Abstract
XynB from Aspergillus niger ATCC1015 (AnXynB) is a mesophilic glycoside hydrolase (GH) family 11 xylanase which holds great potentials in a wide variety of industrial applications. In the present study, the catalytic activity and stability of AnXynB were improved by a combination of computational and experimental approaches. Virtual mutation and molecular dynamics simulations indicated that the introduction of Glu and Asn altered the interaction network at the − 3 subsite. Interestingly, the double mutant S41N/T43E displayed 72% increase in catalytic activity when compared to the wild type (WT). In addition, it also showed a better thermostability than the WT enzyme. Kinetic determination of the T43E and S41N/T43E mutants suggested that the higher xylanase activity is probably due to the increasing binding affinity of enzyme and substrate. Consequently, the enzyme activity and thermostability of AnXynB was both increased by selective site-directed mutagenesis at the − 3 subsite of its active site architecture which provides a good example for a successfully engineered enzyme for potential industrial application. Moreover, the molecular evolution approach adopted in this study led to the design of a library of sequences that captures a meaningful functional diversity in a limited number of protein variants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Xylan is the most abundant hemicellulose compound in the plant cell wall and is composed of a β-1,4-glycosidic bond-linked xylose unit backbone chain that is usually decorated by glucuronic acid, arabinose, xylose, galactose and other groups (methyl, acetyl and feruloyl) (Glass et al. 2013; Scheller and Ulvskov 2010). Endo-β-1,4 xylanase (EC 3.2.1.8) is the key enzyme in the process of degrading xylan into short fragments via the random cleavage of xylan backbone chains (Collins et al. 2005). Xylanase has been widely applied in many industries, including biofuel, fibre and food technologies and pulp and paper industries (Polizeli et al. 2005; Viikari et al. 1994). These industrial treatments require commercial enzymes with high enzyme activity and stability, and protein engineering of enzyme is receiving increased attention.
Directed evolution is a powerful method that can be performed without sufficient knowledge of structure-function relationships. Through constructing random mutant libraries and screening for variants with improved properties, many proteins have been successfully engineered, but the success of this method is highly dependent on the size and quality of the variant library used. Unfortunately, complete coverage of sequence space (Mark et al. 2005; Povolotskaya and Kondrashov 2010) is often unfeasible, even for small proteins. By contrast, engineering proteins by rational design requires a good understanding of the determinants of protein function. Rational design has been proven successful for improving the catalytic activity (Cheng et al. 2015; Huang et al. 2014) and thermostability (Badieyan et al. 2012) of enzymes. However, although some impressive results have been achieved through rational design, identifying appropriate targets based on the enzyme structure and deciding on the design direction remain challenging.
Advances in genomics and structural biology have dramatically expanded our knowledge of protein sequence and structure (Dai et al. 2012; Warnecke et al. 2007). Structural bioinformatics are useful for utilizing this wealth of knowledge to explore structure-function relationships and provide new insight into protein rational design (Tripathi and Varadarajan 2014). Catalysis by glycoside hydrolases (GHs) is closely connected with the active site architecture, where the amino acid residues at different subsites perform enzymatic functions (Halabi et al. 2009; Himmel 2008). In a previous study, determinants of substrate specificity were identified by analysing the sequence profile of the active site architecture of different GH families (Tian et al. 2016). However, despite some progress, learning how to construct a “small but smart” library by utilizing the natural diversity of protein sequences and structures is a major goal.
GH11 xylanases typically adopt a conserved β-jelly-roll fold and degrade the xylan backbone chain randomly via a retaining mechanism (Rye and Withers 2000). The wide range of pH and temperatures tolerated by GH11 xylanases makes them applicable in industrial production (Paës et al. 2012). Thermomyces lanuginosus is the dominant fungus in plant biomass composts and has the ability to degrade hemicellulose during the composting process (Zhang et al. 2015a). TlXynA isolated from T. lanuginosus is a thermostable GH11 xylanase as reported previously (Singh et al. 2003). Aspergillus niger is one of the important industrial species whose growth temperature is 30 °C. AnXynB produced by A. niger as a mesophilic GH11 xylanase is widely used in industry (Andersen et al. 2011). In many previous studies, directed evolution approaches were widely applied to improve biomass-degrading ability (Song et al. 2012), thermostability (Dumon et al. 2008; Miyazaki et al. 2006) and alkaliphilicity (Inami et al. 2003) of a GH11 xylanase. Alternatively, rational design strategies need further practice and improvement. In this study, we investigated structure-function relationships in GH11 family enzymes and proposed a strategy for their engineering by coupling computational and experimental approaches. The feasibility of the strategy was validated using AnXynB as a case study, and both enzyme activity and thermostability were improved significantly by selective site-directed mutagenesis at the − 3 subsite of its active site architecture. Our approach may be generally applicable for future engineering of glucoside hydrolase.
Materials and methods
Bioinformatics analysis
All protein sequences used in the present study were obtained from the National Center for Biotechnology Information (NCBI, http://www.ncbi.nlm.nih.gov/). In total, 74 sequences of GH11 xylanases were selected to construct the dataset. Multiple sequence alignments were performed using CLUSTAL (Larkin et al. 2007). Sequence alignment images were produced using ESPript (Robert and Gouet 2014). The phylogenetic tree was constructed using MEGA (Kumar et al. 2016) and optimized by iTOL (Letunic and Bork 2016).
The TlXynA-substrate complex was constructed by docking the xylohexaose from TrXyn11A (PDB: 4HK8) into the active site cleft of TlXynA (PDB: 1YNA). The root-mean-square deviation (RMSD) between these two structures was 0.59 Å. Based on a cut-off of 5 Å, amino acid residues surrounding the substrate were selected to generate the reference using PyMOL (http://www.pymol.org). The structure of AnXynB was modelled by SWISS-MODEL (Arnold et al. 2006), and all other structures of GH11 xylanases were downloaded from the Protein Data Bank (PDB, http://www.rcsb.org). In total, 25 structures (Supplemental Table S1) were used to perform multiple structural alignments using PyMOL, and the sequence profile of the active site architecture of the whole GH11 family was created by WebLogo (Crooks et al. 2004). The accuracy of the sequence profile was tested using the ConSurf Sever (Ashkenazy et al. 2016).
Gene cloning, site-directed mutagenesis and protein expression
TlXynA (NCBI accession number: AAB94633) and AnXynB (NCBI accession number: ACA24724) were used for the research. A. niger (ATCC1015, American Type Culture Collection) was cultured at 30 °C in the minimal medium (5.95 g/L NaNO3, 0.522 g/L KCl, 1.497 g/L KH2PO4, 0.493 g/L MgSO4·7H2O, 5 g/L yeast extract, 2 g/L casamino acids) (Gong et al. 2016) with 1% xylose. Genomic DNA of A. niger ATCC1015 was used as the template for AnXynB gene cloning. The TlXynA gene was obtained by chemical synthesis (Genewiz, Suzhou, China). The sequences corresponding to the signal peptides of TlXynA and AnXynB were removed, and the remaining sequences were cloned into the pET28a plasmid (TransGen Biotech, Beijing, China) using NdeI and EcoRI restriction enzymes and T4 DNA ligase. AnXynB was used as a WT enzyme, and site-directed mutagenesis was performed using a PCR-based method (Weiner et al. 1994). The sequences of primers used for mutagenesis are listed in Supplemental Table S2. Recombinant plasmids were confirmed by DNA sequencing (Genewiz, Suzhou, China) and transformed into Escherichia coli BL21 (DE3).
E. coli BL21 (DE3) was cultured in LB medium supplemented with 50 μg/mL of kanamycin at 37 °C until the optical density at 600 nm reached 0.6–0.8. Isopropyl-β-D-thiogalactopyranoside of 0.5 mM (IPTG; Solarbio, Beijing, China) was added to the medium for induction, and the culture was incubated for another 20 h in a shaker at 20 °C. E. coli BL21 (DE3) was harvested by centrifugation and resuspended in lysis buffer (50 mM NaH2PO4, 300 mM NaCl, pH 8.0). After ultrasonic fragmentation, nickel column chromatography was used for protein purification. The eluent was replaced by PC buffer (20 mM sodium phosphate, 10 mM citrate, pH 6.0) by ultrafiltration (3-kDa cutoff membrane, Millipore, Billerica, MA) at 4 °C. The protein obtained above was analyzed by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE; Supplemental Fig. S1). Protein concentrations were determined by using the Bradford method (Bradford 1976).
Enzymatic activity assays and thermostability analysis
Xylanase activity was measured by incubating the purified enzymes (0.5 μg/mL) with 1% xylan at 50 °C for 10 min with occasional shaking. For analysis of the optimal temperature, 0.5 μg/mL enzymes was mixed with substrate and incubated for 10 min at different working temperatures. For thermostability analysis, protein samples were incubated at 60 °C for different time intervals of up to 120 min, and then cooled on ice for 5 min before activity measurement at 50 °C. The dinitrosalicylic acid (DNSA) method (Hu et al. 2009) was used to calculate enzyme activity by measuring the production of reducing sugars. Absorbance was read at a wavelength of 550 nm. One unit of xylanase activity corresponds to the amount of enzyme that releases 1 μmol of reducing sugar equivalent from xylose per minute.
Enzyme kinetics
To determine kinetic parameters for the different mutants, enzyme (100 μL, 0.5 μg/mL) was mixed with various concentrations of xylan solution (500 μL) for 3 min, and DNSA solution (400 μL) was added to terminate the reaction. Based on the one site binding equation, kinetic parameters including the turnover rate (k cat), Michaelis constant (K m) and catalytic efficiency (k cat/K m) were nonlinearly fitted using GraphPad Prism 5(Motulsky 2007).
Differential scanning calorimetry
The melting temperature (Tm) of enzymes was monitored using a VP-DSC MicroCalorimeter (MicroCal, Northampton, MA, USA) at a protein concentration of 0.02 mM and a scan rate of 1 °C/min. Protein samples were degassed and added into the 300 μL DSC sample cell. The reference cell was filled with buffer without enzyme. The initial temperature was 20 °C and the final temperature was 100 °C. The discrimination power was measured during the temperature increase.
Time-course analysis of reaction products
Fluorophore-assisted carbohydrate electrophoresis (FACE) was used to assess hydrolysis products of AnXynB and its mutants using xylotetraose (X4) and xylotriose (X3) as substrates. Firstly, substrate solutions (500 μg/mL) were mixed with an equal amount of protein (10 μg/mL) at 65 °C for up to 1 h as described previously (Zhang et al. 2015b) for 0, 4, 8, 15, 30 and 60 min. TIF images were obtained, and band intensity was quantified from peaks using Quantity One software (Bio-Rad Laboratories, Hercules, CA). Grey values for each band corresponded to the peak area value calculated from the peak volume (Gong et al. 2016).
Molecular dynamics simulations
Molecular dynamics simulations of WT AnXynB and its mutants (S41N, T43E and S41N/T43E) were carried out using Gromacs 4.5 as described previously (Jiang et al. 2016; Pronk et al. 2013). The number of contacts between enzyme and ligand were calculated based on the distance of enzyme and ligand. When the distance is less than 5 Å, a contact is formed between enzyme and ligand. In order to explore the influence of mutations on the dynamics of 54Trp, the collective motions of 54Trp in the WT and mutated enzymes were investigated using principal component analysis (PCA) (Jiang et al. 2017; Liu et al. 2014). Trajectory projections of 54Trp on PC1 and PC2 were used to describe its conformational ensembles in different systems. The van der Waals and electrostatic interaction energies between the enzyme and substrate were calculated according to the Amber force field equation (Cornell 1995).
Results
Determinants associated with the catalysis of GH11 xylanases
Protein sequence and structure determine protein function (Clark and Radivojac 2011; Lee et al. 2007). Using phylogenetic tree and active site architecture sequence profile analysis, the determinants associated with the catalysis of GH11 xylanases were investigated. As shown in Fig. 1, based on protein sequence similarity, GH11 xylanases were divided into two separate groups: Bacteria and Eukaryota. GH11 xylanases cannot be clustered in terms of their optimal temperatures. For example, the optimal temperatures of TlXynA and AnXynB were significantly different (Gong et al. 2016; Paës et al. 2012; Singh et al. 2003) despite their close evolutionary relationship. These results indicate that minor differences in protein sequence may mediate variation of the enzyme catalysis.
Phylogenetic tree of the GH11 family in which Bacteria are shown in blue and Eukaryota are shown in pink. The thermophilic TlXynA from T. lanuginosus and the mesophilic AnXynB from A.nigerare labeled withblack solid circles. The text part in the figure is in turn the genus (b, e), EC number, Uniprot ID, optimal pH and temperature separated each other by ‘_’
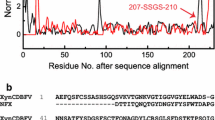
As shown in Supplemental Fig. S2, xylanases of the GH11 family adopt a highly conserved β-sandwich fold in which the active site is the location for xylan hydrolysis. The amino acid residues in the active site architecture were more conserved compared with those in other regions, consistent with the functional importance of the active site architecture. However, the conservation of amino acid residues was not equally distributed over the whole active site architecture (Fig. 2c). Specifically, amino acid residues forming the − 2, − 1 and + 1 subsites were highly conserved, while amino acid residues at distal subsites, and especially at the − 3 subsite, tended to be more variable, indicating that these various amino acid residues might contribute to differences in catalysis between different GH11 xylanases.
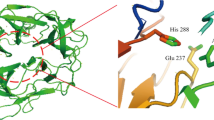
Sequence and structure alignment of TlXynA and AnXynB. a Sequence alignment of TlXynA and AnXynB. Strictly conserved residues are highlighted by a red background, and conservatively substituted residues are boxed. The secondary structure of TlXynA is shown above the aligned sequences. The green coloured number represents the location of the disulfide bond. b Structure alignment of TlXynA (salmon) and AnXynB (cyan). c Sequence profile of the active site architecture of GH11 family enzymes. The size of the letter and the ConSurf score indicates the degree of conservation at a given site. The location of ligand atoms interacting with the amino acid residues at each subsite is marked at the top, where CS stands for cleavage site
Differences in sequence, structure and function between TlXynA and AnXynB
To further investigate how variability at the − 3 subsite could modulate protein function, TlXynA and AnXynB were compared, since they are closely related but display different catalytic properties (Gong et al. 2016; Paës et al. 2012; Singh et al. 2003). The sequence similarity of TlXynA and AnXynB is 52.36%, and sequence differences are mainly concentrated in the loop regions (Fig. 2a), especially in the loop between two β-strands (residues 92–103 in TlXynA vs residues 127–138 in AnXynB) and in another loop between an α-helix and a β-strand (residues 161–171 in TlXynA vs residues 196–205 in AnXynB). The value of RMSD (0.715 Å) between these two structures suggests that TlXynA and AnXynB share similar structures (Chothia and Lesk 1986; Webb and Sali 2014). The catalytic cleft is approximately 30 Å long and contains a thumb structure (Fig. 2b). Among 23 amino acids defining the active site architecture, there are 3 amino acids which are different between TlXynA and AnXynB (Fig. 2c). The surface potential of the active site architecture is intimately related to substrate binding (Julián-Sánchez et al. 2013). As shown in Fig. 3, the surface potential at the − 3 subsite clearly differs between TlXynA and AnXynB. Notably, only two amino acid residues vary at this subsite; according to their predicted protein structures, 5Asn and 7Glu in TlXynA correspond with 41Ser and 43Thr in AnXynB (Fig. 2a). These amino acid residues would therefore be expected to give rise to differences in the surface potential at the − 3 subsite.
Comparison between TlXynA and AnXynB. Electrostatic potential distribution on the surface of TlXynA (a) and AnXynB (b). Electrostatic potential is coloured as a gradient from red (negative) to blue (positive). The local structure at the − 3 subsite is shown to highlight differences between TlXynA (c) and AnXynB (d)
In order to resist the detrimental effects of high temperatures, thermostable enzymes generally have a higher binding affinity for their substrates than mesophilic enzymes (Fields 2001). As shown in Fig. 4a, b, the melting temperature and optimal temperature for TlXynA were 74 and 65 °C, respectively, and the corresponding values for AnXynB were 51 and 47 °C. To verify this hypothesis, kinetic parameters were measured at different temperatures for these two enzymes.
The binding of enzyme and substrate can be a rate-limiting step for enzyme catalysis. As shown in Fig. 4c, the K m value of TlXynA and AnXynB altered minimally until the temperature was increased to the optimal temperature, and the binding affinity of the thermophilic TlXynA for xylan was twice of that of the mesophilic AnXynB. By contrast, the k cat values increased continuously with increasing temperature up to the optimal temperature (Fig. 4d). These results indicate that temperature modulates enzyme activity mainly by affecting the turnover rate (k cat) before enzyme unfolding occurs. Based on the above analysis, the higher binding affinity to xylan for TlXynA may be attributed to its highly polar residues (5Asn and 7Glu) at the − 3 subsite.
Computational analysis of the effects of mutations at the − 3 subsite
To explore the role of amino acid residues at the − 3 subsite in substrate binding, virtual mutants S41N and T43E of AnXynB were constructed. As shown in Fig. 5a, b, according to the result of virtual mutant T43E, one new hydrogen bond should be formed between 43Glu and xylose at the − 2 subsite, and the residue should also form another two new hydrogen bonds with 50Tyr and 52Ser, respectively. When 41Ser was mutated into 41Asn, a steric clash should occur between the side chain of 41Asn and the adjacent residue 54Trp. These results indicate that the mutation at the − 3 subsite altered the hydrogen bonding network within the active site architecture and affected the conformation of functional amino acid residues.
Virtual mutation analysis and molecular dynamics simulations. a, b Effect of S41N and T43E mutations on the local structure of AnXynB. Amino acids are shown in stick or sphere representation. Black dashed lines indicate potential hydrogen bonds. c Number of contacts between enzyme and substrate based on a cut-off distance of 5 Å. d Principal components analysis (PCA) showing conformational ensembles of 54Trp in different systems. PC1 and PC2 are the first two eigenvectors in the motion fluctuation of 54Trp
In order to verify the aforementioned prediction, molecular dynamics simulations were performed for WT AnXynB and its mutants (S41N, T43E and S41N/T43E). The number of contacts between enzyme and ligand was calculated and was ordered S41N/T43E > T43E > WT > S41N from greatest to fewest (Fig. 5c). It is worth mentioning that the average contact number of S41N/T43E and ligand was increased from 500 to 670 compared with that of WT. This result indicates differences in binding affinity to the substrate. We used PCA to investigate the conformation of the active site architecture of WT and enzyme variants S41N, T43E and S41N/T43E. One striking feature of enzymatic catalysis is that the catalytic elements in active site architecture are precisely positioned for their function (Benkovic and Hammes-Schiffer 2003). The conserved aromatic residue 54Trp at the − 2 subsite forms a stacking interaction with the xylose moiety to assist substrate binding (Cheng et al. 2014; Paës et al. 2012). As shown in Fig. 5d, trajectory projections of 54Trp on PC1 and PC2 differed between these enzymes. The mutations S41N and T43E only slightly changed the original conformation of 54Trp seen in the WT enzyme. The S41N/T43E mutation altered the conformation of 54Trp significantly which influenced the binding affinity of AnXynB. Many previous studies pointed out that the catalytic activity of enzymes could be affected by the change of the substrate binding affinity (Bernardi et al. 2014; Zhang et al. 2015c).
Experimental analysis of the effects of mutations at the − 3 subsite
The mutant genes have been synthesized and cloned. The encoded enzyme variants S41N, T43E and S41N/T43E were successfully heterologously expressed and purified. As shown in Fig. 6a, their enzyme activity differed considerably. Compared with WT AnXynB, the enzyme activity of the S41N mutant was slightly decreased, while that of the T43E variant was increased by 20%. Interestingly, the enzyme activity of S41N/T43E was increased by 72%. The specific activities of WT AnXynB and its mutants are shown in Supplemental Table S3. Compared to the other industrial xylanase TlXynA, the enzyme activity of S41N/T43E was increased by 40% (Supplemental Fig. S3). K m values for T43E and S41N/T43E were significantly lower (Table 1), indicating that the binding affinity was higher in these mutants, while S41N displayed a decrease in binding affinity. The turnover rate (k cat) of the three mutants was lower than the WT enzyme, and the T43E variant had the lowest k cat value. Finally, the catalytic efficiency (k cat/K m) of T43E and S41N/T43E mutants was increased, that of S41N was decreased compared to the WT enzyme, consistent with the enzyme activity results described above (Fig. 6a). Additionally, the interaction energy between enzyme and substrate was calculated by molecular dynamics simulation. The results showed that the T43E and S41N/T43E mutants had higher interaction energies than WT AnXynB (Supplemental Fig. S4).
Thermostability of AnXynB and its mutants
As shown in Fig. 6b, the residual enzyme activity of WT AnXynB and its S41N mutant was approximately 10% after incubation for 120 min at 60 °C. By contrast, the enzyme activity of T43E and S41N/T43E mutants decreased more slowly over time, and the residual enzyme activity of the S41N/T43E variant was 35%. In addition, the results of the DSC showed that the melting temperature of S41N/T43E is 3 °C higher than that of the WT (Supplemental Fig. S5). Combined with the virtual mutation results described above (Fig. 5), these results suggest that the increased thermostability might be related to the newly introduced hydrogen bonds within the active site architecture region. A previous study showed the importance of the N-terminus for the stability of proteins with a β-sandwich structure (Jiang et al. 2016), and the introduced hydrogen bond in the S41N/T43E variant is located in the N-terminal region.
Product profiling of AnXynB and its S41N/T43E variant
The enzymatic kinetics can be characterized by detecting the changes in concentration of hydrolysates over a time course using FACE (Gong et al. 2016). This method has been proved to be a simple, sensitive and relatively high-throughput technique (Kosik et al. 2012). Here, the product profiles of the WT AnXynB and its S41N/T43E variant were determined to provide insight into their mechanism of xylotetraose (X4) and xylotriose (X3) hydrolysis. These two enzymes on the degradation of X3 are very weak (Supplemental Fig. S6). As shown in Fig. 7, compared with WT AnXynB, the S41N/T43E mutant possessed a stronger ability to hydrolyze X4, which may contribute to the increased activity of this variant. There were more accumulations of X2 and X3 in the product profiles of S41N/T43E enzyme on xylotetraose. The xylose production (X1) which should be equivalent to the amount of X3 was very little. It is possible that X1 is unstable in the system. So, it suggested that the accumulation of X3 was due to the degradation of X4 by ‘X3 + X1’ mode and the accumulation of xylobiose (X2) was due to the degradation of X4 by ‘X2 + X2’ mode. The above experiments demonstrated that the double mutations increased the ability of the enzyme molecule to bind oligosaccharides.
Time courses of the product profiles of WT (a) and S41N/T43E (c) enzymes on xylotetraose. Time intervals are marked on the top of each lane, and abbreviations of sugars in the markers (M) are as follows; X1 = xylose, X2 = xylobiose, X3 = xylotriose, X4 = xylotetraose. b, d Changes in relative abundance of xylobiose, xylotriose and xylotetraose in hydrolysis products of WT and S41N/T43E, respectively, over time
Discussion
Catalytic activity is an essential feature of industrial enzymes. Active site architecture that makes direct contact with the ligand is an ideal region for the design of catalytic functions (Himmel 2008). The conservation of residues is often not equally distributed over the whole active site architecture. Highly conserved residues in the active site architecture are usually those that directly mediate substrate binding and catalysis, and these are generally similar in members of an enzyme family sharing identical structure and enzymatic mechanism (Paës et al. 2012; Tian et al. 2016). By contrast, more variable are amino acid residues that mediate the functional individuality of enzymes in the same family, and enzymatic properties can often be manipulated by engineering these relatively variable amino acid residues. For example, the mutation of the relatively variable amino acid 216Tyr improved the specific activity of mannanase (Huang et al. 2014). The enzyme activity of TrCel12A was also improved by engineering the relatively variable amino acid 7Trp at the − 4 subsite (Zhang et al. 2015c). In the GH11 family, there are at least two amino acids that differ at the − 3 subsite, but the screening workload of saturation mutagenesis at these positions (19 × 19 = 361 candidates) is laborious. Alternatively, the key steps could be the identification of suitable targets and choosing the appropriate design direction.
The diverse natural enzymes can adjust their sequences, local structures and even protein dynamics to fit their specific function (Henzler-Wildman and Kern 2007). On the basis of the molecular evolution, it becomes feasible to identify the variable amino acid residues which possibly enable the enzyme to catalyze in particular conditions such as high temperature and alkaline solution (Lee et al. 2007). It may be an effective method to create enzyme variants with novel catalytic characters by targeting and mutating these variable amino acid residues in the homologous enzymes. Here, a GH11 xylanase AnXynB was selected as our experimental model. Through probing into the phylogenetic relationship of the GH11 family, we chose the homologous TlXynA as reference model because AnXynB and TlXynA own highly similar sequences and structures but different catalytic properties (Gong et al. 2016; Paës et al. 2012; Singh et al. 2003). The variable amino acid residues in the active site that may affect the enzyme catalysis were found by performing sequence and structural alignments for these two xylanases. The influence of the mutations at these variable sites on enzyme catalysis can be explored by molecular dynamics simulations, which facilitates the rational design of enzymes. Finally, the mutations at the − 3 subsite in AnXynB significantly increased its enzyme activity, which demonstrates the effectiveness of the protocol used in our experiment. Notably, this protocol is also applicable to the protein engineering of other glycoside hydrolases. Of course, more experiments are required to further examine the strategy.
Rational design is already widely used in protein engineering, by which the desirable features can be introduced to target sites based on protein structural analyses. Although successful approaches for improving catalytic activity were reported (Cheng et al. 2015; Huang et al. 2014), the protein engineering strategy aiming at improving catalytic activity is still challenging. The protocols used for designing and screening targets are diverse due to differences in enzyme properties and the demands of different industrial applications. For example, when attempting to improve enzyme thermostability, protein unfolding is often taken into consideration (Zhang et al. 2014). Similarly, understanding the dissociation mechanism of functional amino acid residues in the active site architecture is necessary to design novel enzyme variants active over a wider pH range (Tishkov et al. 2013). Herein, we identified a specific set of sites likely to affect protein function by comparing two closely related GH11 xylanases that possess different enzymatic properties.
The S41N mutant decreased the enzyme activity of AnXynB, whereas the T43E mutant displayed a 20% increase in enzyme activity. Interestingly, the double mutant (S41N/T43E) increased the enzyme activity by 72%. Combining the results of the molecular dynamics simulation, evolutionary covariation (Hopf et al. 2017; Marks et al. 2012) of these two amino acid residues may exist in GH11 xylanases. The coevolved amino acid residues in specific sectors play an important structural and functional role in various protein families (Chakrabarti and Panchenko 2010). In the WT enzyme, Ser41 and Thr43 are located at the distal subsite of the active site and have no direct interaction with the substrate. The side chain of Glu43 in the mutant T43E may provide more hydrogen bonds because it contains one carbonyl group. Glu43 may make a direct hydrogen bond with O3 of the xylose unit at the − 2 subsite (Fig. 5b). The Asn41 and Glu43 residues in the double mutant S41N/T43E altered the conformation of 54Trp significantly which may influence the stacking interaction between the tryptophan and the − 2 sugar, resulting in the change of binding affinity of AnXynB. The increase of the binding affinity in the active site contributes to bind the oligosaccharides above the length of xylotriose, thereby increasing the catalytic efficiency of the enzyme. The results suggested that the interaction network of amino acids in the active site should be given attention to designing enzyme molecules. Compared with T43E, the S41N/T43Evariant has a similar binding affinity (K m) but higher product release efficiency (k cat), which partly demonstrates the importance of substrate binding and product release in enzyme catalysis. Many previous studies about improving the enzyme activity were creating new hydrogen bond or hydrophobic interactions between the enzyme and substrate or adding the non-catalytic carbohydrate binding module to increase the binding affinity of enzymes and substrat e(Thongekkaew et al. 2013; Zhang et al. 2013). But strongly binding to the catalytic pocket can inhibit the enzymatic activity such as by the higher binding energy between cellohexaose and Man5B (Bernardi et al. 2014). Payne et al. (2013) discovered that hydrolytic activity of GH7 cellulases could be enhanced by reducing the binding energy between enzyme and substrate. Therefore, higher catalytic efficiency requires a subtle balance between substrate binding and product release (Tian et al. 2016; Xie et al. 2014). Mutation at the − 3 subsite in AnXynB changed the distribution of binding energy over the whole active site architecture, resulting in an increased ability to bind large xylosaccharides. These results provide insight that will be useful for future glycoside hydrolase engineering.
In conclusion, we rationally engineered variants with higher catalytic activity and stability of the GH11 xylanase AnXynB by a combination of computational and experimental approaches. The results of virtual mutations and molecular dynamic simulations demonstrated that amino acid residues at the − 3 subsite play an important role in substrate binding. The site-directed mutagenesis at the − 3 subsite demonstrated that the binding energy of active site was improved, which accounted for the 72% increase in the catalytic activity of the double mutant S41NT43E. In addition, the thermostability of the enzyme variant S41NT43E was also improved due to the introduction of a polarity interaction in the N-terminus. Our approach provides an effective strategy for establishing a “small but smart” library of sequences that can guide rational design experiments.
References
Andersen MR, Salazar MP, Schaap PJ, van de Vondervoort PJ, Culley D, Thykaer J, Frisvad JC, Nielsen KF, Albang R, Albermann K, Berka RM, Braus GH, Braus-Stromeyer SA, Corrochano LM, Dai Z, van Dijck PW, Hofmann G, Lasure LL, Magnuson JK, Menke H, Meijer M, Meijer SL, Nielsen JB, Nielsen ML, van Ooyen AJ, Pel HJ, Poulsen L, Samson RA, Stam H, Tsang A, van den Brink JM, Atkins A, Aerts A, Shapiro H, Pangilinan J, Salamov A, Lou Y, Lindquist E, Lucas S, Grimwood J, Grigoriev IV, Kubicek CP, Martinez D, van Peij NN, Roubos JA, Nielsen J, Baker SE (2011) Comparative genomics of citric-acid-producing Aspergillus niger ATCC 1015 versus enzyme-producing CBS 513.88. Genome Res 21(6):885–897. https://doi.org/10.1101/gr.112169.110
Arnold K, Bordoli L, Kopp J, Schwede T (2006) The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 22(2):195–201. https://doi.org/10.1093/bioinformatics/bti770
Ashkenazy H, Abadi S, Martz E, Chay O, Mayrose I, Pupko T, Ben-Tal N (2016) ConSurf 2016: an improved methodology to estimate and visualize evolutionary conservation in macromolecules. Nucleic Acids Res 44(W1):W344–W350. https://doi.org/10.1093/nar/gkw408
Badieyan S, Bevan DR, Zhang C (2012) Study and design of stability in GH5 cellulases. Biotechnol Bioeng 109(1):31–44. https://doi.org/10.1002/bit.23280
Benkovic SJ, Hammes-Schiffer S (2003) A perspective on enzyme catalysis. Science 301(5637):1196–1202. https://doi.org/10.1126/science.1085515
Bernardi RC, Cann I, Schulten K (2014) Molecular dynamics study of enhanced Man5B enzymatic activity. Biotechnol Biofuels 7(1):83. https://doi.org/10.1186/1754-6834-7-83
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72(1–2):248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Chakrabarti S, Panchenko AR (2010) Structural and functional roles of coevolved sites in proteins. PLoS One 5(1):e8591. https://doi.org/10.1371/journal.pone.0008591
Cheng YS, Chen CC, Huang CH, Ko TP, Luo W, Huang JW, Liu JR, Guo RT (2014) Structural analysis of a glycoside hydrolase family 11 xylanase from Neocallimastix patriciarum: insights into the molecular basis of a thermophilic enzyme. J Biol Chem 289(16):11020–11028. https://doi.org/10.1074/jbc.M114.550905
Cheng YS, Chen CC, Huang JW, Ko TP, Huang Z, Guo RT (2015) Improving the catalytic performance of a GH11 xylanase by rational protein engineering. Appl Microbiol Biotechnol 99(22):9503–9510. https://doi.org/10.1007/s00253-015-6712-0
Chothia C, Lesk AM (1986) The relation between the divergence of sequence and structure in proteins. EMBO J 5(4):823–826
Clark WT, Radivojac P (2011) Analysis of protein function and its prediction from amino acid sequence. Proteins: Struct Funct Bioinf 79(7):2086–2096. https://doi.org/10.1002/prot.23029
Collins T, Gerday C, Feller G (2005) Xylanases, xylanase families and extremophilic xylanases. FEMS Microbiol Rev 29(1):3–23. https://doi.org/10.1016/j.femsre.2004.06.005
Cornell WD (1995) A second generation force field for the simulation of proteins, nucleic acids, and organic molecules. J Am Chem Soc 117:5179–5197
Crooks GE, Hon G, Chandonia JM, Brenner SE (2004) WebLogo: a sequence logo generator. Genome Res 14(6):1188–1190. https://doi.org/10.1101/gr.849004
Dai X, Zhu Y, Luo Y, Song L, Liu D, Liu L, Chen F, Wang M, Li J, Zeng X, Dong Z, Hu S, Li L, Xu J, Huang L, Dong X (2012) Metagenomic insights into the fibrolytic microbiome in yak rumen. PLoS One 7(7):e40430. https://doi.org/10.1371/journal.pone.0040430
Dumon C, Varvak A, Wall MA, Flint JE, Lewis RJ, Lakey JH, Morland C, Luginbuhl P, Healey S, Todaro T, De Santis G, Sun M, Parra-Gessert L, Tan X, Weiner DP, Gilbert HJ (2008) Engineering hyperthermostability into a GH11 xylanase is mediated by subtle changes to protein structure. J Biol Chem 283(33):22557–22564. https://doi.org/10.1074/jbc.M800936200
Fields PA (2001) Review: protein function at thermal extremes: balancing stability and flexibility. Comp Biochem Physiol Part A: Mol Integr Physiol 129(2):417–431. https://doi.org/10.1016/S1095-6433(00)00359-7
Glass NL, Schmoll M, Cate JHD, Coradetti S (2013) Plant cell wall deconstruction by ascomycete fungi. Annu Rev Microbiol 67:477–498. https://doi.org/10.1146/annurev-micro-092611-150044
Gong W, Zhang H, Tian L, Liu S, Wu X, Li F, Wang L (2016) Determination of the modes of action and synergies of xylanases by analysis of xylooligosaccharide profiles over time using fluorescence-assisted carbohydrate electrophoresis. Electrophoresis 37(12):1640–1650. https://doi.org/10.1002/elps.201600041
Halabi N, Rivoire O, Leibler S, Ranganathan R (2009) Protein sectors: evolutionary units of three-dimensional structure. Cell 138(4):774–786. https://doi.org/10.1016/j.cell.2009.07.038
Henzler-Wildman K, Kern D (2007) Dynamic personalities of proteins. Nature 450(7172):964–972. https://doi.org/10.1038/nature06522
Himmel ME (2008) Biomass recalcitrance: deconstructing the plant cell wall for bioenergy. Wiley-Blackwell, Oxford
Hopf TA, Ingraham JB, Poelwijk FJ, Schärfe CP, Springer M, Sander C, Marks DS (2017) Mutation effects predicted from sequence co-variation. Nat Biotechnol 35(2):128–135. https://doi.org/10.1038/nbt.3769
Hu G, Heitmann JA, Rojas OJ (2009) Quantification of cellulase activity using the quartz crystal microbalance technique. Anal Chem 81(5):1872–1880. https://doi.org/10.1021/ac802318t
Huang JW, Chen CC, Huang CH, Huang TY, Wu TH, Cheng YS, Ko TP, Lin CY, Liu JR, Guo RT (2014) Improving the specific activity of β-mannanase from Aspergillus niger BK01 by structure-based rational design. Biochim Biophys Acta 1844(3):663–669. https://doi.org/10.1016/j.bbapap.2014.01.011
Inami M, Morokuma C, Sugio A, Tamanoi H, Yatsunami R, Nakamura S (2003) Directed evolution of xylanase J from alkaliphilic Bacillus sp. strain 41M-1: restore of alkaliphily of a mutant with an acidic pH optimum. Nucleic Acids Symp Ser (Oxf) 3(1):315–316. https://doi.org/10.1093/nass/3.1.315
Jiang X, Chen G, Wang L (2016) Structural and dynamic evolution of the amphipathic N-terminus diversifies enzyme thermostability in the glycoside hydrolase family 12. Phys Chem Chem Phys 18(31):21340–21350. https://doi.org/10.1039/C6CP02998A
Jiang X, Li W, Chen G, Wang L (2017) Dynamic perturbation of the active site determines reversible thermal inactivation in glycoside hydrolase family 12. J Chem Inf Model 57(2):288–297. https://doi.org/10.1021/acs.jcim.6b00692
Julián-Sánchez A, González-Segura L, Riveros-Rosas H, Díaz-Sánchez ÁG, Muñoz-Clares RA (2013) Electrostatic potential surface in the aldehyde binding site of aldehyde dehydrogenases (ALDHs) is a determinant for substrate specificity. FASEB J 27:lb60
Kosik O, Bromley JR, Busse-Wicher M, Zhang Z, Dupree P (2012) Studies of enzymatic cleavage of cellulose using polysaccharide analysis by carbohydrate gel electrophoresis (PACE). Methods Enzymol 510:51–67. https://doi.org/10.1016/B978-0-112-415931-0.0004-5
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874. https://doi.org/10.1093/molbev/msw054
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23(21):2947–2948. https://doi.org/10.1093/bioinformatics/btm404
Lee D, Redfern O, Orengo C (2007) Predicting protein function from sequence and structure. Nat Rev Mol Cell Biol 8(12):995–1005
Letunic I, Bork P (2016) Interactive tree of life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res 44(W1):W242–W245. https://doi.org/10.1093/nar/gkw290
Liu M, Wang L, Sun X, Zhao X (2014) Investigating the impact of Asp181 point mutations on interactions between PTP1B and phosphotyrosine substrate. Sci Rep 4:5095. https://doi.org/10.1038/srep05095
Mark AD, Weinreich DM, Hartl DL (2005) Missense meanderings in sequence space: a biophysical view of protein evolution. Nat Rev Genet 6(9):678–687
Marks DS, Hopf TA, Sander C (2012) Protein structure prediction from sequence variation. Nat Biotechnol 30(11):1072–1080. https://doi.org/10.1038/nbt.2419
Miyazaki K, Takenouchi M, Kondo H, Noro N, Suzuki M, Tsuda S (2006) Thermal stabilization of Bacillus subtilis Family-11 xylanase by directed evolution. J Biol Chem 281(15):10236–10242. https://doi.org/10.1074/jbc.M511948200
Motulsky H (2007) Prism 5 statistics guide, 2007. GraphPad Software 31:39–42
Paës G, Berrin JG, Beaugrand J (2012) GH11 xylanases: structure/function/properties relationships and applications. Biotechnol Adv 30(3):564–592. https://doi.org/10.1016/j.biotechadv.2011.10.003
Payne CM, Jiang W, Shirts MR, Himmel ME, Crowley MF, Beckham GT (2013) Glycoside hydrolaseprocessivity is directly related to oligosaccharide binding free energy. J Am Chem Soc 135(50):18831–18839. https://doi.org/10.1021/ja407287f
Polizeli ML, Rizzatti AC, Monti R, Terenzi HF, Jorge JA, Amorim DS (2005) Xylanases from fungi: properties and industrial applications. Appl Microbiol Biotechnol 67(5):577–591. https://doi.org/10.1007/s00253-005-1904-7
Povolotskaya IS, Kondrashov FA (2010) Sequence space and the ongoing expansion of the protein universe. Nature 465(7300):922–926. https://doi.org/10.1038/nature09105
Pronk S, Páll S, Schulz R, Larsson P, Bjelkmar P, Apostolov R, Shirts MR, Smith JC, Kasson PM, van der Spoel D (2013) GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29(7):845–854. https://doi.org/10.1093/bioinformatics/btt055
Robert X, Gouet P (2014) Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res 42(W1):W320–W324. https://doi.org/10.1093/nar/gku316
Rye CS, Withers SG (2000) Glycosidase mechanisms. Curr Opin Chem Biol 4(5):573–580. https://doi.org/10.1016/S1367-5931(00)00135-6
Scheller HV, Ulvskov P (2010) Hemicelluloses. Annu Rev Plant Biol 61:263–289. https://doi.org/10.1146/annurev-arplant-042809-112315
Singh S, Madlala AM, Prior BA (2003) Thermomyces lanuginosus: properties of strains and their hemicellulases. FEMS Microbiol Rev 27(1):3–16. https://doi.org/10.1016/S0168-6445(03)00018-4
Song L, Siguier B, Dumon C, Bozonnet S, O'Donohue M (2012) Engineering better biomass-degrading ability into a GH11 xylanase using a directed evolution strategy. Biotechnol Biofuels 5(1):3. https://doi.org/10.1186/1754-6834-5-3
Thongekkaew J, Ikeda H, Masaki K, Iefuji H (2013) Fusion of cellulose binding domain from Trichoderma reesei CBHI to Cryptococcus sp. S-2 cellulase enhances its binding affinity and its cellulolytic activity to insoluble cellulosic substrates. Enzym Microb Technol 52(4):241–246. https://doi.org/10.1016/j.enzmictec.2013.02.002
Tian L, Liu S, Wang S, Wang L (2016) Ligand-binding specificity and promiscuity of the main lignocellulolytic enzyme families as revealed by active-site architecture analysis. Sci Rep 6:23605. https://doi.org/10.1038/srep23605
Tishkov VI, Gusakov AV, Cherkashina AS, Sinitsyn AP (2013) Engineering the pH-optimum of activity of the GH12 family endoglucanase by site-directed mutagenesis. Biochimie 95(9):1704–1710. https://doi.org/10.1016/j.biochi.2013.05.018
Tripathi A, Varadarajan R (2014) Residue specific contributions to stability and activity inferred from saturation mutagenesis and deep sequencing. Curr Opin Struct Biol 24:63–71. https://doi.org/10.1016/j.sbi.2013.12.001
Viikari L, Kantelinen A, Sundquist J, Linko M (1994) Xylanases in bleaching: from an idea to the industry. FEMS Microbiol Rev 13(2–3):335–350
Warnecke F, Luginbühl P, Ivanova N, Ghassemian M, Richardson T, Stege J, Cayouette M, McHardy A, Djordjevic G, Aboushadi N, Sorek R, Tringe S, Podar M, Martin H, Kunin V, Dalevi D, Madejska J, Kirton E, Platt D, Szeto E, Salamov A, Barry K, Mikhailova N, Kyrpides N, Matson E, Ottesen E, Zhang X, Hernández M, Murillo C, Acosta L, Rigoutsos I, Tamayo G, Green B, Chang C, Rubin E, Mathur E, Robertson D, Hugenholtz P, Leadbetter J (2007) Metagenomic and functional analysis of hindgut microbiota of a wood-feeding higher termite. Nature 450(7169):560–565. https://doi.org/10.1038/nature06269
Webb B, Sali A (2014) Protein structure modeling with MODELLER. Humana Press, New York
Weiner MP, Costa GL, Schoettlin W, Cline J, Mathur E, Bauer JC (1994) Site-directed mutagenesis of double-stranded DNA by the polymerase chain reaction. Gene 151(1):119–123
Xie Y, An J, Yang G, Wu G, Zhang Y, Cui L, Feng Y (2014) Enhanced enzyme kinetic stability by increasing rigidity within the active site. J Biol Chem 289(11):7994–8006. https://doi.org/10.1074/jbc.M113.536045
Zhang Y, Wang L, Chen J, Wu J (2013) Enhanced activity toward PET by site-directed mutagenesis of Thermobifida fusca cutinase-CBM fusion protein. Carbohydr Polym 97(1):124–129. https://doi.org/10.1016/j.carbpol.2013.04.042
Zhang H, Li J, Wang J, Yang Y, Wu M (2014) Determinants for the improved thermostability of a mesophilic family 11 xylanase predicted by computational methods. Biotechnol Biofuels 7(1):3. https://doi.org/10.1186/1754-6834-7-3
Zhang L, Ma H, Zhang H, Xun L, Chen G, Wang L (2015a) Thermomyces lanuginosus is the dominant fungus in maize straw composts. Bioresour Technol 197:266–275. https://doi.org/10.1016/j.biortech.2015.08.089
Zhang Q, Zhang X, Wang P, Li D, Chen G, Gao P, Wang L (2015b) Determination of the action modes of cellulases from hydrolytic profiles over a time course using fluorescence-assisted carbohydrate electrophoresis. Electrophoresis 36(6):910–917. https://doi.org/10.1002/elps.201400563
Zhang X, Wang S, Wu X, Liu S, Li D, Xu H, Gao P, Chen G, Wang L (2015c) Subsite-specific contributions of different aromatic residues in the active site architecture of glycoside hydrolase family 12. Sci Rep 5:e18357. https://doi.org/10.1038/srep18357
Funding
This study was funded by The Key Technologies R&D Program of Shandong Province (grant number 2015GSF121019) and The National Natural Science Foundation of China (grant number 31370111).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
ESM 1
(PDF 476 kb)
Rights and permissions
About this article
Cite this article
Wu, X., Tian, Z., Jiang, X. et al. Enhancement in catalytic activity of Aspergillus niger XynB by selective site-directed mutagenesis of active site amino acids. Appl Microbiol Biotechnol 102, 249–260 (2018). https://doi.org/10.1007/s00253-017-8607-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-017-8607-8