Abstract
Biofilm formation in drinking water distribution systems (DWDS) has many adverse consequences. Knowledge of microbial community structure of DWDS biofilm can aid in the design of an effective control strategy. However, biofilm bacterial community in real DWDS and the impact of drinking water purification strategy remain unclear. The present study investigated the composition and diversity of biofilm bacterial community in real DWDSs transporting waters with different purification strategies (conventional treatment and integrated treatment). High-throughput Illumina MiSeq sequencing analysis illustrated a large shift in the diversity and structure of biofilm bacterial community in real DWDS. Proteobacteria, Firmicutes, Bacteroidetes, Actinobacteria, Nitrospirae, and Cyanobacteria were the major components of biofilm bacterial community. Proteobacteria (mainly Alphaproteobacteria, Betaproteobacteria, and Gammaproteobacteria) predominated in each DWDS biofilm, but the compositions of the dominant proteobacterial classes and genera and their proportions varied among biofilm samples. Drinking water purification strategy could shape DWDS biofilm bacterial community. Moreover, Pearson’s correlation analysis indicated that Actinobacteria was positively correlated with the levels of total alkalinity and dissolved organic carbon in tap water, while Firmicutes had a significant positive correlation with nitrite nitrogen.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Maintaining a disinfectant (usually chlorine) residual is a routine practice to control bacterial regrowth in drinking water distribution systems (DWDS), although a high diversity of microorganisms (including pathogens or opportunistic pathogens) can still exist in bulk water and pipe materials of DWDS (Berry et al. 2006; Lu et al. 2013; Pavlov et al. 2004; Revetta et al. 2013). Biofilm attached on the internal surface of pipes is a major reservoir of microorganisms in DWDS (Berry et al. 2006; Revetta et al. 2013). Nearly 95 % of total microbial cells in DWDS are present in biofilms on pipe surfaces (Moritz et al. 2010). Many problems in DWDS can be associated with biofilm formation, such as bio-corrosion of metal pipe (Teng et al. 2008), hosting opportunistic pathogens (Sun et al. 2014), and promoting nitrification (Gomez-Alvarez et al. 2014; Regan et al. 2002). Therefore, a sound understanding of microbial community structure of DWDS biofilm and its influential factors is of great importance in designing effective control strategies and thus improving drinking water quality for the consumer (Berry et al. 2006; Liu et al. 2013; Martiny et al. 2003). It has been documented that microbial community structure of DWDS biofilm can be affected by a variety of factors, such as pipe materials (Jang et al. 2011; Lin et al. 2013; Wang et al. 2014), disinfectants (Gomez-Alvarez et al. 2012; Hwang et al. 2012; Krishna et al. 2013; Wang et al. 2014), water age (Wang et al. 2014), and DWDS biofilm age (Liu et al. 2012; Martiny et al. 2003; Revetta et al. 2013). These previous studies usually used simulated DWDS or bioreactors to study biofilm microbial communities. However, several years can be necessary for the achievement of the steady state of biofilm formation (Liu et al. 2012; Martiny et al. 2003), which limits the relevance of short-term model studies (Berry et al. 2006; Martiny et al. 2003). Due to limited access and high cost involved in sampling biofilm within real DWDS, so far, the composition and dynamics of bacterial communities in real DWDS remain poorly understood. In addition, although it has been reported that different water sources (ground water and surface water) can result in a significant difference of bacterial community diversity and composition in real DWDS (Sun et al. 2014), it remains unclear whether or not the application of different purification strategies for the same raw water can play an important role in shaping biofilm microbial community structure in real DWDS.
Molecular microbial ecology tools, such as terminal restriction fragment length polymorphism (TRFLP), denaturing gradient gel electrophoresis (DGGE), and clone library analysis, have greatly contributed to our knowledge of microbial ecology in DWDS (Grigorescu et al. 2012; Lin et al. 2013; Lu et al. 2013; Vaz-Moreira et al. 2013). However, these low-throughput biology tools can underestimate the overall diversity of a microbial community and are usually not able to detect rare species in complicated environmental samples (Liao et al. 2013a). In contrast, high-throughput sequencing, as a next generation sequencing technology, has illustrated its strong potential in elucidating the complicated biofilm microbial community structure in DWDS (Douterelo et al. 2013; Gomez-Alvarez et al. 2014; Liu et al. 2012, 2014a, b; Sun et al. 2014; Wang et al. 2014). Illumina MiSeq platform is a recently developed high-throughput sequencing platform (Caporaso et al. 2012). Illumina-based 16S rRNA gene sequencing has gained increasing popularity due to its lower costs, higher accuracy, and greater throughput (Nelson et al. 2014). Therefore, the main objective of the present study was to investigate the diversity and composition of biofilm bacterial community in real DWDSs transporting waters with different purification strategies. The bacterial community was characterized using high-throughput Illumina MiSeq sequencing analysis.
Materials and methods
Sample collection
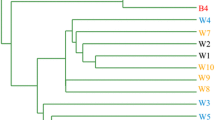
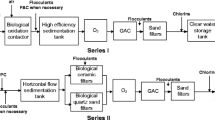
Cast iron pipe sections were collected from a small southeast city of China. River water was used as the sole water source for this city. Two different strategies were adopted for the drinking water purification, namely, conventional treatment process and integrated treatment process. The conventional treatment process was composed of rapid mixing, flocculation, sedimentation, sand filtration, and chlorine disinfection, while integrated treatment process included biological contact oxidation, rapid mixing, flocculation, sedimentation, sand filtration, two-stage ozonation followed by biological activated carbon filtration, and chlorine disinfection. The urban areas are equipped with two sets of DWDSs, one receiving the water after the conventional treatment for cleaning, while another receiving the higher quality water after the integrated treatment for drinking and cooking. In this study, twelve cast iron pipe sections (20–30 years old; diameter of 40–50 cm) were excavated from different DWDS sites from the water treatment works, including six ones (named as BN, XN, SN, GN, RN, and MD) from the DWDS transporting water after conventional treatment, and another six ones (named as BD, XD, SD, GD, RD, and MD) from the DWDS transporting water after integrated treatment. These pipes were distributed in four pipe lines (Fig. 1). These pipe sections were immediately transported back to laboratory after collection. The physicochemical parameters of the tap water at each corresponding DWDS sampling site are shown in Table S1.
Molecular analyses
Biofilm samples were removed from pipes according to the literature (Sun et al., 2014). DNA was extracted using Powersoil DNA extraction kit (Mobio Laboratories) following the manufacturer’s instructions. PCR amplicon libraries were constructed for Illumina MiSeq sequencing using bacterial primers 515F (5′-GTGCCAGCMGCCGCGG-3′) and 806R (5′-GGACTACHVGGGTWTCTAAT-3′) targeting V4 hypervariable regions of bacterial 16S rRNA genes (Caporaso et al. 2012). Quality filtering of reads was performed according to the literature (Caporaso et al. 2012). Sequences were grouped into the operational taxonomic units (OTUs) using a 97 % similarity cutoff. OTU-based community diversity indices (Chao1 estimator and Shannon index) and rarefaction curve of each sample were generated using the MOTHUR program (Schloss et al. 2009). A representative sequence for each OTU was selected, and the RDP classifier was used to assign taxonomic identity to each representative sequence (Wang et al. 2007). Based on the relative abundance of bacterial phyla, unweighted UniFrac with QIIME (http://qiime.org/index.html) was used for unweighted pair group method with arithmetic mean (UPGMA) clustering. In addition, Pearson’s correlation analysis of bacterial community with the water physicochemical parameters and sampling site distance was carried out using SPSS 20.0 software. The gene sequences obtained from high-throughput analysis in the current study were deposited in the NCBI short-read archive under accession number PRJNA255177.
Results
Bacterial diversity
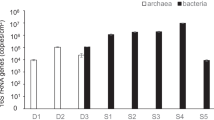
In this study, a total of 6,806–110,401valid bacterial sequences were recovered from each DWDS biofilm sample using Illumina MiSeq sequencing analysis. Each DWDS biofilm bacterial library was composed of 363–582 OTUs at 97 % similarity level (Table 1). The rarefaction curves of all biofilm samples nearly approached a plateau, suggesting that these DWDS biofilm communities had been well sampled (Fig. S1). The values of Chao1 estimator and Shannon index of DWDS biofilm samples were 376–947 and 5.11–7.12, respectively. Therefore, a marked variation of bacterial community diversity occurred in these studied DWDS pipe biofilm samples. Moreover, in the same sampling zone, the biofilm sample from the DWDS transporting water with conventional treatment usually had a higher Shannon community diversity than that from the DWDS transporting water with integrated treatment. This result suggested that the origin of feeding water could affect the bacterial community diversity in DWDS pipe biofilm.
Bacterial community composition
In this study, a total of 23 known and candidate bacterial phyla were found in the 12 DWDS pipe biofilm samples. Acidobacteria, Actinobacteria, Bacteroidetes, Chloroflexi, Cyanobacteria, Firmicutes, Gemmatimonadetes, Nitrospirae, Proteobacteria, and Verrucomicrobia were the frequently detected bacterial phyla among these DWDS biofilm samples (Fig. 2). Proteobacteria (accounting for 66–87 %) predominated in all the biofilm samples, and it mainly consisted of three classes (Alphaproteobacteria, Betaproteobacteria, and Gammaproteobacteria) (Fig. 3). However, a large shift in the proportions of Alphaproteobacteria, Betaproteobacteria, and Gammaproteobacteria was observed in these DWDS pipe biofilm samples. Class Alphaproteobacteria was the largest bacterial group in samples BD, GD, MD, and SN, while class Betaproteobacteria showed the largest relative abundance in samples BN, GN, MN, SD, RN, XD, and SN. Class Gammaproteobacteria predominated in sample RD and was the second largest bacterial group in sample RN. Moreover, phylum Firmicutes occurred in samples RN and RD with a relative abundance of 16.1 and 13.6 %, respectively, but it was detected with a much lower proportion in other DWDS biofilm samples. Phylum Bacteroidetes was highly abundant in samples RD (13.2 %) and XD (8.4 %), while it was a minor component of bacterial community in other biofilm samples. The proportion of Actinobacteria varied greatly in the studied biofilm samples (2.2–10.3 %). In addition, Nitrospirae had a relatively high proportion in samples BN, XN, GN, and SD (4.8–6.4 %). These results indicated a large variation in the bacterial community compositions of biofilm samples from real DWDS pipes.
Based on the relative abundance of bacterial phyla, the result of UPGMA clustering showed that, in the same sampling zone, the pipe biofilm sample from the DWDS transporting water with conventional treatment was usually separated from that from the DWDS transporting water with integrated treatment. This result suggested that the origin of feeding water could affect the compositions of bacterial communities in DWDS pipe biofilm. In addition, samples RN and RD were grouped together, but they were distantly separated from other biofilm samples. This further confirmed a large change of biofilm bacterial community structure in real DWDS pipes.
Table 2 shows the 43 frequently detected genera in DWDS pipe biofilm samples. A number of genera from diverse bacterial phyla were detected in each biofilm sample, implying a high bacterial diversity in DWDS biofilm. At the genus level of taxonomic classifications, the variations among DWDS biofilm samples were evident. Within class Alphaproteobacteria, genera Rhodoplanes and Sphingomonas were distributed in each biofilm sample, while genus Brevundimonas was only detected in samples RN and RD. Within class Betaproteobacteria, genus Sutterella was abundant in samples RN and RD, but it was absent in other biofilm samples. Samples RN and XD had a much higher proportion of Simplicispira than other DWDS biofilm samples. Cupriavidus was in a relatively high proportion in sample MN, while Dok59 in sample SD. Genera Shewanella, Halomonas, and Rhodanobacter (class Gammaproteobacteria) showed the highest proportion in sample RN, while Providencia, Acinetobacter, Pseudomonas, and Stenotrophomonas were the most abundant in sample RD. Moreover, sample GN had the highest proportion of Desulfovibrio (Deltaproteobacteria). Helicobacter (Epsilonproteobacteria) was mainly present in samples RN and RD. These results confirmed a large shift in the genus compositions of proteobacterial communities in DWDS pipe biofilm samples. In addition, a shift in the genus compositions of other bacterial phyla in DWDS biofilm samples was also observed.
Links between bacterial community and water characteristics or sampling site distance
In this study, Pearson’s correlation analysis using SPSS 20.0 software was applied to describe the links between DWDS pipe biofilm bacterial community and the water physicochemical parameters and sampling site distance. OTUs and OTU-based community diversity indices (Chao1 estimator and Shannon index) did not show any significant correlation with each of the measured physicochemical parameters and the distance of sampling site from water works (P > 0.05) (Table 3). The relative abundance of Actinobacteria was found to be positively correlated with the levels of total alkalinity and dissolved organic carbon (DOC) in tap water (P < 0.05). The relative abundance of Firmicutes showed a significant positive correlation with the level of nitrite nitrogen in tap water. However, Proteobacteria and its classes show no significant correlation with the determined water physicochemical parameters. Moreover, no significant correlation was found between the distance of sampling site from water works and the diversity and composition of DWDS biofilm bacterial community.
Discussion
DWDS biofilm bacterial diversity
There have been numerous previous reports on the biofilm bacterial diversity in model DWDS (Gomez-Alvarez et al. 2014; Krishna et al. 2013; Lee et al. 2005; Teng et al. 2008; Wang et al. 2014). In addition, there have been few reports on biofilm bacterial diversity in faucets and water meters. Pyrosequencing analysis showed that the values of Chao1 estimate and OTUs of biofilm bacterial communities in two water meters were 133 (or 208) and 203 (or 341), respectively (Hong et al. 2010). Pyrosequencing analysis of bacterial communities in faucet biofilm indicated 356–1,564 OTUs, Chao1 estimate of 724–2,430, and Shannon index of 3.44–4.49 (Liu et al. 2012), while clone library analysis revealed 17–37 OTUs, Chao1 estimate of 20–83, and Shannon index of 2.11–3.15 (Lin et al. 2013). However, possibly due to limited access to sample pipe biofilm in real DWDS, so far, information on biofilm bacterial diversity of DWDS pipe is very limited. One recent pyrosequencing study revealed 642–1,532 OTUs, Chao1 estimate of 1,079–1,849, and Shannon index of 3.36–5.29 in biofilm bacterial communities on old unlined cast iron pipes in a real DWDS (Sun et al. 2014). In this study, Illumina MiSeq sequencing indicated that biofilm bacterial communities on DWDS pipes had 363–582 OTUs, Chao1 estimator of 376–947, and Shannon index of 5.11–7.12. This confirmed a high biofilm bacterial diversity in real DWDS pipes. In addition, it also illustrated a strong potential of Illumina MiSeq sequencing in elucidating the bacterial diversity of drinking water biofilm.
DWDS biofilm bacterial community composition
The predominance of Proteobacteria has been found in a variety of ecosystems, such as freshwater (Cheng et al. 2014; Vaz-Moreira et al. 2013; Zhang et al. 2014), drinking water biofilter (Feng et al. 2013; Liao et al. 2013a, b), finished drinking water (Pinto et al. 2012; Vaz-Moreira et al. 2013), and tap water in DWDS (Lu et al. 2013; Pinto et al. 2012; Vaz-Moreira et al. 2013). Previous studies usually indicated the predominance of Proteobacteria in DWDS biofilm bacterial community (Gomez-Alvarez et al. 2014; Liu et al. 2013; Sun et al. 2014). The dominance of Betaproteobacteria has been detected in biofilm in real DWDS (Liu et al. 2014a; Sun et al. 2014), in model DWDS (Lee et al. 2005), and on faucet (Liu et al. 2014b). Other earlier studies also showed the dominance of Betaproteobacteria in drinking water biofilm (Batté et al. 2003; Emtiazi et al. 2004; Kalmbach et al. 1997). However, Alphaproteobacteria was found to be the largest component of biofilm bacterial community in real DWDS (Douterelo et al. 2013), in model DWDS (Gomez-Alvarez et al. 2014; Krishna et al. 2013), and tap water (Liu et al. 2014b). Moreover, both Alphaproteobacteria and Betaproteobacteria could be the most dominant biofim bacterial groups in model DWDSs (Wang et al. 2014) and tap waters (Lin et al. 2013; Liu et al. 2012). The bacterial composition in a water meter consisted of two major bacterial populations from the Betaproteobacteria and Alphaproteobacteria, while the Betaproteobacteria population predominated in another water meter microflora (Hong et al. 2010). These previous studies suggested the dominance of Alphaproteobacteria and Betaproteobacteria in the oligotrophic environment of DWDS, while their relative importance remained controversial. In this study, a large shift in the proportions of Alphaproteobacteria, Betaproteobacteria, and Gammaproteobacteria occurred in biofilm samples from cast iron pipes in real DWDSs. Although Alphaproteobacteria and Betaproteobacteria predominated in most of the studied DWDS pipes, Gammaproteobacteria was the largest bacterial group in one pipe. Douterelo et al. (2013) also reported that the dominant bacterial phyla within the biofilms in a laboratory DWDS were Gammaproteobacteria, followed by Betaproteobacteria and Alphaproteobacteria. Therefore, the compositions of the major proteobacterial classes in biofilm bacterial community can vary among different DWDSs.
Previous studies indicated the low abundance of Firmicutes in DWDS pipe biofilm (Liu et al. 2014a; Revetta et al. 2013), faucet biofilm (Lin et al. 2013), and water meter biofilm (Hong et al. 2010). Firmicutes was found to be a minor bacterial group in biofilm on DWDS pipes transporting treated ground waters, while it became a major group in biofilm on DWDS pipes transporting treated surface waters (Sun et al. 2014). In this study, Firmicutes was detected with a very low proportion in most of the studied DWDS biofilm samples, but it was a major component of biofilm bacterial community in two DWDS pipes. Therefore, the importance of Firmicutes could vary among different DWDSs and pipes. Bacteroidetes was usually the rare species in DWDS biofilm (Douterelo et al. 2013, Lin et al. 2013; Liu et al. 2012, 2014a, b; Sun et al. 2014). In this study, Bacteroidetes was a minor component in most of biofilm bacterial communities but showed a high proportion in two biofilm samples. To the authors’ knowledge, this was the first report on the high abundance of Bacteroidetes in DWDS biofilm. Either low or high relative abundance of Actinobacteria in DWDS biofilm has also reported by other previous studies (Krishna et al. 2013; Revetta et al. 2013; Sun et al. 2014), which was in agreement with the result obtained in this study. Nitrospirae was found to be a major component of biofilm bacterial community only in DWDS fed with chloraminated drinking water (Krishna et al. 2013; Gomez-Alvarez et al. 2014). In this study, Nitrospirae had a relatively high proportion (4.8–6.4 %) in four DWDS biofilm samples, which might be attributed to the presence of ammonia nitrogen and nitrite nitrogen in tap waters. This study presented the first evidence for the dominance of Nitrospirae in DWDS fed with chlorinated drinking water.
Factors regulating DWDS biofilm bacterial community
As mentioned above, microbial community structure of DWDS biofilm can be regulated by pipe materials, disinfectants, water age, biofilm age, and hydraulic conditions. However, so far, the composition and dynamics of bacterial communities in real DWDS pipe and their influential factors remain poorly understood. Low dependency of the microbial community structure on the surface material was found in real DWDS used for 20 years (Henne et al. 2012), which was attributed to the mutual influence of adjacent biofilm by the exchange of microorganisms (Liu et al. 2013). Martiny et al. (2003) also reported that the mature biofilms from different sampling points in a pilot DWDS could have a similar microbial community structure (Martiny et al. 2003). In contrast, based on the results of diversity and the bacterial phylum, class, and genus levels, the present study showed a large variation of biofilm bacterial community in the real DWDSs transporting either the water with conventional treatment or the water with integrated treatment, although the studied DWDS pipes had been used for 20–30 years. A recent study also reported the heterogeneity of biofilm bacterial community in real DWDSs (Sun et al. 2014).
Sun et al. (2014) indicated that the difference of water sources (surface water and ground water) could affect DWDS biofilm bacterial community. Pinto et al. (2012) found that water treatment process could shape the bacterial community in bulk water of DWDS. The present study provided the first evidence for the impact of drinking water purification strategy on DWDS biofilm bacterial community. Surface water with conventional treatment or integrated treatment could result in the difference of diversity and structure of bacterial community in DWDS biofilm.
Wang et al. (2014) suggested that water age was a strong factor in shaping biofilm bacterial community structure in a model DWDS. However, in this study, Pearson’s correlation analysis showed no significant correlation between the distance of sampling site from water works and bacterial community in DWDS biofilm. Moreover, Proteobacteria and its classes show no significant correlation with the determined physicochemical parameters. Actinobacteria was positively correlated with the levels of total alkalinity and DOC, while Firmicutes with the level of nitrite nitrogen. In contrast, Sun et al. (2014) found that Proteobacteria was positively correlated with alkalinity and negatively correlated with organic matter. Actinobacteria was positively correlated with water temperature, while Firmicutes was negatively correlated with alkalinity and positively correlated with organic matter (Sun et al. 2014). A myriad of factors may mutually shape the microbial community composition in real DWDS (Vaz-Moreira et al. 2013). Therefore, further work will be necessary in order to elucidate the factors regulating DWDS biofilm bacterial community.
In conclusion, a large variation in the diversity and structure of biofilm bacterial community occurred in real DWDS. Proteobacteria (mainly including Alphaproteobacteria, Betaproteobacteria, and Gammaproteobacteria) predominated in DWDS pipe biofilm microbiota. Drinking water purification strategy could play an important role in shaping DWDS biofilm bacterial community. In addition, Illumina MiSeq sequencing illustrated a strong potential in characterizing bacterial community of drinking water biofilm.
References
Batté M, Mathieu L, Laurent P, Prévost M (2003) Influence of phosphate and disinfection on the composition of biofilms produced from drinking water, as measured by fluorescence in situ hybridization. Can J Microbiol 49:741–753
Berry D, Xi CW, Raskin L (2006) Microbial ecology of drinking water distribution systems. Curr Opin Biotechnol 17:297–302
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Huntley J, Fierer N, Owens SM, Betley J, Fraser L, Bauer M, Gormley N, Gilbert JA, Smith G, Knight R (2012) Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6:1621–1624
Cheng W, Zhang JX, Wang Z, Wang M, Xie SG (2014) Bacterial communities in sediments of a drinking water reservoir. Ann Microbiol 64:875–878
Douterelo I, Sharpe RL, Boxall JB (2013) Influence of hydraulic regimes on bacterial community structure and composition in an experimental drinking water distribution system. Water Res 47:503–516
Emtiazi F, Schwartz T, Marten SM, Krolla-Sidenstein P, Obst U (2004) Investigation of natural biofilms formed during the production of drinking water from surface water embankment filtration. Water Res 38:1197–1206
Feng S, Chen C, Wang QF, Zhang XJ, Yang ZY, Xie SG (2013) Characterization of microbial communities in a granular activated carbon-sand dual media filter for drinking water treatment. Int J Environ Sci Technol 10:917–922
Gomez-Alvarez V, Revetta RP, Domingo JWS (2012) Metagenomic analyses of drinking water receiving different disinfection treatments. Appl Environ Microbiol 78:6095–6102
Gomez-Alvarez V, Schrantz KA, Pressman JG, Wahman DG (2014) Biofilm community dynamics in bench-scale annular reactors simulating arrestment of chloraminated drinking water nitrification. Environ Sci Technol 48:5448–5457
Grigorescu AS, Hozalski RM, LaPara TM (2012) Haloacetic acid-degrading bacterial communities in drinking water systems as determined by cultivation and by terminal restriction fragment length polymorphism of PCR-amplified haloacid dehalogenase gene fragments. J Appl Microbiol 112:809–822
Henne K, Kahlisch L, Brettar I, Höfle MG (2012) Analysis of structure and composition of bacterial core communities in mature drinking water biofilms and bulk water of a citywide network in Germany. Appl Environ Microbiol 78:3530–353
Hong PY, Hwang CC, Ling FQ, Andersen GL, LeChevallier MW, Liu WT (2010) Pyrosequencing analysis of bacterial biofilm communities in water meters of a drinking water distribution system. Appl Environ Microbiol 76:5631–5635
Hwang C, Ling F, Andersen GL, LeChevallier MW, Liu WT (2012) Microbial community dynamics of an urban drinking water distribution system subjected to phases of chloramination and chlorination treatments. Appl Environ Microbiol 78:7856–7865
Jang HJ, Choi YJ, Ka JO (2011) Effects of diverse water pipe materials on bacterial communities and water quality in the annular reactor. J Microbiol Biotechnol 21:115–123
Kalmbach S, Manz W, Szewyk U (1997) Dynamics of biofilm formation in drinking water: phylogenetic affiliation andmetabolic potential of single cells assessed by formazan reduction and in situ hybridization. FEMS Microbiol Ecol 22:265–279
Krishna KCB, Sathasivan A, Ginige MP (2013) Microbial community changes with decaying chloramine residuals in a lab-scale system. Water Res 47:4666–4679
Lee DG, Lee JH, Kim SJ (2005) Diversity and dynamics of bacterial species in a biofilm at the end of the Seoul water distribution system. World J Microbiol Biotechnol 21:155–162
Liao XB, Chen C, Wang Z, Wan R, Chang CH, Zhang XJ, Xie SG (2013a) Pyrosequencing analysis of bacterial communities in drinking water biofilters receiving influents of different types. Process Biochem 48:703–707
Liao XB, Chen C, Wang Z, Wan R, Chang CH, Zhang XJ, Xie SG (2013b) Changes of biomass and bacterial communities in biological activated carbon filters for drinking water treatment. Process Biochem 48:312–316
Lin WF, Yu ZS, Chen X, Liu RY, Zhang HX (2013) Molecular characterization of natural biofilms from household taps with different materials: PVC, stainless steel, and cast iron in drinking water distribution system. Appl Microbiol Biotechnol 97:8393–8401
Liu RY, Yu ZS, Guo HG, Liu MM, Zhang HX, Yang M (2012) Pyrosequencing analysis of eukaryotic and bacterial communities in faucet biofilms. Sci Total Environ 435:124–131
Liu G, Verberk JQJC, Van Dijk JC (2013) Bacteriology of drinking water distribution systems: an integral and multidimensional review. Appl Microbiol Biotechnol 97:9265–9276
Liu G, Bakker GL, Li S, Vreeburg JHG, Verberk JQJC, Medema GJ, Liu WT, Van Dijk JC (2014a) Pyrosequencing reveals bacterial communities in unchlorinated drinking water distribution system: an integral study of bulk water, suspended solids, loose deposits, and pipe wall biofilm. Environ Sci Technol 48:5467–5476
Liu RY, Zhu JG, Yu ZS, Joshi D, Zhang HX, Lin WF, Yang M (2014b) Molecular analysis of long-term biofilm formation on PVC and cast iron surfaces in drinking water distribution system. J Environ Sci 26:865–874
Lu PP, Chen C, Wang QF, Wang Z, Zhang XJ, Xie SG (2013) Phylogenetic diversity of microbial communities in real drinking water distribution systems. Biotechnol Bioprocess Eng 18:119–124
Martiny AC, Jorgensen TM, Albrechtsen HJ, Arvin E, Molin S (2003) Long-term succession of structure and diversity of a biofilm formed in a model drinking water distribution system. Appl Environ Microbiol 69:6899–6907
Moritz MM, Flemming HC, Wingender J (2010) Integration of Pseudomonas aeruginosa and Legionella pneumophila in drinking water biofilms grown on domestic plumbing materials. Int J Hyg Environ Health 213:190–197
Nelson MC, Morrison HG, Benjamino J, Grim SL, Graf J (2014) Analysis, optimization and verification of Illumina-generated 16S rRNA gene amplicon surveys. PLoS ONE 9:e94249
Pavlov D, de Wet CME, Grabow WOK, Ehlers MM (2004) Potentially pathogenic features of heterotrophic plate count bacteria isolated from treated and untreated drinking water. Int J Food Microbiol 92:275–287
Pinto AJ, Xi C, Raskin L (2012) Bacterial community structure in the drinking water microbiome is governed by filtration processes. Environ Sci Technol 46:8851–8859
Regan JM, Harrington GW, Noguera DR (2002) Ammonia- and nitrite-oxidizing bacterial communities in a pilot-scale chloraminated drinking water distribution system. Appl Environ Microbiol 68:73–81
Revetta RP, Gomez-Alvarez V, Gerke TL, Curioso C, Domingo JWS, Ashbolt NJ (2013) Establishment and early succession of bacterial communities in monochloramine-treated drinking water biofilms. FEMS Microbiol Ecol 86:404–414
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541
Sun HF, Shi BY, Bai YH, Wang DS (2014) Bacterial community of biofilms developed under different water supply conditions in a distribution system. Sci Total Environ 472:99–107
Teng F, Guan YT, Zhu WP (2008) Effect of biofilm on cast iron pipe corrosion in drinking water distribution system: Corrosion scales characterization and microbial community structure investigation. Corros Sci 50:2816–2823
Vaz-Moreira I, Egas C, Nunes OC, Manaia CM (2013) Bacterial diversity from the source to the tap: a comparative study based on 16S rRNA gene-DGGE and culture-dependent methods. FEMS Microbiol Ecol 83:361–374
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Wang H, Masters S, Edwards MA, Falkinham JO, Pruden A (2014) Effect of disinfectant, water age, and pipe materials on bacterial and eukaryotic community structure in drinking water biofilm. Environ Sci Technol 48:1426–1435
Zhang JX, Zhang XL, Liu Y, Xie SG, Liu YG (2014) Bacterioplankton communities in a high-altitude freshwater wetland. Ann Microbiol 64:1045–1411
Acknowledgments
This work was financially supported by the State Environmental Protection Key Laboratory of Microorganism Application and Risk Control (No. MARC2012D010), National Water Special Program (No. 2012ZX07404-002), and International Science and Technology Cooperation Program of China (No. 2010DFA91830).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Huiting Wu and Jingxu Zhang contributed equally to this study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 164 kb)
Rights and permissions
About this article
Cite this article
Wu, H., Zhang, J., Mi, Z. et al. Biofilm bacterial communities in urban drinking water distribution systems transporting waters with different purification strategies. Appl Microbiol Biotechnol 99, 1947–1955 (2015). https://doi.org/10.1007/s00253-014-6095-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-014-6095-7