Abstract
Biosurfactants (BSs) are a class of secondary metabolites representing a wide variety of structures that can be produced from renewable feedstock by a wide variety of micro-organisms. They have (potential) applications in the medical world, personal care sector, mining processes, food industry, cosmetics, crop protection, pharmaceuticals, bio-remediation, household detergents, paper and pulp industry, textiles, paint industries, etc. Especially glycolipid BSs like sophorolipids (SLs), rhamnolipids (RLs), mannosylerythritol lipids (MELs) and cellobioselipids (CBLs) have been described to provide significant opportunities to (partially) replace chemical surfactants. The major two factors currently limiting the penetration of BSs into the market are firstly the limited structural variety and secondly the rather high production price linked with the productivity. One of the keys to resolve the abovementioned bottlenecks can be found in the genetic engineering of natural producers. This could not only result in more efficient (economical) recombinant producers, but also in a diversification of the spectrum of available BSs as such resolving both limiting factors at once. Unraveling the genetics behind the biosynthesis of these interesting biological compounds is indispensable for the tinkering, fine tuning and rearrangement of these biological pathways with the aim of obtaining higher yields and a more extensive structural variety. Therefore, this review focuses on recent developments in the investigation of the biosynthesis, genetics and regulation of some important members of the family of the eukaryotic glycolipid BSs (MELs, CBLs and SLs). Moreover, recent biotechnological achievements and the industrial potential of engineered strains are discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
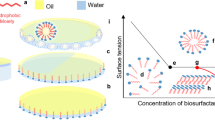
Secondary metabolites produced by microorganisms consist of a wide range of molecules, which have found applications in all aspects of daily human life. One family of secondary metabolites produced by a wide variety of microorganisms are biosurfactants (BSs). These surface-active agents are capable of reducing surface and interfacial tension at the interfaces between immiscible liquids, solids and gases allowing them to disperse readily as emulsions in water or other liquids. Their biodegradability, diverse biological activities and the fact that they can be produced from renewable resources give them an advantage over their chemical counterparts and may therefore make them suitable to (partly) replace chemicals in the future (Banat et al. 2010; Marchant and Banat 2012). However, two major factors still limit the real penetration of BSs into the market: firstly, the limited available structural variety and secondly, the high production price, which is often linked with low productivities of the producing organisms. One of the keys to resolve the abovementioned bottlenecks and to obtain broad application of microbially derived BSs in the industry, is a detailed knowledge of the genetics of the producing organisms. Genetic engineering of natural producers could not only result in more efficient (economical) recombinant producers, but also in a diversification of the spectrum of available BS, thus resolving both limiting factors at once.
BS produced by eukaryotes include polymeric surfactants like liposan (Yarrowia lipolytica) and mannoprotein (Saccharomyces cerevisiae), fatty acids like trachyspic-, decylcitric- and spiculisporic acid (Talaromyces trachyspermus) and polyol lipids (Aerobasidium sp.). However, the best-studied BSs produced by eukaryotes are all members of the family of the glycolipids. Despite the fact that most of these molecules were already discovered more than half a century ago, the genetic background of their production and regulation thereof remained largely unknown for a long time. This is in contrast with other secondary metabolites produced by eukaryotes for which a load of literature is available. Generally spoken the genes encoding the biosynthetic pathways of fungal secondary metabolites can be found in large genomic gene clusters (Brakhage and Schroeckh 2011), which are often found near the telomere(s). These clusters often contain a pathway specific ‘in cluster regulator’ (e.g., aflatoxin; Woloshuk et al. 1994), while for others no pathway specific regulator can be found inside the gene cluster (e.g., penicillin; Martin 2000). Such features were recently also discovered for eukaryotic BSs. Although numerous reviews are available on (eukaryotic) BS production and especially on the possible applications of these molecules (Amaral et al. 2010; Marchant and Banat 2012; Morita et al. 2013b; Rodrigues and Teixeira 2010; Van Bogaert et al. 2007), no reviews specifically dealing with the genetic background and regulatory networks affecting eukaryotic BS production are available yet. However, some important scientific breakthroughs concerning the genetics of eukaryotic glycolipid BS have been achieved the last 5–10 years. Biosynthetic pathways and corresponding gene clusters are now described for mannosylerythritol lipids (MELs), sophorolipids (SLs) and cellobioselipids (CBLs) like ustilagic acid (UA) and flocculosin (FL). Moreover, some important insights concerning the molecular regulation of the biosynthesis of some of these molecules have been obtained and several biotechnological achievements and opportunities have been described. Therefore this mini-review focuses on the recent findings on the genetics and regulation of eukaryotic glycolipid BS. Similar functions in glycolipid biosynthesis are emphasized in the figures by the color of the structural genes in the respective gene clusters. Moreover, biotechnological achievements, opportunities and industrial relevance are discussed.
Eukaryotic glycolipid biosurfactants: genetics and regulation
Mannosylerythritol lipids
MELs are produced by a variety of fungal species, e.g., Schizonella melanogramma (shizonellin), Pseudozyma sp., Ustilago maydis, Kurtzmanomyces sp. and Geotrichum candidum (Fukuoka et al. 2008; Haskins 1950; Kakugawa et al. 2002; Kitamoto et al. 1990; Kurz et al. 2003; Rau et al. 2005b), with highest reported yields of 140 g/l with P. antarctica (Kitamoto et al. 2001a) and 165 g/l with P. aphidis (Rau et al. 2005a), both under optimal fed-batch conditions. Amongst MEL producers, the dimorphic basidiomycete U. maydis and other ustilaginomycetous yeasts like P. aphidis, P. fusiformata and P. graminicola (Kulakovskaya et al. 2005; Morita et al. 2013a) are special, because besides MELs they also produce a second structurally different class of glycolipids (CBLs), which will be described in the next section. MEL (Fig. 1), an extracellular oil heavier than water, generally consists of a mannosylerythritol disaccharide of which the mannosyl moiety is acetylated (position R4 and R6) and acylated with short-chain (C2 to C8) and medium-chain (C10 to C18) fatty acids (positions R2 and R3). Depending on the number of acetyl groups, MELs can be differentiated into four different varieties: MEL-A (fully acetylated), MEL-B and MEL-C (mono-acetylated at R6 and R4, respectively) and the fully deacetylated MEL-D (Kurz et al. 2003). Advanced microbial screening methods recently resulted in the discovery of a novel type of MEL (Morita et al. 2011) and even mannosyl-mannitol–, –arabitol– and –ribitol-lipids (Morita et al. 2012; Morita et al. 2009).
MELs were the first glycolipids produced by fungi or yeast for which a biosynthetic gene cluster was discovered (Hewald et al. 2006). This gene cluster of the smut fungus Ustilago maydis contains five genes (Fig. 2) encoding the complete biosynthetic pathway. The first gene encodes a glucosyltransferase (emt1) responsible for the first step of the biosynthetic pathway: a stereospecific mannosylation (derived from GDP-mannose) of meso-erythritol at the C4 position. A second gene encodes an acetyltransferase (mat1) responsible for acetylation both at the R4 and R6 positions of the mannose moiety. The former occurs after specific acylation by the acyltransferases (mac1 and mac2) at position R2 with a short chain fatty acid (C2 to C8) or R3 with a medium or long chain length fatty acid (C10 to C18). These last two enzymes are highly specific, both in regioselectivity (in contrast to the acetyltransferase Mat1) as in their preference for the length of the acyl-CoA cofactor. The secretion of MELs which are only acetylated at one position indicates that the second acetylation reaction is significantly slower than the first one. Alternatively, these intermediates could be the result of glycolipid catabolism as was demonstrated for the CBL FL (Mimee et al. 2009) and SLs (Roelants 2013). The fifth gene of the cluster encodes a protein of the family of the major facilitators (mmf1). Deletion of the mmf1 gene leads to complete abolishment of extracellular MEL production, indicating that the major facilitator Mmf1 is essential for secretion and that no other secretion systems exist for MELs (Hewald et al. 2006). The lack of MEL production for Δmac1 and Δmac2 mutants was attributed to the selectivity of the transporter protein and acylation at both positions thus appears to be a prerequisite for glycolipid secretion (Hewald et al. 2006). Furthermore, mannosylerythritol has been isolated in significant amounts from MEL-producing cells (Boothroyd et al. 1956), but was not found extracellularly, again suggesting that acylation is necessary for secretion. EST analysis resulted in the identification of the homologue of the emt1 gene of Pseudozyma antarctica (Morita et al. 2010b) and genome sequencing recently resulted in the discovery of the complete MEL biosynthetic gene cluster of this yeast (Morita et al. 2013c). The genes of this cluster show high levels of identity to the corresponding genes in U. maydis and was thus suggested to function in the same way as that of the latter. Further investigation of the genomic analysis and sequence and development of a gene expression system will enable the elucidation of MEL biosynthesis in Pseudozyma yeasts in the future (Morita et al. 2013b).
Expression of all the members of the U. maydis gene cluster is highly induced under conditions of nitrogen starvation. Although no obvious conserved sequence motifs possibly involved in the co-regulation of the genes are present in the promoter regions of the cluster genes, several GATA sequences were found in all of them. This resulted in the suggestion that a GATA factor homologous to the general nitrogen regulator AreA from A. nidulans might be involved in regulation of MEL biosynthesis (Hewald et al. 2006). No in cluster regulator was identified in the U. maydis MEL biosynthetic gene cluster and the borders of the cluster are thus defined by mat1 (um03114) and mac2 (um10636). A candidate transcription factor possibly specifically involved in the regulation of MEL biosynthesis in U. maydis is um02717 (Nit1) (Teichmann et al. 2010) (cfr. CBLs).
Cellobioselipids
A small number of basidiomycetous yeasts, mostly Ustilaginales, secrete CBLs. UA (Fig. 3a) was the first one to be described (Haskins 1950) and consists of a cellobiose moiety linked O-glycosidically to the terminal hydroxyl group of tri- (2,15,16-) or di- (15,16-) hydroxypalmitic acid. The sugar moiety is additionally esterified with an acetyl and a short chain β-hydroxy fatty acid (C6 or C8) (Kulakovskaya et al. 2010). Titers of 16 g/l of UA have been obtained with resting cells of U. maydis (Frautz et al. 1986). Production of CBLs varying in the decorations of the sugar moiety and the hydroxylation pattern of the lipid tail has since been described for other fungal species like Pseudozyma graminicola, P. fusiformata, P. flocculosa, P. aphidis, P. hubeiensis, Trichosporon porosum, Sympodiomycopsis paphiopedili and Cryptococcus humicola (Golubev et al. 2008; Kulakovskaya et al. 2004, 2005, 2010; Mimee et al. 2005; Morita et al. 2013a; Puchkov et al. 2002). Amongst them is the fungal biocontrol agent P. flocculosa, which produces flocculosin (FL) (Fig. 3b), a rare CBL with antifungal activity (Cheng et al. 2003), for which titers of 14 g/l have been reported (Hammami et al. 2008). The lipid tail in this case consists of a (3,15,16-) trihydroxypalmitic acid. FL is acetylated at the 6′ and 3″ position in contrast with UA, which is only acetylated at the 6′ position. Another difference is the acylgroup at the 2″ position, which in the case of FL exclusively consists of a C8 β-hydroxyfatty acid (Teichmann et al. 2011a).
Molecular structure of a ustilagic acid (UA) and b flocculosin (FL), both cellobioselipids (CBL). For UA, the cellobiose moiety is esterified with β-hydroxy-octanoic or β-hydroxy-hexanoic acid and is acetylated at only one position, R = H or OH. The cellobiose moiety of FL is esterified with β-hydroxy-octanoic acid and acetylated at two positions. The respective differences are emphasized in the figure
Shortly after the discovery of the MEL biosynthetic gene cluster of U. maydis, a second gene cluster was discovered in this organism, this time responsible for CBL biosynthesis (Fig. 4a) (Teichmann et al. 2007). This led scientists to suggest the existence of a similar cluster in P. flocculosa (Marchand et al. 2009), which was proven to indeed be the case (Teichmann et al. 2011a) (Fig. 4b). The two gene clusters (45 and 60 kb, respectively) are very similar in gene presence and function and likely evolved from a common origin, which is also the case for the two described MEL biosynthetic gene clusters. The respective genes are all involved in CBL production, decoration and/or secretion. Biosynthesis is thus very similar and will hence be described combined for both organisms. In the following description the gene names of U. maydis will precede the corresponding ones of P. flocculosa (the reader is referred to Fig. 4 for gene names). The first step of the CBL biosynthesis consists of the double hydroxylation of palmitic or β-hydroxypalmitic acid for U. maydis and P. flocculosa, respectively, at the terminal (ω) and subterminal (ω-1) positions (cyp1 and cyp2, respectively, for both organisms). This hydroxylated fatty acid is subsequently glycosylated by a glucosyltransferase (ugt1 vs. fgt1) at the terminal hydroxylgroup by either the sequential addition of two glucose molecules derived from UDP-glucose or of a cellobiose moiety as a whole (the latter remains to be determined) (Teichmann et al. 2011a; Teichmann et al. 2007). This CBL is subsequently further decorated at position 2″ of the cellobiose moiety by the action of an acyltransferase (uat1 vs. fat1). Both clusters contain genes responsible for synthesis (fas2 for both) and β-hydroxylation (uhd1 vs. fhd1) (Teichmann et al. 2011b) of this short-chain fatty acid. Further decoration (acetylation) of the glycolipid at the 6′ position is executed by an acetyltransferase (uat2 vs. fat2). The presence of an additional acetylgroup at the 3″ position of FL is caused by the presence of an additional acetyltransferase gene (fat3) in the FL gene cluster, without homologue in the UA gene cluster. For U. maydis, the last step in the UA biosynthetic pathway consists of hydroxylation (ahd1) at the α-position of the C16 dihydroxy fatty acid of the synthesized CBL, whereas for FL the hydroxyl group at the β-position is already present in the substrate (β-hydroxypalmitic acid) before hydroxylation carried out by Cyp1 occurs. The gene responsible for this β-hydroxylation remains to be discovered, but it was suggested that β-hydroxy fatty acids could be derived from de novo fatty acid synthesis (Teichmann et al. 2011a), as is described for rhamnolipid biosynthesis trough the action of the PhaG enzyme (Deziel et al. 2003; Rehm et al. 1998). Finally, both gene clusters contain an ABC transporter (atr1 for both) responsible for the export of the respective CBLs, which is in contrast to the MELs, where the transporter is a member of the family of the major facilitators. The only two differences between the two CBL gene clusters — the unique presence of ahd1 and fat3 in the UA and FL gene clusters, respectively — thus account for the structural differences between the produced CBL molecules. Two putative homologues of cyp1 of U. maydis were recently isolated from the newly identified CBL/MEL producers P. aphidis and P. hubeiensis (Morita et al. 2013a). Moreover, a screening method to activate silent gene clusters recently resulted in the discovery of a new kind of CBLs produced by U. maydis (Yang et al. 2013).
The CBL biosynthetic gene cluster from a U. maydis (40 kb) and b P. flocculosa (60 kb) containing two cyp monooxygenase genes cyp1 and cyp2, one glucosyltransferase gene ugt1 vs. fgt1, an acetyltransferase gene for acetylation at position 6′ uat2 vs. fat2 and a second acetyltransferase gene fat3 for acetylation at position 3′ of FL. A fatty acid synthase gene fas2 is responsible for synthesis of the short chain length fatty acid and uhd1 vs. fhd1 for hydroxylation thereof. The hydroxylated fatty acid is subsequently attached at position 2″ by an acyltransferase (uat1 vs. fat1). The ahd1 gene is responsible for α-hydroxylation of the long chain length fatty acid of UA. An ABC transporter gene (atr1) is responsible for CBL transport and rua1 vs. rfl1 are pathway specific ’in cluster’ regulators
UA biosynthesis in U. maydis occurs under nitrogen starvation. Since expression of the first gene of the pathway (cyp1) was found to be strongly induced under such conditions (Hewald et al. 2005) and several GATA DNA motifs were also found in the promoter sequences of the UA cluster genes, regulation of UA biosynthesis was again (~MELs) suggested to involve direct or indirect regulation by a GATA factor like AreA (Hewald et al. 2005). However, a very interesting finding was done by Teichmann et al. (2010) when they demonstrated the leftmost gene of the UA gene cluster (rua1) to be an in cluster regulatory protein similar as for other fungal secondary metabolites. Deletion of rua1 leads to complete loss of UA production, whereas overexpression promotes increased UA synthesis, even in the presence of a good nitrogen source (Teichmann et al. 2010). Rua1 is thus both necessary and sufficient to trigger UA biosynthesis, which indicates that all environmental signals that affect UA biosynthesis are integrated at the rua1 promoter. Moreover, Rua1 acts as a cluster-specific regulator as MEL biosynthesis is not affected. Rua1 was found to bind directly to a conserved sequence element present in all promoters of the UA cluster (except for rua1 itself), which mediates Rua1-dependent expression of the UA biosynthetic gene cluster in a nitrogen dependent way. A positive feedback regulation for Rua1 was suggested, as a point mutation in Rua1 not only results in an abolishment of UA production, but also in the abolishment of Rua1 expression. On another level, it was suggested that rua1 expression could be subject to posttranscriptional control as is described for the global nitrogen regulator AreA in A. nidulans where AreA mRNA is specifically degraded in response to intracellular glutamine (Caddick et al. 2006). The FL gene cluster is flanked by a gene (rfl1) which contains a C2H2 zinc finger region highly similar to that of rua1 and this gene is thus most likely an in cluster regulator for FL biosynthesis (Teichmann et al. 2011a). The exact mechanisms regulating the expression of rua1 (and rfl1) remain to be elucidated, but regulatory proteins that sense global and more specific nitrogen availability like the U. maydis AreA/Nit2 homolog were suggested. However, the U. maydis Nit2 homolog was recently shown not to be involved in UA biosynthesis (Horst et al. 2012), so other regulatory proteins, e.g., the homolog of the pathway specific activator Nit4, are likely to be involved. Another candidate transcription factor possibly specifically involved in the nutrient control of secondary metabolism is Nit1 (Teichmann et al. 2010). Nit1 was suggested to be involved in nitrogen dependent transcription of Rua1. Interestingly, deletion of this gene affects both UA and MEL biosynthesis in U. maydis. Besides nitrogen limitation, other factors affecting CBL biosynthesis are the carbon source (Haskins 1950; Spoeckner et al. 1999) and the cellular stage of the culture (Hammami et al. 2008; Kitamoto et al. 1992).
Sophorolipids
Glycolipids consisting of a sophorose molecule linked O-glycosidically to a fatty acid, i.e., SLs (Fig. 5), were first described in 1961 as glycolipids produced by the yeast Torulopsis magnolia (Gorin et al. 1961). The hydrophobic part was identified as 17-hydroxystearate (C18:0) and 17-hydroxyoleate (C18:1), but also intermediates with 16 C-atoms were detected (Tulloch et al. 1962). These SLs contain acetate groups at the 6′ and 6″ positions (Tulloch et al. 1967) and a macrocyclic lactone structure formed between the 4″ hydroxyl group of the terminal glucose molecule and the hydroxyl acid carboxyl group can also be present (Tulloch et al. 1967) (Fig. 5b). Several closely related SL producing yeasts like Starmerella bombicola (formerly known as Candida bombicola), C. batistae, C. riodocensis, C. apicola, C. stellata and Candida sp. NRRL Y-27208 (Kurtzman et al. 2010), but also less related organisms like Wickerhamiella domercqiae (Chen et al. 2006) produce SLs. The highest reported titers are 422 g/l in a two stage process with S. bombicola and Cryptococcus curvatus (Daniel et al. 1999) and over 300 g/l using S. bombicola alone (Davila et al. 1997; Kim et al. 2009; Pekin et al. 2005). Although these yeasts all produce SLs mainly consisting of a C18:1 hydroxyfatty acid (sub)terminally linked to a sophorose molecule, a clear diversity of the produced SL mixture was demonstrated (Kurtzman et al. 2010). Moreover, a different kind of SLs for which the fatty acid consists of docosanoic acid (C22:0), attached to the sophorose moiety by internal hydroxylation of the fatty acid (Fig. 5c) were found to be produced by Rhodotorula bogoriensis (formerly Candida bogoriensis) (Tulloch et al. 1968). These similarities and respective differences are most probably a reflection of the genetic background of the producing organisms as was demonstrated to be the reason for the structural differences between UA and FL (cfr. CBLs). Recent screening for SL producing yeasts resulted in the discovery of a new species Candida sp. NRRL Y-27208 that produces significant amounts of novel SLs (Price et al. 2012).
Although the involvement of glucosyl- and acetyltranferases in SL biosynthesis of R. bogoriensis was already described in the early seventies (Esders and Light 1972b), it was only in 2009 that the discovery of a gene involved in SL biosynthesis by the industrially important yeast S. bombicola was reported (Van Bogaert et al. 2009). Today, the complete SL biosynthetic pathway of S. bombicola is described (Saerens et al. 2011a,b,c) and not unexpectedly the responsible genes were recently also described to be grouped in one large subtelomeric gene cluster (Van Bogaert et al. 2013b) (Fig. 6). Only the gene responsible for the lactonisation of SLs was found elsewhere in the genome (Ciesielska et al. 2014) and expression of this gene was also found to be differentially regulated as the other genes of the biosynthetic pathway (Ciesielska et al. 2013; Roelants 2013). The first step in SL biosynthesis consists of (sub)terminal hydroxylation of a fatty acid (cyp52M1), preferably oleic acid (C18:1). In contrast to CBL biosynthesis, two glucosylstransferases instead of one are responsible for subsequent glycosylation of the hydroxylated fatty acid. The first glucosyltransferase (ugtA1) (Saerens et al. 2011a) is responsible for the transfer of a first glucose molecule from UDP-glucose, while the second one (ugtB1) (Saerens et al. 2011c) specifically transfers a second glucose molecule from UDP-glucose to the formed glucolipid and not to the hydroxylated fatty acid as deletion of ugtA1 results in a complete abolishment of SL production. The formed SLs are subsequently acetylated by the action of an acetyltransferase (at) (Saerens et al. 2011b) and secreted by an ABC transporter (mdr) (Van Bogaert et al. 2013b) into the extracellular space, where they are further lactonised by the action of an only recently identified cell wall-bound lactonesterase (le) (Ciesielska et al. 2014). The involvement of a lipase-like enzyme in lactonisation of SLs was already suggested for SLs produced by Candida apicola (Hommel et al. 1994b) and a similar gene is probably present in this and other yeasts producing lactonic SLs, while it is probably absent or not activated for others, solely producing acidic SLs.
The SL biosynthetic gene cluster of S. bombicola (11 kb) containing a cyp52M1 monooxygenase gene, two glucosyltransferase genes ugtA1 and ugtB1, an acetyltransferase gene at, an ABC transporter gene mdr. The lactonesterase (le) responsible for lactonisation of SLs is not present in the SL biosynthetic gene cluster, but is located at the other side of the same chromosome
Similarly, as described above for CBL biosynthesis, the nitrogen and carbon source (hydrophobic and hydrophilic) and their ratio, as well as the cellular stage of the culture, influence the production of SLs (Albrecht et al. 1996; Cutler and Light 1979; Davila et al. 1997; Hommel and Huse 1993; Hommel et al. 1994a). However, also pH, pO2, the culture method and the presence of citrate in the culture medium were shown to affect SL biosynthesis (Davila et al. 1997; Guilmanov et al. 2002; Roelants 2013; Stüwer et al. 1987). Although SL production by S. bombicola is often described as a two stage process for which growth and production are clearly separated (Cooper and Paddock 1984), both the glucosyltransferases (UgtA1 and UgtB1) and the acetyltransferase (At) are already active in mid exponential S. bombicola (Saerens 2012) and R. bogoriensis (Esders and Light 1972a; Esders and Light 1972b) cell lysates. These results are in line with the conclusions of Albrecht et al. (1996), which suggested that the enzymes involved in SL formation are, at least at a low basal level, constitutive and that nitrogen (and/or phosphate) depletion indirectly leads to enhanced SL synthesis through an intracellular citrate accumulation, which is needed to supply acetyl-CoA for fatty acid biosynthesis. Clear upregulation of the SL biosynthetic gene cluster of S. bombicola in the stationary growth phase was recently demonstrated by transcriptomics (Roelants 2013) and proteomics (Ciesielska et al. 2013), so probably a combined effect leads to enhanced production of SLs in the stationary growth phase. Furthermore, similarly as for CBLs the C/N ratio regulates SL biosynthesis in Rhodotorula bogoriensis (Cutler and Light 1979) and S. bombicola (Casas and Garcia-Ochoa 1999). For C. apicola, not only the C/N ratio, but also the absolute quantity of N was suggested to regulate SL biosynthesis (Hommel et al. 1994a), whereas for Wickerhamiella domercqiae a clear regulatory effect of the type of nitrogen source on SL biosynthesis was demonstrated (Ma et al. 2011). High glucose concentrations (or C/N ratio) were shown to be absolutely necessary for upregulation of the genes of the SL gene cluster of S. bombicola at the transcriptome level (RNA sequencing) (Roelants et al. 2013). Interestingly, the expression of the gene responsible for lactonisation (le) was not significantly regulated by absolute glucose concentrations, which indicates independent regulation of the le gene in contrast to coregulation of the SL cluster genes. No ‘in cluster’ regulator is present in the SL gene cluster of S. bombicola (Van Bogaert et al. 2013b). However, an SL cluster specific regulator might still exist and several candidates were found in the S. bombicola genome (Roelants 2013) and are the subject of continuing research.
Applications and biotechnological achievements/opportunities for eukaryotic BS
The expanding knowledge of the genetics of BS-producing organisms is of significant importance as this knowledge represents the necessary base for the genetic engineering of BS producers. The latter is indispensable for the development of enhanced recombinant strains, which may well become industrial strains. Some of the efforts made in this respect for the abovementioned BSs in terms of structural variety and productivity are discussed below. The reader is referred to the sections above for the respective gene names. The suggested applications for these (new-to-nature) BSs are only briefly discussed, and the reader is referred to more extensive reviews for more information.
Mannosylerythritol lipids
In recent years, the interest in MELs has increased due to their high biodegradability, mild production conditions and a variety of biological functions, which will broaden their application in new technology areas (Kitamoto et al. 2009; Morita et al. 2013b). For example, Japanese companies like Kanebo, Daito Kasei Kogyo Co., Ltd., and Toyobo (SurfMellow®) have commercialized and patented these molecules as cosmetic ingredients (Kitagawa et al. 2012; Kitagawa and Yamamoto 2009; Morita et al. 2013b). Moreover, Aventis Pharma Deutschland GmbH published a patent concerning a MEL (ustilipid) produced by U. maydis DSM 11494 to be used in the treatment of schizophrenia or diseases caused by dopamine metabolic dysfunction (Vertesy et al. 1998). Other applications of MEL, such as chemical tools for purification of proteins or as anti-agglomeration agents of ice-slurry are known (Kitamoto et al. 2000, 2001b; Im et al. 2003). The first example of successful metabolic engineering of extracellular glycolipids produced by yeasts or fungi was the exclusive production of fully deacetylated MELs (MEL-D) by deletion of one gene (mat1) of the MEL biosynthetic pathway (Hewald et al. 2006). An alternative way of producing this compound (and its diastereomer) was recently shown by (in vitro) lipase-catalyzed hydrolysis of MEL-B and its respective diastereomer (Fukuoka et al. 2011, 2012). Furthermore, lipase-catalyzed acylations of MEL-A and MEL-B with uncommon fatty acids from other microbial glycolipids yielded functionalized products at the C-1 position of erythritol (Recke et al. 2013a). Hewald et al. (2006) suggested production of fully acetylated MEL-A through overexpression of this acetyltransferase as a large fraction of MELs acetylated at only one position is secreted by wild type U. maydis cells. The importance of such achievements was stressed as MEL-A has some interesting properties not shared by the other less acetylated variants, e.g., a dramatic increase in gene transfection efficiency of liposomes (Igarishi et al. 2006; Yonetani et al. 2005) and the formation of large vesicles called coacervates (Imura et al. 2004). Secretion of non-acetylated MELs (Δmat1) and a large fraction of MELs acetylated at only one position (wild type) indicates limited specificity of the MEL transporter, which is typical for members of the large family of major facilitators and interesting for the production of tailor-made MELs. Hewald et al. (2005) also attempted to stimulate MEL production by overexpressing the glycosyltransferase gene (emt1) with the arabinose-inducible crg promoter. Although this is a strong promotor, only weak MEL production could be observed, which was suggested to be attributed to low glycolipid production in the presence of arabinose as carbon source (Hewald et al. 2005). Improved MEL production was later achieved through overexpression of a mitochondrial ADP/ATP carrier (aac1), which was found to be highly expressed in MEL producing conditions (Morita et al. 2010a). Mannosylerythritol compounds have been used for the production of diverse products (Desai and Banat 1997) and the Emt1 protein was therefore also suggested to have some interesting biotechnological applications (Hewald et al. 2005). Deletion of this gene in U. maydis furthermore results in strains unable to produce MELs (Hewald et al. 2005), which is very important for the exclusive production of CBLs with this fungus (and others producing both BSs).
Cellobioselipids
CBLs are of interest with respect to the development of novel biological antifungal preparations (Kulakovskaya et al. 2010). A biocontrol product (Sporodex ®) based on the conidia of the basidiomycetous P. flocculosa active against powdery mildews is already commercialized (Jarvis et al. 2007). A patent of Unilever deals with the use of CBLs (and other glycolipids) in laundry detergents (Hall and Haverkamp 1991). Only U. maydis has hitherto been subject to genetic engineering approaches as no good transformation protocol exists yet for P. flocculosa (Teichmann et al. 2007, 2011a). Most research aims for the production of tailor-made CBLs. Expression of the P. flocculosa fat3 gene in U. maydis for example results in the production of four additional UA derivatives corresponding to molecules carrying an additional acetyl group as compared to the wild type molecules (Teichmann et al. 2011a). Furthermore, deletion of ahd1 responsible for α-hydroxylation leads to secretion of the naturally occurring UA derivatives lacking the α-hydroxyl group on the long chain fatty acid, whereas deletion of the cyp2 gene leads to the secretion of novel UA variants lacking the subterminal hydroxyl group. Deletion of uhd1 on the other hand only results in the absence of the β-hydroxyl group of the short chain fatty acid attached to the distal glucose molecule (Teichmann et al. 2011b), whereas deletion of uat2 leads to the production of UA derivatives lacking the acetyl group on the proximal glucose molecule (Teichmann et al. 2011a). The Atr1 transporter appears to be quite unspecific, as many of the UA derivatives that are produced by the mutants are readily exported, which is important when the production of tailored molecules is aimed for. Finally, a codon optimized version of the glucosyltransferase of U. maydis (ugt1) has been used for tailored CBL production in the SL producer S. bombicola (Roelants et al. 2013). The high selectivity of cyp1 and cyp2 for hydroxylation either at the ω or ω-1 position, respectively, was suggested to be attractive for biotechnological applications that require regio-specific introduction of hydroxyl groups (Teichmann et al. 2007). One could use them to specifically synthesize hydroxylated fatty acids, which have been reported to be valuable compounds in the chemical and medical industry (Bitto et al. 2009). The same holds true for the gene products of uhd1 and fhd1, which exhibit specific substrate specificity for β-hydroxylation of short-chain fatty acids. The most exciting finding is the fact that constitutive expression of the regulator of UA biosynthesis of U. maydis (Rua1) gives rise to production of CBLs even under non-inducing conditions (Teichmann et al. 2010). Such a specific regulator could represent a powerful tool for industrial production of CBLs.
Sophorolipids
The efficient production of SLs by S. bombicola resulted in commercialization of SLs for applications in, e.g., cosmetics produced by Soliance (Sopholiance), the ecological cleaning products of Ecover and Wheatoleo (Sophoclean), dishwasher products of Saraya (Sophoron) and the filing of numerous patent applications. A review of 255 worldwide patents on BSs (Shete et al. 2006) demonstrated that 24 % of those acted on SLs, clearly demonstrating the international commercial interest in SLs. Due to their biological activity they also find potential applications as antimicrobial, immune modulating, antiviral and anticancer agents (Van Bogaert et al. 2007). Enzymatic or chemo-enzymatic synthesis and/or modification of SLs to produce customized SL derivatives and/or biopolymers have been the subject of numerous research papers (Azim et al. 2006; Bisht et al. 1999, 2000; Carr and Bisht 2003; Gross et al. 1999; Imura et al. 2010; Nunez et al. 2004; Rau et al. 2001; Recke et al. 2013b; Singh et al. 2003), and several patent applications dealing with such modified SLs can be found in the patent literature (Giessler-Blank et al. 2009; Gross and Thavasi 2004; Tao 2011). When the complete SL biosynthetic pathway of S. bombicola was discovered, the genetic modification approach was initiated. Whereas deletion of the gene responsible for the first (cyp52M1 gene) or second (ugtA1 gene) step in SL biosynthesis results in a complete abolishment of SL production without any effect on the viability or growth rate of the yeast cells, deletion of the second glucosyltransferase (ugtB1) results in the secretion of (acetylated) glucolipids (Saerens et al. 2011c). Acidic glucolipids (and SLs) especially attract attention because they are asymmetrical bola-amphiphiles that, in addition to the supramolecular structures they typically form, have increased chemical versatility as compared to the chemically synthesized symmetrical ones (Zhou et al. 2004). The ΔugtB1 deletion mutant was suggested to be an interesting strain as it offers in vivo production of these biomolecules starting from cheap renewable substrates (Saerens et al. 2011c). Deletion of the acetyltransferase gene (at) results in the production of exclusively non-acetylated SLs in both the acidic as lactonic conformation (Saerens et al. 2011b). Such non-acetylated SLs have attracted attention for applications as antiviral drugs (Shah et al. 2005) or as starting molecules for the synthesis of dispersible nanoparticles (Kasture et al. 2007). However, the yields for these two SL intermediates (glucolipids and non-acetylated SLs) are a lot lower than those for the wild type (10 %). The latter could be attributed to regulatory and/or transport effects and further engineering of the pathway can possibly alleviate these issues. The described strains are subject of a patent application (Soetaert et al. 2010) and Evonik-Degussa also filed a patent dealing with the engineering of yeast strains for the production of tailored SL derivatives (Schaffer et al. 2009). Deletion of the gene responsible for lactonisation of SLs (Δle) results in the formation of exclusively acidic SLs with varying degrees of acetylation (Ciesielska et al. 2014). An overexpression strain of this gene (oele) almost exclusively produces lactonic SLs (Roelants et al. 2013, unpublished results). In contrast to the abovementioned recombinant strains (Δat and ΔugtB1), the yields for the latter two are comparable to the wild type SL production. Both strains thus represent significant industrial opportunities as for the wild type SLs post-fermentative purification steps are required for certain applications where only one of the SL compounds is specifically required (Marchant and Banat 2012). Both strains are the subject of a patent application which was recently published (Van Bogaert et al. 2013a). Another modified S. bombicola strain is protected by Ecover (Van Bogaert et al. 2011). This strain (Δmfe-2) is blocked in the β-oxidation of fatty acids to enable the production of medium-chain SLs.
Conclusion
This mini-review demonstrates that eukaryotic glycolipid BS producers show high similarity with each other and with other fungal secondary metabolites producers, of which the biosynthetic genes are found in large clusters often located near the telomeres. The latter was proven to represent some regulatory effects (Shaaban et al. 2010), which might also be the case for BS gene clusters located near the telomeres. Moreover, for some of these gene clusters a specific ‘in cluster’ regulator was discovered. Future advances in genomics, transcriptomics, proteomics and metabolomics will enable the further elucidation of the complete biosynthetic pathways and more specifically of the regulatory networks affecting eukaryotic BS biosynthesis. A profound knowledge of the genetics behind BS biosynthesis is part of the necessary base to enable engineering of the respective producers. This was demonstrated by the specific production of tailored glycolipids and by the development of strains with enhanced productivity. As stated in the Introduction section, these two factors (enhancement of structural variety and productivity) are indispensable for the further industrialization of BSs. Although a lot of work still remains to be done before many of these strains can become an industrial reality, some of the results described in this review represent real industrial opportunities. Further elucidation of biosynthetic mechanisms and regulatory networks involved in yeast glycolipid production will enable more advanced endogenous and heterologous metabolic engineering for the economical production of tailored glycolipids. The latter in combination with the advanced investigation of the physicochemical properties and possible chemo-enzymatic modifications of the produced molecules will facilitate further commercialization of these promising biomolecules. The fact that large companies like BASF, Evonik-Degussa, Unilever, Henkel, and Cargill have initiated R&D projects focusing on green alternatives of chemical surfactants is indicative of the recent movements in this field and it is expected that the BS market will grow substantially in the following years (Anonymous 2012).
References
Albrecht A, Rau U, Wagner F (1996) Initial steps of sophoroselipid biosynthesis by Candida bombicola ATCC 22214 grown on glucose. Appl Microbiol Biotechnol 46(1):67–73. doi:10.1007/s002530050784
Amaral PFF, Coelho AAZ, Marrucho IM, Coutinho JAP (2010) Biosurfactants from yeasts: characteristics, production and application. In: Sen R (ed) Biosurfactants. Adv Exp Med and Biol, vol 672, 2010 edn. Springer Science, New York, pp 236–249
Anonymous (2012) Biosurfactants market — global scenario, raw material and consumption trends, industry analysis, size, share and forecasts, 2011–2018. http://www.researchandmarkets.com/
Azim A, Shah V, Doncel GF, Peterson N, Gao W, Gross R (2006) Amino acid conjugated sophorolipids: A new family of biologically active functionalized glycolipids. Bioconjug Chem 17(6):1523–1529. doi:10.1021/bc060094n
Banat IM, Franzetti A, Gandolfi I, Bestetti G, Martinotti MG, Fracchia L, Smyth TJ, Marchant R (2010) Microbial biosurfactants production, applications and future potential. Appl Microbiol Biotechnol 87:427–444. doi:10.1007/s00253-010-2589-0
Bisht KS, Gross RA, Kaplan DL (1999) Enzyme-mediated regioselective acylations of sophorolipids. J Org Chem 64:780–789. doi:10.1021/jo981497m
Bisht KS, Gao W, Gross RA (2000) Glycolipids from Candida bombicola: polymerization of a 6-O-acryloyl sophorolipid derivative. Macromolecules 33(17):6208–6210. doi:10.1021/ma0001537
Bitto NJ, Graichen FHM, Monahan BJ (2009) Functionality at the end of a fatty acid chain chemical and biological routes to omega-hydroxylated fatty acids. Lipid Technol 21(10):216–219. doi:10.1002/lite.200900055
Boothroyd B, Thorn JA, Haskins RH (1956) Biochemistry of the ustilaginales: 12. Characterization of extracellular glycolipids produced by Ustilago sp. Can J Biochem Physiol 34(1):10–14. doi:10.1139/o56-003
Brakhage AA, Schroeckh V (2011) Fungal secondary metabolites — strategies to activate silent gene clusters. Fungal Genet Biol 48(1):15–22. doi:10.1016/j.fgb.2010.04.004
Caddick MX, Jones MG, van Tonder JM, Le Cordier H, Narendja F, Strauss J, Morozov IY (2006) Opposing signals differentially regulate transcript stability in Aspergillus nidulans. Mol Microbiol 62(2):509–519. doi:10.1111/j.1365-2958.2006.05383.x
Carr JA, Bisht KS (2003) Enzyme-catalyzed regioselective transesterification of peracylated sophorolipids. Tetrahedron 59(39):7713–7724. doi:10.1016/s0040-4020(03)01213-4
Casas JA, Garcia-Ochoa F (1999) Sophorolipid production by Candida bombicola: medium composition and culture methods. J Biosci Bioeng 88(5):488–494. doi:10.2478/v10026-010-0011-4
Chen J, Song X, Zhang H, Qu YB, Miao JY (2006) Sophorolipid produced from the new yeast strain Wickerhamiella domercqiae induces apoptosis in H7402 human liver cancer cells. Appl Microbiol Biotechnol 72(1):52–59. doi:10.1007/s00253-005-0243-z
Cheng YL, McNally DJ, Labbe C, Voyer N, Belzile F, Belanger RR (2003) Insertional mutagenesis of a fungal biocontrol agent led to discovery of a rare cellobiose lipid with antifungal activity. Appl Environ Microbiol 69(5):2595–2602. doi:10.1128/aem.69.5.2595-2602.2003
Ciesielska K, Li B, Groeneboer S, Van Bogaert I, Lin Y, Soetaert W, Van de Peer Y, Devreese B (2013) SILAC-based proteome analysis of Starmerella bombicola sophorolipid production. J Proteome Res 12(10):4376–4392. doi:10.1021/pr400392a
Ciesielska K, Van Bogaert I, Chevineau S, Li B, Groeneboer S, Soetaert W, Van de Peer Y, Devreese B (2014) Exoproteome analysis of Starmerella bombicola results in the discovery of an esterase required for lactonization of sophorolipids. J Proteomics 98:159–174. doi:10.1016/j.jprot.2013.12.026
Cooper DG, Paddock DA (1984) Production of a biosurfactant from Torulopsis bombicola. Appl Environ Microbiol 47(1):173–176
Cutler AJ, Light RJ (1979) Regulation of hydroxydocosanoic acid sophoroside production in Candida bogoriensis by the levels of glucose and yeast extract in the growth medium. J Biol Chem 254(6):1944–1950. doi:10.1002/jmr.2188
Daniel HJ, Otto RT, Binder M, Reuss M, Syldatk C (1999) Production of sophorolipids from whey: development of a two-stage process with Cryptococcus curvatus ATCC 20509 and Candida bombicola ATCC 22214 using deproteinized whey concentrates as substrates. Appl Microbiol Biotechnol 51(1):40–45. doi:10.1007/s002530051360
Davila AM, Marchal R, Vandecasteele JP (1997) Sophorose lipid fermentation with differentiated substrate supply for growth and production phases. Appl Microbiol Biotechnol 47(5):496–501. doi:10.1007/s002530050962
Desai JD, Banat IM (1997) Microbial production of surfactants and their commercial potential. Microbiol Mol Biol Rev 61(1):47–64. doi:10.1099/mic.0.26154-0
Deziel E, Lepine F, Milot S, Villemur R (2003) RhlA is required for the production of a novel biosurfactant promoting swarming motility in Pseudomonas aeruginosa: 3-(3-hydroxyalkanoyloxy) alkanoic acids (HAAs), the precursors of rhamnolipids. Microbiology-Sgm 149:2005–2013. doi:10.1099/mic.0.26154-0
Esders TW, Light RJ (1972a) Characterization and in-vivo production of three glycolipids from Candida bogoriensis — 13-glucopyranosylglucopyranosyloxydocosanoic acid and its monoacetylated and diacetylated derivatives. J Lipid Res 13(5):663–671
Esders TW, Light RJ (1972b) Glucosyltransferases and acetyltransferases involved in biosynthesis of glycolipids from Candida bogoriensis. J Biol Chem 247(5):1375–1386
Frautz B, Lang S, Wagner F (1986) Formation of cellobioselipids by growing and resting cells of Ustilago maydis. Biotechnol Lett 8(11):757–762. doi:10.1007/bf01020817
Fukuoka T, Kawamura M, Morita T, Imura T, Sakai H, Abe M, Kitamoto D (2008) A basidiomycetous yeast, Pseudozyma crassa, produces novel diastereomers of conventional mannosylerythritol lipids as glycolipid biosurfactants. Carbohydr Res 343(17):2947–2955. doi:10.1016/j.carres.2008.08.034
Fukuoka T, Yanagihara T, Imura T, Morita T, Sakai H, Abe M, Kitamoto D (2011) Enzymatic synthesis of a novel glycolipid biosurfactant, mannosylerythritol lipid-D and its aqueous phase behavior. Carbohydr Res 346:266–271. doi:10.1016/j.carres.2010.11.025
Fukuoka T, Yanagihara T, Imura T, Morita T, Sakai H, Abe M, Kitamoto D (2012) The diastereomers of mannosylerythritol lipids have different interfacial properties and aquous phase behavior, reflecting the erythritol configuration. Carbohydr Res 351:81–86. doi:10.1016/j.carres.2012.01.019
Giessler-Blank S, M. S., Thum O, Sieverding E (2009) Use of sophorolipids and derivatives thereof in combination with pesticides as adjuvant/additive for plant protection and the industrial non-crop field. MX2012003495 (A)
Golubev WI, Kulakovskaya TV, Shashkov AS, Kulakovskaya EV, Golubev NV (2008) Antifungal cellobiose lipid secreted by the epiphytic yeast Pseudozyma graminicola. Microbiology 77(2):171–175. doi:10.1134/s0026261708020082
Gorin PAJ, Spencer JFT, Tulloch AP (1961) Hydroxy fatty acid glycosides of sophorose from Torulopsis magnoliae. Can J Chem 39:846–855. doi:10.1139/v61-104
Gross RA, Thavasi TR (2004) Modified sophorolipids combinations as antimicrobial agents. US2013142855 (A1)
Gross RA, Guilmanov V, Scholz C (1999) Glycolipids from Torulopsis bombicola: biosynthesis, lipase-selective modifications and anti-cancer activity. Abstr Pap Am Chem Soc 217:1–2
Guilmanov V, Ballistreri A, Impallomeni G, Gross RA (2002) Oxygen transfer rate and sophorose lipid production by Candida bombicola. Biotechnol Bioeng 77(5):489–494. doi:10.1002/bit.10177
Hall P, Haverkamp J (1991) Detergent compositions. EP0499434 (A1)
Hammami W, Labbe C, Chain F, Mimee B, Belanger RR (2008) Nutritional regulation and kinetics of flocculosin synthesis by Pseudozyma flocculosa. Appl Microbiol Biotechnol 80(2):307–315. doi:10.1007/s00253-008-1541-z
Haskins RH (1950) Biochemistry of the ustilaginales: I. Preliminary cultural studies of Ustilago zeae. Can J Res 28c:213–223. doi:10.1139/cjr50c-012
Hewald S, Josephs K, Bolker M (2005) Genetic analysis of biosurfactant production in Ustilago maydis. Appl Environ Microbiol 71(6):3033–3040. doi:10.1128/AEM.71.6.3033-3040.2005
Hewald S, Linne U, Scherer M, Marahiel MA, Kamper J, Bolker M (2006) Identification of a gene cluster for biosynthesis of mannosylerythritol lipids in the basidiomycetous fungus Ustilago maydis. Appl Environ Microbiol 72(8):5469–5477. doi:10.1128/aem.00506-06
Hommel RK, Huse K (1993) Regulation of sophorose lipid production by Candida (Torulopsis) apicola. Biotechnol Lett 15(8):853–858. doi:10.1007/BF00180154
Hommel RK, Stegner S, Weber L, Kleber HP (1994a) Effect of ammonium ions on glycolipid production by Candida (Torulopsis) apicola. Appl Microbiol Biotechnol 42(2–3):192–197. doi:10.1007/BF00902716
Hommel RK, Weber L, Weiss A, Himmelreich U, Rilke O, Kleber HP (1994b) Production of sophorose lipid by Candida (Torulopsis) apicola grown on glucose. J Biotechnol 33(2):147–155. doi:10.1016/0168-1656(94)90107-4
Horst RJ, Zeh C, Saur A, Sonnewald S, Sonnewald U, Voll LM (2012) The Ustilago maydis Nit2 homolog regulates nitrogen utilization and is required for efficient induction of filamentous growth. Eukaryot Cell 11:368–380. doi:10.1128/ec.05191-11
Igarishi S, Hattori Y, Maitani Y (2006) Biosurfactant MEL-A enhances cellular association and gene transfection by cationic liposome. J Control Release 112:362–368. doi:10.1016/j.jconrel.2006.03.003
Im JH, Yanagishita H, Ikegami T, Takeyama Y, Idemoto Y, Koura N, Kitamoto D (2003) Mannosylerythritol lipids, yeast glycolipid biosurfactants, are potential affinity ligand materials for human immunoglobulin. J Biomed Mat Res A 65A:379–385. doi:10.1002/jbm.a.10491
Imura T, Yanagihara H, Kitamoto D (2004) Coacervate formation from natural glycolipid: one acetyl group on the headgroup triggers coacervate-to-vesicle transition. J Am Chem Community 289:57–61. doi:10.1021/ja0400281
Imura T, Masuda Y, Minamikawa H, Fukuoka T, Konishi M, Morita T, Sakai H, Abe M, Kitamoto D (2010) Enzymatic conversion of diacetylated sophoroselipid into acetylated glucoselipid: surface-active properties of novel bolaform biosurfactants. J Oleo Sci 59(9):495–501. doi:10.5650/jos.59.495
Jarvis WR, Traquair JA, Bélanger RR (2007) Sporodex®, fungal biocontrol for powdery mildew in greenhouse crops. In: Vincent C, Goettel MS and Lazarovits G (ed) Biological control: a global perspective. CABI, p 224
Kakugawa K, Tamai M, Imamura K, Miyamoto K, Miyoshi S, Morinaga Y, Suzuki O, Miyakawa T (2002) Isolation of yeast Kurtzmanomyces sp I-11, novel producer of mannosylerythritol lipid. Biosci Biotechnol Biochem 66(1):188–191. doi:10.1271/bbb.66.188
Kasture M, Singh S, Patel P, Joy PA, Prabhune AA, Ramana CV, Prasad BLV (2007) Multiutility sophorolipids as nanoparticle capping agents: synthesis of stable and water dispersible Co nanoparticles. Langmuir 23(23):11409–11412. doi:10.1021/la702931j
Kim YB, Yun HS, Kim E (2009) Enhanced sophorolipid production by feeding-rate-controlled fed-batch culture. Bioresour Technol 100(23):6028–6032. doi:10.1016/j.biortech.2009.06.053
Kitagawa M, Yamamoto S (2009) Biosurfactant-containing oil-in-water type emulsion cosmetic composition. JP2009275017 (A)
Kitagawa M, Nishimoto K, Tanaka T (2012) Cosmetic pigments, their production method and cosmetics containing the cosmetic pigments. US2012183592 (A1)
Kitamoto D, Haneishi K, Nakahara T, Tabuchi T (1990) Production of mannosylerythritol lipids by Candida antarctica from vegetable oils. Agric Biol Chem 54(1):37–40. doi:10.1271/bbb1961.54.37
Kitamoto D, Fuzishiro T, Yanagishita H, Nakane T, Nakahara T (1992) Production of mannosylerythritol lipids as biosurfactants by resting cells of Candida antarctica. Biotechnol Lett 14(4):305–310. doi:10.1007/bf01022329
Kitamoto D, Ghosh S, Ourisson G, Nakatani Y (2000) Formation of giant vesicles from diacylmannosylerythritols, and their binding to concanavalin A. Chem Commun 10:861–862. doi:10.1039/b000968g
Kitamoto D, Ikegami T, Suzuki GT, Sasaki A, Takeyama Y, Idemoto Y, Koura N, Yanagishita H (2001a) Microbial conversion of n-alkanes into glycolipid biosurfactants, mannosylerythritol lipids, by Pseudozyma (Candida antarctica). Biotechnol Lett 23(20):1709–1714. doi:10.1023/a:1012464717259
Kitamoto D, Yanagishita H, Endo A, Nakaiwa M, Nakane T, Akiya T (2001b) Remarkable antiagglomeration effect of a yeast biosurfactant, diacylmannosylerythritol, on ice-water slurry for cold thermal storage. Biotechnol Prog 17:362–365. doi:10.1021/bp000159f
Kitamoto D, Morita T, Fukuoka T, Konishi M, Imura T (2009) Self-assembling properties of glycolipid biosurfactants and their potential applications. Curr Opin Colloid Interface 14:315–328. doi:10.1016/j.cocis.2009.05.009
Kulakovskaya TV, Shashkov AS, Kulakovskaya EV, Golubev WI (2004) Characterization of an antifungal glycolipid secreted by the yeast Sympodiomycopsis paphiopedili. FEMS Yeast Res 5:247–252. doi:10.1016/j.femsyr.2004.07.008
Kulakovskaya TV, Shashkov AS, Kulakovskaya EV, Golubev WI (2005) Ustilagic acid secretion by Pseudozyma fusiformata strains. FEMS Yeast Res 5:919–923. doi:10.1016/j.femsyr.2005.04.006
Kulakovskaya TV, Golubev WI, Tomashevskaya MA, Kulakovskaya EV, Shashkov AS, Grachev AA, Chizhov AS, Nifantiev NE (2010) Production of antifungal cellobiose lipids by Trichosporon porosum. Mycopathologia 169(2):117–123. doi:10.1007/s11046-009-9236-2
Kurtzman CP, Price NPJ, Ray KJ, Kuo TM (2010) Production of sophorolipid biosurfactants by multiple species of the Starmerella (Candida) bombicola yeast clade. FEMS Microbiol Lett 311:140–146. doi:10.1111/j.1574-6968.2010.02082.x
Kurz M, Eder C, Isert D, Li ZY, Paulus EE, Schiell M, Toti L, Vertesy L, Wink J, Seibert G (2003) Ustilipids, acylated beta-d-mannopyranosyl d-erythritols from Ustilago maydis and Geotrichum candidum. J Antibiot 56(2):91–101
Ma XJ, Li H, Shao LJ, Shen J, Song X (2011) Effects of nitrogen sources on production and composition of sophorolipids by Wickerhamiella domercqiae var. sophorolipid. CGMCC 1576. Appl Microbiol Biotechnol 91(6):1623–1632. doi:10.1007/s00253-011-3327-y
Marchand G, Remus-Borel W, Chain F, Hammami W, Belzile F, Belanger RR (2009) Identification of genes potentially involved in the biocontrol activity of Pseudozyma flocculosa. Phytopathology 99(10):1142–1149. doi:10.1094/phyto-99-10-1142
Marchant R, Banat I (2012) Biosurfactants: a sustainable replacement for chemical surfactants? Biotechnol Lett 34(9):1597–1605. doi:10.1007/s10529-012-0956-x
Martin JF (2000) Molecular control of expression of penicillin biosynthesis genes in fungi: regulatory proteins interact with a bidirectional promoter region. J Bacteriol 182(9):2355–2362. doi:10.1128/jb.182.9.2355-2362.2000
Mimee B, Labbe B, Pelletier R, Belanger RR (2005) Antifungal activity of flocculosin, a novel glycolipid isolated from Pseudozyma flocculosa. Antimicrob Agents Chemother 49(4):1597–1599. doi:10.1128/aac.49.4.1597-1599.2005
Mimee B, Labbe C, Belanger RR (2009) Catabolism of flocculosin, an antimicrobial metabolite produced by Pseudozyma flocculosa. Glycobiology 19(9):995–1001. doi:10.1093/glycob/cwp078
Morita T, Fukuoka T, Konishi M, Imura T, Yamamoto S, Kitagawa M, Sogabe A, Kitamoto D (2009) Production of a novel glycolipid biosurfactant, mannosylmannitol lipid, by Pseudozyma parantarctica and its interfacial properties. Appl Microbiol Biotechnol 83(6):1017–1025. doi:10.1007/s00253-009-1945-4
Morita T, Ito E, Fukuoka T, Imura T, Kitamoto D (2010a) The role of PaAAC1 encoding a mitochondrial ADP/ATP carrier in the biosynthesis of extracellular glycolipids, mannosylerythritol lipids, in the basidiomycetous yeast Pseudozyma antarctica. Yeast 27:379–388. doi:10.1002/yea.1761
Morita T, Ito E, Kitamoto H, Takegawa K, Fukuoka T, Konishi M, Imura T, Kitamoto D (2010b) Identificiation of the gene PaEMT1 for biosynthesis of mannosylerythritol lipids in the basidiomycetous yeast Pseudozyma antarctica. Yeast 27:905–917. doi:10.1002/yea.1794
Morita T, Ogura Y, Takashima M, Hirose N, Fukuoka T, Imura T, Kondo Y, Kitamoto D (2011) Isolation of Pseudozyma churashimaensis sp.nov., a novel ulstilaginomycetous yeast species as a producer of glycolipid biosurfactants, mannosylerythritol lipids. J Biosci Eng 112:137–144. doi:10.1016/j.jbiosc.2011.04.008
Morita T, Fukuoka T, Imura T, Kitamoto D (2012) Formation of the two novel glycolipid biosurfactants, mannosylribitol lipid and mannosylarabitol lipid by Pseudozyma parantarctica JCM 11752(T). Appl Microbiol Biotechnol 96:931–938. doi:10.1007/s00253-012-4230-x
Morita T, Fukuoka T, Imura T, Kitamoto D (2013a) Accumulation of cellobiose lipids under nitrogen-limiting conditions by two ustilaginomycetous yeasts, Pseudozyma aphidis and Pseudozyma hubeiensis. FEMS Yeast Res 13(1):44–49. doi:10.1111/1567-1364.12005
Morita T, Fukuoka T, Imura T, Kitamoto D (2013b) Production of mannosylerythritol lipids and their application in cosmetics. Appl Microbiol Biotechnol 97(11):4691–4700. doi:10.1007/s00253-013-4858-1
Morita T, Koike H, Koyama Y, Hagiwara H, Ito E, Fukuoka T, Imura T, Machida M, Kitamoto D (2013c) Genome sequence of the basidiomycetous yeast Pseudozyma antarctica T-34, a producer of the glycolipid biosurfactants mannosylerythritol lipids. Genome Announc 1(2):1–2. doi:10.1128/genomeA.00064-13
Nunez A, Ashby R, Foglia TA, Solaiman DKY (2004) LC/MS analysis and lipase modification of the sophorolipids produced by Rhodotorula bogoriensis. Biotechnol Lett 26(13):1087–1093. doi:10.1023/B:BILE.0000032970.95603.6d
Pekin G, Vardar-Sukan F, Kosaric N (2005) Production of sophorolipids from Candida bombicola ATCC 22214 using Turkish corn oil and honey. Eng Life Sci 5(4):357–362. doi:10.1002/elsc.200520086
Price NPJ, Ray KJ, Vermillion KE, Dunlap CA, Kurtzman CP (2012) Structural characterization of novel sophorolipid biosurfactants from a newly identified species of Candida yeast. Carbohydr Res 348:33–41. doi:10.1016/j.carres.2011.07.016
Puchkov EO, Zähringer U, Lindner B, Kulakovskaya TV, Seydel U, Wiese A (2002) The mycocidal, membrane-active complex of Cryptotoccus humicola is a new type of cellobiose lipid with detergent features. Biochim Biophys Acta 1558:161–170
Rau U, Hammen S, Heckmann R, Wray V, Lang S (2001) Sophorolipids: a source for novel compounds. Ind Crop Prod 13:85–92. doi:10.1016/S0926-6690(00)00055-8
Rau U, Nguyen LA, Roeper H, Koch H, Lang S (2005a) Fed-batch bioreactor production of mannosylerythritol lipids secreted by Pseudozyma aphidis. Appl Microbiol Biotechnol 68(5):607–613. doi:10.1007/s00253-005-1906-5
Rau U, Nguyen LA, Schulz S, Wray V, Nimtz M, Roeper H, Koch H, Lang S (2005b) Formation and analysis of mannosylerythritol lipids secreted by Pseudozyma aphidis. Appl Microbiol Biotechnol 66(5):551–559. doi:10.1007/s00253-004-1672-9
Recke VK, Beyrle C, Gerlitzki M, Hausmann R, Syldatk C, Wray V, Tokuda H, Suzuki N, Lang S (2013a) Lipase-catalyzed acylation of microbial mannosylerythritol lipids (biosurfactants) and their characterization. Carbohydr Res 373:82–88. doi:10.1016/j.carres.2013.03.013
Recke VK, Gerlitzki M, Hausmann R, Syldatk C, Wray V, Tokuda H, Suzuki N, Lang S (2013b) Enzymatic production of modified 2-dodecyl-sophorosides (biosurfactants) and their characterization. Eur J Lipid Sci Technol 115(4):452–463. doi:10.1002/ejlt.201300012
Rehm BHA, Kruger N, Steinbuchel A (1998) A new metabolic link between fatty acid de novo synthesis and polyhydroxyalkanoic acid synthesis — the phaG gene from Pseudomonas putida KT2440 encodes a 3-hydroxyacyl-acyl carrier protein coenzyme A transferase. J Biol Chem 273(37):24044–24051. doi:10.1074/jbc.273.37.24044
Rodrigues LR, Teixeira JA (2010) Biomedical and therapeutic applications of biosurfactants. In: Sen R (ed) Biosurfactants, vol 672, Advances in experimental medicine and biology. Springer Science, New York, pp 75–87
Roelants S (2013) Starmerella bombicola as a platform organism for the production of biobased compounds. PhD, Ghent University
Roelants SLKW, Saerens K, Derycke T, Li B, Lin Y-C, Van de Peer Y, De Maeseneire SL, Van Bogaert INA, Soetaert W (2013) Candida bombicola as a platform organism for the production of tailor-made biomolecules. Biotechnol Bioeng 110(9):494–503. doi:10.1002/bit.24895
Saerens K (2012) Synthesis of glycolipids by Candida bombicola. Ghent University, Ghent
Saerens KMJ, Roelants SLKW, Van Bogaert INA, Soetaert W (2011a) Identification of the UDP-glucosyltransferase gene UGTA1, responsible for the first glucosylation step in the sophorolipid biosynthetic pathway of Candida bombicola ATCC 22214. FEMS Yeast Res 11(1):123–132. doi:10.1111/j.1567-1364.2010.00695.x
Saerens KMJ, Saey L, Soetaert W (2011b) One-step production of unacetylated sophorolipids by an acetyltransferase negative Candida bombicola. Biotechnol Bioeng 108(12):2923–2931. doi:10.1002/bit.23248
Saerens KMJ, Zhang J, Van Bogaert I, Soetaert W (2011c) Cloning and functional characterization of the UDP-glucosyltransferase UgtB1 involved in sophorolipid production by Candida bombicola and creation of a glucolipid-producing yeast strain. Yeast 28:279–292. doi:10.1002/yea.1838
Schaffer S, Wessel M, Thiessenhusen A (2009) Cells, nucleic acids, enzymes and use thereof and methods for the production of sophorolipids. US2013035403 (A1)
Shaaban M, Palmer JM, El-Naggar WA, El-Sokkary MA, Habib EE, Keller NP (2010) Involvement of transposon-like elements in penicillin gene cluster regulation. Fungal Genet Biol 47(5):423–432. doi:10.1016/j.fgb.2010.02.006
Shah V, Doncel GF, Seyoum T, Eaton KM, Zalenskaya I, Hagver R, Azim A, Gross R (2005) Sophorolipids, microbial glycolipids with anti-human immunodeficiency virus and sperm-immobilizing activities. Antimicrob Agents Chemother 49(10):4093–4100. doi:10.1128/aac.49.10.4093-4100.2005
Shete AM, Wadhawa G, Banat IM, Chopade BA (2006) Mapping of patents on bioemulsifier and biosurfactant: a review. J Sci Ind Res 65(2):91–115
Singh SK, Felse AP, Nunez A, Foglia TA, Gross RA (2003) Regioselective enzyme-catalyzed synthesis of sophorolipid esters, amides, and multifunctional monomers. J Org Chem 68(14):5466–5477. doi:10.1021/jo0204395
Soetaert W, De Maeseneire SL, Saerens K, Roelants S, Van Bogaert I (2010) Yeast strains modified in their sophorolipid production and uses thereof. WO2011154523 (A1)
Spoeckner S, Wray V, Nimtz M, Lang S (1999) Glycolipids of the smut fungus Ustilago maydis from cultivation on renewable resources. Appl Microbiol Biotechnol 51:33–39. doi:10.1007/s002530051359
Stüwer O, Hommel R, Haferburg D, Kleber HP (1987) Production of crystalline surface-active glycolipids by a strain of Torulopsis apicola. J Biotechnol 6(4):259–269. doi:10.1016/0168-1656(87)90057-5
Tao HT (2011) Chemically modified sophorolipids and uses thereof. WO2013003291 (A2)
Teichmann B, Linne U, Hewald S, Marahiel MA, Bolker M (2007) A biosynthetic gene cluster for a secreted cellobiose lipid with antifungal activity from Ustilago maydis. Mol Microbiol 66(2):525–533. doi:10.1111/j.1365-2958.2007.05941.x
Teichmann B, Liu LD, Schink KO, Bolker M (2010) Activation of the ustilagic acid biosynthesis gene cluster in Ustilago maydis by the C2H2 Zinc finger transcription factor Rua1. Appl Environ Microbiol 76(8):2633–2640. doi:10.1128/aem.02211-09
Teichmann B, Labbe C, Lefebvre F, Bolker M, Linne U, Belanger RR (2011a) Identification of a biosynthesis gene cluster for flocculosin a cellobiose lipid produced by the biocontrol agent Pseudozyma flocculosa. Mol Microbiol 79(6):1483–1495. doi:10.1111/j.1365-2958.2010.07533.x
Teichmann B, Lefebvre F, Labbe C, Bolker M, Linne U, Belanger RR (2011b) Beta hydroxylation of glycolipids from Ustilago maydis and Pseudozyma flocculosa by an NADPH-dependent beta-hydroxylase. Appl Environ Microbiol 77(21):7823–7829. doi:10.1128/aem.05822-11
Tulloch AP, Spencer JFT, Gorin PAJ (1962) The fermentation of long-chain compounds by Torulopsis magnoliae. Can J Chem 40:1326–1338. doi:10.1139/v62-203
Tulloch AP, Hill A, Spencer JFT (1967) A new type of macrocycli lactone from Torulopsis apicola. Chem Commun 12:584–586. doi:10.1039/c19670000584
Tulloch AP, Spencer JFT, Deinema MH (1968) A new hydroxy fatty acid sophoroside from Candida bogoriensis. Can J Chem 46:345–348. doi:10.1139/v68-057
Van Bogaert INA, Saerens K, De Muynck C, Develter D, Soetaert W, Vandamme EJ (2007) Microbial production and application of sophorolipids. Appl Microbiol Biotechnol 76(1):23–34. doi:10.1007/s00253-007-0988-7
Van Bogaert INA, Demey M, Develter D, Soetaert W, Vandamme EJ (2009) Importance of the cytochrome P450 monooxygenase CYP52 family for the sophorolipid-producing yeast Candida bombicola. FEMS Yeast Res 9(1):87–94. doi:10.1111/j.1567-1364.2008.00454.x
Van Bogaert I, Develter D, Fleurackers S (2011) Method for the production of medium-chain sophorolipids. US2011136110
Van Bogaert I, Ciesielska K, Devreese B, Soetaert W, Roelants S (2013a) A lactonase derived from Candida bombicola and uses thereof. WO 2013/092421 A1
Van Bogaert INA, Holvoet K, Roelants S, Li B, Lin YC, Van de Peer Y, Soetaert W (2013b) The biosynthetic gene cluster for sophorolipids: a biotechnological interesting biosurfactant produced by Starmerella bombicola. Mol Microbiol 88(3):501–509. doi:10.1111/mmi.12200
Vertesy L, Kurz M, Noelken G, Wink J (1998) Ustilipides, method for the production and the use thereof. US6472158 (B1)
Woloshuk CP, Foutz KR, Brewer JF, Bhatnagar D, Cleveland TE, Payne GA (1994) Molecular characterization of AflR, a regulatory locus for aflatoxin biosynthesis. Appl Environ Microbiol 60(7):2408–2414
Yang XL, Awakawa T, Wakimoto T, Abe I (2013) Induced production of the novel glycolipid ustilagic acid C in the plant pathogen Ustilago maydis. Tetrahedron Lett 54(28):3655–3657. doi:10.1016/j.tetlet.2013.04.131
Yonetani Y, Hattori Y, Igarashi S, Kitamoto M, Kitagawa M, Sogabe A (2005) Gene transfer carrier consisting of liposome. JP2005281146 (A)
Zhou SQ, Xu C, Wang J, Gao W, Akhverdiyeva R, Shah V, Gross R (2004) Supramolecular assemblies of a naturally derived sophorolipid. Langmuir 20(19):7926–7932. doi:10.1021/la048590s
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Roelants, S.L.K.W., De Maeseneire, S.L., Ciesielska, K. et al. Biosurfactant gene clusters in eukaryotes: regulation and biotechnological potential. Appl Microbiol Biotechnol 98, 3449–3461 (2014). https://doi.org/10.1007/s00253-014-5547-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-014-5547-4