Abstract
Only limited studies are available on the molecular-level biosynthesis of cyclic lipopeptides (cyclic and hybrid molecules consisting of peptide and fatty acid moieties) in filamentous fungi. Here, we identified and characterized biosynthetic genes of the cyclic lipopeptides, known as verlamelins. Only four genes, coding for non-ribosomal peptide synthetase (NRPS), fatty acid hydroxylase, thioesterase, and AMP-dependent ligase, were found to be involved in verlamelin biosynthesis by the analysis of corresponding gene knockouts. Surprisingly, no gene(s) coding for fatty acid synthase or polyketide synthase was present in the cluster, while verlamelin A/B contained a 5-hydroxytetradecanoic acid moiety. Precursor feeding experiment indicated that both fatty acid hydroxylase and thioesterase are involved to supply 5-hydroxytetradecanoic acid. The results suggested that 5-hydroxytetradecanoic acid was supplied from primary metabolism via fatty acid hydroxylase and loaded onto NRPS. Elongation of the peptide and final cyclization were accomplished by NRPS. The knowledge obtained through this study should provide new insight into fungal lipopeptide biosynthesis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Verlamelin is a peptide antibiotic that was initially isolated from Verticillium lamellicola (Onishi et al. 1980) and shown to exhibit antifungal activity against plant pathogenic fungi, including morphological changes to fungal cells such as swelling or bulging (Kim et al. 2002; Rowin et al. 1986). Structurally, verlamelin is a cyclic hexadepsipeptide and is bridged by ester bonding between a hydroxy group on a fatty acid moiety and a carboxyl group on the terminal Val of amide-bonded tetradecanoyl-hexapeptide (D-alloThr-D-Ala-L-Pro-L-Gln-D-Tyr-L-Val). In our last study, the absolute structures of two verlamelins, verlamelin A and its new derivative (verlamelin B) produced by Lecanicillium sp. HF627, were definitively determined as shown in Fig. 1 (Ishidoh et al. 2014).
Stereochemical structure of verlamelin A and B in this study (Ishidoh et al. 2014)
Most cyclic depsipeptides of fungal origin are synthesized by non-ribosomal peptide synthethases (NRPS) and composed of amino acids, but some contain fatty acid moieties: destruxins, insecticidal hexadepsipeptides isolated from entomopathogenic fungus Metarhizium anisopliae, are composed of five amino acids and an α-hydroxyisocaproic acid, the moiety of which is occasionally modified (Wang et al. 2012); aureobasidine A, an antifungal depsipeptide isolated from the black yeast Aureobasidium pullulans, contains eight amino acids and a 2-hydroxy-3-methylpentanoic acid moiety (Slightom et al. 2009); beauvericine/eniatins contain amino acids and D-hydroxyisovalerate (Hornbogen et al. 2007; Xu et al. 2008). However, the majority of these fatty acids are derived from common amino acids, such as α-hydroxyisocaproic acid, which is converted from Leu (Wang et al. 2012), and only a few of them (e.g., emericellamide or FR901469) contain linear or branched-chain fatty acids, which may be derived from fatty acid synthase (FAS) or polyketide synthase (PKS) (Chiang et al. 2008; Fujie et al. 2000). As for genetic studies on the biosynthesis of peptides in filamentous fungi, many of these reports have focused on genes encoding NRPS, which is a multifunctional enzyme composed of modules corresponding to the order of amino acids in the structure of cyclic peptides (Marahiel et al. 1997). Although a number of biosynthetic clusters have been clarified, there have been only a few detailed biosyntheses of cyclic peptides which contain a long/middle-chain fatty acid moiety, such as apicidin or echinocandin (Cacho et al. 2012; Jin et al. 2010).
Here, we identified and characterized the NRPS gene responsible for the biosynthesis of verlamelin. Further analysis of genes flanking the NRPS gene together with gene disruptions revealed the presence of a verlamelin biosynthetic gene cluster, and we described the probable biosynthetic pathway/mechanism of verlamelin based on the cluster analysis.
Materials and methods
Strains and media
Lecanicillium sp. HF627 (deposited in The National Institute of Agrobiological Sciences Genebank under strain number MAFF635047), isolated from a chillie thrips cadaver, was used as a wild-type strain. Ki6 (pyrG − ku80 −) and Ki2p (ΔpyrG ku80 −) strains created from HF627 (Ishidoh et al. 2014) were used as the parental strains for the functional analysis of verlamelin biosynthetic genes. Escherichia coli DH5α was used for the construction and propagation of plasmids. YES medium (8 % sucrose, 4 % yeast extract) and SMY medium (4 % maltose, 1 % yeast extract, 1 % peptone) were used as production media of verlamelin. Conidia were collected from strains grown on potato dextrose agar (PDA) (Difco), supplemented with 10 mM of uridine as required. CD+ medium was prepared by the addition of 0.2 % of (NH4)2SO4 to Czapek-Dox (CD) medium (Oxoid, Hampshire UK) (with 1.5 % of agar for plate) and was used as a limited medium of uridine. CD+ medium supplemented with 0.8 M KCl was used for transformation of Ki6 and Ki2p.
Plasmids
pLSPYRG containing pyrG from HF627 was constructed on pUC19 as described by Ishidoh et al. (2014). For constructing disruption vectors, a 1.2-kbp region downstream of pyrG was amplified by PCR with a primer pair (Table S1; Online Resource, pyrgfoa1-pyrgfoa2) and was cloned into the SphI/HindIII sites of pLSPYRG, yielding pLSPYRG1200R, with which pyrG with the promoter region can be eliminated from the transformants via homologous recombination.
Search for NRPS gene fragments
The wild-type strain was cultivated statically in 20 ml of YES medium in 100-ml Erlenmeyer flasks at 25 °C for 4 days. Total RNA was isolated from a mycelial mat using an RNeasy plant minikit (Qiagen) and treated with DNase I (Takara). cDNA was synthesized using Superscript III RNaseH− (Invitrogen) with random primers (Invitrogen) according to the manufacturer’s instruction. Degenerated primers for RT-PCR were designed from the conserved regions of the A domain (coa1, coa4, A0S1, A2, A2C, A3) and PCP domain (T1S1). RT-PCR was carried out using TaKaRa Ex Taq® (Takara) with the indicated primer sets under the following conditions: 5 min at 94 °C and 35 (primer, coa1-coa4) or 40 (primer, A0S1-A2) cycles of 94 °C for 30 s, 51 °C for 30 s, and 72 °C for 60 s, followed by a final elongation step at 72 °C for 7 min. Subsequently, nested RT-PCR was performed using TaKaRa Ex Taq® with primer sets (A0S1-T1S1 for the first round, A2C-A3 for the second round) under the following conditions for both the first and second round: 94 °C for 5 min and 30 cycles of 94 °C for 30 s, 51 °C for 30 s, and 72 °C for 90 s (for first round) or 30 s (for second round)], followed by a final elongation step at 72 °C for 7 min. PCR fragments were cloned into the EcoRI or XbaI site of pUC19 to analyze their nucleotide sequences.
Vector insertion into a locus of the NRPS gene
A 1.6-kbp fragment of the NRPS gene was amplified by PCR using PrimeSTAR® HS DNA Polymerase (Takara) with a primer pair (nrpsDdis1–nrpsDdis2). The PCR fragment was cloned into the XhoI/SacI site of pLSPYRG, yielding the NRPS-targeting vector (pDISNRPSD).
Strain Ki6 was transformed as described by Ishidoh et al. (2014) using the constructed vector (pDISNRPSD). Emerging colonies on the selection plates were screened by PCR using an FTA® classic card (Whatman®) with a primer pair (nrpsDdischeck-M13fw). The genotypes of the candidate strains were further analyzed by Southern hybridization with a probe prepared by PCR (nrpsD1–nrpsD2) against EcoRI-digested genomic DNA.
Transcriptional analysis of vlmS-proximal genes
The wild-type strain was statically cultivated in 20 ml of YES and SMY media in 100-ml Erlenmeyer flasks at 25 °C for 5 days. Total RNA was isolated from a mycelial mat using an RNeasy plant minikit (Qiagen) and treated with DNase I (Takara). The cDNA was synthesized using Superscript III RNaseH− (Invitrogen) with random primers (Invitrogen) according to the manufacturer’s instruction. RT-PCR was performed using GoTaq green master mix (Promega KK) with primer pairs (orfA1–orfA2, orfB1–orfB2, orfC1–orfC2, orfD1–orfD2, orfE1–orfE2, orfF1–orfF2, orfG1–orfG2, orfH1–orfH2, orfI1–orfI2, orfJ1–orfJ2, act1–act2, nrpsD1–nrpsD2; Table S1) under the following conditions: 94 °C for 5 min and 30 cycles of 94 °C for 30 s, 55 °C for 30 s, and 72 °C for 30 s, followed by a final elongation step at 72 °C for 7 min.
Disruption of vlmS-proximal genes
Each of five genes (orf-a, orf-b, orf-c, orf-d, and orf-g, clustering around vlmS) was disrupted by targeted replacement with pyrG. Each of the 5′-upstream fragments was amplified by PCR using PrimeSTAR® HS DNA Polymerase (Takara) or TaKaRa Ex Taq® (Takara) with primer pairs (orfAdisN1–orfAdisN2, orfBdisN1–orfBdisN2, orfCdisN1–orfCdisN2, orfDdisN1–orfDdisN2, and orfGdisN1–orfGdisN2, respectively), yielding 1.6-, 1.5-, 1.5-, 1.6-, and 1.4-kbp fragments, respectively. Similarly, 3′-downstream fragments were also PCR-amplified with primer pairs (orfAdisC1–orfAdisC2, orfBdisC1–orfBdisC2, orfCdisC1–orfCdisC2, orfDdisC1–orfDdisC2, and orfGdisC1–orfGdisC2, respectively), yielding 1.5-, 1.5-, 1.5-, 1.5-, and 1.8-kbp fragments, respectively. The cognate 5′- and 3′-fragments were cloned on pLSPYRG (for orf-b and orf-g) or pLSPYRG1200R (for orf-a, orf-c, and orf-d), making the pyrG gene flanked by the 5′- and 3′-fragments. The disruption vector was linearized by digestion with HindIII (for the orf-a, orf-b, and orf-d disruption vectors) or SacI (for the orf-c and orf-g disruption vectors) prior to transformation. Transformation of Ki2p was carried out as described previously (Ishidoh et al. 2014). Emerging colonies on the selection plates were screened by PCR using an FTA® classic card (Whatman®) with primer pairs (orfA2–pprv, orfF1–pprv, nrpsDup–pp1200fw, orfB1–pp1200fw, and orfC1–pp1200fw, respectively). The genotypes of candidate strains were confirmed by Southern hybridization using probes prepared by PCR with primer pairs (orfB1–orfB2 for orf-a and orf-b; orfC1–orfC2 for orf-c and orf-d; and orfH1–orfH2 for orf-g).
Verlamelin production in disruptants
The production profile of low molecular weight compounds was analyzed by HPLC. Cultures of the wild type and transformants were prepared by static cultivation with 20 ml of YES medium in 100-ml Erlenmeyer flasks at 25 °C for 5, 10, and 15 days. Whole cultures including the mycelial mat were extracted with n-butanol (10 ml) by mild shaking for 1 h, followed by vigorous shaking for 1 min. The n-butanol layer was collected after centrifugation (3,000 rpm, 10 min) and evaporated in vacuo, and the residue was dissolved in methanol or dimethyl sulfoxide (DMSO) (for the preparation of further concentrated samples) for HPLC analysis using an Imtakt Cadenza CD-C18 (Ø4.6 × 75 mm) column with a stepwise gradient of CH3CN from 15 to 85 % in 0.1 % formic acid aqueous solution at a flow rate of 1.2 ml/min (3 min at 15 %, 3 min from 15 to 40 %, 6 min at 40 %, 7 min from 40 to 45 %, 3 min from 45 to 85 %, 7 min at 85 %, 3 min from 85 to 15 %).
Feeding experiment of 5-hydroxytetradecanoic acid
5-Hydroxytetradecanoic acid was obtained by cleaving the lactone ring of δ-tetradecanolactone (TCI, Japan) by the following procedures. δ-Tetradecanolactone (50 mg) was dissolved in 1 M NaOH in methanol/water (4/1) and stirred overnight at room temperature. After neutralization with 1 M HCl, the reactant was extracted with n-hexane and concentrated in vacuo, followed by purification with silica gel chromatography using a hexane-AcOEt solvent system (4/1, 2/1, 1/1, 0/1). Fractions containing 5-hydroxytetradecanoic acid (Rf value 0.17, versus 0.5 for δ-tetradecanolactone) on silica gel TLC (Hexane/AcOEt; 2/1) were pooled and dried in vacuo, yielding 5-hydroxytetradecanoic acid (10 mg, m/z 245 by CI+MS). Disruptants of vlmA and vlmB were grown statically in YES liquid medium at 25 °C. After 4 days of cultivation, 5-hydroxytetradecanoic acid was added at a final concentration of 0.2, 0.4, or 0.8 mM, and the culture was harvested after an additional 6 days of cultivation for verlamelin analysis.
Accession numbers
The sequences of vlmS, vlmA, vlmB, and vlmC have been deposited in DNA Data Bank of Japan (DDBJ) under accession numbers AB862312, AB862313, AB862314, and AB862315, respectively.
Results
Identification of a NRPS gene responsible for verlamelin biosynthesis
The fact that the peptidyl moiety of verlamelin A and B was composed of D-amino acids as well as L-amino acids indicates that this moiety is probably synthesized by a nonribosomal peptide synthetase (NRPS). Based on the conserved sequences of the adenylation domains and peptidyl carrier protein domains in NRPSs (Marahiel et al. 1997), PCR primers (Table S1) were designed, and RT-PCR was conducted with RNA isolated from the Lecanicillium mycelia grown under verlamelin production conditions, resulting in five different PCR fragments encoding putative adenylation domains of NRPS. To identify the NRPS gene responsible for verlamelin biosynthesis, each adenylation domain was disrupted by inserting a pyrG gene through homologous recombination. HPLC analysis of low molecular weight metabolites from each transformant demonstrated that disruption of an adenylation domain from one of the PCR fragments caused a complete loss of verlamelin production (Fig. 2a), while disruption of other adenylation domains did not affect the production of verlamelin (data not shown).
HPLC analysis of the vlmS disruptant (a) and disruptants of orf-a, orf-b, orf-c, orf-d, and orf-g (b). Control strains were obtained by non-targeted integration of pLSPYRG (single asterisk) to the ki6 strain (Ishidoh et al. 2014) and non-targeted (double asterisks) integration of the orf-c disruption vector to the ki2p strain (Fig. S1c, lane 4), respectively. The chromatograms were recorded with detection at 210 nm. Arrows indicate the elution position of verlamelin A
Based on the sequence information causing the loss of verlamelin, the whole coding sequence of the NRPS gene was identified in the draft genomic sequence of Lecanicillium sp. HF627, and the NRPS gene responsible for verlamelin biosynthesis was named vlmS (AB862312). The vlmS gene containing one intron (50 n.t.) was 26,762 n.t., encoding an 8,903-amino-acid protein which showed highest similarity (90 %) to NRPS from Fusarium pseudograminearum (EKJ70673). Domain analysis with Pfam using the deduced amino acid sequence as query detected seven modules composed of seven peptidyl carrier protein domains, six adenylation domains, seven condensation domains, and four epimerization domains (Fig. 3b). Among the seven modules, only the initial module at the N-terminus lacked the adenylation domain. The epimerization domain in the initial module appeared to be inactive because the catalytic His residue was substituted to Ala. The number of modules and the presence of epimerization domains in the second, third, and sixth modules matched the structure of the peptidyl moiety of verlamelin A and B.
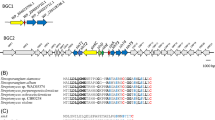
Verlamelin biosynthetic gene cluster. a Gene organization around vlmS. The direction of the white arrows corresponds to that of the reading frame. b Domain structure of VlmS. The white boxes labeled A represent adenylation domains. The striped boxes labeled C represent condensation domains. The dotted boxes labeled E represent epimerization domains. Black boxes represent peptidyl carrier protein domains. c RT-PCR analysis of ORFs found in the 50-kbp neighboring region of vlmS under the verlamelin-producing conditions [YES medium (upper panel) and SMY medium (lower panel)]. a, orf-a (vlmA); b, orf-b (vlmB); c, orf-c (vlmC); d, orf-d; e, orf-e; f, orf-f; g, orf-g; h, orf-h; i, orf-i; j, orf-j; act, actin gene as a positive control; s, vlmS. Pictures of gels were taken under UV light after staining with ethidium bromide. No transcription of orf-e, orf-f, orf-I, or orf-j was detected, although several PCR conditions and different primer sets were tested (data not shown)
Biosynthetic genes for verlamelin in the flanking regions of vlmS
Because all the genes necessary for the biosynthesis of a particular secondary metabolite are almost always clustered, both the 5′- and 3′-flanking regions of vlmS were analyzed to identify the remaining genes essential for verlamelin biosynthesis. Sequence analysis and a database search with BLASTX detected ten flanking genes (Fig. 3a and Table 1). Transcriptional analysis of the ten genes by RT-PCR using RNA isolated from cells grown under verlamelin production conditions revealed that six genes (orf-a, orf-b, orf-c, orf-d, orf-g, and orf-h) among the ten were transcribed (Fig. 3c), suggesting that vlmS and the six genes may constitute the verlamelin biosynthetic gene cluster, although orf-h (encoding AAA ATPase) should be eliminated from the putative cluster considering that ATPase seems unlikely to be involved in verlamelin biosynthesis.
To further examine which of the five genes are involved in verlamelin biosynthesis, each of the five genes was disrupted by homologous recombination with pyrG insertion as the marker. After confirming the genotype of uridine prototroph by Southern hybridization (Fig. S1; Online Resource), the disruptant of each gene was cultivated under verlamelin production conditions, and verlamelin production was analyzed by C18 HPLC (Fig. 2b). Verlamelin production was mostly/completely lost by disruption of orf-a, orf-b, and orf-c but remained intact by disruption of orf-d or orf-g, demonstrating that only three genes (orf-a, orf-b, and orf-c) together with vlmS are involved in verlamelin biosynthesis. Although the productivity of verlamelin was completely lost in both the orf-a and orf-c disruptants, it was slightly retained in the orf-b disruptant (less than 5 % of that in the wild type). As for the morphology and growth, no significant differences were observed between the disruptants (data not shown).
Based on these results, three genes (orf-a, orf-b, and orf-c) were concluded to be components of the verlamelin biosynthetic pathway and were named vlmA (AB862313), vlmB (AB862314), and vlmC (AB862315), respectively. VlmA showed similarity to proteins belonging to the fatty acid hydroxylase superfamily, including C-5 sterol desaturase ERG3 (NP_013157, 67 % similarity to VlmA) and methylsterol methyl monooxygenase ERG25 (NP_011574, 63 % similarity to VlmA) of Saccharomyces cerevisiae. VlmA contains three histidine-rich motifs (essential for metal binding) that are conserved among other proteins in this family. VlmC was deduced to be AMP-dependent ligase, which is classified into a group containing acyl-CoA synthetase and the NRPS adenylation domain, indicating that they catalyze the thioester formation via activation of carboxylic acid by adenylation (Gulick 2009). The genes with the highest similarity in the fungal secondary metabolism were EasD (XP_660153, 76 % similarity to VlmC) and EcdI (AFT91380 76 % similarity to VlmC), both of which are fatty acid-activating enzymes in the lipopeptide biosynthesis of emericellamide and echinocandin (Cacho et al. 2012; Chiang et al. 2008). As for VlmB, although three proteins belonging to the thioesterase superfamily showed high similarity (AET11897, EGU84193, and ENH63667, 83-92 %,), no information on their functions, substrates, or products was available.
Feeding experiment of 5-hydroxytetradecanoic acid
To identify the gene(s) involved in the supply of 5-hydroxytetradecanoic acid (an immediate precursor for verlamelin biosynthesis), a feeding experiment was conducted on the disruptants of vlmA and vlmB. The loss/decrease of verlamelin production was complemented both in vlmA (7.7, 9.6, and 23.4 % of the wild-type strain by 0.2, 0.4, and 0.8 mM precursor addition) and vlmB (9.3 and 12.4 % of the wild-type strain by 0.2 and 0.4 mM precursor addition), although the productivity was lower than that of the wild-type strain (Fig. 4). Thus, vlmA and vlmB were demonstrated to function in the 5-hydroxytetradecanoic acid synthesis.
Feeding experiment of 5-hydroxytetradecanoic acid. The chromatograms were recorded with detection at 210 nm. Arrows indicate the elution position of verlamelin A. Verlamelin A production level in the no-feeding control culture of vlmB disruptant was 3.3 % of wild-type level. The size of the peak eluted immediately before verlamelin A was much larger than that detected in Fig. 2 probably due to the difference of organic solvent (DMSO instead methanol) used to dissolve the culture extracts
Discussion
The NRPS gene responsible for verlamelin biosynthesis was identified and named vlmS (AB862312). Analysis with the Conserved Domain Database (CDD) revealed that VlmS is composed of seven modules, among which the initial module at its N-terminus lacks an adenylation domain (Fig. 3b). The lack of an adenylation domain in the initial module is consistent with the cases of EcdA (AFT91378) and EasA (XP_660149), which are NRPSs for biosynthesis of the lipopeptides echinocandin and emericellamide, respectively (Cacho et al. 2012; Chiang et al. 2008). In both cases, the initial module is considered to accept the fatty-acyl intermediate as the starter compound for further peptide elongation, and thus VlmS should be considered to accept a fatty-acyl intermediate onto the initial module to further elongate amino acid residues by the downstream modules. In addition, in the last module at its C-terminus, VlmS contains a surplus condensation (C) domain that should be active based on the presence of the conserved catalytic His residue and the high similarity with other C domains in the database (65–75 %) to each of seven C domains in EcdA (AFT91378)]. Because a set of domains in the typical domain arrangement (C domain–A domain–PCP domain) that is necessary for simple elongation of the last amino acid already exists in this module, the surplus C domain may have another function, such as cyclization as the last step to form verlamelin, from the analogy that most fungal NRPSs utilize an additional C-terminal thioesterase domain (Gao et al. 2012; Haynes et al. 2011) to cyclize linearly mature peptides.
Regarding the gene cluster, four genes [vlmS (AB862312), vlmA (AB862313), vlmB, (AB862314) and vlmC (AB862315)] were identified as the cluster components responsible for verlamelin biosynthesis. Among the ten genes existing in the vicinity of vlmS, five genes were initially selected from their coincidental transcriptions under the verlamelin production conditions and their putative functions in the biosynthesis of secondary metabolites (Fig. 3a and Table 1), but two (orf-d and orf-g) were eliminated based on the unaffected verlamelin production in the corresponding disruptants. It was unexpected that orf-g is not involved because it encodes a member of the Zn(II)2Cys6-type transcriptional regulators, which are commonly present in fungal secondary metabolic gene clusters (Baba et al. 2006; Shimizu et al. 2007; Yu et al. 1996), where they function to ascertain the synchronized expression of necessary structure genes. Because no other regulatory genes are present in the vicinity of vlmS, but direct/inverted repeats of GTAC-N1-3-GTA(C/G) are present in the 150–300 bp region upstream of only the four verlamelin biosynthetic genes, it remains to be clarified how verlamelin production is regulated.
Apart from the peptidyl moiety in the verlamelin structure, 5-hydroxytetradecanoic acid is the major component to be incorporated. However, no gene(s) encoding fatty acid synthase or polyketide synthase was present in the 50-kbp vicinity of vlmS, although fungal gene clusters for the fatty acid-containing compounds often possess a set of fatty acid synthase genes specific for the relevant component (Balakrishnan et al. 2013; Bhatnagar et al. 2003). Due to the fact that only a single set of genes encoding primary fatty acid synthase is present in the whole genome sequence of HF627, but no single PKS gene corresponding to tetradecanoic acid synthesis is present, the primary fatty acid synthase seems to synthesize the tetradecanoic acid, which should then be hydroxylated at position C5 by one of the vlm cluster enzymes to provide the immediate precursor, 5-hydroxytetradecanoic acid. The most likely candidate is VlmA (Fig. 5), which is highly similar (63 % similarity) to ERG25 (NP_011574), which catalyzes hydroxylation of the C4-methyl group of 4,4-dimethylzymosterol (Bard et al. 1996). Indeed, feeding of 5-hydroxytetradecanoic acid to the vlmA disruptant could partially restore the production of verlamelin A in a dose-dependent manner to 7.7-23.4 % of that in the wild-type, demonstrating that vlmA is involved in the synthesis of 5-hydroxytetradecanoic acid (Fig. 4). Similarly, feeding of 5-hydroxytetradecanoic acid to the vlmB disruptant could restore verlamelin production. VlmB shows similarity to acyl-CoA thioesterase in 4HBT family, and can be estimated to hydrolyze fatty acyl-CoA, synthesized by fungal primary FAS (Tehlivets et al. 2007). However, when tetradecanoic acid was fed to vlmB disruptant, verlamelin production was not restored (data not shown), suggesting that VlmB is not functioning in the supply of tetradecanoic acid. Both the products of vlmA and vlmB are altogether regarded as essential components in the biosynthesis of 5-hydroxytetradecanoic acid, while the precise function of VlmB still remains to be solved.
Scheme of verlamelin biosynthesis proposed from this study. The white circles labeled A represent adenylation domains. The black circles labeled C represent condensation domains. The dotted circles labeled E represent epimerization domains. The black boxes indicate peptidyl carrier protein (PCP) domains. Asterisk the E domain placed in the initial modules is considered to be inactive
To be loaded onto the waiting NRPS, 5-hydroxytetradecanoic acid should either be activated in the form of a CoA derivative by acyl-CoA ligase and transferred in the manner of EasD (acyl-CoA ligase) and EasC (acyltransferase) in the emelliceramide biosynthesis (Chiang et al. 2008) or activated in the form of acyladenylate by an AMP-dependent ligase, in the manner of EcdI in echinocandin biosynthesis (Cacho et al. 2012). Because in the vlm cluster no acyltransferase-like genes are present, but only VlmC, which is homologous to AMP-dependent ligase, verlamelin biosynthesis can be estimated to proceed via formation and loading of 5-hydroxytetradecanoyl adenylate onto VlmS.
Among the lipopeptide biosynthesis clarified in filamentous fungi, there are two pathways reported to supply the fatty acid moieties, as the cases of echicocandin B and apicidin, which contained C18 and C10 fatty acid moiety, respectively (Cacho et al. 2012; Jin et al. 2010). The in vitro experiment indicated that the C18 fatty acid moiety in echinocandin B comes from linoleic acid produced in primary metabolism, while the decanoic acid used for the fatty amino acid moiety in apicidins was deduced to be synthesized by specific FAS complex, followed by oxidation and transamination to supply acyl amino acids as immediate substrate for the NRPS. Similar to the case of echinocandin B, tetradecanoyl moiety in verlamelins seems to be synthesized by the primary FAS, whereas further hydroxylation by VlmA and VlmB is needed to provide immediate precursor to be incorporated. This study indicated a biosynthetic example of the fatty acid precursor for NRPS by partial modification of a fatty acid from primary metabolism, which has not been identified in fungal lipopeptide biosynthesis so far.
In this study, we isolated the vlm biosynthetic gene cluster. Gene disruption and precursor feeding suggested that verlamelin biosynthesis may have the unique characteristic of a fatty acid moiety derived from primary metabolism. The knowledge obtained in this study should provide new insight into the biosynthetic pathway or mechanism of fungal lipopeptides.
References
Baba S, Abe Y, Ono C, Hosobuchi M (2006) Targeted disruption of the genes, mlcR and ariB, which encode GAL4-type proteins in Penicillium citrinum. Biochim Biophys Acta 1759(8):410–416
Balakrishnan B, Karki S, Chiu SH, Kim HJ, Suh JW, Nam B, Yoon YM, Chen CC, Kwon HJ (2013) Genetic localization and in vivo characterization of a Monascus azaphilone pigment biosynthetic gene cluster. Appl Microbiol Biotechnol 97:6337–6345
Bard M, Bruner DA, Pierson CA, Lees ND, Biermann B, Frye L, Koegel C, Barbuch R (1996) Cloning and characterization of ERG25, the Saccharomyces cerevisiae gene encoding C-4 sterol methyl oxidase. Proc Natl Acad Sci U S A 93(1):186–190
Bhatnagar D, Ehrlich KC, Cleveland TE (2003) Molecular genetic analysis and regulation of aflatoxin biosynthesis. Appl Microbiol Biotechnol 61(2):83–93
Cacho RA, Jiang W, Chooi YH, Walsh CT, Tang Y (2012) Identification and characterization of the echinocandin B biosynthetic gene cluster from Emericella rugulosa NRRL 11440. J Am Chem Soc 134(40):16781–16790
Chiang YM, Szewczyk E, Nayak T, Davidson AD, Sanchez JF, Lo HC, Ho WY, Simityan H, Kuo E, Praseuth A, Watanabe K, Oakley BR, Wang CC (2008) Molecular genetic mining of the Aspergillus secondary metabolome: discovery of the emericellamide biosynthetic pathway. Chem Biol 15(6):527–532
Fujie A, Iwamoto T, Muramatsu H, Okudaira T, Nitta K, Nakanishi T, Sakamoto K, Hori Y, Hino M, Hashimoto S, Okuhara M (2000) FR901469, a novel antifungal antibiotic from an unidentified fungus No. 11243. I. Taxonomy, fermentation, isolation, physico-chemical properties and biological properties. J Antibiot 53(9):912
Gao X, Haynes SW, Ames BD, Wang P, Vien LP, Walsh CT, Tang Y (2012) Cyclization of fungal nonribosomal peptides by a terminal condensation-like domain. Nat Chem Biol 8:823–830
Gulick AM (2009) Conformational dynamics in the Acyl-CoA synthetases, adenylation domains of non-ribosomal peptide synthetases, and firefly luciferase. ACS Chem Biol 4(10):811–827
Haynes SW, Ames BD, Gao X, Tang Y, Walsh CT (2011) Unraveling terminal C-domain-mediated condensation in fungal biosynthesis of imidazoindolone metabolites. Biochemistry 50(25):5668–5679
Hornbogen T, Riechers SP, Prinz B, Schultchen J, Lang C, Schmidt S, Mügge C, Turkanovic S, Süssmuth RD, Tauberger E, Zocher R (2007) Functional characterization of the recombinant N-methyltransferase domain from the multienzyme enniatin synthetase. Chembiochem 8(9):1048–1054
Ishidoh KI, Kinoshita H, Ihara F, Nihira T (2014) Efficient and versatile transformation systems in entomopathogenic fungus Lecanicillium species. Curr Genet 60(2):99–108
Jin JM, Lee S, Lee J, Baek SR, Kim JC, Yun SH, Park SY, Kang S, Lee YW (2010) Functional characterization and manipulation of the apicidin biosynthetic pathway in Fusarium semitectum. Mol Microbiol 76(2):456–466
Kim JC, Choi GJ, Kim HJ, Kim HT, Ahn JW, Cho KY (2002) Verlamelin, an antifungal compound produced by a mycoparasite, Acremonium strictum. Plant Pathol J 18(2):102–105
Marahiel MA, Stachelhaus T, Mootz HD (1997) Modular peptide synthetases involved in nonribosomal peptide synthesis. Chem Rev 97(7):2651–2674
Onishi JC, Rowin GL, Miller JE (1980) Antibiotic A43F. United States Patent 4201771. May 6, 1980
Rowin GL, Miller JE, Albers-Schönberg G, Onishi JC, Davis D, Dulaney EL (1986) Verlamelin, a new antifungal agent. J Antibiot 39(12):1772–1775
Shimizu T, Kinoshita H, Nihira T (2007) Identification and in vivo functional analysis by gene disruption of ctnA, an activator gene involved in citrinin biosynthesis in Monascus purpureus. Appl Environ Microbiol 73(16):5097–5103
Slightom JL, Metzger BP, Luu HT, Elhammer AP (2009) Cloning and molecular characterization of the gene encoding the aureobasidin A biosynthesis complex in Aureobasidium pullulans BP-1938. Gene 431(1):67–79
Tehlivets O, Scheuringer K, Kohlwein SD (2007) Fatty acid synthesis and elongation in yeast. Biochim Biophys Acta 1771(3):255–270
Wang B, Kang Q, Lu Y, Bai L, Wang C (2012) Unveiling the biosynthetic puzzle of destruxins in Metarhizium species. Proc Natl Acad Sci U S A 109(4):1287–1292
Xu Y, Orozco R, Wijeratne EM, Gunatilaka AA, Stock SP, Molnár I (2008) Biosynthesis of the cyclooligomer depsipeptide beauvericin, a virulence factor of the entomopathogenic fungus Beauveria bassiana. Chem Biol 15(9):898–907
Yu JH, Butchko RA, Fernandes M, Keller NP, Leonard TJ, Adams TH (1996) Conservation of structure and function of the aflatoxin regulatory gene aflR from Aspergillus nidulans and A. flavus. Curr Genet 29(6):549–555
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 158 kb)
Rights and permissions
About this article
Cite this article
Ishidoh, Ki., Kinoshita, H. & Nihira, T. Identification of a gene cluster responsible for the biosynthesis of cyclic lipopeptide verlamelin. Appl Microbiol Biotechnol 98, 7501–7510 (2014). https://doi.org/10.1007/s00253-014-5803-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-014-5803-7