Abstract
This paper investigates a novel sulfur-oxidizing autotrophic denitrifying anaerobic fluidized bed membrane bioreactor (AnFB-MBR) that has the potential to overcome the limitations of conventional sulfur-oxidizing autotrophic denitrification systems. The AnFB-MBR produced consistent high-quality product water when fed by a synthetic groundwater with NO3 −–N ranging 25–80 mg/L and operated at hydraulic retention times of 0.5–5.0 h. A nitrate removal rate of up to 4.0 g NO3 −-N/Lreactord was attained by the bioreactor, which exceeded any reported removal capacity. The flux of AnFB-MBR was maintained in the range of 1.5–15 L m−2 h−1. Successful membrane cleaning was practiced with cleaning cycles of 35–81 days, which had no obvious effect on the AnFB-MBR performance. The 15 N-tracer analyses elucidated that nitrogen was converted into 15 N2-N and 15 N-biomass accounting for 88.1–93.1 % and 6.4–11.6 % of the total nitrogen produced, respectively. Only 0.3–0.5 % of removed nitrogen was in form of 15N2O-N in sulfur-oxidizing autotrophic denitrification process, reducing potential risks of a significant amount of N2O emissions. The sulfur-oxidizing autotrophic denitrifying bacterial consortium was composed mainly of bacteria from Proteobacteria, Chlorobi, and Chloroflexi phyla, with genera Thiobacillus, Sulfurimonas, and Ignavibacteriales dominating the consortium. The pyrosequencing assays also suggested that the stable microbial communities corresponded to the elevated performance of the AnFB-MBR. Overall, this research described relatively high nitrate removal, acceptable flux, indicating future potential for the technology in practice.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Nitrate contamination of groundwater is now widespread due to the increased use of synthetic nitrogen fertilizers, concentrated animal feeding operations, and industrial waste effluent discharge (Puckett 1995). A high level of nitrate in drinking water, producing nitrite in the human gut, can cause methemoglobinemia or blue baby syndrome infants and gastrointestinal cancer in adults (Ward et al. 2005). Nitrate pollution is widely considered to be one of the main threats to groundwater quality, requiring urgent and intensive monitoring and strong policies. The United States Environmental Protection Agency set the maximum allowable concentrations of 10 mg NO3 −–N L−1 and 1.0 mg NO2 −–N L−1 for drinking water (Lee and Rittmann 2002).
Groundwater polluted with nitrates contains almost no organic matter. Thus, autotrophic sulfur-oxidizing denitrification is considered to be an alternative process for nitrate removal from drinking water (Sierra-Alvarez, et al. 2007; Puig et al. 2012; Sahinkaya and Kilic 2014). In such systems, sulfur and nitrate act as an electron donor and an acceptor, respectively (reaction 1). This process has four major advantages: (1) Inorganic substances are utilized as electron donors, eliminating potential problems associated with residual organics; (2) external organic carbon is not necessary, which reduces the cost and risk of operation and the release of carbon dioxide; (3) lower cell yield reduces microbial contamination; and (4) autotrophic sulfur-oxidizing denitrifying bacteria produce less nitrous oxide (N2O) (a greenhouse gas) than heterotrophic denitrifying bacteria. Elemental sulfur is water-insoluble and stable under normal conditions and could be physically removed from effluents for reuse. Thus, elemental sulfur is an attractive source of energy for autotrophic denitrification of nitrate-contaminated groundwater (Moon et al. 2008; Sahinkaya et al. 2011).
A number of researches have used reduced sulfur compounds for biological denitrification of ground water (Zhang and Zeng 2006; Sierra-Alvarez et al. 2007; Sahinkaya et al. 2011; Sun and Nemati 2012). These were dominantly packed bed processes and were often subject to mass transfer problems associated with channeling (caused by clogging) and nitrite accumulation due to biofilm control issues. Post-treatment is generally required in these systems to remove sloughed biomass from the product water, resulting in high capital investment. Additionally, the full-scale applications of these packed processes are hampered by the low nitrate-loading rates. Membrane bioreactors (MBRs), which can facilitate complete retention of the biomass, offer a more integrated approach, although development for drinking water application is still in its infancy and only limited research has been conducted (McAdam and Judd 2007). Liao et al. (2006) recommended more research on combining membranes with existing high-rate reactor configurations already determined to be suitable for dilute wastewaters. For this purpose, the anaerobic fluidized bed bioreactor is proposed to be particularly advantageous due to good mass transfer characteristics. Kim et al. (2011) evaluated the performance of anaerobic fluidized bed MBR for low-strength wastewater, which are capable of achieving high effluent quality. Laboratory-scale extractive and diffusive MBRs have been evaluated on heterotrophic denitrification of drinking water (Ergas and Rheinheimer 2004; McAdam and Judd 2006). However, to our knowledge, elemental sulfur-oxidizing autotrophic denitrifying AnFB-MBR is not reported for treatment of nitrate-contaminated drinking water so far.
For sulfur-oxidizing autotrophic denitrification processes, a significant research effort has been devoted to their application in environment technology. Previous authors have investigated process design criteria such as nitrate loading rate (Zhang and Zeng 2006), hydraulic residence time (Ahmed et al. 2012), pH and alkalinity (Kim and Bae 2000), sulfur particle size (Koenig and Liu 2001), sulfur/limestone ratio (Zhang and Lampe 1999), and addition of supplemental organic substances (Sahinkaya et al. 2011). Some studies focused on fundamental issues such as kinetics, stoichiometric relationships, modeling, etc. (Zeng and Zhang 2005; Cardoso et al. 2006; Sierra-Alvarez et al. 2007). However, knowledge of bacterial communities contributing to sulfur-oxidizing autotrophic denitrification is still very limited (Koenig et al. 2005; Fernández et al. 2008; Moon et al. 2010). The underlying communities of autotrophic denitrifying bacteria and their response to environmental conditions need to be further understood.
Some culture-independent molecular ecology techniques such as fluorescence in situ hybridization, denaturing gradient gel electrophoresis, genetic libraries, and quantitative real-time polymerase chain reaction (PCR) have been applied in a number of studies concerned with the heterotrophic denitrifying communities in wastewater treatment systems (Throbäk et al. 2004; Hallin et al. 2006; Warneke et al. 2011). In addition, barcoded pyrosequencing method was recently used for analyzing bacterial community structural changes of these key denitrifying genes (Mao et al. 2011; Zhao et al. 2011). In contrast, the detailed investigation about microbial diversity and functional genes of sulfur-oxidizing autotrophic denitrifying biofilm using the above molecular ecology techniques has often been ignored.
This paper develops a novel sulfur-oxidizing autotrophic denitrifying AnFB-MBR system, which integrates membranes and an elemental sulfur-based autotrophic denitrifying anaerobic fluidized bed for treatment of nitrate-contaminated groundwater. This goal of this study is to systematically evaluate the denitrification performance of the novel sulfur-based AnFB-MBR system, investigate the N mass balance and kinetics of denitrification, and demonstrate the feasibility of autotrophic denitrification by the sulfur-based AnFB-MBR system. An additional objective is to determine the microbial ecology of the sulfur-oxidizing autotrophic denitrification. We monitored changes in the microbial community according to specific group: total bacteria by 16S RNA gene and denitrifiers by the functional genes nirK, nirS, and nosZ. We also analyzed bacterial communities in the sulfur-based AnFB-MBR through pyrosequencing to better understand of the “black-box” of sulfur-oxidizing autotrophic denitrification, and result in better reactor operation and control.
Materials and methods
Experimental setup
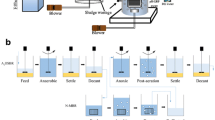
The setup of the AnFB-MBR system is shown in Fig. 1. The AnFB-MBR was a 2.3-L continuous-flow system consisting of an acrylic column and a submerged self-made membrane module. The internal diameter of the column reactor was 10 cm and the height, 40 cm. The membrane module contained 200 polyvinylidenefluoride ultrafiltration hollow-fiber membranes (2.2 mm inner diameter, nominal pore size of 0.1 μm, and representing a total membrane surface area of 0.3768 m2; provided by Xiamen Kunpeng, Ltd., China), each with a 0.3-m active length. 200 g of sulfur particle (50–200 μm) was initially added as source of electron donor of sulfur-based autotrophic denitrification. Additional 100 g of the same sulfur particle was supplemented each 60-day operation to maintain a relatively stable sulfur content in the AnFB-MBR. An overhead stirrer with a rotating speed of 450 rpm was used to maintain fluidization of the sulfur particle. A peristaltic pump (BT300-2 J, Baoding Longer Precision Pump Co., Ltd., China) was used to deliver the feed solution to the reactor. A liquid level indicator was connected to the peristaltic pump to maintain a constant water level at the top of the AnFB-MBR. The same kind peristaltic pump (as above) was connected to the top open sections of the membrane fibers in the AnFB-MBR to achieve the effluent. The transmembrane pressure (TMP) required to maintain flow was monitored with a vacuum pressure meter. The AnFB-MBR was operated at 28 ± 3 °C.
The feed solution for the system was tap water amended with a concentrated stock solution of KNO3, NaHCO3, phosphate buffer (Na2HPO4 + KH2PO4), and trace element solution. The purpose of adding NaHCO3 was to ensure an ample supply of inorganic carbon for autotrophic growth, while the phosphate and trace element solution were added to ensure sufficient nutrient for bacterial growth. The influent concentrations of NO3 −-N were systematically changed from 25 to 80 mg/L according to the operational stages. The mineral salts utilized in all experiments were composed of (milligrams per liter): NaH2PO4 · 2H2O (150), K2HPO4 · 2H2O (220), NaHCO3 (500), MgCl2 · 7H2O (5), CaCl2 (1), and trace element solution supplied at 0.1 mL/L. The trace element solution contained (milligrams per liter): MnCl2 (64), CoCl2 · 7H2O (200), (NH4)6Mo7O24 · 4H2O (55), ZnCl2 (70), NiCl2 · 6H2O (24), CuSO4 (2), FeSO4 · 7H2O (500), H3BO3 (6), EDTA (500). The pH was adjusted to 7.5–7.7 with sodium hydroxide. N2 sparge system was used to exclude oxygen from the feed solution.
Continuous experiments
Different periods of operation can be distinguished based on the empty bed hydraulic retention time (HRT) and the influent nitrate concentration maintained in the reactor (Table S1). Each steady state had a duration of 5–10 days.
The conversion of substrates was monitored periodically by measuring the influent concentration of nitrate: the concentration of nitrate, nitrite, and sulfate in the effluent. Fresh effluent samples were collected from the effluent line.
Batch experiments
The conversion of NO3 −–N and N-balance of sulfur-oxidizing autotrophic denitrifying process were elucidated with a 15 N-tracer technique. The bath assays were conducted in glass serum flasks (500 mL) supplemented with 200 mL medium lacking NH4 +-N. 15 N-NO3 − expresses the nitrogen isotope 15 N of the nitrate (NO3 −) as a sole nitrogen substrate. The mineral medium (pH 7.5) contained variable concentrations of 15 N-labelling potassium nitrate (K15NO3) with 99.7 atom% 15 N. The experiments were inoculated with 0.20 g SS/L of the denitrifying culture and elemental sulfur particles from the AnFB-MBR. The headspace was flushed with high-purity helium to exclude oxygen and background nitrogen from the assays. All flasks were sealed with butyl rubber stoppers and aluminum crimp seals. Controls lacking inoculum were run in parallel to monitor the possible abiotic degradation of the electron donor and electron acceptor. All bioassays were carried out in duplicate in an orbital shaker (150 rpm).
Headspace samples in the bioassays were analyzed for 15 N2 and 15N2O content at regular intervals throughout the experiment. Liquid samples were withdrawn periodically to monitor the conversion of substrates and the formation of degradation products. Solid samples were finally withdrawn to determine the nitrogen converted into 15 N-biomass. The N-balance of sulfur-oxidizing autotrophic denitrification was determined by measuring the amounts of consumed 15NO3 −–N that were recovered as 15 N-N2, 15 N-N2O, and 15 N-biomass.
Cleaning protocol
When the AnFB-MBR was continuously operated until the TMP was increased by 0.3 bar, the membrane module was taken out from the bioreactor and replaced with a clean membrane. The fouled membrane was soaked in 0.5 % NaOCl solution overnight and followed by thorough flushing with deionized water.
Analytical measurements
Nitrate, nitrite, and sulfate were determined by ion chromatography (IC) (Dionex ICS 2000). The IC had an AS19 column (4 × 250 mm) and AG 19 precolumn (4 × 50 mm), an eluent of 30 mM KOH, and an eluent flow rate of 1 mL/min. The injection volume was 20 μL. Liquid samples were membrane-filtered (0.2 μm) prior to chromatographic analysis. The retention time of NO3 −, NO2 −, and SO4 2− were 10.2, 6.3, and 7.9 min, respectively.
Dinitrogen (N2) content was determined by gas chromatography (GC) using a 6890 N system (Agilent Technologies, USA) equipped with thermal conductivity detection. The gas chromatograph was fitted with a stainless steel packed column (2 m × 3 mm, Porapak T, 80-100 mesh). The temperature of the column, the injector port, and the thermal conductivity detector were 60 ℃, 110 ℃, and 100 °C, respectively. The carrier gas was helium at a flow rate of 19.1 mL/min. Headspace samples (150 μL) was collected using a pressure-lock gas syringe. Nitrous oxide (N2O) was measured by GC analysis of the headspace. The 20-mL headspace sample was injected into a SP3410 gas chromatograph (Beijing Beifen-Ruili Analytical Instrument Co. Ltd., China) with a conductivity detector and a stainless steel packed column (2 m × 3 mm, Porapak T, 80-100 mesh). High-purity nitrogen at a flow rate of 30 mL/min was the carrier gas. The column and detector temperature were both set at 72 °C.
The 15 N fractions of 15 N-labeled N2 and N2O were determined by a Delta V Advantage Isotope Ratio Mass Spectrometer (IRMS, Thermo Fisher Scientific, Inc., USA) in the Stable Isotope Ratio Mass Spectrometry Laboratory, Chinese Academy of Forestry Sciences. A gas sample is flushed from a vial by a He carrier stream. Carbon dioxide, NO, and water are chemically removed prior to chromatographic separation of N2 from N2O. The potential contribution to N2 and N2O formation of NO3 − denitrification was calculated from the produced 29 N2 and 30 N2. For analysis of 15 N fractions in the biomass solid, aliquots of solid biomass were weighed into tin capsules. Samples were combusted at 1,020 °C with excess oxygen in an elemental analyzer (Thermo Fisher Scientific, Inc., USA), and the resulting combustion gases passed through a reduction furnace at 650 °C. After removal of water with magnesium perchlorate and purification in a gas chromatographic column, N2 was measured with the above IRMS. Nitrogen isotope ratios are reported in the conventional delta notation with respect to atmospheric N2 (AIR). The analytical reproducibility is 0.2‰ for δ15N.
DNA extraction
The sludge samples including elemental sulfur were collected for all stages when the reactor reached a steady state in terms of different NO3 − concentrations. The DNA was extracted according to the procedures described in the Powersoil® DNA Isolation kit (MOBIO laboratories, Inc.). All samples were extracted in duplicate. The DNA concentrations were measured with a UV-vis spectrophotometer (NanoDrop Technologies, USA), and its yield and purity was documented based on the 260/280 nm absorbance ratio. Then the samples were stored in −80 freezer for the following molecular applications.
Real-time PCR
Quantitative real-time PCR was used to enumerate nirS, nirK, nosZ, and 16S rRNA genes of sulfur-oxidizing denitrifying bacteria. Quantification was performed in triplicate on an ABI Prism 7300 sequence detection system (Life Technologies, Carlsbad, CA) according to the manufacturer’s instructions using SYBR-green based detection. The 20-μL reaction mixture contained 10 μL of 2 × SYBR Premix EX TaqTM (Takara Bio Inc., Japan), 0.4 μL of Rox Reference Dye (50×), 0.4 μL of each primer (10 μM), and 5 ng of DNA template, 7.8 μL ddH2O. The nirS, nirK, nosZ, and 16S rRNA genes were PCR-amplified from extracted DNA with the primers and conditions listed in Table S2.
Following amplification, dissociation curves were determined across a temperature range of 85.7–88.0 °C. Ct values for each well were calculated using the manufacturer’s software. The PCR products were then cloned into the pGEM-T Easy Vector (Promega Madison, USA) to construct plasmids containing target fragments. Plasmids used for standards for quantitative analyses were extracted from the correct insert clones of each target gene. The concentration of plasmid DNA was determined on a Nanodrop® ND-1000 UV-vis Spectrophotometer (NanoDrop Technologies, USA). The standard curves based on serial dilutions containing between 1010 and 104 target gene copies were generated. The copy numbers of the sludge samples were calculated by comparison to standard curves. Negative controls with sterilized water instead of DNA template were included in every PCR setup. The triplicate PCR reactions were performed for all samples and negative controls.
Barcoded pyrosequencing
To analyze the taxonomic composition of bacterial communities contributing to sulfur-oxidizing autotrophic denitrification, the V3-V1 region of the 16S rRNA gene was chosen for the amplification and subsequent pyrosequencing of the PCR product. The 16S rRNA genes were amplified using the primers 27 F/533R. The forward primer contained the A adapter (CCA TCT CAT CCC TGC GTG TCT CCG AC), and the reverse primer contained the B adapter (CCT ATC CCC TGT GTG CCT TGG CAG TC). Amplicon pyrosequencing was performed on a 454 GS FLX Titanium platform (Roche Inc., USA) (Majorbio Bio-Pharm Technology Co., Ltd., Shanghai, China) according to standard protocols.
Raw pyrosequencing data that were obtained from this study were deposited to the NCBI Sequence Read Archive with accession no. SRP045436. Sequences were first extracted from the raw data according to the Genome Sequencer Data Analysis Software Manual (Software Version 2.0.00, October 2008). To minimize the effects of random sequencing errors, low-quality sequences were removed from the analysis if they were <200 bp, contained ambiguous characters, had homopolymers longer than 8 bp, more than one MID mismatch, or more than two mismatches to the reverse primer sequence. The sequences that matched with the reverse primer were converted to their reverse complement counterparts using BioEdit to make all the sequences forward-oriented. Finally, pyrosequencing produced 5246 (S30), 4982 (S50), and 6045 (S80) high-quality V1–V3 tags of the 16S rRNA gene with an average length of 483 bp.
The 16S rRNA gene sequences were aligned by NAST (Greengenes). Operational taxonomic units (OUT) were obtained from the quality sequences using the MOTHUR program by setting a 0.03 distance limit (equivalent to 97 % similarity). Rarefaction curves, Shannon diversity index (http://www.mothur.org/wiki/Shannon), and species richness estimator of Chao1 (http://www.mothur.org/wiki/Chao) were generated in MOTHUR for each sample. Sequences were classified using the Greengenes database at an 80 % confidence threshold with MOTHUR. After phylogenetic allocation of the sequences down to the phylum, class, and genus levels, relative abundance of a given phylogenetic group was set as the number of sequences affiliated with that group divided by the total number of sequences per sample.
Results
Reactor performance
To assess long-term treatment performance, the AnFB-MBR system was operated for around 170 days treating simulated nitrate-contaminated groundwater. The whole operation was divided into five periods based on the HRT. The HRT was decreased from 5.0 h in period I to 0.5 h in period V. For each operational period, the concentration of nitrate in the bioreactor feed was increased stepwise from around 25 to 80 mg NO3 −-N/L. The performance of the bioreactor as a function of time is illustrated in Fig. 2.
Inoculation of the bioreactor with the sulfur-oxidizing denitrifying enrichment culture permitted the rapid start-up of the sulfur-based denitrification process. Complete removal of nitrate was observed during the first 30 days (period Ι) when the reactor treated an influent with 25–80 mg/L NO3 −-N at a HRT of 5.0 h. Nitrate was also removed to non-detectable levels during the other periods (days 31–170), although the HRT was decreased stepwise from 5 to 0.5 h. High nitrate removal efficiency (100 %) was maintained throughout the operation. Nitrite accumulation has been shown to be reliable marker of reactor overloading in sulfur-based denitrification processes (Flere and Zhang 1999). In periods Ι–ΙΙΙ, nitrite was not detected in the effluent. On day 110, the influent NO3 −-N concentration was increased to 80 mg/L at a HRT of 1 h. The effluent nitrite concentration increased briefly then returned to <non-detectable levels. During the final part of phase V, nitrite was detected at concentrations of up to 1.5–4.0 mg/L when the influent NO3 −-N concentration was increased to around 80 mg/L at a HRT of 0.5 h. However, the effluent NO2 −-N concentration decreased to <the regulatory levels (0.07 mM or 1 mg/L NO2 −-N) about 10 days after the incident. These results suggested that the reactor was operated below its maximum loading capacity, especially at relatively low nitrate concentration (40 mg/L NO3 −-N or lower). However, in the present study, the HRT was not further decreased to a shorter time because of the constraints of membrane flux. Further decrease in HRT may be accomplished by increasing the number of membrane fibers.
Sulfate was the only product of the sulfur oxidation detected in the effluent during the denitrification course of the study. The sulfate production could be taken as an indicator for the denitrification. The observed sulfate concentrations were in good agreement with the theoretical sulfate production calculated according to the stoichiometry of the complete sulfur-oxidizing denitrification (Fig. 3), indicating that nitrate was completely reduced for all tested concentrations. However, sulfate production is the main disadvantage of sulfur autotrophic denitrification process. As shown in Fig. 2, only when presenting a low nitrate concentration of less than 30 mg /L NO3 −-N, the produced sulfate was below limit value of 250 mg/L. Theoretically, around 33 mg/L NO3 −-N could be removed using sulfur-oxidizing autotrophic denitrification without exceeding sulfate regulatory level of 250 mg/L, set by US EPA. Thus, sulfur-oxidizing autotrophic denitrification process was especially feasible for treating contaminated groundwater with low nitrate concentrations (<30 mg/L NO3 −-N). For the waters with higher nitrate, some additional strategies should be considered to control the sulfate formation.
Membrane performance
Fresh membrane was installed at the beginning of the experiments. Performance efficiency of the membrane module was assessed by monitoring flux and transmembrane pressure (TMP). Results of flux and TMP analyses for long-term operation are illustrated in Fig. 4. During the whole operation the flux was maintained between 1.5 and 15 L m−2 h−1, and TMP remained in the range of 0.03–0.30 bar at the HRTs of 0.5-5.0 h. In periods Ι and ΙΙ (days 1–54), the AnFB-MBR did not show evidence of serious membrane fouling as the TMP remained at 0.03–0.08 bar under the low flux of 1.5–2.5 L m−2 h−1. This result suggests that the AnFB-MBR is being operated at low flow conditions, which is expected to maintain a reliable flux before irreversible fouling occurs. From day 55 on, the flux was increased to around 3.8 L m−2 h−1 by reducing HRT to 2 h. The TMP increased gradually to 0.3 bar on day 81, and then the ex situ chemical cleaning of the membrane was carried out to avoid irreversible fouling. In periods IV (days 82–120), a flux of 7.5 L m−2 h−1 was applied at a HRT of 1.0 h. The AnFB-MBR was continuously for 43 days when the TMP increased to >0.3 bar. During the operation of phase V, the HRT was further decreased to 0.5 h, resulting in a higher flux up to 15 L m−2 h−1. The boundary of membrane cleaning of 0.3 bar was reached in 35 day for this phase, indicating a higher flux led to a shorter fouling cycle. It has generally observed that the duration of the slow fouling rate decrease when fluxes approach the critical flux (Guglielmi et al. 2007).
15 N-labelling denitrification process
[15 N] nitrate-labelling experiments were conducted to investigate nitrate utilization and product formation by mixed liquor of the bioreactor. The 15NO3 −-N concentrations (30–60 mg/L 15NO3 −-N) are in the range of those applied in the continuous sulfur-based bioreactor. Figure 5 shows the conversion of 15NO3 −-N, the evolution of 15NO2 −-N and 15N2O−-N, and the generation of 15 N2-N in the bioassays, respectively, as function of time. The nitrate was completely consumed within 5–8 h at concentrations of nitrate ranging from 30 to 60 mg/L 15NO3 −-N. Nitrite and nitrous oxide were detected as the intermediate degradation products during the sulfur-oxidizing denitrification process. Maximum of nitrite was accumulated from nitrate reduction in the first 3–6.5 h and completely removed afterward. Nitrite was sequentially reduced via nitric oxide (NO) and N2O to N2 during denitrification. NO gas could not be detected in this experiment. However, as shown in Fig. 5, N2O was another main intermediate except for nitrite during the sulfur-oxidizing denitrification. Peak 15N2O-N concentrations (0.088–0.123 mM) were observed in the first 4–8 h, and thereafter decreased steadily to <131 ppb after 16–26 h. N2 gas tended to increase steadily following the complete denitrification of nitrate. The final 15 N-N2 concentrations achieved were 0.409, 0.525, 0.670, and 0.783 mM for different initial 15NO3 −-N concentrations of 30, 40, 50, and 60 mg/L, respectively. The culture was stopped when no more N2 was released, which coincided with the total consumption of nitrate.
Nitrogen balances were determined based on the above tests for further elucidating the removal mechanisms of sulfur-oxidizing autotrophic denitrification. Table 1 summarizes the 15NO3 −-N consumed, the amounts of 15 N2-N and 15N2O-N released, and the 15 N-biomass formed. The nitrogen recovery as 15 N2-N, 15N2O-N, and 15 N-biomass were calculated by taking into account the elementary compositions of the substrate and products. A nitrogen recovery of 98.9–105.6 % was obtained for nitrate denitrification. Nitrogen converted into 15 N2-N and 15 N-biomass accounted for 88.1–93.1 % and 6.4–11.6 % of the total nitrogen produced, respectively. These results indicated that nitrate was completely reduced to N2 during the sulfur-oxidizing autotrophic denitrification. Meanwhile, the 15NO3 −-N was immobilized as 15 N-biomass, implying nitrate as a sole nitrogen source supported microbial growth. N2O was a main intermediate product during denitrification. Recently, N2O generation and emission have been paid more attention because N2O is a potent greenhouse gas and major ozone-depleting substance in the atmosphere (Zhu and Chen 2011; Pan et al. 2012). In the present study, only 0.3–0.6 % of removed nitrogen was in form of 15N2O-N, i.e., <131 ppb of N2O was released from the sulfur-oxidizing autotrophic denitrification.
Functional gene abundance
Figure 6a shows the 16S rRNA, nirS, nirK, and nosZ gene copy numbers during the enrichment culture of activated sludge. The 16S rRNA gene continuously decreased through the enrichment culture stage, indicating a decrease of microbial biomass. The autotrophic sulfur-oxidizing denitrifiers were cultured and enriched during the acclimatization period. However, the autotrophic bacteria growing on inorganic carbons had a lower growth rate compared with the heterotrophic bacteria (from the activated sludge), resulting in a decreasing microbial biomass with the acclimatization of the inocula. The copy numbers of the nirS and nosZ gradually increased from 6.92 × 1011 and 1.48 × 109 to 7.94 × 1013 and 6.31 × 1010 copies/g SS at the end of the acclimatization period. Presumably, the increases represented accumulation of more autotrophic sulfur-oxidizing denitrifiers over time. The copy numbers of the nirK gene (copper-based NO2-reductase) did not change significantly, probably because nirK was not a good marker for denitrifiers in the autotrophic sulfur-oxidizing denitrifying biomass.
Figure 6b summarizes the real-time PCR results as a function of different NO3 − removal flux during the stable operation. Overall, the abundance of nirS and nosZ tended to increase when NO3 − removal flux increased, suggesting the nirS and nosZ genes possibly tracked the NO3 − removal capacity of the bioreactor. However, during the stable operation, the detected genes indicative of nirS outnumbered genes indicative nirK by approximately 104–107 orders of magnitude, further implying nirS-type denitrifiers are possibly are important for the autotrophic sulfur-oxidizing denitrification process.
Bacterial communities
We used pyrosequencing to analyze the bacterial communities of the autotrophic sulfur-oxidizing denitrifying microbial biomass for dominant phyla, classes, and genera. Pyrosequencing yielded a total of 16,870 high-quality sequences for the 16S rRNA gene for all samples, and Fig. S1 in the supporting information shows the rarefaction curves based on an OUT definition of 97 % sequence similarity. Figure 7, which summarizes bacterial diversity at the phyla, class, and genus levels, shows that the dominant bacterial structures in the bioreactor for steady state at the different influent nitrate concentrations of 30 (S30), 50 (S50), and 80 mg/L NO3 −-N (S80) were relatively stable. In total, eight identified phyla in each sample were detected (Fig. 7a). Proteobacteria was by far the largest phyla in the bioreactor (73.1 %, 67.3 %, and 72.2 % for S30, S50, and S80, respectively). Chlorobi was the second abundant phyla, which was approximately 20.4 % (S30), 22.3 % (S50), and 22.6 % (S80) of the total reads. The sum of the two largest phyla accounted for over 90 % for all the three samples. Chloroflexi as the third largest phyla, whose relative abundance in S30, S50, and S80 was 3.2 %, 4.2 %, and 3.4 %, respectively.
The class level identification of the bacterial communities in S30, S50, and S80 is illustrated in Fig. 7b. Pyrosequencing detected 11 bacterial classes in all the three samples. The majority of sequences belonged to Betaproteobacteria, Chlorobi, and Epsilonproteobacteria. Betaproteobacteria was the dominant classes, with the percentage of 60.9 %, 56.1 %, and 54.2 % for S30, S50, and S80, respectively. Chlorobi and Epsilonproteobacteria were the second and third dominant classes, whose abundances were in the ranges of 20.4–22.6 % and 10.9–16.5 %, respectively. About 8.8 % (S30), 9.8 % (S50), and 6.9 % (S80) of the total reads in each sample were not classified at the class level, indicating that these bacteria are unknown.
Figure 7c shows the relative proportions of the most abundant bacterial genera for the steady sate at the different influent NO3 −-N concentrations. Clearly, the AFB-MBR had similar dominant bacterial genera, although the influent NO3 −-N concentrations were gradually increased from 30 to 80 mg/L during the stable operation. The most numerous genera were Thiobacillus (58.6 %, 52.7 %, and 51.4 %), Ignavibacteriales (18.9 %, 20.6 %, and 20.7 %), and Sulfurimonas (10.7 %, 10.9 %, and 16.3 %), whose sum accounted for 88.2 %, 84 %, and 88.4 % for S30, S50, and S80, respectively. The unclassified bacteria at a genus class accounted for 1.2 % (S30), 1.74 % (S50), and 1.1 % (S80) of the total reads.
Discussion
Sulfur-oxidizing autotrophic denitrification has been mostly used for drinking production from nitrate-contaminated ground water. Most of the autotrophic denitrification applications were carried out through sulfur-packed bed reactor (Zhang and Zeng 2006; Sierra-Alvarez et al. 2007; Wan 2009; Sahinkaya et al. 2011; Sun and Nemati 2012). In the current study, a sulfur-oxidizing autotrophic denitrifying AnFB-MBR was developed and specifically tested for groundwater denitrification. The results obtained from this study and those reported by other authors are summarized in Table 2. The nitrate removal rates obtained in our study was up to 1.25–4.0 g NO3 −-N/Lreactord when the bioreactor treated 25–80 mg/L NO3 −-N at a HRT of 0.5 h. The HRT attained in our study during the treatment of 25–80 mg/L NO3 −-N, 0.5 h, is relatively low compared with literature studies. It should be noted from Fig. 2 that the removal capacity of the AnFB-MBR may be further enhanced by increasing the number of membrane fibers through the optimization of module and reactor configurations. Therefore, it could be deduced that the AnFB-MBR system in this study greatly exceeded any reported removal capacity. The relatively high nitrate loading rates were achieved in the present study because it led to nearly absolute biomass retention and had good mass transfer characteristics. Also, the greater nitrate removal may result from the greater working volume and more efficient sulfur utilization of the AnFB-MBR compared with the packed bed reactors. Finally, in contrast to all other packed beds and conventional biological systems, the AnFB-MBR proved to have a good potential as a sulfur-oxidizing autotrophic denitrification technology also because it prevents secondary contamination of drinking water by bacteria, which was emitted with the treated water in the conventional biological processes. In fact, the AnFB-MBR has been determined to been suitable for dilute wastewater or wastewater treatment (Liao et al. 2006; Kim et al. 2011).
The maximum flux of AnFB-MBR achieved was higher than those between 5 and 10 L m−2 h−1 observed in anaerobic membrane bioreactors (Ho and Sung 2010; Spagni et al. 2010) but lower than those typically reported in aerobic MBRs, being between 20 and 25 L m−2 h−1 (Judd 2002). Anaerobic membrane bioreactors have potential for energy-efficient treatment of wastewater treatment, membrane fouling being a major hurdle to application (Kim et al. 2011). In this current case, the AnFB-MBR maintained a reliable flux before irreversible fouling occurs with a frequent ex situ chemical cleaning. The membrane cleaning cycles required were 35–81 days under the different flux at the HRTs of 0.5–5 h when no automated backwash was performed in the membrane. The autotrophic sulfur-oxidizing denitrifying bacteria having low cell yields resulted in a low biomass concentration, which might prolong membrane operation. In addition, the use of sulfur particles as a carrier may be beneficial for enhancing membrane performance. The positive effects of carries on membrane fouling were also reported previously by other authors in MBR systems (Liu et al. 2010; Kim et al. 2011). These results suggest that the application of AnFB-MBR appears to be feasible for treatment of nitrate-contaminated groundwater if an appropriate fouling control protocol is applied.
The NO3 −-N conversion and N-balance of sulfur-oxidizing autotrophic denitrifying process were elucidated with a 15 N-tracer technique. These results indicated that nitrate was completely reduced to N2 during the sulfur-oxidizing autotrophic denitrification. Meanwhile, the 15NO3 −-N was immobilized as 15 N-biomass, implying nitrate as a sole nitrogen source supported microbial growth. Total 15N2O-N release of 0.3–0.6 % of removed 15NO3 −-N was yielded in the sulfur-oxidizing autotrophic denitrifying processes, indicating a negligible release of N2O. However, heterotrophic denitrification is implicated as a potential source of global N2O emission during biological nitrogen removal operations (Lu and Chandran 2010; Wunderlin et al. 2012). Autotrophic sulfur-oxidizing denitrifying bacteria were considered to produce less N2O than heterotrophic denitrifying bacteria (Park et al. 2002). It implied that the sulfur-oxidizing autotrophic denitrifying process produced less N2O than the conventional heterotrophic denitrifying systems, which greatly reduced the potential risks caused by a significant amount of N2O emissions.
In this study, except for the AnFB-MBR performance, we also characterize the bacterial phylogenetic membership of sulfur-oxidizing autotrophic denitrifiers during the stable operation period. The traditional approaches (e.g., clone library, DGGE, and T-RFLP) used in previous studies may largely underestimate the diversity of microbial communities involved in sulfur-oxidizing autotrophic denitrification. Thus, high-throughput deep-sequencing approaches are essential to improve our knowledge of the diversity of these microbial communities. The rarefaction curves of these genes were close to saturation after sequencing ∼5,000 for each sample, indicating that such a sequencing effort is sufficient to elucidate the diversity and structure of the microbial communities (Fig. S1). Pyrosequencing results confirmed the dominance of Proteobacteria, Chlorobi, and Chloroflexi in our reactors, signifying that future research dedicated to the functional importance of these bacteria is warranted. The enrichment of Proteobacteria, Chlorobi, and Chloroflexi has been detected in previous studies characterizing denitrification and even chemolithotrophic denitrification (Knowles 1982; Koenig et al. 2005; Fernández et al. 2008). Compared with the Proteobacteria and Chlorobi, much less is known regarding the role of Chloroflexi in denitrification systems. In our reactor system, members of the Chloroflexi accounted for 3.2–4.2 % of the dominant community in the samples. Despite their abundance in molecular surveys, knowledge about Chloroflexi physiology is scarce (Krangelund et al. 2007), yet isolated members of the Chloroflexi have been shown to reduce nitrate to nitrite (Kohno et al. 2002).
From major subdivisions, our AnFB-MBRs were mainly enriched in classes Betaproteobacteria, Chlorobi, and Epsilonproteobacteria. This finding was slight different from 16S rDNA-based DGGE results conducted on autosulfurotrophic denitrifying biofilm, which revealed the enrichment related to four subdivisions (α, β, γ, and δ) of the kingdom Proteobacteria and from the kingdom Chlorobia (green sulfur bacteria) (Koenig et al. 2005). However, a functional role for members of Betaproteobacteria in cathodic denitrifying biofilms has been suggested by Wrighton et al. (2010).
Standing on the genus level allows us to infer the functions of the communities, based on the assumption that members that are closely related phylogenetically share similar metabolic capabilities. In this sulfur-oxidizing autotrophic denitrification system, Thiobacillus (51.4–58.6 %), Sulfurimonas (18.9–20.7 %), and Ignavibacteriales (18.9–20.7 %) formed a dominant community structure. Thiobacillus was the first autotrophic denitrifier, whose autotrophic denitrification may outcompete its heterotrophic counterpart and play a dominant role in nitrogen cycle in certain ecosystems (Brettar and Rheinheimer 1991). Up to date, Thiobacillus and Sulfurimonas were the most commonly reported autotrophic denitrifiers, which reduced nitrate to N2 while oxidizing elemental sulfur or reduced sulfur compounds to sulfate (Koenig et al. 2005; Shao et al. 2010). Ignavibacteriales, as the second dominant genus, seemed to be associated with the sulfur-based autotrophic denitrification. However, Ignavibacteriales as a dominant species has not been reported in the sulfur-based autotrophic denitrifying process to date. Ignavibacteriales was previously reported to be collected from a sulfide-rich hot spring at Yumata, Nagano, Japan, and was identified as a member of Chlorobi phyla (Takao et al. 2010). Thus, future research dedicated to the functional importance of the novel genus Ignavibacteriales needs to be warranted.
Our study also evaluated differences in community structure (richness and abundance) in response to performance. These findings also suggest that the bacterial community may have implications on the functional performance of the AnFB-MBR. The great stability in the community structure resulted in a great stability and high-efficiency performance of the reactor, although influent nitrate was increased from 25 to 80 mg NO3 −-N/L. These findings show, for the first time in the AFB-MBR, potential link between community structure and function. Further studies elucidating the relationships between bacterial OUT richness and evenness, phylogenetic diversity, and system performance are necessary to optimize the bacterial component of these systems.
In summary, AnFB-MBR is an effective and innovative technology for the treatment of nitrate from drinking water with the superiority of the performance compared with conventional packed bed processes while concurrently having nearly absolute biomass retention and avoiding sloughed biomass from the product water. The 15 N-tracer analyses demonstrated 15NO3 −-N was recovered mainly as 15 N2-N and 15 N-biomass in the sulfur-oxidizing autotrophic denitrification. N2O as a main intermediate was further reduced to N2, implying this process reduced potential risks of a significant amount of N2O emissions. Our results provided the detailed characterization of bacterial communities in sulfur-oxidizing autotrophic denitrifying system, suggesting the stable microbial community structure corresponded to the elevated performance of the AnFB-MBR. To be noted is that this is an initial feasibility study only to evaluate whether fluidized bed operation in conjunction with membrane filtration had potential benefits for applications in the sulfur-oxidizing autotrophic denitrification. Although AnFB-MBR appears promising for the treatment of nitrate from drinking water, significant advancements and large-scale demonstration are necessary for AnFB-MBR to be considered over more established treatment technologies. Further study is required to optimize AnFB-MBR performance, which might be accomplished by maximizing membrane flux and alternative efficient fouling control strategies. This research also provides a framework for further ecological and physiological microbial research in the sulfur-oxidizing autotrophic denitrifying system.
References
Ahmed Z, Kim SM, Kim IS, Bum MS, Chae KJ, Joo JH, Ok YS, Oh SE (2012) Nitrification and denitrification using biofilters packed with sulfur and limestone at a pilot-scale municipal wastewater treatment plant. Environ Technol 33(11):1271–1278
Brettar I, Rheinheimer G (1991) Denitrification in the Central Baltic: evidence for H2S-oxidation as motor of denitrification at the oxic–anoxic interface. Mar Ecol Prog Ser 77:157–196
Cardoso RB, Sierra-Alvarez R, Rowlette P, Flores ER, Gómez J, Field JA (2006) Sulfide oxidation under chemolithoautotrophic denitrifying conditions. Biotechnol Bioeng 95(6):1149–1156
Ergas SJ, Rheinheimer DE (2004) Drinking water denitrification using a membrane bioreactor. Water Res 38:3225–3232
Fernández N, Sierra-Alvarez R, Field JA, Amils R, Sanz JL (2008) Microbial community dynamics in a chemolithotrophic denitrification reactor inoculated with methanogenic granular sludge. Chemosphere 70:462–474
Guglielmi G, Chiarani D, Judd SJ, Andreottola G (2007) Flux criticality and sustainability in a hollow fibre submerged membrane bioreactor for municipal wastewater treatment. J Memb Sci 289(1-2):241–248
Hallin S, Throbäk IN, Disksved J, Pell M (2006) Metabolic profiles and genetic diversity of denitrifying communities in activated sludge after addition of methanol or ethanol. Appl Environ Microbiol 72:5445–5452
Ho J, Sung S (2010) Methanogenic activities in anaerobic membrane bioreactors (AnMBR) treating synthetic municipal wastewater. Bioresour Technol 101(7):2191–2196
Judd SJ (2002) Submerged membrane bioreactors: flat plate or hollow fiber? Filtr Sep 39(5):30–31
Kim EW, Bae JH (2000) Alkalinity requirements and the possibility of simultaneous heterotrophic denitrification during sulfur-utilizing autotrophic denitrification. Water Sci Technol 42(3-4):233–238
Kim J, Kim K, Ye H, Lee E, Shin C, Mccarty PL, Bae J (2011) Anaerobic fluidized bed membrane bioreactor for wastewater treatment. Environ Sci Technol 45:576–581
Kimura K, Nakamura M, Watanabe Y (2002) Nitrate removal by a combination of elemental sulphur-based denitrification and membrane filtration. Water Res 36:1758–1766
Knowles R (1982) Denitrification. Microbiol Rev 46:43–70
Koenig A, Liu LH (2001) Kinetic model of autotrophic denitrification in sulphur packed-bed reactors. Water Res 35:1969–1978
Koenig A, Zhang T, Liu LH, Fang HHP (2005) Microbial community and biochemistry process in autosulfurotrophic denitrifying biofilm. Chemosphere 58:1041–1047
Kohno T, Sei K, Mori K (2002) Characterization of type 1851 organisms isolated from activated sludge samples. Water Sci Technol 46:111–114
Krangelund C, Levantesi C, Borger A, Thelen K, Eikelboom D, Tandoi V, Kong Y, Van Der Waarde J, Krooneman J, Rossetti S, Thomsen TR, Nielsen PH (2007) Identity, abundance and ecophysiology of filamentous Chloroflexi species present in activated sludge treatment plants. FEMS Microbial Ecol 59:671–682
Lee KC, Rittmann BE (2002) Applying a novel autohydrogenotrophic hollow-fiber membrane biofilm reactor for denitrification of drinking water. Water Res 36:2040–2052
Liao BQ, Kraemer JT, Bagley DM (2006) Anaerobic membrane bioreactors: application and research directions. Crit Rev Environ Sci Technol 36(6):489–530
Liu Q, Wang XC, Liu Y, Yuan H, Du Y (2010) Performance of a hybrid membrane bioreactor in municipal wastewater treatment. Desalination 258:143–147
Lu HJ, Chandran K (2010) Factors promoting emissions of nitrous oxide and nitric oxide from denitrifying sequencing batch reactors operated with methanol and ethanol as electron donors. Biotechnol Bioengin 106(3):390–398
Mao YJ, Yannarell AC, Mackie RI (2011) Changes in N-transforming archaea and bacteria in soil during the establishment of bioenergy crops. PLoS One 6(9), e24750:1–12
McAdam EJ, Judd SJ (2006) A review of membrane bioreactor potential for nitrate removal from drinking water. Desalination 196(1-3):135–148
McAdam EJ, Judd SJ (2007) Denitrification from drinking water using a membrane bioreactor: chemical and biochemical feasibility. Water Res 41:4242–4250
Moon HS, Shin DY, Nam K, Kim JY (2008) A long-term performance test on an autotrophic denitrification column for application as a permeable reactive barrier. Chemosphere 73:723–728
Moon HS, Shin D, Nam K, Kim JA, ASCE AM (2010) Distribution of the microbial community structure in sulfur-based autotrophic denitrification columns. J Environ Eng 136:481–486
Pan YT, Liu Y, Bj N, Yuan ZG (2012) Effect of pH and N2O reduction and accumulation during denitrification by methanol utilizing denitrifiers. Water Res 46:4832–4840
Park JH, Shin HS, Lee IS, Bae JH (2002) Denitrification of high NO3 −-N containing wastewater using elemental sulfur: nitrogen loading rate and N2O production. Environ Technol 23(1):53–65
Puckett LJ (1995) Identifying the major sources of nutrient water pollution. Environ Sci Technol 29:408A–414A
Puig S, Coma M, Desloover J, Boon N, Colprim J, Balaguer MD (2012) Autotrophic denitrification in microbial fuel cells treating low ionic strength waters. Environ Sci Technol 46:2309–2315
Sahinkaya E, Dursun N (2012) Sulfur-oxidizing autotrophic and mixotrophic denitrification processes for drinking water treatment: elimination of excess sulfate production and alkalinity requirement. Chemosphere 89:144–149
Sahinkaya E, Kilic A (2014) Heterotrophic and elemental-sulfur-based autotrophic denitrification processes for simultaneous nitrate and Cr(VI) reduction. Water Res 50:278–286
Sahinkaya E, Dursun N, Kilic A, Demirel S, Uyanik S, Cinar O (2011) Simultaneous heterotrophic and sulfur-oxidizing autotrophic denitrification process for drinking water treatment: control of sulfate production. Water Res 45:6661–6667
Shao MF, Zhang T, Fang HHP (2010) Sulfur-driven autotrophic denitrification: diversity, biochemistry, and engineering applications. Appl Microbiol Biotechnol 88:1027–1042
Sierra-Alvarez R, Beristain-Cardoso R, Salazar M, Gómez J, Razo-Flores E, Field JA (2007) Chemolithotrophic denitrification with elemental sulfur for groundwater treatment. Water Res 41:1253–1262
Spagni A, Casu S, Crispino NA, Farina R, Mattioli D (2010) Filterability in a submerged anaerobic membrane bioreactor. Desalination 250:787–792
Sun YM, Nemati M (2012) Evaluation of sulfur-based autotrophic denitrification and denitritation for biological removal of nitrate and nitrite from contaminated waters. Bioresour Technol 114:207–216
Takao L, Mori K, Uchino Y, Nakagawa T, Harayama S, Suzuki KI (2010) Ignavibacterium album gen. nov., sp. nov., a moderately thermophilic anaerobic bacterium isolated from microbial mats at a terrestrial hot spring and proposal of Ignavibacteria classis nov., for a novel lineage at the periphery of green sulfur bacteria. Int J Syst Evol Microbiol 60:1376–1382
Throbäk IN, Enwall K, Jarvis Å, Hallin S (2004) Reassessing PCR primers targeting nirS, nirK and nosZ genes for community surveys of denitrifying bacteria with DGGE. FEMS Microbiol Ecol 49:401–417
Wan DJ (2009) Researches on the combined bioelectrochemical hydrogen with sulfur autotrophic denitrification process for nitrate removal [D]. Research Center for Eco-Environmental Science, Chinese Academy of Science
Ward MH, deKok TM, Levallois P, Brender J, Gulis G, Nolan BT, VanDerslice J (2005) Workgroup report: drinking water nitrate and health-recent findings and research needs. Environ Health Perspec 113:1607–1614
Warneke S, Schipper LA, Matiasek MG, Scow KM, Cameron S, Bruesewitz DA, McDonald IR (2011) Nitrate removal, communities of denitrifiers and adverse effects in different carbon substrates for use in denitrification beds. Water Res 45:5463–5475
Wrighton K, Virdis B, Clauwaert P, Read ST, Daly RA, Boon N, Piceno Y, Andersen GL, Coates JD, Rabaey K (2010) Bacterial community structure corresponds to performance during cathodic nitrate reduction. ISME J 4:1443–1455
Wunderlin P, Mohn J, Joss A, Emmenegger L, Siegrist H (2012) Mechanisms of N2O production in biological wastewater treatment under nitrifying and denitrifying conditions. Water Res 46:1027–1037
Zeng H, Zhang TC (2005) Evaluation of kinetic parameters of a sulfur-limestone autotrophic denitrification biofilm process. Water Res 39:4941–4962
Zhang TC, Lampe DG (1999) Sulfur: limestone autotrophic denitrification processes for treatment of nitrate-contaminated water: batch experiments. Water Res 33(3):599–608
Zhang TC, Zeng H (2006) Development of a response surface for prediction of nitrate removal in sulfur-limestone autotrophic denitrification fixed-bed reactors. J Environ Eng 132:1068–1072
Zhao HP, Ginkel SV, Tang Y, Kang DW, Rittmann B, Krajmalnik-Brown R (2011) Environ Sci Technol 45:10155–10162
Zhu XY, Chen YG (2011) Reduction of N2O and NO generation in anaerobic-aerobic (low dissolved oxygen) biological wastewater treatment process by using sludge alkaline fermentation liquid. Environ Sci Technol 45:2137–2143
Acknowledgments
The authors are grateful for financial support from the National Natural Science Foundation of China (No. 41171377) and the Major Science and Technology Program for Water Pollution Control and Treatment (2012ZX07408001).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 80 kb)
Rights and permissions
About this article
Cite this article
Zhang, L., Zhang, C., Hu, C. et al. Denitrification of groundwater using a sulfur-oxidizing autotrophic denitrifying anaerobic fluidized-bed MBR: performance and bacterial community structure. Appl Microbiol Biotechnol 99, 2815–2827 (2015). https://doi.org/10.1007/s00253-014-6113-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-014-6113-9