Abstract
The production of amylolytic enzymes in Aspergillus oryzae is induced in the presence of starch or maltose, and two Zn2Cys6-type transcription factors, AmyR and MalR, are involved in this regulation. AmyR directly regulates the expression of amylase genes, and MalR controls the expression of maltose-utilizing (MAL) cluster genes. Deletion of malR gene resulted in poor growth on starch medium and reduction in α-amylase production level. To elucidate the activation mechanisms of these two transcription factors in amylase production, the expression profiles of amylases and MAL cluster genes under carbon catabolite derepression condition and subcellular localization of these transcription factors fused with a green fluorescent protein (GFP) were examined. Glucose, maltose, and isomaltose induced the expression of amylase genes, and GFP-AmyR was translocated from the cytoplasm to nucleus after the addition of these sugars. Rapid induction of amylase gene expression and nuclear localization of GFP-AmyR by isomaltose suggested that this sugar was the strongest inducer for AmyR activation. In contrast, GFP-MalR was constitutively localized in the nucleus and the expression of MAL cluster genes was induced by maltose, but not by glucose or isomaltose. In the presence of maltose, the expression of amylase genes was preceded by MAL cluster gene expression. Furthermore, deletion of the malR gene resulted in a significant decrease in the α-amylase activity induced by maltose, but had apparently no effect on the expression of α-amylase genes in the presence of isomaltose. These results suggested that activation of AmyR and MalR is regulated in a different manner, and the preceding activation of MalR is essential for the utilization of maltose as an inducer for AmyR activation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A filamentous fungus, Aspergillus oryzae, has the ability to produce copious amounts of enzymes such as amylolytic enzymes and has been utilized in Japan for the production of traditional Japanese fermented foods and beverages such as sake, soy sauce, and miso (soybean paste) for over a thousand years (Machida et al. 2008). The production of amylolytic enzymes by A. oryzae is induced in the presence of starch or maltooligosaccharides (Tonomura et al. 1961), and the induction of the corresponding amylolytic genes is regulated by a fungal-specific Zn2Cys6 transcription activator, AmyR (Gomi et al. 2000; Petersen et al. 1999). The AmyR activation mechanism has been well studied in Aspergillus nidulans by subcellular localization analysis using a green fluorescent protein (GFP)-fused AmyR. In A. nidulans, isomaltose induces amylase synthesis (Kato et al. 2002b) and triggers rapid nuclear localization of GFP-AmyR, whereas the absence of inducing sugars has been found to result in the distribution of GFP-AmyR in the cytoplasm (Makita et al. 2009; Murakoshi et al. 2012). Although the expression of amylolytic genes is strongly repressed by a C2H2-type transcription factor, CreA, under glucose-containing condition, the nuclear localization of GFP-AmyR is also triggered by glucose. However, glucose and maltose require higher concentration and longer time, when compared with isomaltose, to induce nuclear localization of GFP-AmyR (Murakoshi et al. 2012). These observations indicate that isomaltose is the strongest inducer for the activation of A. nidulans AmyR.
Both A. oryzae and A. nidulans amyR disruption strains showed poor growth on the starch medium (Gomi et al. 2000; Tani et al. 2001). However, an A. oryzae amyR disruption strain could grow normally in the maltose medium, whereas an A. nidulans amyR disruption strain exhibited a significantly limited growth (Hasegawa et al. 2010; Tani et al. 2001). This result suggests that regulation of the maltose utilization system in A. oryzae is independent of AmyR. In a previous study, we identified a Zn2Cys6-type transcription activator MalR, the ortholog of yeast maltose-utilizing (MAL) activator, in the A. oryzae genome (Hasegawa et al. 2010). Similar to the yeast MAL activator, the malR gene was found to constitute the cluster together with genes encoding putative maltose permease (MalP) and maltase (MalT). The transcription of malP and malT was noted to be regulated by MalR, and the expression of MAL cluster genes, including malR, was not affected by the amyR disruption. The disruption mutants of malR and malP exhibited a growth defect in the maltose medium, suggesting that maltose utilization in A. oryzae is controlled by the MAL cluster independent of AmyR (Hasegawa et al. 2010). The expression of the agdF gene, an ortholog of malT in A. nidulans, is regulated by AmyR (Nakamura et al. 2006). These observations suggest that the mechanism for the regulation of maltose utilization in A. oryzae appreciably differs from that in A. nidulans. In addition, involvement of MalR in α-amylase production indicates that the mechanism of amylolytic enzyme production differs between A. oryzae and A. nidulans.
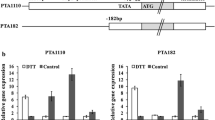
Expression profiles of amylase genes and MAL cluster genes in the ∆creA strain. The wild-type and ∆creA mutant strains were grown in liquid MM + 0.1 % polypeptone medium containing 1 % glycerol as the carbon source for 24 h, followed by transfer to liquid MM containing 1 % glucose or maltose. The mycelia were harvested at the time points indicated, and the total RNA was extracted from the harvested mycelia. Approximately 20 μg of the total RNA was subjected to northern blot analysis and the digoxigenin (DIG)-labeled fragments of each gene were used as probes. The loading control used was 18S rRNA
In our recent study, A. oryzae ∆creA mutant strains showed a high α-amylase activity in a complete medium containing glucose, suggesting that glucose could act as an inducer for α-amylase production in A. oryzae (Ichinose et al. 2014). However, there is no information on the activation mechanisms of AmyR and MalR in A. oryzae. In the present study, we investigated the activation mechanism of A. oryzae AmyR and MalR through expression analysis of amylases and MAL cluster genes in the ∆creA strain, and performed subcellular localization analysis of these transcription activators using GFP-fused proteins.
Materials and methods
Strains and media
An A. oryzae ∆ligD::loxP pyrG-deficient strain (∆ligD::loxP, niaD −, sC −, pyrG −), which was derived from a ∆ligD::loxP strain (Mizutani et al. 2012), was used as the recipient strain for malR deletion. The ∆ligD::loxP strain was constructed from A. oryzae NS4 strain (niaD −, sC −) (Yamada et al. 1997) derived from the wild-type strain, A. oryzae RIB40 (National Research Institute of Brewing Stock Culture, Higashi-hiroshima, Japan). The A. oryzae strains used in this study are listed in Table 1. The ∆ligD::loxP pyrG::niaD strain (Ichinose et al. 2014) was defined as the wild-type strain in this study. Escherichia coli DH5α (Promega, Madison, WI, USA) was used for the construction and propagation of plasmid DNAs. The minimal medium (MM) for A. oryzae culture was Czapek–Dox (CD) medium that contained 0.5 % (NH4)2SO4; 0.05 % KCl; 0.2 % KH2PO4; 0.05 % MgSO4; trace amounts of FeSO4, ZnSO4, CuSO4, MnSO4, Na2B4O7, and (NH4)6Mo7O24; and 1 % sugar, supplemented with 0.0003 % (0.02 mM) methionine. For the cultivation of strain ∆ligD::loxP pyrG −, 0.2 % uracil was added to the medium. YPM medium (0.5 % yeast extract, 1 % Bacto-peptone, and 1 % maltose) was used to assess the \( \boldsymbol{\upalpha} \)-amylase production of the ∆amyR strains that express intact or C-terminal truncated AmyR of A. oryzae and A. nidulans.
Constructions of plasmid DNAs and DNA fragment for gene deletion
The plasmids for the GFP-fused AmyR and MalR expression driven by the thiA promoter were constructed as follows. The thiA promoter region was amplified by PCR with a primer set, PthiAsenPstI and PthiAantiSalI, using the genomic DNA of A. oryzae RIB40 as the template. The amplified fragment was digested with PstI and SalI, and replaced with the glaA142 promoter of the plasmid pNGA142 (Tamalampudi et al. 2007), which contains the niaD gene as a selectable marker, yielding the plasmid pNthiA. Next, the gfp gene was amplified by PCR with a primer set, GFPsenSalI and GFP5GAantiNotI, using the plasmid pAGAR-F (Makita et al. 2009; provided by Prof. Kobayashi of Nagoya University, Japan) as the template. To construct functional GFP fusion proteins, five Gly-Ala repeats were attached to the C terminus of GFP as a linker, as reported previously (Yang et al. 2004). The SalI/NotI-digested gfp fragment was inserted into pNthiA, yielding the plasmid pNthiA-GFP5GA. Then, the amyR and malR genes were amplified by PCR using primer sets, amyRsenNotI + amyRantiSphI and malRsenNotI + malRantiSpeI, respectively, and these fragments were inserted into pNthiA-GFP5GA to be fused in-frame at the 3′ terminus of the gfp gene through codons for five Gly-Ala repeats. The 3′-terminally truncated amyR gene encoding the AmyR1–511 variant was also amplified by PCR using a primer set, amyRsenNotI + amyRDCantiSphI, and inserted into NotI/SphI-digested pNthiA-GFP5GA. The plasmids carrying a codon for an alanine substitution in the putative nuclear localization signals (NLSs) were generated by QuikChange site-directed mutagenesis (Agilent Technologies, Santa Clara, CA, USA) with primer sets AmyRNLS1m + AmyRNLS1m-r, AmyRNLS2m + AmyRNLS2m-r, and MalRNLSm + MalRNLSm-r.
The plasmids for expression of the A. nidulans amyR gene were constructed as follows. The intact and 3′-truncated amyR genes were amplified by PCR with primer set ANamyRsenNotI + ANamyRantiSphI, using the plasmids pAGAR-F and pAR-Zn3 (Makita et al. 2009; provided by Prof. Kobayashi of Nagoya University, Japan) as templates, respectively. The obtained PCR fragments were digested with NotI and SphI and inserted into the plasmid pNE, which was constructed by replacement of the glaA142 promoter of the plasmid pNGA142 with the enolase gene promoter derived from the plasmid pNGEG (Tsuboi et al. 2005). In a similar manner, the A. oryzae intact and 3′-truncated amyR genes were amplified by PCR with primer sets amyRsenNotI + amyRantiSphI and amyRsenNotI + amyR∆CantiSphI, using the A. oryzae genomic DNA as the template, and inserted into the plasmid pNE.
The DNA fragment for malR deletion using pyrG as a selectable marker was amplified by PCR with the genomic DNA of the A. oryzae malR deletion strain (provided by Dr. Yasuji Koyama of Noda Institute for Scientific Research, Japan) using primers malRsen and malRanti. The nucleotide sequences of all the primers used in this study are shown in Supplemental Table S1.
Fungal transformation
Transformation of A. oryzae was performed according to the method described by Gomi et al. (1987).
Southern and northern blot analyses
The preparation of A. oryzae genomic DNA and total RNA, Southern blot analysis, and northern blot analysis were performed as described previously (Tanaka et al. 2012).
Fluorescence microscopy
Approximately 5 × 103 A. oryzae conidiospores were inoculated onto coverslips (18 × 18 mm) dipped in 500 μl of MM containing 1 % casamino acids as the carbon source with or without sugars. The hyphal cells grown on the coverslips were examined by fluorescence microscopy. To examine subcellular localization of GFP-AmyR in a relatively short period after sugar addition, the medium was removed from the coverslip following incubation at 30 °C for 12 h, and the hyphae on the coverslip were dipped in fresh MM containing 1 or 0.1 % sugar with thiamine at a final concentration of 10 μM. Then, the fluorescence of the hyphae was examined after incubation at 30 °C for 10 or 30 min. For nuclear fluorescence staining, Hoechst33342 was added at a final concentration of 0.1 mg/ml for 15 min before microscopy imaging. A confocal laser scanning microscope (FV1000-D IX81; Olympus, Tokyo, Japan) was used for microscopy and image preparations. Fluorescence images were uniformly adjusted for clarity using Adobe Photoshop software (Adobe Systems, San Jose, CA, USA).
Intracellular protein extraction
The mycelium grown in the liquid MM was harvested by filtration through Miracloth (EMD Millipore Corporation, Billerica, MA, USA), washed with distilled water, and frozen in liquid nitrogen. Then, the mycelium was ground to a fine powder in liquid nitrogen using a mortar and pestle. The powdered mycelium was suspended in protein extraction buffer [25 mM Tris–HCl (pH 8.0), 0.25 % CHAPS, 100 mM NaCl, 2 mM phenylmethylsulfonyl fluoride (PMSF), 15 μM pepstatin A, and complete EDTA-free protease inhibitor (Roche, Indianapolis, IN, USA)] and incubated on ice for 15 min, followed by centrifugation at 20,400×g for 10 min at 4 °C. The protein concentration in the supernatant was measured by the Bradford method (Bradford 1976) using a Coomassie protein assay kit and a microplate photometer Multiskan FC (Thermo Fisher Scientific Inc., Waltham, MA, USA), and approximately 100 μg of proteins was concentrated by precipitation with 10 % trichloroacetic acid (TCA).
Western blot analysis
Approximately 10 μg of intracellular proteins was subjected to SDS–PAGE and transferred onto an Immobilon P polyvinylidene difluoride membrane (EMD Millipore Corporation, Billerica, MA, USA) with Towbin buffer. Anti-GFP (mFX75; Wako, Osaka, Japan) antibody was used for the detection of GFP-fused proteins. Signal detection was performed using Chemi-Lumi One L kit (Nacalai Tesque, Kyoto, Japan), according to the manufacturer’s instructions, and the chemiluminescence signal was detected using an image analyzer ImageQuant LAS-4000 (GE Healthcare, Piscataway, NJ, USA).
α-Amylase activity assay
The α-amylase activity was measured as described previously (Ichinose et al. 2014; Sato et al. 2011).
Results
Expression analysis of amylolytic genes and MAL cluster genes in ∆creA strain
As reported previously (Hasegawa et al. 2010), to verify the involvement of MalR in amylolytic enzyme production, we generated a deletion mutant of malR by replacement with the A. nidulans pyrG gene. The deletion of malR gene was confirmed by Southern blot analysis (Supplemental Fig. S1). To examine the effect of malR deletion on amylolytic enzyme production, the mycelia of the wild-type and ∆malR strains grown in liquid MM with casamino acids as the carbon source were transferred to the maltose medium, and the α-amylase activities in the culture supernatants were measured. As shown in Table 2, the α-amylase activity of the ∆malR strain was significantly reduced, when compared with that of the wild-type strain, suggesting that MalR essentially contributed to the amylolytic enzyme production in A. oryzae.
To verify whether glucose could induce the transcription of amylolytic genes in A. oryzae, the expression of α-amylase genes (amyA/B/C) and a glucoamylase gene (glaA) in the ∆creA strain was examined (Fig. 1). The mycelia of the wild-type and ∆creA strains grown in casamino acids medium were transferred to glucose and maltose media, and the gene expression level was monitored by northern blot analysis. In the wild-type strain, the expression of amyA/B/C and glaA was strongly induced after transfer to the maltose medium, whereas induction of these genes was not observed after transfer to the glucose medium. However, in the ∆creA strain, the expression of these genes was induced by maltose as well as glucose. These results indicated that glucose could induce the amylolytic gene expression under CreA-deficient condition in A. oryzae. Furthermore, we found that the expression of MAL cluster genes (malP and malT) was strongly induced by maltose in both wild-type and ∆creA strains. However, the transcripts of these MAL cluster genes were not detectable in the ∆creA strain after transfer to the glucose medium. These results suggested that glucose could act as an inducer for AmyR activation, but not for MalR activation.
Subcellular localization of GFP-AmyR and GFP-MalR
To investigate the subcellular localization of AmyR and MalR in A. oryzae, GFP-fused AmyR and MalR were expressed under the control of the thiA promoter. In this experiment, the expression level of both GFP-fused regulators was controlled by thiamine addition (Kubodera et al. 2003). The growth of the amyR or malR disruption mutant on the starch medium was restored by the introduction of the expression plasmid for GFP-AmyR or GFP-MalR, respectively, suggesting that the N-terminal fusion of GFP to these regulators had no apparent effect on the transcriptional activation potential of both AmyR and MalR (Fig. 2a, d). To observe GFP fluorescence, the transformant expressing GFP-AmyR was cultured in liquid MM containing casamino acids with or without sugars for 12 h, and the fluorescence of GFP was observed by confocal fluorescence microscopy. In the absence of sugars, GFP fluorescence was observed in the cytoplasm of the hyphae (Fig. 2b), whereas in the presence of glucose or maltose, GFP fluorescence was observed in organelles stained with Hoechst dye, indicating that GFP-AmyR was accumulated in the nucleus. In addition, when we examined the effect of sugars on subcellular localization of GFP-AmyR in a short period, the GFP fluorescence was detected in the nucleus 30 min after the addition of glucose or maltose (Fig. 2c). These results suggested that inducing sugars, such as maltose and glucose, for amylolytic gene expression can trigger the nuclear localization of AmyR in A. oryzae, as observed for AmyR in A. nidulans (Makita et al. 2009; Murakoshi et al. 2012). On the contrary, GFP fluorescence was observed in the nucleus when the transformant expressing GFP-MalR was cultured in the liquid MM using casamino acids as the carbon source (Fig. 2e). The nuclear localization of GFP-MalR was unaffected by the addition of sugars, indicating that MalR was constitutively localized in the nucleus (Fig. 2e).
Subcellular localization of GFP-AmyR and GFP-MalR. a Complementation of GFP-AmyR. Approximately 1 × 103 conidiospores of wild-type, ∆amyR, and GFP-AmyR-expressing strains were grown on MM agar plates containing 0.1 % glucose or starch as the sole carbon source at 30 °C for 3 days. b GFP fluorescence of GFP-AmyR after 12-h cultivation in MM using 1 % casamino acids as the carbon source, with or without 1 % sugars. The hyphal cells were examined by confocal microscopy at ×1000 magnification. c GFP fluorescence 30 min after the addition of sugars with thiamine. The hyphae grown in MM using 1 % casamino acids as the carbon source for 12 h were dipped in fresh MM containing 1 % sugars with thiamine at a final concentration of 10 μM. d Complementation of GFP-MalR. Approximately 100 conidiospores of the ∆malR and GFP-MalR-expressing strains were grown on MM agar plates containing 0.1 % glucose or starch as the sole carbon source at 30 °C for 3 days. e GFP fluorescence of the GFP-MalR after 12-h cultivation in MM containing 1 % casamino acids with or without sugars
Effects of mutations in putative NLSs on AmyR and MalR subcellular localization
In A. nidulans AmyR, two NLSs situated within the zinc binuclear motif have been identified (Makita et al. 2009). As two basic amino acid clusters also existed within the zinc binuclear motif of A. oryzae AmyR (Fig. 3a), these amino acid clusters in GFP-AmyR were replaced with alanine residues. After transfer to the maltose medium, in both single and double NLSs-mutated GFP-AmyRs, GFP fluorescence was not observed in the nucleus, but was detectable around the nuclear envelope (Fig. 3b). In addition, the degradation products of NLSs-mutated AmyR proteins could not be detected by western blot analysis (Supplemental Fig. S2a). These observations indicated that both the basic amino acid clusters are functional NLSs of A. oryzae AmyR.
Subcellular localization of NLS-mutated AmyR and MalR. a Schematic representation of the AmyR domain structure. The zinc binuclear motif and four MH domains are represented as black boxes. The amino acid sequence of the zinc binuclear motif is shown and two putative NLSs are underlined. b GFP fluorescence of NLS-mutated GFP-AmyR after 12-h cultivation in the medium containing 1 % maltose. The hyphae were examined at ×3000 magnification. c Schematic representation of the MalR domain structure. The zinc binuclear motif and three MH domains are represented as black boxes. The amino acid sequence of the zinc binuclear motif is shown and a basic amino acid cluster is underlined. d GFP fluorescence of NLS-mutated GFP-MalR after 12-h cultivation in the medium containing 1 % maltose. The hyphae were examined at ×3000 magnification
A single basic amino acid cluster, RRK, was noted within the zinc binuclear motif of MalR (Fig. 3c). To examine whether this basic amino acid cluster acts as a NLS, RRK was replaced with three alanine residues (AAA) in the GFP-MalR, and subcellular localization of the mutated GFP-MalR (GFP-MalRNLSm) was observed by fluorescence microscopy. The GFP fluorescence of GFP-MalRNLSm was diffused in the cytoplasm under any culture conditions, although it was still observed in the nucleus (Fig. 3d). Western blot analysis using anti-GFP showed that intact GFP-MalRNLSm protein primarily existed in the cells, suggesting that GFP fluorescence in the cytoplasm was not derived from free GFP produced by the degradation of GFP-MalRNLSm protein (Supplemental Fig. S2b). These results indicated that the basic amino cluster was a functional NLS of MalR, although nuclear localization of MalR could not be regulated by this signal alone.
Effects of C-terminal deletion on AmyR subcellular localization
The A. oryzae AmyR protein structure closely resembled that of A. nidulans AmyR and could be also divided into five regions, including a zinc binuclear motif and four MH regions (Fig. 3a; Tani et al. 2001). In contrast, although the entire amino acid sequence of MalR showed homologies to that of AmyR, MalR presented no identity to the MH4 domain of AmyR (Fig. 3c and Supplemental Fig. S3). It has been shown that the deletion of the C-terminal region, including MH4, caused constitutive nuclear localization of A. nidulans AmyR and resulted in constitutive expression of amylolytic genes (Makita et al. 2009). Thus, to examine the role of the C-terminal region of A. oryzae AmyR in its subcellular localization, the truncated AmyR (AmyR1–511) that lacks the C-terminal 93 amino acid residues, including the MH4 region, was fused to the GFP protein and expressed in the amyR disruption strain. In the absence of inducing sugars, the fluorescence of GFP-AmyR1–511 was observed in the nucleus, suggesting that GFP-AmyR1–511 was constitutively localized to the nucleus (Fig. 4a). However, the GFP-AmyR1–511-expressing strain showed a little α-amylase activity in the liquid medium even in the presence of maltose (Fig. 4b). Western blot analysis using anti-GFP antibody revealed that the amount of intact GFP-AmyR1–511 protein was comparable with that of GFP-AmyR (Supplemental Fig. S2c), suggesting that the decrease in α-amylase production may not have been caused by the proteolytic degradation of GFP-AmyR1–511. These results suggested that the C-terminal region of A. oryzae AmyR is required for transcriptional activity and intracellular localization.
Subcellular localization of the C-terminal truncated GFP-AmyR. a GFP fluorescence of the C-terminal truncated GFP-AmyR. The hyphae were examined as described in Fig. 3b. b The α-amylase activity of intact and C-terminal truncated AmyRs-expressing strains. Approximately 1 × 107 conidiospores of each strain were grown in the casamino acids medium with or without 1 % maltose at 30 °C for 24 h. The harvested mycelia were incubated in 100 mM phosphate buffer for 60 min to release the α-amylase bound to the cell wall, and the α-amylase activities in the culture broth and phosphate buffer were measured. The total activity was divided by the mycelial dry weight. Error bars indicate the standard errors of three independent experiments
Comparison of the ability of maltose and isomaltose to activate AmyR and MalR
To investigate whether maltose or isomaltose could act as a stronger inducer for AmyR and MalR, the expression profiles of AmyR- and MalR-regulated genes in the ∆creA strain were monitored by northern blot analysis after the addition of each sugar (Fig. 5a). The expression of amyA/B/C genes was induced 10 min after the addition of isomaltose. In contrast, these genes were not expressed at least for 10 min, but were strongly induced 30 min after the addition of maltose. In agreement with amyA/B/C expression profiles, GFP-AmyR was accumulated in the nucleus 10 min after isomaltose addition, and it remained in the cytoplasm for the same period after maltose induction (Fig. 5b). Conversely, malT and malP genes were not expressed even after 60 min of incubation following isomaltose addition. These results clearly indicated that isomaltose could act as a strong inducer for AmyR activation, but not for MalR activation. Contrary to the observation that the amyA/B/C genes were expressed 30 min after induction, the expression of malT and malP was induced within 10 min after maltose addition, suggesting that MalR was activated prior to AmyR activation in the presence of maltose as an inducing sugar. Finally, we examined the expression profiles of amyA/B/C genes in the ∆malR strain after the addition of maltose or isomaltose. In the ∆malR strain, the expression of amyA/B/C genes was normally induced after isomaltose addition, but was strongly repressed after maltose addition (Fig. 5c). This result suggested that MalR was essential for the utilization of maltose as an inducer for AmyR activation.
Comparison of the ability of maltose and isomaltose to activate AmyR and MalR. a Expression profiles of amylase and MAL cluster genes in the ∆creA strain after the addition of maltose or isomaltose. The ∆creA strain was grown in MM containing 1 % casamino acids for 24 h, followed by transfer to MM containing 0.1 % maltose or isomaltose as the sole carbon source. The mycelia were harvested and subjected to northern blot analysis as described Fig. 1. b GFP fluorescence of GFP-AmyR in 10 min after the addition of maltose or isomaltose with thiamine. The hyphae were examined as described in Fig. 2c. The final concentration of maltose and isomaltose added to the medium is 0.1 %. c Expression of α-amylase genes in the wild-type and ∆malR strains after the addition of maltose or isomaltose
Discussion
The amylolytic enzyme production by A. oryzae is strongly induced in the presence of maltose, and the expression of amylolytic genes and MAL genes are regulated by two distinct Zn2Cys6 transcription activators, AmyR and MalR, respectively. Deletion of the malR gene resulted in a reduced α-amylase activity, poor growth on the starch medium, and retardation of α-amylase gene expression in the maltose medium (Table 2, Figs. 4a and 5c), indicating that MalR also contributes to the regulation of amylolytic gene expression. In the present study, we examined the expression profiles of these genes in ∆creA strain and subcellular localization of the transcription activators to understand the mechanism of amylolytic gene expression in A. oryzae.
The results obtained in this study clearly indicated that the activation mechanism of AmyR and MalR drastically differed from one another. Subcellular localization analysis using GFP-AmyR revealed that AmyR was translocated from the cytoplasm to the nucleus in response to the presence of glucose, maltose, and isomaltose (Fig. 3b). In addition, the expression of α-amylase genes was induced by the addition of these sugars in the carbon catabolite derepression mutant ∆creA and isomaltose could rapidly trigger the α-amylase gene expression (approximately 10 min after induction). These results suggested that activation of A. oryzae AmyR is regulated by its translocation in the nucleus and that isomaltose is the strongest inducer for AmyR activation, as reported for A. nidulans AmyR (Kato et al. 2002a; Makita et al. 2009). In contrast to AmyR, MalR was constitutively localized to the nucleus and the expression of MAL cluster genes involved in maltose utilization was induced only by maltose and not by glucose or isomaltose. Conversely, the α-amylase gene expression was induced within 30 min of incubation after the addition of maltose, and the expression of MAL cluster genes was induced within 10 min after maltose addition. These results indicated that activation of AmyR was preceded by MalR activation in the presence of maltose.
In A. nidulans, the transglycosylation activity of two α-glucosidases (AgdA and AgdB) with a signal peptide is involved in the conversion from maltose to isomaltose for α-amylase gene induction, although other α-glucosidase(s) with isomaltose-forming activity may exist (Kato et al. 2002a, b). The nuclear localization of A. nidulans GFP-AmyR after maltose addition was inhibited by the addition of a α-glucosidase inhibitor, castanospermine (Murakoshi et al. 2012). However, in A. oryzae, induction of α-amylase gene expression after maltose addition was not inhibited by the addition of α-glucosidase inhibitors, castanospermine and 1-deoxynojirimycin (data not shown). It should be noted that the malT gene in the MAL cluster encodes the functional intracellular α-glucosidase (Hasegawa et al. 2010). Therefore, we hypothesized that the conversion of maltose to isomaltose in A. oryzae is executed intracellularly, whereas that in A. nidulans is completed extracellularly by the transglycosylation activity of AgdA and AgdB (Kato et al. 2002a, b). The normal expression of α-amylase genes after isomaltose addition in the ∆malR strain could support this idea (Fig. 5c). To assess our hypothesis, construction of the malT deletion mutant and assay for transglycosylation activity of MalT are currently underway. However, understandably, the possibility of involvement of other intercellular and/or extracellular α-glucosidase(s) in isomaltose formation should not be ruled out. In this context, according to the Carbohydrate-Active enZYme (CAZy) database (http://www.cazy.org/), ten α-glucosidases that belong to glycoside hydrolase family 31 (GH31), including extracellular AgdA and AgdB, are found in the A. oryzae genome. Furthermore, additional four enzymes highly homologous to MalT, which belongs to GH13, are present in the A. oryzae genome. It will be necessary to examine whose gene(s) other than the malT gene is directly regulated by MalR.
Based on our experimental results, we proposed the following regulation mechanism of A. oryzae α-amylase gene expression in the presence of maltose. First, the maltose transporter MalP whose gene is expressed at the basal level incorporates a small amount of maltose into the A. oryzae cell, triggering the activation of MalR. Then, transcription of MAL cluster genes is quickly induced by the activated MalR, and a large amount of maltose is incorporated into the cell by MalP. Finally, the transglycosylation activity of intracellular α-glucosidase(s) converts maltose into isomaltose, which simultaneously triggers the activation and translocation of AmyR into the nucleus.
The nuclear localization of AmyR was blocked by mutation of basic amino acid clusters present within their Zn(II)2Cys6 DNA binuclear domain, suggesting that these basic amino acid clusters are functional NLSs. The recognition of NLS by nuclear importins mediates protein transport from the cytoplasm to the nucleus (Lott and Cingolani 2011; Marfori et al. 2011; Xu et al. 2010). The GFP fluorescence of the mutated AmyRs harboring mutation in single NLS was observed around the nuclear envelope, whereas double NLS mutation resulted in the distribution of GFP-AmyR in the cytoplasm. We speculated that although intact NLS of a single NLS-mutated AmyR was recognized by nuclear importins, the interaction force of the mutated AmyRs with nuclear importins was not sufficient for the import of AmyR into the nucleus. On the other hand, mutation of a basic amino acid cluster located within the zinc binuclear domain of MalR led to the diffusion of GFP fluorescence in the cytoplasm, suggesting that this basic amino acid cluster was also a functional NLS. However, strong GFP fluorescence of the NLS-mutated MalR was still observed in the nucleus, indicating that the basic amino acid cluster of MalR was not essential for nuclear localization, and that other sequences required for nuclear import of MalR may exist. In fact, amino acids 220–226, PTERARR, in MalR has also been predicted as NLS by pSORT program (http://psort.hgc.jp/); therefore, we are planning to examine whether this putative NLS actually functions as another NLS in MalR by site-directed mutagenesis.
The MH4 domain of A. nidulans AmyR has been reported to be essential for its cytoplasmic location (Makita et al. 2009). Accordingly, in the present study, deletion of the C-terminal region containing the MH4 domain resulted in constitutive nuclear localization of A. oryzae AmyR (Fig. 3b). By contrast, MalR showed no identity to the MH4 domain of AmyR (Supplemental Fig. S3), which might contribute to the difference in subcellular localization under inducer-deficient condition between AmyR and MalR. It has been reported that the constitutive nuclear localization of the C-terminal deleted A. nidulans AmyR resulted in constitutive amylase production (Makita et al. 2009). In the present study, deletion of the C-terminal region of A. oryzae AmyR led to a marked loss of α-amylase production even in the presence of maltose (Fig. 3c). In addition, the expression of the C-terminal truncated A. nidulans AmyR in the A. oryzae ∆amyR strain restored α-amylase production (Supplemental Fig. S4). These results suggested that the C-terminal region of A. oryzae AmyR is essential for transcriptional activation potential, probably either by functioning as an activation domain or by functioning to make the protein properly folded. Although the C-terminal region of the yeast MAL activator also led to marked loss of transcriptional activation potential (Danzi et al. 2003), it has been proposed that the C-terminal region of the MAL activator possesses both positive and negative regulatory roles (Danzi et al. 2003; Hu et al. 1999). Therefore, the function of the C-terminal regions of AmyRs may be considerably complicated and further analysis is required.
The process of MalR activation remains completely unclear. It is known that phosphorylation of transcription factors regulates their stability, localization, protein–protein interaction, DNA binding, and transcriptional activity (Holmberg et al. 2002). For instance, it has been reported that the activation of A. oryzae XlnR, the transcriptional activator of xylanolytic and cellulolytic genes, is controlled by reversible phosphorylation (Noguchi et al. 2011). In addition, it has been indicated that the activation of yeast MAL activator is regulated by interaction with several chaperone proteins (Bali et al. 2003; Ran et al. 2008, 2010). Thus, studies on post-translational modifications and interaction with chaperone proteins of AmyR and MalR are necessary for a more detailed understanding of their activation mechanism.
References
Bali M, Zhang B, Morano KA, Michels CA (2003) The Hsp90 molecular chaperone complex regulates maltose induction and stability of the Saccharomyces MAL gene transcription activator Mal63p. J Biol Chem 278:47441–47448
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein–dye binding. Anal Biochem 72:248–254
Danzi SE, Bali M, Michels CA (2003) Clustered-charge to alanine scanning mutagenesis of the Mal63 MAL-activator C-terminal regulatory domain. Curr Genet 44:173–183
Gomi K, Iimura Y, Hara S (1987) Integrative transformation of Aspergillus oryzae with a plasmid containing the Aspergillus nidulans argB gene. Agric Biol Chem 51:2549–2555
Gomi K, Akeno T, Minetoki T, Ozeki K, Kumagai C, Okazaki N, Iimura Y (2000) Molecular cloning and characterization of a transcriptional activator gene, amyR, involved in the amylolytic gene expression in Aspergillus oryzae. Biosci Biotechnol Biochem 64:816–827
Hasegawa S, Takizawa M, Suyama H, Shintani T, Gomi K (2010) Characterization and expression analysis of a maltose-utilizing (MAL) cluster in Aspergillus oryzae. Fungal Genet Biol 47:1–9
Holmberg CI, Tran SE, Eriksson JE, Sistonen L (2002) Multisite phosphorylation provides sophisticated regulation of transcription factors. Trends Biochem Sci 27:619–627
Hu Z, Gibson AW, Kim JH, Wojciechowicz LA, Zhang B, Michels CA (1999) Functional domain analysis of the Saccharomyces MAL-activator. Curr Genet 36:1–12
Ichinose S, Tanaka M, Shintani T, Gomi K (2014) Improved α-amylase production by Aspergillus oryzae after a double deletion of genes involved in carbon catabolite repression. Appl Microbiol Biotechnol 98:335–343
Kato N, Murakoshi Y, Kato M, Kobayashi T, Tsukagoshi N (2002a) Isomaltose formed by α-glucosidases triggers amylase induction in Aspergillus nidulans. Curr Genet 42:43–50
Kato N, Suyama S, Shirokane M, Kato M, Kobayashi T, Tsukagoshi N (2002b) Novel α-glucosidase from Aspergillus nidulans with strong transglycosylation activity. Appl Environ Microbiol 68:1250–1256
Kubodera T, Watanabe M, Yoshiuchi K, Yamashita N, Nishimura A, Nakai S, Gomi K, Hanamoto H (2003) Thiamine-regulated gene expression of Aspergillus oryzae thiA requires splicing of the intron containing a riboswitch-like domain in the 5′-UTR. FEBS Lett 555:516–520
Lott K, Cingolani G (2011) The importin β binding domain as a master regulator of nucleocytoplasmic transport. Biochim Biophys Acta 1813:1578–1592
Machida M, Yamada O, Gomi K (2008) Genomics of Aspergillus oryzae: learning from the history of koji mold and exploration of its future. DNA Res 15:173–183
Makita T, Katsuyama Y, Tani S, Suzuki H, Kato N, Todd RB, Hynes MJ, Tsukagoshi N, Kato M, Kobayashi T (2009) Inducer-dependent nuclear localization of a Zn(II)2Cys6 transcriptional activator, AmyR, in Aspergillus nidulans. Biosci Biotechnol Biochem 73:391–399
Marfori M, Mynott A, Ellis JJ, Mehdi AM, Saunders NF, Curmi PM, Forwood JK, Bodén M, Kobe B (2011) Molecular basis for specificity of nuclear import and prediction of nuclear localization. Biochim Biophys Acta 1813:1562–1577
Mizutani O, Masaki K, Gomi K, Iefuji H (2012) Modified Cre-loxP recombination in Aspergillus oryzae by direct introduction of Cre recombinase for marker gene rescue. Appl Environ Microbiol 78:4126–4133
Murakoshi Y, Makita T, Kato M, Kobayashi T (2012) Comparison and characterization of α-amylase inducers in Aspergillus nidulans based on nuclear localization of AmyR. Appl Microbiol Biotechnol 94:1629–1635
Nakamura T, Maeda Y, Tanoue N, Makita T, Kato M, Kobayashi T (2006) Expression profile of amylolytic genes in Aspergillus nidulans. Biosci Biotechnol Biochem 70:2363–2370
Noguchi Y, Tanaka H, Kanamaru K, Kato M, Kobayashi T (2011) Xylose triggers reversible phosphorylation of XlnR, the fungal transcriptional activator of xylanolytic and cellulolytic genes in Aspergillus oryzae. Biosci Biotechnol Biochem 75:953–959
Petersen KL, Lehmbeck J, Christensen J (1999) A new transcriptional activator for amylase genes in Aspergillus. Mol Gen Genet 262:668–676
Ran F, Bali M, Michels CA (2008) Hsp90/Hsp70 chaperone machine regulation of the Saccharomyces MAL-activator as determined in vivo using noninducible and constitutive mutant alleles. Genetics 179:331–343
Ran F, Gadura N, Michels CA (2010) Hsp90 cochaperone Aha1 is a negative regulator of the Saccharomyces MAL activator and acts early in the chaperone activation pathway. J Biol Chem 285:13850–13862
Sato H, Toyoshima Y, Shintani T, Gomi K (2011) Identification of potential cell wall component that allows Taka-amylase A adsorption in submerged cultures of Aspergillus oryzae. Appl Microbiol Biotechnol 92:961–969
Tamalampudi S, Talukder MM, Hama S, Tanino T, Suzuki Y, Kondo A, Fukuda H (2007) Development of recombinant Aspergillus oryzae whole-cell biocatalyst expressing lipase-encoding gene from Candida antarctica. Appl Microbiol Biotechnol 75:387–395
Tanaka M, Tokuoka M, Shintani T, Gomi K (2012) Transcripts of a heterologous gene encoding mite allergen Der f 7 are stabilized by codon optimization in Aspergillus oryzae. Appl Microbiol Biotechnol 96:1275–1282
Tani S, Katsuyama Y, Hayashi T, Suzuki H, Kato M, Gomi K, Kobayashi T, Tsukagoshi N (2001) Characterization of the amyR gene encoding a transcriptional activator for the amylase genes in Aspergillus nidulans. Curr Genet 39:10–15
Tonomura K, Suzuki H, Nakamura N, Kuraya K, Tanabe O (1961) On the inducers of α-amylase formation in Aspergillus oryzae. Agric Biol Chem 25:1–6
Tsuboi H, Koda A, Toda T, Minetoki T, Hirotsune M, Machida M (2005) Improvement of the Aspergillus oryzae enolase promoter (P-enoA) by the introduction of cis-element repeats. Biosci Biotechnol Biochem 69:206–208
Xu D, Farmer A, Chook YM (2010) Recognition of nuclear targeting signals by Karyopherin-β proteins. Curr Opin Struct Biol 20:782–790
Yamada O, Lee BR, Gomi K (1997) Transformation system for Aspergillus oryzae with double auxortophic mutations, niaD and sC. Biosci Biotechnol Biochem 61:1367–1369
Yang L, Ukil L, Osmani A, Nahm F, Davies J, De Souza CP, Dou X, Perez-Balaguer A, Osmani SA (2004) Rapid production of gene replacement constructs and generation of a green fluorescent protein-tagged centromeric marker in Aspergillus nidulans. Eukaryot Cell 3:1359–1362
Acknowledgments
We thank Tetsuo Kobayashi for providing the plasmids pAGAR-F and pAR-Zn3 and for technical advice about subcellular localization analysis. We also thank Yasuji Koyama and Osamu Mizutani for providing ∆malR::pyrG and ∆ligD::loxP pyrG − strains, respectively. This study was supported by Program for Promotion of Basic and Applied Researches for Innovations in Bio-oriented Industry, Science and Technology Research Promotion Program for Agriculture, Forestry, Fisheries and Food Industry, and JSPS KAKENHI (Grant No. 22248007, 25292044).
Author information
Authors and Affiliations
Corresponding author
Additional information
Kuta Suzuki and Mizuki Tanaka contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 338 kb)
Rights and permissions
About this article
Cite this article
Suzuki, K., Tanaka, M., Konno, Y. et al. Distinct mechanism of activation of two transcription factors, AmyR and MalR, involved in amylolytic enzyme production in Aspergillus oryzae . Appl Microbiol Biotechnol 99, 1805–1815 (2015). https://doi.org/10.1007/s00253-014-6264-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-014-6264-8