Abstract
Domestic wastes, ranging from sewage and sludge to municipal solid waste, are usually treated in bioprocessing systems. These systems are regarded as main conduits for the elevated levels of antibiotic resistance genes (ARGs) observed in the environment. This paper mainly reviews recent studies on the occurrence and dynamics of ARGs in wastewater bio-treatment systems and discusses the ins and outs of ARG dissemination from the perspective of the microbial community. Our analysis shows that concentration of antibiotics through adsorption to microbial aggregates triggers the bacteria to acquire ARGs, which can be facilitated by the presence of mobile genetic elements. Notably, the acquisition and flow of ARGs during the rapid dissemination process is directed towards and for the best interests of the microbial community as a whole, and is influenced by surrounding nutrient levels, toxicant types, and sensitivities of the species in the prevailing antibiotic-stressed conditions. Furthermore, our review argues that predation of ARG-carrying bacteria by bacteriophages does periodically enhance the accessibility of ARGs to bacteria, which indirectly facilitates the recruitment of ARGs into environmental microbial communities.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The classic story of how Alexander Fleming discovered antibiotic-producing microorganisms tells us that nature itself harbors reservoirs of antibiotics. Previous studies have also shown that antibiotic resistance genes (ARGs) are ancient DNA fragments (D’Costa et al. 2011), predating the use of antibiotics by Homo sapiens by tens of thousands of years (Hardy et al. 2012). But notably, by analyzing soil samples spanning the last 70 years, environmental scientists have shown that the basal levels of ARGs are increasing in parallel with the modern massive production of antibiotics (Graham et al. 2016; Knapp et al. 2010). From the local to the transcontinental scale, the disposal of anthropogenic wastes is considered as a major factor fueling ARG dissemination (Pehrsson et al. 2016; Zhu et al. 2017).
The rapid anthropogenic dissemination of antimicrobial resistant bacteria (ARB) and ARGs across the world challenges the concept that antimicrobial resistance (AMR) is just a “natural feature of diverse microbial ecosystems” (Crofts et al. 2017; Wellington et al. 2013). This review aims to integrate studies addressing chemical (antibiotics/metals) selection pressures versus the cost-effect balance of acquiring ARGs (being resistant) from the perspective of bacteria and bacterial community. Our conclusions are expected to serve as starting points for the development of ecological approaches to reduce the spread of AMR issues in the environment.
Dissemination of ARGs is not random
General benefits and costs of being antibiotic resistant
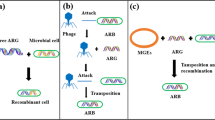
In natural ecosystems, antibiotics produced by intrinsically resistant bacteria (in pristine environments) are playing various functional roles, like signaling (mutual beneficial signals), aiding dispersal, and acting as toxicants (Ratcliff and Denison 2011), which give antibiotics-producing bacteria advantages over their competitors in the microbial community. Current studies generally presume that the genes for antibiotic resistance initially emerged in the wastes and body (e.g., intestinal microenvironments) under selective pressures of antibiotics-feeding animals or humans, or stem from intrinsic resistant bacteria (Pehrsson et al. 2016), whereas the subsequent dissemination of ARGs via horizontal gene transfer (HGT) is characterized with more randomness (Czekalski et al. 2014; Gaze et al. 2011). It seems plausible that ARGs or their fragments preserved in environments could be incidentally acquired by bacteria via HGT (Guo et al. 2017; Mao et al. 2014), and thereby facilitate the propagation of AMR. Previous studies have consistently pointed out that mobile genetic elements (MGEs) including integrons, insertion sequences, transposons, and plasmids are responsible for the “promiscuity” of ARGs (Yu et al. 2016; Zhu et al. 2017). According to the evolution theory of Darwin, the overall increasing levels of ARGs in anthropogenic wastes and their impacted environments, whether in farming lands (Su et al. 2015), wastewater treatment plants (Yang et al. 2014), municipal landfills (Sun et al. 2016), or contaminated river (Jiang et al. 2013; Stepanauskas et al. 2006), should also confer a benefit to the bacteria that have acquired ARGs.
It is recognized that HGT allows distantly related bacteria (different species) to transfer genes (Koonin et al. 2001). This evolutionary phenomenon usually takes place during a period of adaptation to new environmental conditions, where HGT requires extra energy to capture extracellular DNA fragments, replicate and maintain these genetic materials (Baltrus 2013). In a long-term, compared to conventional, bacterial reproduction (Fig. 1), microorganisms will experience additional detrimental side effects known as “metabolic burdens” that are induced by increases in gene expressions of the HGT-related DNA regions (Park and Zhang 2012). Therefore, if newly acquired ARGs are not fitted to an antimicrobial role in none antibiotic-stressed conditions (Fig. 1), the HGT-related costs will be net energy losses. In this case, bacteria that host ARGs will have comparatively less energy in reproduction. This could inherently decrease the proportion of ARGs in the gene pools of any type of environment (Bjorkman and Andersson 2000). It is common to observe the loss and decreasing abundance of HGT-related genes in environmental samples (Mao et al. 2014; Rysz et al. 2013). Thus, to unravel the puzzle why ARGs are massively sprawling from human to natural environments, we need to better understand how bacteria benefit from being antibiotic resistant.
HGT requires considerable extra energy (yellow stars) and increases metabolic burdens of the bacterial cells (bacteria in pink color). This will decrease the proportion of ARGs in environmental gene pools without the imposed stresses from antibiotics. The dark yellow circles represent the bacterial plasmids, and the red rods (Abs) represent antibiotics in surrounding environment
Impacts of discharge and bio-adsorption of antibiotics
The total amount of antibiotics consumed by humans increased by 46% between 2000 and 2010 (Van Boeckel et al. 2014), reaching an annual consumption of 15 g per capita (Zhang et al. 2008). Kümmerer et al. (2000) estimated that domestic sewer networks and sewage treatment plants receive 20–40% of all outdated medicaments. Although pharmaceuticals, classified as hazardous wastes (Environment Agency 2013), are not permitted to be disposed in sewer now, a wide range of pharmaceuticals is still found in anthropogenic wastes and waste treatment systems (Bound and Voulvoulis 2005). Studies show that 75% of antibiotics fed to animals (200,000 tons/year are used in livestock farming systems) and up to 50% of total amount prescribed to humans are discharged (e.g., urinary excretion) in an active form (Kemper 2008; Van Boeckel et al. 2014).
In general, the levels of antibiotics detected in domestic wastewater and solid wastes are at the nanogram per liter or per kilogram level (Marx et al. 2015; Miao et al. 2004; Sun et al. 2016; Wu et al. 2015; Zhou et al. 2013). This is one order of magnitude lower than the minimum selective concentration (MSC), which can cause inhibition (AMR mutations) on drug-hypersensitive strains (Andersson and Hughes, 2014). This seems to suggest that the acquisition of ARGs would not confer an advantage, being resistant to antibiotics, to the recipient bacteria (Fig. 1). However, conventional bio-treatment facilities utilizing densely agglomerated bacteria (active sludge flocs or biofilms) were not originally designed for the reduction of antibiotics (Marx et al. 2015). The observed reduction of most antibiotics in wastewater treatment effluent is more indicative of their “disappearance” via bio-adsorption than of them being biodegraded (Kümmerer 2008). This is because bacteria compete to rapidly adsorb organic matter from wastewater and then conduct a slower metabolic assimilation (Chua and Hua 1996). Consequently, a distinct gradient of organic matters (e.g., antibiotics) is formed around the radius of the sludge flocs (Fig. 2). The research by Guellil et al. (2001) shows that sludge’s biosorption rates of organic matters from range from 5 to 15 mgCODgTSS −1 min−1.
Active sludge aggregated in conventional bio-treatment systems can rapidly adsorb organic matters from wastewater. The high contents of antibiotics and MGEs formed on the periphery of bio-flocs promoted the ARGs dissemination across the whole bio-treatment systems. The dark yellow circles represent the bacterial plasmids, and the red rods (Abs) represent antibiotics
In terms of antibiotics, compounds that are featured with high octanolewater partition coefficient (K ow > 2.5), such as tetracyclines, macrolides (MLs), and fluoroquinolones (FQs), usually show high sorption potentials onto solids or sludge (Michael et al. 2013). From a wider perspective, a mass balance analysis conducted by Zhou et al. (2013) shows that on average 70% of the tetracycline entering a domestic wastewater treatment plant were redistributed to the biosolids (sludge) phase. Lindberg et al. (2006) pointed out that, for FQs (norfloxacin and ciprofloxacin), this value can increase to 80%. A higher adsorption rate in wastewater treatment systems can be promoted by increasing the sludge concentration, which is achieved through increasing the sludge retention time (SRT) in practice (Abegglen et al. 2009). Similarly, extending SRT from 3 to 40 days resulted in a 50% higher transfer of MLs and FQs to sludge (Li et al. 2013).
According to Mao et al. (2014), the addition of plasmids doubled the ARG-uptake rate from the extracellular matrix under an antibiotics-stressed condition (kanamycin; 20 mg/L), relative to the unstressed condition; however, under the same antibiotic-stressed condition, no ARGs transfer was observed when plasmids were not added. These observations suggest that a rapid ARG propagation through natural transformation from extracellular DNA to indigenous bacteria needs the (i) sufficient amount of MGEs and (ii) presence of antibiotics that confer selective pressures. As shown in Fig. 2, the prevailing bio-adsorption of antibiotics by sludge flocs provides these two prerequisites. This may imply that the widely disseminated ARGs, instead of being incidentally involved in HGT, do function in antibiotic resistance in wastewater bio-treatment systems. Luczkiewicz et al. (2010) reported that the abundances of tetracycline-resistant E. faecalis and penicillin-resistant E. coliforms increased by 10 and 15%, respectively, in activate sludge. Similarly, Szczepanowski et al. (2009) showed that bacteria in sludge from a domestic wastewater treatment plant exhibited a reduced susceptibility to antibiotics and meanwhile 140 clinically important ARGs, situated on MGEs, were enriched in the system.
Mediation of microbial community conditions
Bacterial communities in either natural environments or bio-treatment systems have the tendency to act as a unit, where each species contributes to the function of the whole consortium. From this perspective, the dissemination of ARGs needs to fit the best interests of individual bacteria as well as the whole microbial consortium. For example, a metagenomic analysis shows that ARGs are rarely associated with MGEs (Chen et al. 2016) in pristine Tibetan soil samples, which are not contaminated by antibiotics. Also, environmental resistome is observed to structure with composition of microbial community, which is commonly defined by surrounding carbon or secondary nutrient contents (Forsberg et al. 2014). These findings suggest that the dissemination of ARGs in those environments is not facilitated by random or unregulated associations with MGEs (Fig. 1).
Serving best interests
The currently identified mechanisms of antibiotic resistance are comprised of efflux pumps, target modification (resistance mutations that modify the target protein), inactivation of the antibiotic, and target bypass (e.g., impermeable cell membrane) or lack of target (Allen et al. 2010). Compared to other bacteria, the ARGs-carried ones appear to have a wider choice of substrates if the integrated ARGs allow them to consume antibiotics via their inactivation functions (Dantas et al. 2008). Nevertheless, it is important to note that antibiotics are not a good source of carbon and energy. For example, the presence of acetate can reduce the biodegradation rate of antibiotics (Drillia et al. 2005).
In the case of domestic wastes and treatment systems, the easily degradable organics are rarely depleted and therefore the microorganisms in these systems are reluctant to switch to antibiotics as substrate. Metagenomics analysis also shows that the most prevalent mechanism of antibiotic resistance is efflux pumps (Christgen et al. 2015; Yang et al. 2014), which keeps antibiotics out of the cell, instead of decomposing them. But notably, the dissemination of ARGs is coordinated by a whole microbial community. Cordero et al. (2012) and Nguyen et al. (2011) state that bacterial populations are not driven by gene-centric and “selfish” dynamics but represent socially “cohesive units” regarding antimicrobial resistance in common microhabitats. For example, a previous study has shown that, in an antibiotics-stressed domestic waste treatment system, denitrification by denitrifying bacteria proceeded smoothly thanks to the presence and activity of ARGs in the non-denitrifying party of the population (Wu et al. 2017). This means that not all bacteria affected by antibiotics need to and indeed will acquire ARGs, and become resistant in the environment. In mixed-species biofilms, Lee et al. (2014) has shown that the increased antibiotic resistance/tolerance was conferred via cross-protection (Fig. 3), which was offered by one resistant species (Pseudomonas protegens Pf-5) to all members in the microbial assembly; this study also found that the role of protector (resistant strains) can be switched to other species, under different cultivation (antibiotic types and nutrition levels) conditions, for the highest growth rate of biofilms. All these observations suggest that the aggregated bacteria are prone to serve the best interests of microbial communities, where the dissemination direction of ARGs is systematically oriented by antibiotic types, bacterial sensitivity, and the most needed functions for microbial community to expand (Fig. 3).
The importance of metals
Apart from antibiotics, (heavy) metals are considered to impose relatively strong selection pressure on surrounding bacteria (Hu et al. 2017). Metals have physico-chemical characteristics that promote the build-up of resistance-selective concentrations, e.g., bio-adsorption, complexation, and physical adsorption in the environment (Seiler and Berendonk 2012), more so than the other AMR selective agents (Czekalski et al. 2014). The links between metals and increasing tolerance to antibiotics have been studied extensively. Baker-Austin et al. (2006) systematically grouped the resistance mechanisms induced by metals into three classes. Briefly, they are (i) cross-selection resistance, where one gene encodes resistance to both antibiotics and metals; (ii) co-selection resistance, where ARGs and metal resistant genes (MRGs) encode resistances to antibiotics and metals respectively but are physically located closely (generally in MGEs, plasmids for example); and (iii) co-regulatory resistance, where genetic transcription links metals and antibiotic resistance together, observed as unselective, active efflux of inhibitory toxicants, regardless of exposure to either metals or antibiotics. Among these resistance mechanisms, the co-selection of antibiotic and metal resistance was most commonly observed in bio-treatment systems, probably due to the high abundances of MGEs (Di Cesare et al. 2016; Yu et al. 2016).
Li et al. (2017) have revealed that (i) ARGs and MRGs are genetically associated with MGEs (signal of co-selection) in all environmental samples and (ii) HGT of ARGs and MRGs does rarely result in same ARG-MRG couples being shared across different environments. These two findings suggest that co-selection of ARGs and MRGs is a pervasive mechanism in all types of environments on one hand, but also imply that environmental conditions, rather than MGEs, determine which ARG-MRG pair shall be coupled on the other. This is presumably because that, as a defensive function (similar to the AMR) whose acquisition costs extra energy and risks low reproduction rates of bacteria (Fig. 1), the metal resistance (mechanism) has evolved to benefit the resistance gene(s) carriers and/or the whole microbial community (Miao et al. 2015). Additionally, Li et al. (2017) have shown that the multidrug resistance type(s) are most frequently observed to co-occur with MRGs (25% of all samples). Metagenomic analyses have shown that efflux pumps, known as multi-resistance to metals and antibiotics, are the prevalent mechanism (50–80% of resistance types) in domestic waste impacted areas and bio-treatment systems (Chen et al. 2013; Christgen et al. 2015; Su et al. 2015). Thus, future studies aiming to contain the rapid dissemination of ARGs should pay attention to the distribution of multidrug resistance and co-conferring antibiotic resistances of MRGs.
What if bacteria are victims?
In the previous sections, we have discussed the dissemination of ARGs in domestic wastes and waste treatment systems from bacterial perspectives as beneficiaries that master the game-theory in gaining maximal energy-gains. The published studies have also shown that bacteria can actively compromise resistance-sensitive species for the interest of the whole community (Lee et al. 2014; Wu et al. 2017). However, bacteria are not the microbes at the end of multi-hierarchical “food chains.” They can be “consumed” by bacteriophages (or phages). Compared to other life forms, phages are the most abundant entities on earth (Frost et al. 2005). Their high multiplication rates (~ 1 × 1025 per second) and concomitant lysis rates of the bacterial hosts greatly facilitate the reproduction and release of genetic elements to other bacterial cells, namely, transduction (Canchaya et al. 2003).
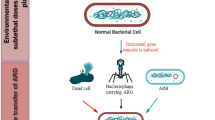
The genome size of phages ranges from a few to several 100 kb (Wu and Liu 2009), which is equivalent to the size of plasmids. This indicates that the phage-head is spacious enough to accommodate ARGs (Colomer-Lluch et al. 2011). During transduction, DNA fragments from an infected donor (host) can be accidentally loaded into the phage and then transferred to a recipient cell (Fig. 4). The aggregated biomass in waste and waste treatment systems is densely populated with phages (Colomer-Lluch et al. 2014; Withey et al. 2005). Moreover, according to Shapiro and Kushmaro (2011), some phages favor the introduction of antibiotics or metals in that toxicants may facilitate lysis of bacterial populations in domestic wastes. Rather than establishing a mutualistic relationship with bacteria, phages sometimes function as bacterial population manipulators (Rodriguez-Valera et al. 2009; Thingstad 2000). Phages are capable of attenuating the proliferation of some ARB strains in activated sludge (Yu et al. 2017). Presently, we have very limited knowledge of the benefits for phages by integrating ARGs or attacking ARB. But, phages were commonly observed to infect the dominant bacterial strains in the waste treatment systems (Shapiro et al. 2010), where bacteria could benefit from the acquisition of ARGs and then become major species, the “winners of microbial competition” (Fig. 1).
In bio-treatment systems, phages acting as population manipulators prefer to infect the dominant ARB (in the presence of antibiotics). The phage-fraction ARGs are more resistant to disinfection (e.g., UV and chlorination) than the bacteria-fraction. This facilitates the dissemination of ARGs in wastes receiving environments. Red rods represent antibiotics (Abs), and red and green segments represent two different ARGs
However, phages may not be used for removal of AMR genes from bio-treatment systems. Calero-Caceres and Muniesa (2016) have shown that ARGs (bla TEM, bla CTX-M and sul1) could persist longer when they were in phages than when in the bacterial fraction. Importantly, the commonly used disinfection technologies, including UV-irradiation and chlorination, have no significant effects on the inactivation of the phage-fraction ARGs (Brown-Jaque et al. 2015; Calero-Caceres and Muniesa 2016; Colomer-Lluch et al. 2014). This suggests that predation of ARG-carrying bacteria by phages increase the retention time of ARGs in waste treatment systems (Chen et al. 2016). Figure 4 shows that these ARG-containing phages will be released into the environment along with the treated wastewater (waste) being discharged. They might re-infect bacterial populations in receiving environments and transduct ARGs from phage head into the, possibly dominant (Thingstad 2000), bacteria carrying no ARGs. Therefore, the phage-mediated ARG transduction could also indirectly promote propagation of ARGs. To validate this assumption, future studies should focus on the direction of the flow of the phage-fraction ARGs, possibly via DNA isotopic or fluorescent labeling.
Conclusions
Domestic wastes are hotspots of antibiotics and ARGs. Their treatment systems conventionally utilizing dense microbial communities facilitate the dissemination of ARGs within these aggregates and exacerbate AMR issues in the environments receiving the wastes. Thus, to contain the propagation of ARGs, we need to consider the salient characteristics of ARGs from the perspective of the bacteria, and then determine the possibility to reduce ARGs’ contents. Our literature review underlines that dissemination of ARGs is not a random, rapid process, but requires the presence of selective pressure (metals and antibiotics) and mobile genetic elements (e.g., plasmids), the two prerequisites that are provided in the bio-treatment system due to bio-adsorption. Here, we propose a theory that the propagation of ARGs is mediated by the bacteria at the microbial community level: the community controls which species shall carry ARGs to counter the pressure of antibiotics, metals, or other toxicants, and thereby maximize its expansion rates and survival chances. Furthermore, we have conceptually analyzed potentials of AMR dissemination by bacterial phages, where a quantitative insight in the integration and release flux of the bacteriophage-fraction ARGs is yet to be explored.
References
Abegglen C, Joss A, McArdell CS, Fink G, Schlusener MP, Ternes TA, Siegrist H (2009) The fate of selected micropollutants in a single-house MBR. Water Res 43(7):2036–2046. 10.1016/j.watres.2009.02.005
Allen HK, Donato J, Wang HH, Cloud-Hansen KA, Davies J, Handelsman J (2010) Call of the wild: antibiotic resistance genes in natural environments. Nat Rev Microbiol 8(4):251–259. 10.1038/nrmicro2312
Baker-Austin C, Wright MS, Stepanauskas R, McArthur JV (2006) Co-selection of antibiotic and metal resistance. Trends Microbiol 14(4):176–182. 10.1016/j.tim.2006.02.006
Baltrus DA (2013) Exploring the costs of horizontal gene transfer. Trends Ecol Evol 28(8):489–495 doi:10.1016/j.tree.2013.04.002
Bjorkman J, Andersson DI (2000) The cost of antibiotic resistance from a bacterial perspective. Drug Resis Updat 3(4):237–245. 10.1054/drup.2000.0147
Bound JP, Voulvoulis N (2005) Household disposal of pharmaceuticals as a pathway for aquatic contamination in the United Kingdom. Environ Health Perspect 113(12):1705–1711. 10.1289/ehp.8315
Brown-Jaque M, Calero-Caceres W, Muniesa M (2015) Transfer of antibiotic-resistance genes via phage-related mobile elements. Plasmid 79:1–7. 10.1016/j.plasmid.2015.01.001
Calero-Caceres W, Muniesa M (2016) Persistence of naturally occurring antibiotic resistance genes in the bacteria and bacteriophage fractions of wastewater. Water Res 95:11–18. 10.1016/j.watres.2016.03.006
Canchaya C, Fournous G, Chibani-Chennoufi S, Dillmann M-L, Brüssow H (2003) Phage as agents of lateral gene transfer. Curr Opin Microbiol 6(4):417–424. 10.1016/s1369-5274(03)00086-9
Chen B, Yang Y, Liang X, Yu K, Zhang T, Li X (2013) Metagenomic profiles of antibiotic resistance genes (ARGs) between human impacted estuary and deep ocean sediments. Environ Sci Technol 47(22):12753–12760. 10.1021/es403818e
Chen B, Yuan K, Chen X, Yang Y, Zhang T, Wang Y, Luan T, Zou S, Li X (2016) Metagenomic analysis revealing antibiotic resistance genes (ARGs) and their genetic compartments in the Tibetan environment. Environ Sci Technol 50(13):6670–6679. 10.1021/acs.est.6b00619
Christgen B, Yang Y, Ahammad SZ, Li B, Rodriquez DC, Zhang T, Graham DW (2015) Metagenomics shows that low-energy anaerobic-aerobic treatment reactors reduce antibiotic resistance gene levels from domestic wastewater. Environ Sci Technol 49(4):2577–2584. 10.1021/es505521w
Chua H, Hua FL (1996) Effects of a heavy metal (zinc) on organic adsorption capacity and organic removal in activated sludge. Appl Biochem Biotechnol 57-8:845-849 doi:Doi 10.1007/Bf02941764
Colomer-Lluch M, Calero-Caceres W, Jebri S, Hmaied F, Muniesa M, Jofre J (2014) Antibiotic resistance genes in bacterial and bacteriophage fractions of Tunisian and Spanish wastewaters as markers to compare the antibiotic resistance patterns in each population. Environ Int 73:167–175. 10.1016/j.envint.2014.07.003
Colomer-Lluch M, Imamovic L, Jofre J, Muniesa M (2011) Bacteriophages carrying antibiotic resistance genes in fecal waste from cattle, pigs, and poultry. Antimicrob Agents Ch 55(10):4908–4911. 10.1128/AAC.00535-11
Cordero OX, Wildschutte H, Kirkup B, Proehl S, Ngo L, Hussain F, Le Roux F, Mincer T, Polz MF (2012) Ecological populations of bacteria act as socially cohesive units of antibiotic production and resistance. Science 337(6099):1228–1231. 10.1126/science.1219385
Crofts TS, Gasparrini AJ, Dantas G (2017) Next-generation approaches to understand and combat the antibiotic resistome. Nature Rev Microbiol 15(7):422–434. 10.1038/nrmicro.2017.28
Czekalski N, Gascon Diez E, Burgmann H (2014) Wastewater as a point source of antibiotic-resistance genes in the sediment of a freshwater lake. ISME J 8(7):1381–1390. 10.1038/ismej.2014.8
D'Costa VM, King CE, Kalan L, Morar M, Sung WW, Schwarz C, Froese D, Zazula G, Calmels F, Debruyne R, Golding GB, Poinar HN, Wright GD (2011) Antibiotic resistance is ancient. Nature 477(7365):457–461. 10.1038/nature10388
Dantas G, Sommer MOA, Oluwasegun RD, Church GM (2008) Bacteria subsisting on antibiotics. Science 320(5872):100–103. 10.1126/science.1155157
Di Cesare A, Eckert EM, D'Urso S, Bertoni R, Gillan DC, Wattiez R, Corno G (2016) Co-occurrence of integrase 1, antibiotic and heavy metal resistance genes in municipal wastewater treatment plants. Water Res 94:208–214. 10.1016/j.watres.2016.02.049
Drillia P, Dokianakis SN, Fountoulakis MS, Kornaros M, Stamatelatou K, Lyberatos G (2005) On the occasional biodegradation of pharmaceuticals in the activated sludge process: the example of the antibiotic sulfamethoxazole. J Hazard Mater 122(3):259–265. 10.1016/j.jhazmat.2005.03.009
Environment Agency (2013) Waste management 2013—England. UK
Forsberg KJ, Patel S, Gibson MK, Lauber CL, Knight R, Fierer N, Dantas G (2014) Bacterial phylogeny structures soil resistomes across habitats. Nature 509(7502):612–616. 10.1038/nature13377
Frost LS, Leplae R, Summers AO, Toussaint A (2005) Mobile genetic elements: the agents of open source evolution. Nat Rev Microbiol 3(9):722–732. 10.1038/nrmicro1235
Gaze WH, Zhang L, Abdouslam NA, Hawkey PM, Calvo-Bado L, Royle J, Brown H, Davis S, Kay P, Boxall AB, Wellington EM (2011) Impacts of anthropogenic activity on the ecology of class 1 integrons and integron-associated genes in the environment. ISME J 5(8):1253–1261. 10.1038/ismej.2011.15
Graham DW, Knapp CW, Christensen BT, McCluskey S, Dolfing J (2016) Appearance of beta-lactam resistance genes in agricultural soils and clinical isolates over the 20(th) century. Sci Rep 6(1):21550. 10.1038/srep21550
Guellil A, Thomas F, Block JC, Bersillon JL, Ginestet P (2001) Transfer of organic matter between wastewater and activated sludge flocs. Water Res 35(1):143-150 doi:Doi 10.1016/S0043-1354(00)00240-2
Guo J, Li J, Chen H, Bond PL, Yuan Z (2017) Metagenomic analysis reveals wastewater treatment plants as hotspots of antibiotic resistance genes and mobile genetic elements. Water Res 123:468–478. 10.1016/j.watres.2017.07.002
Hardy K, Buckley S, Collins MJ, Estalrrich A, Brothwell D, Copeland L, Garcia-Tabernero A, Garcia-Vargas S, de la Rasilla M, Lalueza-Fox C, Huguet R, Bastir M, Santamaria D, Madella M, Wilson J, Cortes AF, Rosas A (2012) Neanderthal medics? Evidence for food, cooking, and medicinal plants entrapped in dental calculus. Naturwissenschaften 99(8):617–626. 10.1007/s00114-012-0942-0
Hu HW, Wang JT, Li J, Shi XZ, Ma YB, Chen D, He JZ (2017) Long-term nickel contamination increases the occurrence of antibiotic resistance genes in agricultural soils. Environ Sci Technol 51(2):790–800. 10.1021/acs.est.6b03383
Jiang L, Hu X, Xu T, Zhang H, Sheng D, Yin D (2013) Prevalence of antibiotic resistance genes and their relationship with antibiotics in the Huangpu River and the drinking water sources, Shanghai, China. Sci Total Environ 458-460:267–272. 10.1016/j.scitotenv.2013.04.038
Kümmerer K (2008) Pharmaceuticals in the Environment: sources, fate, effects and risks, 3rd edn. Springer-Verlag Berlin, Heidelberg. 10.1007/978-3-540-74664-5
Kümmerer K, Al-Ahmad A, Mersch-Sundermann V (2000) Biodegradability of some antibiotics, elimination of the genotoxicity and affection of wastewater bacteria in a simple test. Chemosphere 40(7):701–710. 10.1016/S0045-6535(99)00439-7
Kemper N (2008) Veterinary antibiotics in the aquatic and terrestrial environment. Ecol Indic 8(1):1–13. 10.1016/j.ecolind.2007.06.002
Knapp CW, Dolfing J, Ehlert PA, Graham DW (2010) Evidence of increasing antibiotic resistance gene abundances in archived soils since 1940. Environ Sci Technol 44(2):580–587. 10.1021/es901221x
Koonin EV, Makarova KS, Aravind L (2001) Horizontal gene transfer in prokaryotes: quantification and classification. Annu Rev Microbiol 55(1):709–742. 10.1146/annurev.micro.55.1.709
Lee KW, Periasamy S, Mukherjee M, Xie C, Kjelleberg S, Rice SA (2014) Biofilm development and enhanced stress resistance of a model, mixed-species community biofilm. ISME J 8(4):894–907. 10.1038/ismej.2013.194
Li LG, Xia Y, Zhang T (2017) Co-occurrence of antibiotic and metal resistance genes revealed in complete genome collection. ISME J 11(3):651–662. 10.1038/ismej.2016.155
Li W, Shi Y, Gao L, Liu J, Cai Y (2013) Occurrence, distribution and potential affecting factors of antibiotics in sewage sludge of wastewater treatment plants in China. Sci Total Environ 445-446:306–313. 10.1016/j.scitotenv.2012.12.050
Lindberg RH, Olofsson U, Rendahl P, Johansson MI, Tysklind M, Andersson BAV (2006) Behavior of fluoroquinolones and trimethoprim during mechanical, chemical, and active sludge treatment of sewage water and digestion of sludge. Environ Sci Technol 40(3):1042–1048. 10.1021/es0516211
Luczkiewicz A, Jankowska K, Fudala-Ksiazek S, Olanczuk-Neyman K (2010) Antimicrobial resistance of fecal indicators in municipal wastewater treatment plant. Water Res 44(17):5089–5097. 10.1016/j.watres.2010.08.007
Mao D, Luo Y, Mathieu J, Wang Q, Feng L, Mu Q, Feng C, Alvarez PJ (2014) Persistence of extracellular DNA in river sediment facilitates antibiotic resistance gene propagation. Environ Sci Technol 48(1):71–78. 10.1021/es404280v
Marx C, Gunther N, Schubert S, Oertel R, Ahnert M, Krebs P, Kuehn V (2015) Mass flow of antibiotics in a wastewater treatment plant focusing on removal variations due to operational parameters. Sci Total Environ 538:779–788. 10.1016/j.scitotenv.2015.08.112
Miao XS, Bishay F, Chen M, Metcalfe CD (2004) Occurrence of antimicrobials in the final effluents of wastewater treatment plants in Canada. Environ Sci Technol 38(13):3533–3541. 10.1021/es030653q
Miao Y, Liao R, Zhang XX, Wang Y, Wang Z, Shi P, Liu B, Li A (2015) Metagenomic insights into Cr(VI) effect on microbial communities and functional genes of an expanded granular sludge bed reactor treating high-nitrate wastewater. Water Res 76:43–52. 10.1016/j.watres.2015.02.042
Michael I, Rizzo L, McArdell CS, Manaia CM, Merlin C, Schwartz T, Dagot C, Fatta-Kassinos D (2013) Urban wastewater treatment plants as hotspots for the release of antibiotics in the environment: a review. Water Res 47(3):957–995. 10.1016/j.watres.2012.11.027
Nguyen D, Joshi-Datar A, Lepine F, Bauerle E, Olakanmi O, Beer K, McKay G, Siehnel R, Schafhauser J, Wang Y, Britigan BE, Singh PK (2011) Active starvation responses mediate antibiotic tolerance in biofilms and nutrient-limited bacteria. Science 334(6058):982–986. 10.1126/science.1211037
Park C, Zhang J (2012) High expression hampers horizontal gene transfer. Genome biology and evolution 4(4):523–532. 10.1093/gbe/evs030
Pehrsson EC, Tsukayama P, Patel S, Mejia-Bautista M, Sosa-Soto G, Navarrete KM, Calderon M, Cabrera L, Hoyos-Arango W, Bertoli MT, Berg DE, Gilman RH, Dantas G (2016) Interconnected microbiomes and resistomes in low-income human habitats. Nature 533(7602):212–216. 10.1038/nature17672
Ratcliff WC, Denison RF (2011) Microbiology. Alternative actions for antibiotics. Science 332(6029):547–548. 10.1126/science.1205970
Rodriguez-Valera F, Martin-Cuadrado AB, Rodriguez-Brito B, Pasic L, Thingstad TF, Rohwer F, Mira A (2009) Explaining microbial population genomics through phage predation. Nat Rev Microbiol 7(11):828–836. 10.1038/nrmicro2235
Rysz M, Mansfield WR, Fortner JD, Alvarez PJ (2013) Tetracycline resistance gene maintenance under varying bacterial growth rate, substrate and oxygen availability, and tetracycline concentration. Environ Sci Technol 47(13):6995–7001. 10.1021/es3035329
Seiler C, Berendonk TU (2012) Heavy metal driven co-selection of antibiotic resistance in soil and water bodies impacted by agriculture and aquaculture. Frontiers Microbiol 3:399. 10.3389/fmicb.2012.00399
Shapiro OH, Kushmaro A (2011) Bacteriophage ecology in environmental biotechnology processes. Curr Opin Biotech 22(3):449–455. 10.1016/j.copbio.2011.01.012
Shapiro OH, Kushmaro A, Brenner A (2010) Bacteriophage predation regulates microbial abundance and diversity in a full-scale bioreactor treating industrial wastewater. ISME J 4(3):327–336. 10.1038/ismej.2009.118
Stepanauskas R, Glenn TC, Jagoe CH, Tuckfield RC, Lindell AH, King CJ, McArthur JV (2006) Coselection for microbial resistance to metals and antibiotics in freshwater microcosms. Environ Microbiol 8(9):1510–1514. 10.1111/j.1462-2920.2006.01091.x
Su JQ, Wei B, Ou-Yang WY, Huang FY, Zhao Y, HJ X, Zhu YG (2015) Antibiotic resistome and its association with bacterial communities during sewage sludge composting. Environ Sci Technol 49(12):7356–7363. 10.1021/acs.est.5b01012
Sun M, Ye M, Schwab AP, Li X, Wan J, Wei Z, Wu J, Friman VP, Liu K, Tian D, Liu M, Li H, Hu F, Jiang X (2016) Human migration activities drive the fluctuation of ARGs: case study of landfills in Nanjing, eastern China. J Hazard Mater 315:93–101. 10.1016/j.jhazmat.2016.04.077
Szczepanowski R, Linke B, Krahn I, Gartemann KH, Gutzkow T, Eichler W, Puhler A, Schluter A (2009) Detection of 140 clinically relevant antibiotic-resistance genes in the plasmid metagenome of wastewater treatment plant bacteria showing reduced susceptibility to selected antibiotics. Microbiology 155(Pt 7):2306–2319. 10.1099/mic.0.028233-0
Thingstad TF (2000) Elements of a theory for the mechanisms controlling abundance, diversity, and biogeochemical role of lytic bacterial viruses in aquatic systems. Limnol Oceanogr 45(6):1320–1328. 10.4319/lo.2000.45.6.1320
Van Boeckel TP, Gandra S, Ashok A, Caudron Q, Grenfell BT, Levin SA, Laxminarayan R (2014) Global antibiotic consumption 2000 to 2010: an analysis of national pharmaceutical sales data. Lancet Infect Dis 14(8):742–750. 10.1016/s1473-3099(14)70780-7
Wellington EMH, Boxall ABA, Cross P, Feil EJ, Gaze WH, Hawkey PM, Johnson-Rollings AS, Jones DL, Lee NM, Otten W, Thomas CM, Williams AP (2013) The role of the natural environment in the emergence of antibiotic resistance in gram-negative bacteria. Lancet Infect Dis 13(2):155–165. 10.1016/s1473-3099(12)70317-1
Withey S, Cartmell E, Avery LM, Stephenson T (2005) Bacteriophages—potential for application in wastewater treatment processes. Sci Total Environ 339(1–3):1–18. 10.1016/j.scitotenv.2004.09.021
Wu D, Chen G, Zhang X, Yang K, Xie B (2017) Change in microbial community in landfill refuse contaminated with antibiotics facilitates denitrification more than the increase in ARG over long-term. Sci Rep 7:41230. 10.1038/srep41230
Wu D, Huang Z, Yang K, Graham D, Xie B (2015) Relationships between antibiotics and antibiotic resistance gene levels in municipal solid waste leachates in Shanghai, China. Environ Sci Technol 49(7):4122–4128. 10.1021/es506081z
Wu Q, Liu WT (2009) Determination of virus abundance, diversity and distribution in a municipal wastewater treatment plant. Water Res 43(4):1101–1109. 10.1016/j.watres.2008.11.039
Yang Y, Li B, Zou S, Fang HH, Zhang T (2014) Fate of antibiotic resistance genes in sewage treatment plant revealed by metagenomic approach. Water Res 62:97–106. 10.1016/j.watres.2014.05.019
Yu P, Mathieu J, GW L, Gabiatti N, Alvarez PJ (2017) Control of antibiotic-resistant bacteria in activated sludge using polyvalent phages in conjunction with a production host. Environ Sci Technol Lett 4(4):137–142. 10.1021/acs.estlett.7b00045
Yu Z, He P, Shao L, Zhang H, Lu F (2016) Co-occurrence of mobile genetic elements and antibiotic resistance genes in municipal solid waste landfill leachates: a preliminary insight into the role of landfill age. Water Res 106:583–592. 10.1016/j.watres.2016.10.042
Zhang Y, Geissen SU, Gal C (2008) Carbamazepine and diclofenac: removal in wastewater treatment plants and occurrence in water bodies. Chemosphere 73(8):1151–1161. 10.1016/j.chemosphere.2008.07.086
Zhou LJ, Ying GG, Liu S, Zhao JL, Yang B, Chen ZF, Lai HJ (2013) Occurrence and fate of eleven classes of antibiotics in two typical wastewater treatment plants in South China. Sci Total Environ 452-453:365–376. 10.1016/j.scitotenv.2013.03.010
Zhu YG, Zhao Y, Li B, Huang CL, Zhang SY, Yu S, Chen YS, Zhang T, Gillings MR, JQ S (2017) Continental-scale pollution of estuaries with antibiotic resistance genes. Nat Microbiol 2:16270. 10.1038/nmicrobiol.2016.270
Acknowledgments
This work was supported by the Natural Science Foundation of China (21577038,31370510) and East China Normal University: Outstanding doctoral dissertation cultivation plan of action (PY2015034). J.D. acknowledges an international exchange grant from the Royal Society (Grant IE131283) for work on the treatment of landfill leachate. D.W. acknowledges the support from Distinguished Young PhD student Fellowship granted by Shanghai Tongji Gao-Tingyao Environmental Science & Technology Development Foundation (STGEF2017).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical statement
There is no ethical or legal conflict involved in this article.
Consent for publication
The manuscript has not been published elsewhere and all authors have seen the manuscript and approved to submit to your journal with mutual consent.
Rights and permissions
About this article
Cite this article
Wu, D., Dolfing, J. & Xie, B. Bacterial perspectives on the dissemination of antibiotic resistance genes in domestic wastewater bio-treatment systems: beneficiary to victim. Appl Microbiol Biotechnol 102, 597–604 (2018). https://doi.org/10.1007/s00253-017-8665-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-017-8665-y