berim et al. 2014). Secondly, the localization of ObF8H-1 in yeast mitochondria as compared to cytosolic FOMTs entails a necessity for the substrate SALV to be transported across compartments. Importantly, 8HS did not accumulate in the GB strain, suggesting that its export back into the cytosol is not limiting but leaving open the question of SALV import.
The rates of overall API conversion differed considerably across strains. For example, the CIRM strain produced 12 mg CIRM and over 0.5 mg GENK, while the SALV strain only yielded 5 mg SALV and 1.5 mg AdM (Supplementary Fig. S1; Supplementary Table S2). Changes in the system’s productivity upon component exchange or addition are not unusual and can have multiple reasons (Rodriguez et al. 2017; Santos et al. 2011). For a better evaluation, an assessment of the individual enzymes’ transcripts and/or peptides in different strains is necessary. To pinpoint the rate-limiting reactions in the different strains, it would be instructive to attempt balancing the flux (Jones et al. 2015; Santos et al. 2011), e.g., by fine-tuning the expression of the individual genes/enzymes. One prime candidate for overexpression is ObF6H-1, as a higher 6-hydroxylation rate would capture the nascent GENK and reduce the formation of AdM. Another important optimization point not addressed in this study and highly relevant and specific to the production of methylated flavonoids in a heterologous host is the availability of the endogenous co-substrate, S-adenosyl-l-methionine. For E. coli, its supply has been shown to be limiting for the efficiency of anthocyanidin methylation (Cress et al. 2017). Upon optimizing the conversions downstream of API, our system can be grafted within the yeast strains optimized for the production of NAR (Koopman et al. 2012; Rodriguez et al. 2017), which can be converted into API in a single enzymatic step.
Production of diverse compounds using alternative substrates is limited by the substrate specificities of both FOMTs and hydroxylases
One of the goals of the present study was to establish whether and how the enzymes isolated from basil act upon alternative substrates. Such precursor-directed biosynthesis is a successful approach for the biotechnological production of novel compounds (Chemler et al. 2007; Eudes et al. 2015), which critically depends on the plasticity of the employed enzymes. Basil FOMTs methylate a variety of flavonoids in vitro while showing strong preference for their physiological substrates (Berim et al. 2012). The basil F6H is active with several flavonoid classes including flavanones and flavonols (Berim and Gang 2013b). The basil F8H was only tested with basil flavones and exhibited significant activity only with SALV (Berim et al. 2014). Based on these biochemical studies, the strains would be able to produce at least some amounts of 6-hydroxylated, methoxylated derivatives of LUT, NAR, and KAEM.
As expected from data collected earlier in vitro, the accumulation of 7- and 7,4′-O-methylated derivatives was readily detectable with all fed precursors and in all relevant strains. ObFOMT1 is only moderately active with NAR in vitro (Berim et al. 2012). This appears to be relevant for the whole-cell setup as the overall flavonoid flux (estimated as the sum of product abundances) is considerably lower for NAR (Supplementary Fig. S5) than for API (Supplementary Fig. S1) and LUT (Supplementary Fig. S7) for each strain. KAEM was not previously evaluated but appears to be a good substrate for ObFOMT1 (Supplementary Fig. S9). With NAR and LUT as fed precursors, the abundance of the dimethylated by-products NdM and LdM is lower than that of the 7-monomethylated derivatives SAK and L7Me, respectively. This pattern is due to the low activity of basil F4′OMTs ObFOMT3/5 with NAR and LUT (Berim et al. 2012). The sensitivity of these FOMTs to the 3′-OH residue and the non-planar flavanone backbone also leads to accumulation of C7Me and PED. The high accumulation of 7-O-methylated derivatives with alternative substrates is also indicative of insufficient 6-hydroxylation rates by basil F6H. This high abundance of intermediates suggests that either pathway balancing or, more likely, alternative catalysts should be employed to increase the yields of the target products with precursors other than API. An alternative F7OMT could be SaOMT-2 from Streptomyces avermitilis with high relative activity with NAR, KAEM, and quercetin (Kim et al. 2006a). It would also be instructive to test the stereoselective NAR 7OMT from rice (Shimizu et al. 2012). Only one enantiomer of SAK would circulate in the cells, and its further conversion may reveal if the basil F6H and F4′OMTs also possess stereoselectivity. An alternative F4′OMT would be ShOMT2 from Solanum lycopersicum (Schmidt et al. 2011). This enzyme is highly active with quercetin-7-methyl ether and is thus a good candidate for conversion of the derivatives of both LUT and KAEM. SOMT-2 from soybean (Kim et al. 2005) may be tested for 4′-O-methylation of NAR derivatives. An alternative F6H might be CYP82D66 from peppermint, which possesses somewhat broader substrate specificity compared to ObF6H-1 (Berim and Gang 2013b). The F6H from soybean is most active with NAR but is sensitive to 7-O-methylation of the substrate (Latunde-Dada et al. 2001). For the choice of alternative catalysts, it is necessary to consider how they may alter the pathway intermediate landscape and fit into the existing catalytic modules. For example, the F6H from Scutellaria baicalensis was reported to be active with API (Zhao et al. 2018). However, the resulting scutellarein would be an unfavorable pathway intermediate as all of the employed basil FOMTs are only moderately active with this substrate.
The experiments presented here did not lead to production of significant amounts of 8-hydroxylated flavonoids other than 8HS and GB. Taking into account the low yields of 8HS and GB in cultures fed with API, tests with alternative precursors should be repeated after optimizing the performance of ObF8H-1. Alternative F8Hs are currently scarce. The F8H from S. baicalensis is very specific for chrysin (Zhao et al. 2018). The few other enzymes reported to catalyze flavonoid 8-hydroxylations are equally unlikely to fit the constructed strains and the pathway employed (Lee et al. 2014; Sordon et al. 2016).
The total yields with the same fed flavonoid can vary between strains, such as the outputs of the CIRM strain versus other strains with KAEM as substrate (Supplementary Fig. S10). As concluded above for the bioconversions with API, data regarding the abundance of individual transcripts and/or proteins are needed to begin to discuss this phenomenon.
Unequal distribution of products and intermediates poses numerous questions to be addressed in the future
Our study revealed unequal distribution of the flavonoid pathway intermediates and products in the various strains, with strong intracellular enrichment of a number of compounds (Fig. 4; Supplementary Table S3). S7Me and LAD as well as their dihydro- and 3′-OH analogs presented a notable deviation from the norm with their medium-localized fraction being substantially higher compared to other methylated flavonoids. Indeed, while it is too early to draw conclusions, our data suggest that the free 6-hydroxyl residue might be the structural feature shifting the equilibrium towards the extracellular enrichment. Data concerning flavonoid localization during fermentation are limited. NAR was enriched in medium both when produced by S. cerevisiae (Jiang et al. 2005; Koopman et al. 2012; Leonard et al. 2005) and when fed to S. cerevisiae, as were its glycosides and 2′-hydroxylated derivative (Brazier-Hicks and Edwards 2013; Werner and Morgan 2009). By contrast, pinocembrin (NAR lacking the 4′-OH residue) was stored in the yeast cells (Jiang et al. 2005). In E. coli, 25% of produced API and GENK (Leonard et al. 2006a) and 50% of anthocyanidins (Yan et al. 2008) were retained in cells. Over 95% of methylated flavonoids were retrieved from E. coli cells while glycosylated flavonoids were found in the spent medium (Willits et al. 2004). Notably, methylated and glycosylated anthocyanidin peonidin-3-O-glucoside appears to be secreted into the medium by E. coli (Cress et al. 2017). Unfortunately, studies aimed at production of several other flavonoids did not address product localization (Trantas et al. 2009). The method often used for product retrieval involves mixing the cell suspension with solvent and subsequent cell removal by centrifugation (Eichenberger et al. 2017; Koopman et al. 2012; Rodriguez et al. 2017). In our hands, this procedure retrieves both intra- and extracellular flavonoids and hence does not allow conclusions regarding their distribution.
Incorporating the distribution of involved compounds into the reaction scheme may improve our interpretation of the observed outcomes. The predominantly extracellular localization of the biosynthesis intermediates such as PED and C7Me poses the question of whether the rates of their export and re-import have an immediate effect upon the flux and the yields of downstream products. The high extracellular proportion of SALV and AdM in SF as compared to RF cultures might indicate that cell death and lysis, and consequently product release into medium, are occurring under SF conditions with limited nutrient supply. Alternatively, it is possible that translocation activities are, to some degree, medium- or cell density-dependent. To answer these questions, it is necessary to shed light on the mechanisms of the underlying translocation processes, which are currently entirely unknown. In general, secretion of xenobiotics by yeast is catalyzed by a family of multi-drug ATP-binding cassette (ABC) transporters (Ernst et al. 2010; Rogers et al. 2001). While the lack of pronounced specificity is an intrinsic feature of these proteins, their substrate spectra are not identical. In a study examining the effect of several flavonoids on yeast strains lacking three transporters, individually or in combinations, the ΔYOR1 mutant displayed the highest susceptibility to all tested compounds as compared to other mutants, suggesting that YOR1p is responsible for their export out of the cells (Rogers et al. 2001). In another study, the Pdr5p transporter was differentially inhibited by certain prenylated flavonoids (Conseil et al. 2000). The uptake of flavonoids by yeast also remains to be studied. Additionally, the localization of the substrate and the (by)products is relevant for the process optimization. On the one hand, extracellularly accumulated products can be collected with the spent medium during nutrient replenishment. On the other hand, intracellular accumulation of products and intermediates might pose an impediment for the application of a modular polyculture approach to our system. The use of co- and polycultures is one of the most promising developments in metabolic engineering (Jones et al. 2017; Zhang and Wang 2016), and it would be a very useful tool for optimization of the strains presented here. Given the nature of the proteins involved in the pathway, developing two-species consortia (e.g., baker’s yeast—E. coli) may be of benefit (Jia et al. 2016; Johns et al. 2016), as shown in a study where plant terpenes were the target products (Zhou et al. 2015). Whether a single- or a two-species consortium is to be employed, this approach depends on efficient translocation of appropriate intermediates between cells expressing different catalytic modules (Jones et al. 2017; Zhang and Wang 2016). There is good evidence that at least some flavonoids are continuously translocated between the medium and the cells. Despite the predominantly intracellular storage of methylated products, co-culture of E. coli cells expressing individual FOMTs led to successful production of di- and tri-methylated flavonoids (Willits et al. 2004). Overexpression of the E. coli transporter, yadH, increased the extracellular accumulation of the anthocyanin cyanidin-3-O-glucoside, whereas another efflux pump, tolC, was tentatively identified as being responsible for the secretion of the fed substrate catechin (Lim et al. 2015). Exogenously applied and predominantly extracellularly localized NAR is taken up and re-exported by S. cerevisiae as it is rapidly and, in some cases, completely converted by intracellular enzymes (Amor et al. 2010; Brazier-Hicks and Edwards 2013; Werner and Morgan 2009). An immediate way to test the polyculture suitability for the strains presented in this work will be to co-culture any of them with the yeast cells expressing just basil F6H (Berim and Gang 2013b) and evaluate the absolute or relative increase in the abundance of 6-substituted products. Moreover, identification and inclusion of relevant plant transporter proteins, e.g., from sweet basil peltate glandular trichomes, will enable incorporation of an additional layer of engineering effort that can be optimized for enhanced production of specific compounds or (re-)direction of flux through the flavonoid network in a fermentation system. Those transporter proteins are yet to be discovered, but once identified will play important roles in future engineering efforts.
References
Amor IL-B, Hehn A, Guedon E, Ghedira K, Engasser J-M, Chekir-Ghedrira L, Ghoul M (2010) Biotransformation of naringenin to eriodictyol by Saccharomyces cerevisiae functionally expressing flavonoid 3′ hydroxylase. Nat Prod Commun 5:1893–1898
Anarat-Cappillino G, Sattely ES (2014) The chemical logic of plant natural product biosynthesis. Curr Opin Plant Biol 19:51–58
Berim A, Gang DR (2013a) Characterization of two candidate flavone 8-O-methyltransferases suggests the existence of two potential routes to nevadensin in sweet basil. Phytochemistry 92:33–41
Berim A, Gang DR (2013b) The roles of a flavone 6-hydroxylase and 7-O-demethylation in the flavone biosynthetic network of sweet basil. J Biol Chem 288:1795–1805
Berim A, Gang DR (2016) Methoxylated flavones: occurrence, importance, biosynthesis. Phytochem Rev 15:363–390
Berim A, Hyatt DC, Gang DR (2012) A set of regioselective O-methyltransferases gives rise to the complex pattern of methoxylated flavones in sweet basil. Plant Physiol 160:1052–1069
Berim A, Park JJ, Gang DR (2014) Unexpected roles for ancient proteins: flavone 8-hydroxylase in sweet basil trichomes is a Rieske-type, PAO-family oxygenase. Plant J 80:385–395
Brazier-Hicks M, Edwards R (2013) Metabolic engineering of the flavone-C-glycoside pathway using polyprotein technology. Metab Eng 16:11–20
Chemler JA, Yan YJ, Leonard E, Koffas MAG (2007) Combinatorial mutasynthesis of flavonoid analogues from acrylic acids in microorganisms. Org Lett 9:1855–1858
Choi K-Y, T-j K, Koh S-K, Roh C-H, Pandey BP, Lee N, Kim B-G (2009) A-ring ortho-specific monohydroxylation of daidzein by cytochrome P450s of Nocardia farcinica IFM10152. Biotechnol J 4:1586–1595
Chouhan S, Sharma K, Zha J, Guleria S, Koffas MAG (2017) Recent advances in the recombinant biosynthesis of polyphenols. Front Microbiol 8
Conseil G, Decottignies A, Jault JM, Comte G, Barron D, Goffeau A, Di Pietro A (2000) Prenyl-flavonoids as potent inhibitors of the Pdr5p multidrug ABC transporter from Saccharomyces cerevisiae. Biochemistry 39:6910–6917
Cress BF, Leitz QD, Kim DC, Amore TD, Suzuki JY, Linhardt RJ, Koffas MAG (2017) CRISPRi-mediated metabolic engineering of E. coli for O-methylated anthocyanin production. Microb Cell Factories 16:10. https://doi.org/10.1186/s12934-016-0623-3
Eichenberger M, Lehka BJ, Folly C, Fischer D, Martens S, Simon E, Naesby M (2017) Metabolic engineering of Saccharomyces cerevisiae for de novo production of dihydrochalcones with known antioxidant, antidiabetic, and sweet tasting properties. Metab Eng 39:80–89
Ernst R, Kueppers P, Stindt J, Kuchler K, Schmitt L (2010) Multidrug efflux pumps: substrate selection in ATP-binding cassette multidrug efflux pumps—first come, first served? FEBS J 277:540–549
Eudes A, Benites VT, Wang G, Baidoo EEK, Lee TS, Keasling JD, Loque D (2015) Precursor-directed combinatorial biosynthesis of cinnamoyl, dihydrocinnamoyl, and benzoyl anthranilates in Saccharomyces cerevisiae. PLoS One 10:e0138972. https://doi.org/10.1371/journal.pone.0138972
Grayer RJ, Bryan SE, Veitch NC, Goldstone FJ, Paton A, Wollenweber E (1996) External flavones in sweet basil, Ocimum basilicum, and related taxa. Phytochemistry 43:1041–1047
Grayer RJ, Veitch NC, Kite GC, Price AM, Kokubun T (2001) Distribution of 8-oxygenated leaf-surface flavones in the genus Ocimum. Phytochemistry 56:559–567
Hovland P, Flick J, Johnston M, Sclafani RA (1989) Galactose as gratuitous inducer of GAL gene expression in yeasts growing on glucose. Gene 83:57–64
Jeon YM, Kim BG, Ahn JH (2009) Biological synthesis of 7-O-methyl apigenin from naringenin using Escherichia coli expressing two genes. J Microbiol Biotechnol 19:491–494
Jia X, Liu C, Song H, Ding M, Du J, Ma Q, Yuan Y (2016) Design, analysis and application of synthetic microbial consortia. Synth Syst Biotechnol 1:109–117
Jiang HX, Morgan JA (2004) Optimization of an in vivo plant P450 monooxygenase system in Saccharomyces cerevisiae. Biotechnol Bioeng 85:130–137
Jiang HX, Wood KV, Morgan JA (2005) Metabolic engineering of the phenylpropanoid pathway in Saccharomyces cerevisiae. Appl Environ Microbiol 71:2962–2969
Johns NI, Blazejewski T, Gomes ALC, Wang HH (2016) Principles for designing synthetic microbial communities. Curr Opin Microbiol 31:146–153
Jones JA, Toparlak OD, Koffas MAG (2015) Metabolic pathway balancing and its role in the production of biofuels and chemicals. Curr Opin Biotechnol 33:52–59
Jones JA, Vernacchio VR, Collins SM, Shirke AN, Xiu Y, Englaender JA, Cress BF, McCutcheon CC, Linhardt RJ, Gross RA, Koffas MAG (2017) Complete biosynthesis of anthocyanins using E. coli polycultures. MBio 8:e00621–e00617. https://doi.org/10.1128/mBio.00621-17
Kim DH, Kim BG, Lee Y, Ryu JY, Lim Y, Hur HG, Ahn JH (2005) Regiospecific methylation of naringenin to ponciretin by soybean O-methyltransferase expressed in Escherichia coli. J Biotechnol 119:155–162
Kim BG, Jung BR, Lee Y, Hur HG, Lim Y, Ahn JH (2006a) Regiospecific flavonoid 7-O-methylation with Streptomyces avermitilis O-methyltransferase expressed in Escherichia coli. J Agric Food Chem 54:823–828
Kim JH, Kim BG, Park Y, Han JH, Lim Y, Ahn J-H (2006b) Production of 5,7-dihydroxy, 3′,4′,5′-trimethoxyflavone from 5,7,3′,4′,5′-pentahydroxyflavone using two O-methyltransferases expressed in Escherichia coli. Agric Chem Biotechnol 49:114–116
Kim B-G, Lee YJ, Lee S, Lim Y, Cheong Y, Ahn J-H (2008) Altered regioselectivity of a poplar O-methyltransferase, POMT-7. J Biotechnol 138:107–111
Koirala N, Pandey RP, Parajuli P, Jung HJ, Sohng JK (2014) Methylation and subsequent glycosylation of 7,8-dihydroxyflavone. J Biotechnol 184:128–137
Koirala N, Thuan NH, Ghimire GP, Thang DV, Sohng JK (2016) Methylation of flavonoids: chemical structures, bioactivities, progress and perspectives for biotechnological production. Enzym Microb Technol 86:103–116
Koopman F, Beekwilder J, Crimi B, van Houwelingen A, Hall RD, Bosch D, van Maris AJA, Pronk JT, Daran J-M (2012) De novo production of the flavonoid naringenin in engineered Saccharomyces cerevisiae. Microb Cell Factories 11:155. https://doi.org/10.1186/1475-2859-11-155
Latunde-Dada AO, Cabello-Hurtado F, Czittrich N, Didierjean L, Schopfer C, Hertkorn N, Werck-Reichhart D, Ebel J (2001) Flavonoid 6-hydroxylase from soybean (Glycine max L.), a novel plant P-450 monooxygenase. J Biol Chem 276:1688–1695
Lee H, Kim BG, Ahn JH (2014) Production of bioactive hydroxyflavones by using monooxygenase from Saccharothrix espanaensis. J Biotechnol 176:11–17
Lee H, Kim BG, Kim M, Ahn JH (2015) Biosynthesis of two flavones, apigenin and genkwanin, in Escherichia coli. J Microbiol Biotechnol 25:1442–1448
Lee D, Park HL, Lee SW, Bhoo SH, Cho MH (2017) Biotechnological production of dimethoxyflavonoids using a fusion flavonoid O-methyltransferase possessing both 3′- and 7-O-methyltransferase activities. J Nat Prod 80:1467–1474
Leonard E, Yan YJ, Lim KH, Koffas MAG (2005) Investigation of two distinct flavone synthases for plant-specific flavone biosynthesis in Saccharomyces cerevisiae. Appl Environ Microbiol 71:8241–8248
Leonard E, Chemler J, Lim KH, Koffas MAG (2006a) Expression of a soluble flavone synthase allows the biosynthesis of phytoestrogen derivatives in Escherichia coli. Appl Microbiol Biotechnol 70:85–91
Leonard E, Yan YJ, Koffas MAG (2006b) Functional expression of a P450 flavonoid hydroxylase for the biosynthesis of plant-specific hydroxylated flavonols in Escherichia coli. Metab Eng 8:172–181
Lim CG, Wong L, Bhan N, Dvora H, Xu P, Venkiteswaran S, Koffas MAG (2015) Development of a recombinant Escherichia coli strain for overproduction of the plant pigment anthocyanin. Appl Environ Microbiol 81:6276–6284
Malla S, Koffas MAG, Kazlauskas RJ, Kim BG (2012) Production of 7-O-methyl aromadendrin, a medicinally valuable flavonoid, in Escherichia coli. Appl Environ Microbiol 78:684–694
Pandey RP, Parajuli P, Koffas MAG, Sohng JK (2016) Microbial production of natural and non-natural flavonoids: pathway engineering, directed evolution and systems/synthetic biology. Biotechnol Adv 34:634–662
Rodriguez A, Strucko T, Stahlhut SG, Kristensen M, Svenssen DK, Forster J, Nielsen J, Borodina I (2017) Metabolic engineering of yeast for fermentative production of flavonoids. Bioresour Technol 245:1645–1654
Rogers B, Decottignies A, Kolaczkowski M, Carvajal E, Balzi E, Goffeau A (2001) The pleitropic drug ABC transporters from Saccharomyces cerevisiae. J Mol Microbiol Biotechnol 3:207–214
Santos CNS, Koffas M, Stephanopoulos G (2011) Optimization of a heterologous pathway for the production of flavonoids from glucose. Metab Eng 13:392–400
Schmidt A, Li C, Shi F, Jones AD, Pichersky E (2011) Polymethylated myricetin in trichomes of the wild tomato species Solanum habrochaites and characterization of trichome-specific 3 '/5 '- and 7/4 '-myricetin O-methyltransferases. Plant Physiol 155:1999–2009
Shimizu T, Lin F, Hasegawa M, Okada K, Nojiri H, Yamane H (2012) Purification and identification of naringenin 7-O-methyltransferase, a key enzyme in biosynthesis of flavonoid phytoalexin sakuranetin in rice. J Biol Chem 287:19315–19325
Sordon S, Madej A, Poplonski J, Bartmanska A, Tronina T, Brzezowska E, Juszczyk P, Huszcza E (2016) Regioselective ortho-hydroxylations of flavonoids by yeast. J Agric Food Chem 64:5525–5530
Trantas E, Panopoulos N, Ververidis F (2009) Metabolic engineering of the complete pathway leading to heterologous biosynthesis of various flavonoids and stilbenoids in Saccharomyces cerevisiae. Metab Eng 11:355–366
Trantas EA, Koffas MAG, Xu P, Ververidis F (2015) When plants produce not enough or at all: metabolic engineering of flavonoids in microbial hosts. Front Plant Sci 6. https://doi.org/10.3389/fpls.2015.00007
Ververidis F, Trantas E, Douglas C, Vollmer G, Kretzschmar G, Panopoulos N (2007) Biotechnology of flavonoids and other phenylpropanoid-derived natural products. Part I: chemical diversity, impacts on plant biology and human health. Biotechnol J 2:1214–1234
Wang Y, Halls C, Zhang J, Matsuno M, Zhang Y, Yu O (2011) Stepwise increase of resveratrol biosynthesis in yeast Saccharomyces cerevisiae by metabolic engineering. Metab Eng 13:455–463
Werner SR, Morgan JA (2009) Expression of a Dianthus flavonoid glucosyltransferase in Saccharomyces cerevisiae for whole-cell biocatalysis. J Biotechnol 142:233–241
Williams IS, Chib S, Nuthakki VK, Gatchie L, Joshi P, Narkhede NA, Vishwakarma RA, Bharate SB, Saran S, Chaudhuri B (2017) Biotransformation of chrysin to baicalein: selective C6-hydroxylation of 5,7-dihydroxyflavone using whole yeast cells stably expressing human CYP1A1 enzyme. J Agric Food Chem 65:7440–7446
Willits MG, Giovanni M, Prata RTN, Kramer CM, De Luca V, Steffens JC, Graser G (2004) Bio-fermentation of modified flavonoids: an example of in vivo diversification of secondary metabolites. Phytochemistry 65:31–41
Yan YJ, Kohli A, Koffas MAG (2005) Biosynthesis of natural flavanones in Saccharomyces cerevisiae. Appl Environ Microbiol 71:5610–5613
Yan YJ, Li Z, Koffas MAG (2008) High-yield anthocyanin biosynthesis in engineered Escherichia coli. Biotechnol Bioeng 100:126–140
Zhang H, Wang X (2016) Modular co-culture engineering, a new approach for metabolic engineering. Metab Eng 37:114–121
Zhao Q, Cui M-Y, Levsh O, Yang D, Liu J, Li J, Hill L, Yang L, Hu Y, Weng J-K, Chen X-Y, Martin C (2018) Two CYP82D enzymes function as flavone hydroxylases in the biosynthesis of root-specific 4′-deoxyflavones in Scutellaria baicalensis. Mol Plant 11:135–148
Zhou K, Qiao KJ, Edgar S, Stephanopoulos G (2015) Distributing a metabolic pathway among a microbial consortium enhances production of natural products. Nat Biotechnol 33:377–383
Acknowledgements
AB would like to thank Dr. G. O. Berim (SUNY at Buffalo, NY) for help with statistical analysis.
Funding
This work was in part supported by the US Department of Energy Biological and Environmental Research Program (grant DE-SC0001728 to D.R.G.).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals
Electronic supplementary material
ESM 1
(PDF 948 kb)
Rights and permissions
About this article
Cite this article
Berim, A., Gang, D.R. Production of methoxylated flavonoids in yeast using ring A hydroxylases and flavonoid O-methyltransferases from sweet basil. Appl Microbiol Biotechnol 102, 5585–5598 (2018). https://doi.org/10.1007/s00253-018-9043-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-9043-0