Abstract
Odorant-binding proteins (OBPs) are small soluble polypeptides found in sensory organs of vertebrates and insects as well as in secretory glands and are dedicated to detection and release of chemical stimuli. OBPs of vertebrates belong to the family of lipocalin proteins, while those of insects are folded into α-helical domains. Both types of architectures are extremely stable to temperature, organic solvents and proteolytic digestion. These characteristics make OBPs suitable elements for fabricating biosensors to be used in the environment, as well as for other biotechnological applications. The affinity of OBPs for small volatile organic compounds is in the micromolar range, and they have broad specificity to a range of ligands. For biotechnological applications, OBPs can be expressed in bacterial systems at low cost and are easily purified. The large amount of information available on their structures and affinities to different molecules should allow the design of specific mutants with desired characteristics and represent a solid base for tailoring OBPs for different applications.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
This report describes the structure of odorant-binding proteins (OBPs) with a focus on their actual and potential technological uses. Several reviews during the last two decades (Leal 2013; Pelosi 1994, 1996, 1998, 2001; Pelosi and Maida 1990, 1995; Pelosi et al. 2006; Tegoni et al. 2000, 2004; Steinbrecht 1998; Vogt 2003, 2005) have summarised structural and physiological aspects of OBPs, but none of these have taken into account technological applications of these proteins, which, thanks to their exceptional stability, have potential as new materials for sensing devices and other uses. On the other hand, reviews have discussed the design and fabrication of sensors for odours of different kinds (Bailey 2011; Baldwin et al. 2011; Bar-Cohen. 2011; Craven et al. 1996; Gardner 2004), and attempts have been made to incorporate olfactory receptors into such devices (Yoon et al. 2009; Lee et al. 2012), but only few researchers have considered the use of OBPs.
We believe, instead, that OBPs could represent ideal tools not only for robust, cheap, and versatile sensing elements, but also in other applications, such as in the programmed release of chemicals in the environment or as scavengers for pollutants and other noxious compounds.
These different types of applications are suggested by the dual role of OBPs in biological system: (1) in detecting pheromones and odours in sensory organs and (2) in storing semiochemicals for their controlled release from glands and other non-sensory organs.
In the first part of this review, we briefly introduce the structure and function of OBPs, focusing on those characteristics that make them most suitable for artificial devices. In the second part, we discuss their actual and potential technological applications.
OBPs are ligand-binding proteins of exceptional stability
Introduction
Odorant-binding proteins refer to two classes of proteins, those found in vertebrates and those from insects, which are different in structures but similar in function. They are associated to olfaction and chemoreception and are found highly concentrated in the nasal mucus of vertebrates and in the lymph of chemosensilla in insects (Pelosi 1994, 1998; Pelosi and Maida 1995; Steinbrecht 1998; Tegoni et al. 2000; Vogt 2005; Pelosi et al. 2006; Leal 2013). By coincidence, they were discovered independently at the same time in vertebrates (Pelosi et al. 1981, 1982) and in insects (Vogt and Riddiford 1981). Thanks to many genome research projects, a large number of DNA sequences encoding OBPs are known in several species of vertebrates and more than a hundred species of insects.
Although this review is focused on OBPs, we cannot avoid briefly mentioning another class of small soluble binding proteins, reported only in insects and named chemosensory proteins (CSPs). These polypeptides are structurally different from OBPs but, in several cases, are endowed with similar functions (Wanner et al. 2004; Pelosi et al. 2006).
Structure of OBPs
OBPs of vertebrates (Bignetti et al. 1985; Dal Monte et al. 1991; Paolini et al. 1998) belong to the superfamily of lipocalins (Flower 1996; Flower et al. 2000) that include carrier proteins, such as retinol-binding protein, β-lactoglobulin and other proteins of different functions. Their three-dimensional structures (Böcskei et al. 1992; Bianchet et al. 1996; Tegoni et al. 1996; Spinelli et al. 1998, 2002; Vincent et al. 2000) follow the typical folding of lipocalins, with eight antiparallel β-sheets and a short α-helical segment close to the C terminus. Cysteine residues do not play a major role in the stability of vertebrate OBPs. In fact, some of them, such as the bovine OBP, are devoid of cysteines, while others, like the pig OBP, present only a single disulphide bridge.
We can also observe that the bovine OBP is a homodimer, in which the subunits interact in a “domain swapping” fashion (Bianchet et al. 1996; Tegoni et al. 1996), while the pig OBP, like most proteins of this family, is a monomer (Spinelli et al. 1998). Figure 1 reports the amino acid sequence and the three-dimensional structures of these two OBPs.
Three-dimensional structures of bovine (Bianchet et al. 1996; PDB: 1PBO) and pig (Vincent et al. 2000; PDB: 1A3Y) OBPs. Both proteins are folded in the typical β-barrel motif, common to all lipocalins. The bovine OBP, which does not contain cysteines, is a homodimer interacting in the typical “domain swapping” fashion. Pig OBP, instead, which presents a disulphide bridge, is a monomer. The two cysteine residues of this protein are in bold font. The ligand inside the cavity of pig OBP is thymol. Identical residues between the two proteins are shaded. Structures have been visualised using the DeepView software (Guex and Peitsch 1997)
OBPs of insects present a completely different folding pattern, well conserved within the protein family. It is constituted by six α-helical domains arranged in a very compact structure, which encloses a hydrophobic cavity (Sandler et al. 2000) (Fig. 2). The structure of insect OBPs is further stabilised by three interlocked disulphide bridges (Scaloni et al. 1999; Leal et al. 1999). This structural feature is a landmark characterising all OBPs of insects, which otherwise may have very little in common when comparing amino acid sequences not only between species, but also within the same species. Despite large variability in amino acid sequences, the structure of insect OBPs is remarkably well conserved even between members of different Orders of insects (Tegoni et al. 2004).
Structure of the B. mori PBP at neutral (Sandler et al. 2000; PDB: 1DQE) and low (Horst et al. 2001; PDB: 1GM0) pH. In acidic conditions, the glutamate and aspartate residues of the C-terminal segment (underlined) lose their negative charges, thus allowing the formation of a seventh α-helix, which enters the binding pocket, thus causing the bound pheromone to be displaced. This mechanism can be exploited in biosensors as a molecular switch or to rapidly restore the binding capacity of the protein. Structures have been visualised using the DeepView software (Guex and Peitsch 1997)
OBPs are also involved in delivering semiochemicals
Besides being expressed in the nasal mucus of vertebrates and in the chemosensilla of insects, OBPs have been identified in glands and organs involved in the synthesis and/or delivery of pheromones. Examples from vertebrates include the urine of mice and rats (Bacchini et al. 1992; Böcskei et al. 1992), the vaginal secretion of hamsters (Vincent et al. 2001), the saliva of pigs (Marchese et al. 1998; Spinelli et al. 2002) and the sweat of horses (D’Innocenzo et al. 2006). These proteins, in most cases identical to those found in the nose, are often loaded with specific pheromones when isolated from such secretions. Other proteins, belonging to the family of OBPs, have been described as allergens. They are mostly present in the saliva and sweat of several mammals and most likely perform the same carrier functions of OBPs. Although the allergenic properties of OBPs and other lipocalins have not been clarified in detail (Virtanen et al. 2012), certainly they indicate high stability to temperature and proteolytic degradation of such proteins, making them suitable for technological uses.
A similar phenomenon has been also documented in insects, where OBPs and CSPs have been identified in pheromone glands and in reproductive organs, often associated with pheromone molecules (Li et al. 2008; Zhou et al. 2013; Sun et al. 2012a; Dani et al. 2011; Ban et al. 2013).
Thermal stability
Thermal stability is an important characteristic for proteins to be used in biotechnological applications, such as biosensors for environmental monitoring, cartridges for cleaning waste waters or deodorizers. Both insect and vertebrate OBPs can withstand high temperatures before undergoing denaturation, and, if unfolding occurs, this phenomenon is reversed after restoring the initial conditions (Paolini et al. 1999; Ban et al. 2003). In a first study (Paolini et al. 1999), Fourier-transform infrared spectroscopy (FT-IR) was used to monitor conformational changes in the pig OBP caused by temperature. Two transitions occurring at 65–70 and 80–85 °C were related to molten globule states of the β-barrel, which, however, maintained its integrity at such temperatures. The stability of the protein was further increased in the presence of good ligands, such as 2-isobutyl-3-methoxypyrazine and 3,7-dimethyloctanol, but not for poor ligands as 2-phenylethanol. Similar studies have confirmed the high stability of vertebrate OBPs and provided additional information (Marabotti et al. 2008a, b; Scire et al. 2009).
Insect OBPs are even more resistant to denaturation and degradation. Their very compact structure is further stabilised, with respect to vertebrate OBPs, by the presence of three interlocked disulphide bridges, which confer limited flexibility to the structure, thus preventing thermal denaturation and attack by proteolytic enzymes. The only study specifically addressing this topic is a very recent one, where FT-IR and CD (circular dichroism) have been used to monitor denaturation of honeybee OBP14, a protein presenting only two of the typical disulphide bridges. Transitions in the IR bands were observed around 55 °C, probably corresponding to a relaxed conformation of the structure rather than denaturation. Such transition temperature is increased by about 10 °C in the presence of strong ligands (Schwaighofer A, Kotlowski, C, Amaran C, Chu N, Mastrogiacomo R, Becker C, Pelosi P, Larisika M, Nowak C, submitted). Moreover, the exceptional thermal stability of insect OBPs and their ability to refold in the correct way after denaturation has been verified in several cases, when recombinant proteins had to be reduced and denatured in order to be solubilised, to be refolded by spontaneous oxidation. These procedures always afforded fully functional proteins (Ban et al. 2003; Calvello et al. 2003).
Mechanisms of ligand binding and release
OBPs bind small molecules
OBPs were discovered due to their property of binding molecules of odorants and pheromones. This feature, combined with their expression in sensory organs, can still help to distinguish OBPs from other proteins of similar structures.
Measuring binding affinity and specificity is important for characterising OBPs and relating their expression in specific organs to physiological functions. In the past, affinity constants have been measured with methods involving the use of radioactive-labelled odorants or pheromones and separation of bound from free ligands by gel filtration, electrophoresis or other techniques (Pelosi et al. 1981; Topazzini et al. 1985; Dal Monte et al. 1993; Pelosi and Maida 1995; Maida et al. 2003; Nespoulous et al. 2004). Cleaner and faster protocols, introduced more recently, use fluorescent probes as reporters (Paolini et al. 1999; Campanacci et al. 2001; Ban et al. 2002). The emission spectrum of a fluorescent compound undergoes a strong increase in intensity and a blue shift when it is bound inside the hydrophobic pocket of a protein. In this way, the signal produced by the bound probe can be evaluated without the need of separation steps, thus allowing more accurate measurements. However, the concentration of the bound fluorescent ligand and consequently the dissociation constant can be calculated only assuming a simple 1:1 stoichiometry of the interaction involved. Affinities of other ligands are calculated from their capability of competing with the fluorescent compound in the binding pocket. Two probes, 1-aminoanthracene (1-AMA) and N-phenyl-1-naphthylamine (1-NPN) are most commonly used, but other fluorescent compounds have also been tested (Riviere et al. 2003; Briand et al. 2002a; Mastrogiacomo R, Iovinella I, Napolitano E, Pelosi P, submitted).
Dissociation constants for most ligands are in the micromolar range. OBPs differ largely for their selectivity towards different chemical compounds. Some, like the pig OBP (Dal Monte et al. 1993), bind a variety of different structures, including terpenoids (3,7-dimethyloctanol), linear aldehydes (decanal) and aromatic compounds (benzyl benzoate). Three-dimensional structures of pig OBP complexed with the above ligands has shown that, although exhibiting similar affinities, each of the three compounds sits in the same binding pocket with a different orientation (Vincent et al. 2000). Other proteins are more narrowly tuned to few odorant structures, but very high specificity has never been associated with OBPs (Calvello et al. 2003; Sun et al. 2012b). Within the same species, different OBPs present different spectra of ligand-binding activities, as in the case of the three rat OBPs, each recognising a different subset of odorants (Löbel et al. 2002). A special case is the human OBP (Briand et al. 2002b), shown to bind medium-length aliphatic aldehydes with a specific mechanism involving action of a lysine in the binding pocket (Tcatchoff et al. 2006). This last study provides an interesting example on the potentialities of site-directed mutagenesis to modify the ligand selectivity of OBPs.
Such broad specificity is required for a detection device, like the olfactory system, using a limited number of proteins, which have to interact with a very large number of potential ligands. When using OBPs in artificial instruments for odour analysis, this characteristic allows the assembly of multisensor arrays, utilising different OBPs with overlapping specificities. The large variety of OBPs available with respect to their binding selectivity can be further increased with the synthesis of specific mutants designed for desired needs. As an example, Wei et al. (2008) modified the pig OBP by replacing phenylalanine residue 88, present inside the binding pocket with a tryptophan. The mutant allowed measuring the binding of aromatic compounds without the need of an additional fluorescent probe, by monitoring the quenching of tryptophan intrinsic fluorescence. At the same time, the presence of a tryptophan in the binding pocket increased the affinity of the protein for polycyclic aromatic hydrocarbons.
A detailed study on the structures of bovine OBP has shown that access of a ligand inside the binding cavity of the protein is controlled by a phenylalanine (Phe89) that opens like a door by rotation around a carbon–carbon single bond (Bianchet et al. 1996). Similar conformational changes involving aromatic residues have been shown to occur in molecular modelisation with pig and rat OBPs (Golebiowski et al. 2007). These mechanisms have been related to the exceptionally slow rate of release of the ligand (Pevsner et al. 1990; Yabuki et al. 2011). Such property may represent a problem if we want to use OBPs in biosensors, which need rapid recovery, but could be an advantage in other applications, like dosimeters for environmental pollutants.
A molecular switch
Insect OBPs, on the contrary, do not have the gate-controlled entrance of ligands reported above for vertebrate OBPs, and, as a consequence, dissociation rates are relatively high. This is the reason why attempts to use separation methods for the evaluation of binding affinities have failed when applied to OBPs of insects.
Instead, another interesting mechanism, a sort of molecular switch, has been discovered with the PBP1 of Bombyx mori and found to be active with several other proteins of similar structure (Damberger et al. 2000; Horst et al. 2001). At low pH, the C terminus of the protein, which bears several acidic residues, looses most of its negative charges and folds into a seventh α-helix. This helix, in turn, enters the binding cavity, thus displacing any ligand present inside. This mechanism has been proposed to be active in picking up the pheromone from the environment and handling it to the membrane-bound receptor. Figure 2 reports the three-dimensional structures of the B. mori PBP1 in the two conformational states. Although criticism has been raised on the possibility that such mechanism may occur in vivo, nevertheless, such “molecular switch” could prove very interesting when using this OBP and other similar proteins as biosensing elements, providing a fast, efficient way of releasing the ligand and restoring the functionality of the sensor. The same mechanism is active with all those insect OBPs presenting a C terminus long enough to form the seventh α-helix and rich of acidic residues (Leal 2013). Moreover, it is conceivable that this strategy could be applied also to other OBPs by opportunely modifying the C-terminal segment.
Dual role in detection and delivery suggests different applications
Despite the large amount of structural information on OBPs, both of vertebrates and insects, their role and mode of action in detection of odours and pheromones is still a matter of debate. Recently, a few reports have provided experimental evidence that in insect OBPs are required for olfaction and are involved in recognising different semiochemicals (Xu et al. 2005; Matsuo et al. 2007; Swarup et al. 2011; Sun et al. 2012b). Whatever the specific physiological function, the large structural variety of OBPs reflects different specificities towards diverse organic compounds and represent a strong basis for building a multisensor device for odour discrimination.
When looking for technological applications of OBPs, biological systems can provide, once again, interesting suggestions. In fact, both in vertebrates and insects, OBPs perform two major roles: detection and delivery of chemical stimuli.
The lesson to learn is that OBPs could be used in sensors to detect volatile molecules, but could also act as reservoir for odorants to obtain a slow release in the environment, or else as traps to clean closed spaces from obnoxious odours.
OBPs as biosensing elements for odours
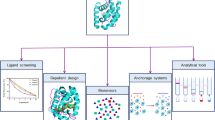
Despite the interesting characteristics of OBPs making them ideal tools for biosensors, few studies have used these proteins in artificial devices for the detection of odorants. Generally speaking, there are only a few ways of detecting binding of a ligand to immobilised proteins that do not have catalytic activity. In a biosensor, the binding can modify electrical properties of a protein, which can be part of a transistor or a capacitor; the extra mass of the ligand bound to the protein can be measured with a quartz microbalance; the binding can modify the refractive index of the protein; and a ligand can displace a fluorescent probe from the binding site of a protein. In some cases, calorimetry can be used to measure thermal properties changing when a ligand binds to the protein. Figure 3 summarises some of these methods of transduction together with other technological applications of OBPs.
Technological uses of OBPs. Due to their exceptional stability, OBPs can be used in several applications, such as in sensing devices to monitor odorants and volatile compounds, as reservoirs for small organic compounds to be released into or removed from the environment, and as active chromatographic phases for chemical analysis
Electronic devices
Using the recombinant rat OBP1F, Hou et al. (2005) were able to form stable Langmuir–Blodgett films and to transfer them to gold electrodes. Then they measured electric resistance by using non-faradaic electrochemical spectroscopy and found a large difference between the resistance of untreated film (1.18 MΩ) and that of a film that had been exposed to vapours of isoamyl acetate (25 kΩ). Based on this single experiment, it is not clear whether the effect is specific and related to selective binding to the protein, or else is the consequence of a physical effect of the odorant on the structure of the film. Besides, there is no indication on the reversibility of the binding or on the lifetime of such sensors.
An apparently more robust and simple device has been assembled by Liu et al. (2013), who immobilised a CSP of the honeybee (ASP3c) onto interdigitated electrodes and measured the impedance both in the absence and in the presence of odorants over a concentration range. They managed to detect ligands down to concentrations of 0.1–1 μM. The method is simple and fast enough to represent an interesting alternative to currently used protocols for ligand-binding measurements. The sensitivity is not satisfactory when compared to biological olfactory systems, but could be suitable for applications in environmental monitoring, for example, for water quality assessment.
Reduced graphene oxide transistors containing immobilised honeybee OBP14 have been fabricated and tested in odour sensing (Larisika M, Kotlowski C, Mastrogiacomo R, Pelosi P, Kleber C, Knoll W, Nowak C, submitted). Variations in the drain current are reproducible and linear with concentration of the odorant. Some volatile compounds, such as eugenol and derivatives, proved to be strong ligands with dissociation constants in the micromolar range, while others, such as citral, presented lower affinity, with a trend that reproduces the specificity of OBP14 in solution (Iovinella et al. 2011).
Quartz microbalances
Quartz microbalances measure the change in mass produced by anything bound to the quartz. Bulk absorption of analyte molecules such as volatile organic compounds (VOCs) increases the mass of the quartz crystal, resulting in a decrease of the resonance frequency. The absorption process is fully reversible. Since very thin layers are used, the diffusion-limited process is very fast, in the order of milliseconds. By selecting different coatings on the quartz crystal, the sensors can be sensitised to particular molecular properties such as polarity, polarizability, shape and volume. If a binding protein is immobilised onto the quartz, any ligand that is bound will produce a change in the resonant frequency. The suitability of such devices as sensors relies on the exceptional accuracy in measuring frequencies. In practice, we can measure a difference in the weight of a typical crystal as low as 0.1 ng depending on the frequency of oscillation of the crystal. The main drawback limiting the use of quartz balances for gas sensing is the difficulty in controlling humidity and temperature. Wyszynski and Nakamoto (2009) have reviewed the use of quartz microbalances in biomimetic or bioinspired applications especially for “electronic noses”. Rodgers et al. (2010) developed a method of immobilising MUPs on a quartz microbalance using self-assembled monolayers that could be used for immobilising histidine-tagged proteins. They reported the operation of the quartz crystal in a solution containing ligands and demonstrated that the bound protein produced a signal upon binding 2-isobutyl-3-methoxypyrazine.
Sankaran et al. (2011) reported on the fabrication of a biosensor for alcohols using a particular Drosophila melanogaster OBP called LUSH. This protein was first reported as specific for binding low molecular weight alcohols (Kruse et al. 2003), but this was proved to be wrong and due to artefacts (Zhou et al. 2004). In fact, in subsequent papers, the same authors reported on the affinity of LUSH for vaccenyl acetate a 20-carbon male pheromone (Xu et al. 2005). To set up their sensor, Sankaran et al. (2011) synthesised a peptide reproducing the protein binding site that was supposed to bind alcohols and immobilised it on a quartz microbalance. This device responded to 3-methyl-1-butanol and to 1-hexanol down to concentrations of few ppm. The specificity of the sensor was not checked for other organic compounds of different nature and size, nor using different peptides or proteins. Most likely, the immobilised peptide acted as absorbing matrix, just like any other protein or synthetic polymer. Although based on a wrong assumption and lacking all the necessary controls, the paper shows that the technique of quartz microbalances can be explored to make biosensors and is likely to provide good results if the correct proteins and relative ligands are used.
Persaud et al. (2009) achieved immobilisation of a number of OBPs from various species ranging from the pig to the wasp onto quartz microbalances (QMBs) and showed that they reversibly responded to a number of odorants with varying selectivity. A change in resonant frequency of the crystal is observed when exposed to a pulse of odorant. Pig OBP was immobilised on to a QMB, and pulses of eight odorants were applied at almost equal concentrations. Figure 4 shows the dynamic response obtained by pig OBP to a pulse of 350 ppm of 2-isobutyl-3-methoxypyrazine in air. Responses were seen to all compounds chosen from pyrazine families, but the highest sensitivity was to pyridine.
Dynamic response of a QMB coated with pig OBP to a pulse of 2-isobutyl methoxy pyrazine (350 ppm in air). The y-axis shows the change in frequency of oscillation observed as the ligand adsorbs on to the biosensor. This is reflected as a decrease in oscillatory frequency as the mass of the QMB increases due to the ligand binding (from Persaud et al. 2009)
Still, using quartz crystals and measuring surface acoustic wave (SAW), Di Pietrantonio et al. (2013) presented a first version of a multisensor array based on OBPs. The three proteins used were the pig OBP, the bovine OBP and a mutant of the latter. The two bovine OBPs showed very similar responses to 1-octen-3-ol and carvone, as expected, but the pig OBP gave a much stronger signal in the presence of octenol. This simple system allows discrimination between the two odorants and shows the way for a more complex multisensor array. The sensors appear to be robust with good reproducibility and a sensitivity in the range of few ppm.
Optical methods
Optical detection of ligand binding has not been adopted in biosensors, apart from a couple of reports. This is surprising, given the wide applications and popularity of surface plasmon resonance measurements adopted in commercial Biacore instruments. Although such analytical instruments are very reliable and accurate, their size and complexity of operation represent practical problems when transferring this technology to small, portable and easy to operate biosensors. Moreover, the Biacore instruments are more suitable to measure binding of large ligands, such as proteins, rather than small organic compounds, like odorants.
Using a similar type of approach, D’Auria et al. (2006) used the dog OBP to develop a biosensor for detecting odours, based on the measurement of refractive index. Unfortunately, experimental details are scarce, and this work was not followed by other reports using the same method.
It is also rather surprising that, given the wide popularity of the fluorescent-based protocol for measuring binding to OBPs, such approach has not been applied to sensor devices. This method, in fact, offers several advantages in terms of simplicity of use and equipment. Optical signals can be easily measured using simple devices, such as photodiodes, and can be efficiently transmitted through fibre optics. If suitable probes are adopted, the fluorescence changes can be immediately appreciated by direct observation, thus allowing a rapid screening of samples. The method can be also applied to the design and construction of dosimetres, taking advantage of the slow release of ligands by some OBPs, such as those of vertebrates, resulting in accumulation of the analyte inside the protein. Applications of this kind would include measuring the amount of degradation products in foods or accumulation of pollutants in the environment.
Other actual and potential applications
Controlled release of odorants and pheromones
The presence of OBPs in glands and organs producing pheromones suggests a use of these proteins to control the release of odorants in the environment. Currently, cyclodextrins can efficiently perform such task and are also used to reduce the concentration of objectional odours in closed spaces or on the human body (Damodaran and Arora 2013; Reineccius et al. 2002). We can reasonably predict that OBPs could perform the same tasks, with some additional advantages. In fact, proteins can provide better selectivity when needed, for example, by reducing the release of more volatile components of a blend, thus assuring a constant and balanced composition of a complex bouquet. The main objection against the use of OBPs are their allergenic properties, which could be answered by using recombinant human OBPs, which could be opportunely modified in their binding pocket, to meet required binding characteristics, without affecting their antigenic regions.
Trapping odours in the environment (deodorisers)
An interesting application of OBPs in purifying waste waters was suggested by Bianchi et al. (2013), who used bovine OBP to fabricate cartridges for removing the herbicide atrazine and similar compounds from water. Although the affinity of this compound for the protein is very weak, with a dissociation constant around 0.3 mM, their cartridges seem to be efficient in removing triazines from water even if these pollutants are present at micromolar levels. The authors themselves were not able to explain such behaviour; however, the idea is original and worth investigating for other possible applications. For example, aromatic chlorinated compounds, such as dioxin and several other derivatives, are expected to have high affinities for OBPs, based on the similarity of their structures with those of other good ligands (such as 1-aminoanthracene), as well as on the presence of chlorine atoms, which increase the hydrophobicity and thus the binding strength.
A very recent study explores the possibility of fabrics for clothes functionalised with OBPs, which could slowly release pleasant fragrances or reduce cigarette smoke and other unpleasant odours. The approach needs to be further investigated for practical application, but the idea is new and appealing (Silva et al. 2013).
Affinity columns for analysis and purification
The affinity and specificity of OBPs for small organic molecules can be exploited in a variety of analytical applications. Affinity columns derivatised with OBPs are easy to prepare and, due to the high stability of these proteins, can be stable after several uses and extensive washes with buffers that may also contain organic solvents. An example of such an application is reported by Margaryan et al. (2006), who covalently linked B. mori PBP1 and Culex quinquefasciatus OBP1 to affinity columns. The immobilisation chemistry did not affect the performance of the proteins, which retained their affinity to ligands. In particular, using a column derivatised with B. mori PBP1, different compounds were eluted with rates correlated to their known affinities in solution. Moreover, the same protein retained its property of a chemical switch, which is the change of conformation induced by pH, which could be exploited to release tightly bound ligand from the column. The use of OBPs to prepare affinity columns presents different and interesting applications, which would be worth investigating in more detail. For instance, proteins are ideal substrates to differentiate enantiomers, a task still far from being solved in analytical chemistry. Discrimination of enantiomers by OBPs has been poorly investigated, but a few examples indicate that this aspect should be better documented (Plettner et al. 2000; Cavaggioni et al. 2003). In fact, one of the main difficulties when using proteins for analytical columns is their poor stability to temperature and solvents. Once again, OBPs can provide the solution in such respect. OBP-derivatised columns can be directly used in resolving racemate mixtures of pheromones or other natural compounds. In fact, both classes of chemicals represent natural ligands for these proteins. Finally, the structures of OBPs, relative to the binding pockets, can be easily modified to meet other analytical chemistry requirements.
Conclusions
The exceptional stability of OBPs to thermal denaturation and proteolytic degradation is the basic characteristic which makes these proteins ideal elements in sensing devices as well as in other biotechnological applications. In addition, the large structural information on these proteins allows easy design and synthesis of mutants. Finally, OBPs are easy and economical to make and purify.
Only a few papers describe the potential of OBPs as biosensors for odours or describe their versatility in matching the complexity and variety of environmental odours. However, an efficient transduction system to obtain electric signals from ligand-binding events still seems to be lacking. Optical systems, using either external fluorescent probes or the tryptophan intrinsic fluorescence of the protein seem so far the most promising approach, although, as with other systems, the sensitivity is orders of magnitudes lower than in biological systems.
Other applications, using OBPs as reservoirs for slow release of odorants or as scavengers for pollutants, or else as active chromatographic phases for enantiomers separations, appear now more realistic and practical. In this case, the main issue is the cost of the protein, which has to be used in relatively large amounts. Once again, OBPs meet such requirements quite satisfactorily, as they can be expressed both in bacterial and eucaryotic systems with good yield and low cost.
References
Bacchini A, Gaetani E, Cavaggioni A (1992) Pheromone-binding proteins in the mouse Mus musculus. Experientia 48:419–421
Bailey K (2011) Making sense of it all: a review of olfactory biosensing. In: Kane D, Micolich A, Rabeau J (eds) Nanotechnology in Australia: Showcase of Early Career Research. Pan Stanford Publishing, Singapore, pp 375–408
Baldwin EA, Bai J, Plotto A, Dea S (2011) Electronic noses and tongues: applications for the food and pharmaceutical industries. Sensors 11:4744–4766
Ban LP, Zhang L, Yan YH, Pelosi P (2002) Binding properties of a locust’s chemosensory protein. Biochem Biophys Res Commun 293:50–54
Ban LP, Scaloni A, D’Ambrosio C, Zhang L, Yan YH, Pelosi P (2003) Biochemical characterisation and bacterial expression of an odorant-binding protein from Locusta migratoria. Cell Mol Life Sci 60:390–400
Ban L, Napolitano E, Serra A, Zhou X, Iovinella I, Pelosi P (2013) Identification of pheromone-like compounds in male reproductive organs of the oriental locust Locusta migratoria. Biochem Biophys Res Commun 437:620–624
Bar-Cohen Y (2011) Biological senses as inspiring model for biomimetic sensors. IEEE Sensors J 11:3194–3201
Bianchet MA, Bains G, Pelosi P, Pevsner J, Snyder SH, Monaco HL, Amzel LM (1996) The three dimensional structure of bovine odorant-binding protein and its mechanism of odor recognition. Nat Struct Biol 3:934–939
Bianchi F, Basini G, Grolli S, Conti V, Bianchi F, Grasselli F, Careri M, Ramoni R (2013) An innovative bovine odorant binding protein-based filtering cartridge for the removal of triazine herbicides from water. Anal Bioanal Chem 405:1067–1075
Bignetti E, Cavaggioni A, Pelosi P, Persaud KC, Sorbi RT, Tirindelli R (1985) Purification and characterization of an odorant binding protein from cow nasal tissue. Eur J Biochem 149:227–231
Böcskei Z, Groom CR, Flower DR, Wright CE, Phillips EV, Cavaggioni A, Findlay JBC, North ACT (1992) Pheromone binding to two rodent urinary proteins revealed by X-ray crystallography. Nature 360:186–188
Briand L, Nespoulous C, Huet JC, Takahashi M, Pernollet JC (2002a) Characterization of a chemosensory protein (ASP3c) from honeybee (Apis mellifera L.) as a brood pheromone carrier. Eur J Biochem 269:4586–4596
Briand L, Eloit C, Nespoulous C, Bézirard V, Huet JC, Henry C, Blon F, Trotier D, Pernollet JC (2002b) Evidence of an odorant-binding protein in the human olfactory mucus: location, structural characterization, and odorant-binding properties. Biochemistry 41:7241–7252
Calvello M, Guerra N, Brandazza A, D’Ambrosio C, Scaloni A, Dani FR, Turillazzi S, Pelosi P (2003) Soluble proteins of chemical communication in the social wasp Polistes dominulus. Cell Mol Life Sci 60:1933–1943
Campanacci V, Krieger J, Bette S, Sturgis JN, Lartigue A, Cambillau C, Breer H, Tegoni M (2001) Revisiting the specificity of Mamestra brassicae and Antheraea polyphemus pheromone-binding proteins with a fluorescence binding assay. J Biol Chem 276:20078–20084
Cavaggioni A, Mucignat-Caretta C, Zagotto G (2003) Absolute configuration of 2-sec-butyl-4,5-dihydrothiazole in male mouse urine. Chem Senses 28:791–797
Craven MA, Gardner JW, Bartlett PN (1996) Electronic noses—development and future prospects. Trends Anal Chem 15:486–493
Dal Monte M, Andreini I, Revoltella R, Pelosi P (1991) Purification and characterization of two odorant binding proteins from nasal tissue of rabbit and pig. Comp Biochem Physiol 99B:445–451
Dal Monte M, Centini M, Anselmi C, Pelosi P (1993) Binding of selected odorants to bovine and porcine odorant-binding proteins. Chem Senses 18:713–721
Damberger F, Nikonova L, Horst R, Peng G, Leal WS, Wuthrich K (2000) NMR characterization of a pH-dependent equilibrium between two folded solution conformations of the pheromone-binding protein from Bombyx mori. Protein Sci 9:1038–1041
Damodaran S, Arora A (2013) Off-flavor precursors in soy protein isolate and novel strategies for their removal. Ann Rev Food Sci Technol 4:327–346
Dani FR, Michelucci E, Francese S, Mastrobuoni G, Cappellozza S, La Marca G, Niccolini A, Felicioli A, Moneti G, Pelosi P (2011) Odorant-binding proteins and chemosensory proteins in pheromone detection and release in the silkmoth Bombyx mori. Chem Senses 36:335–344
D’Auria S, Staiano M, Varriale A, Scognamiglio V, Rossi M, Parracino A, Campopiano S, Cennamo N, Zeni L (2006) The odorant-binding protein from Canis familiaris: purification, characterization and new perspectives in biohazard assessment. Protein Pept Lett 13:349–352
Di Pietrantonio F, Cannatà D, Benetti M, Verona E, Varriale A, Staiano M, D’Auria S (2013) Detection of odorant molecules via surface acoustic wave biosensor array based on odorant-binding proteins. Biosens Bioelectron 41:328–334
D’Innocenzo B, Salzano AM, D’Ambrosio C, Gazzano A, Niccolini A, Sorce C, Dani FR, Scaloni A, Pelosi P (2006) Secretory proteins as potential semiochemical carriers in the horse. Biochemistry 45:13418–13428
Flower DR (1996) The lipocalin protein family: structure and function. Biochem J 318:1–14
Flower DR, North AC, Sansom CE (2000) The lipocalin protein family: structural and sequence overview. Biochim Biophys Acta 1482:9–24
Gardner JW (2004) Review of conventional electronic noses and their possible application to the detection of explosives, Electronic noses & sensors for the detection of explosives. Nato Science series II Mathematics, Physics and Chemistry. Springer, Netherlands, pp 1–28
Golebiowski J, Antonczak S, Fiorucci S, Cabrol-Bass D (2007) Mechanistic events underlying odorant binding protein chemoreception. Proteins 67:448–458
Guex N, Peitsch MC (1997) SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis 18:2714–2723
Horst R, Damberger F, Luginbuhl P, Guntert P, Peng G, Nikonova L, Leal WS, Wuthrich K (2001) NMR structure reveals intramolecular regulation mechanism for pheromone binding and release. Proc Natl Acad Sci U S A 98:14374–14379
Hou Y, Jaffrezic-Renault N, Martelet C, Tlili C, Zhang A, Pernollet J-C, Briand L, Gomila G, Errachid A, Samitier J, Salvagnac L, Torbiéro B, Temple-Boyer P (2005) Study of Langmuir and Langmuir–Blodgett films of odorant-binding protein/amphiphile for odorant biosensors. Langmuir 21:4058–4065
Iovinella I, Dani FR, Niccolini A, Sagona S, Michelucci E, Gazzano A, Turillazzi S, Felicioli A, Pelosi P (2011) Differential expression of odorant-binding proteins in the mandibular glands of the honey bee according to caste and age. J Proteome Res 10:3439–3449
Kruse SW, Zhao R, Smith DP, Jones DN (2003) Structure of a specific alcohol-binding site defined by the odorant binding protein LUSH from Drosophila melanogaster. Nat Struct Biol 10:694–700
Leal WS (2013) Odorant reception in insects: roles of receptors, binding proteins, and degrading enzymes. Annu Rev Entomol 58:373–391
Leal WS, Nikonova L, Peng G (1999) Disulfide structure of the pheromone binding protein from the silkworm moth, Bombyx mori. FEBS Lett 464:85–90
Lee SH, Jin HJ, Song HS, Hong S, Park TH (2012) Bioelectronic nose with high sensitivity and selectivity using chemically functionalized carbon nanotube combined with human olfactory receptor. J Biotech 157:467–472
Li S, Picimbon J-F, Ji SD, Kan YC, Qiao CL, Zhou J-J, Pelosi P (2008) Multiple functions of an odorant-binding protein in the mosquito Aedes aegypti. Biochem Biophys Res Commun 372:464–468
Liu Q, Wang H, Li H, Zhang J, Zhuang S, Zhang F, Hsia KJ, Wang P (2013) Impedance sensing and molecular modeling of an olfactory biosensor based on chemosensory proteins of honeybee. Biosens Bioelectron 40:174–179
Löbel D, Jacob M, Völkner M, Breer H (2002) Odorants of different chemical classes interact with distinct odorant binding protein subtypes. Chem Senses 27:39–44
Maida R, Ziegelberger G, Kaissling KE (2003) Ligand binding to six recombinant pheromone-binding proteins of Antheraea polyphemus and Antheraea pernyi. J Comp Physiol 173:565–573
Marabotti A, Lefevre T, Staiano M, Crescenzo R, Varriale A, Rossi M, Pezolet M, D’Auria S (2008a) Mutant bovine odorant-binding protein: temperature affects the protein stability and dynamics as revealed by infrared spectroscopy and molecular dynamics simulations. Proteins Struct Funct Genet 72:769–778
Marabotti A, Scire A, Staiano M, Crescenzo R, Aurilla V, Tanfani F, D’Auria S (2008b) Wild-type and mutant bovine odorant-binding proteins to probe the role of the quaternary structure organization in the protein thermal stability. J Proteome Res 7:5221–5229
Marchese S, Pes D, Scaloni A, Carbone V, Pelosi P (1998) Lipocalins of boar salivary glands binding odours and pheromones. Eur J Biochem 252:563–568
Margaryan A, Moaddel R, Aldrich JR, Tsuruda JM, Chen AM, Leal WS, Wainer IW (2006) Synthesis of an immobilized Bombyx mori pheromone-binding protein liquid chromatography stationary phase. Talanta 70:752–755
Matsuo T, Sugaya S, Yasukawa J, Aigaki T, Fuyama Y (2007) Odorant-binding proteins OBP57d and OBP57e affect taste perception and host-plant preference in Drosophila sechellia. PLoS Biol 5:e118
Nespoulous C, Briand L, Delage MM, Tran V, Pernollet JC (2004) Odorant binding and conformational changes of a rat odorant-binding protein. Chem Senses 29:189–198
Paolini S, Scaloni A, Amoresano A, Marchese S, Napolitano E, Pelosi P (1998) Amino acid sequence, post-translational modifications, binding and labelling of porcine odorant-binding protein. Chem Senses 23:689–698
Paolini S, Tanfani F, Fini C, Bertoli E, Pelosi P (1999) Porcine odorant-binding protein: structural stability and ligand affinities measured by fourier-transform infrared spectroscopy and fluorescence spectroscopy. Biochim Biophys Acta 1431:179–188
Pelosi P (1994) Odorant-binding proteins. Crit Rev Biochem Mol Biol 29:199–228
Pelosi P (1996) Perireceptor events in olfaction. J Neurobiol 30:3–19
Pelosi P (1998) Odorant-binding proteins: structural aspects. Ann N Y Acad Sci 855:281–293
Pelosi P (2001) The role of perireceptor events in vertebrate olfaction. Cell Mol Life Sci 58:503–509
Pelosi P, Maida R (1990) Odorant binding proteins in vertebrates and insects: similarities and possible common function. Chem Senses 15:205–215
Pelosi P, Maida R (1995) Odorant-binding proteins in insects. Comp Biochem Physiol 111B:503–514
Pelosi P, Pisanelli AM, Baldaccini NE, Gagliardo A (1981) Binding of [3H]-2-isobutyl-3-methoxypyrazine to cow olfactory mucosa. Chem Senses 6:77–85
Pelosi P, Baldaccini NE, Pisanelli AM (1982) Identification of a specific olfactory receptor for 2-isobutyl-3-methoxypyrazine. Biochem J 201:245–248
Pelosi P, Zhou J-J, Ban LP, Calvello M (2006) Soluble proteins in insect chemical communication. Cell Mol Life Sci 63:1658–1676
Persaud KC, Ng SM, Mucignat C, Pelosi P (2009) Odorant binding proteins and mouse urinary proteins: potential biomimetic sensing systems. Chem Senses 34:E36–E37
Pevsner J, Hou V, Snowman AM, Snyder SH (1990) Odorant-binding protein. Characterization of ligand binding. J Biol Chem 265:6118–6125
Plettner E, Lazar J, Prestwich EG, Prestwich GD (2000) Discrimination of pheromone enantiomers by two pheromone binding proteins from the gypsy moth Lymantria dispar. Biochemistry 39:8953–8962
Reineccius TA, Reineccius GA, Peppard TL (2002) Encapsulation of flavors using cyclodextrins: comparisonof flavor retention in alpha, beta, and gamma types. J Food Sci 67:3271–3279
Riviere S, Lartigue A, Quennedey B, Campanacci V, Farine JP, Tegoni M, Cambillau C, Brossut R (2003) A pheromone-binding protein from the cockroach Leucophaea maderae: cloning, expression and pheromone binding. Biochem J 371:573–579
Rodgers M, Findlay J, Millner P (2010) Lipocalin based biosensors for low mass hydrophobic analytes; development of a novel SAM for polyhistidine tagged proteins. Sensors Actuators B Chem 150:12–18
Sandler BH, Nikonova L, Leal WS, Clardy J (2000) Sexual attraction in the silkworm moth: structure of the pheromone-binding-protein-bombykol complex. Chem Biol 7:143–151
Sankaran S, Panigrahi S, Mallik S (2011) Odorant binding protein based biomimetic sensors for detection of alcohols associated with Salmonella contamination in packaged beef. Biosens Bioelectron 26:3103–3109
Scaloni A, Monti M, Angeli S, Pelosi P (1999) Structural analyses and disulfide-bridge pairing of two odorant-binding proteins from Bombyx mori. Biochem Biophys Res Commun 266:386–391
Scire A, Marabotti A, Staiano M, Briand L, Varriale A, Bertoli E, Tanfani F, DAuria S (2009) Structure and stability of a rat odorant-binding protein: another brick in the wall. J Proteome Res 8:4005–4013
Silva C, Matamá T, Azoia NG, Mansilha C, Casal M, Cavaco-Paulo A (2013) Odorant binding proteins: a biotechnological tool for odour control. Appl Microbiol Biotechnol. doi:10.1007/s00253-013-5243-5249
Spinelli S, Ramoni R, Grolli S, Bonicel J, Cambillau C, Tegoni M (1998) The structure of the monomeric porcine odorant binding protein sheds light on the domain swapping mechanism. Biochemistry 37:7913–7918
Spinelli S, Vincent F, Pelosi P, Tegoni M, Cambillau C (2002) Boar salivary lipocalin. Three-dimensional X-ray structure and androsterol/androstenone docking simulations. Eur J Biochem 269:2449–2456
Steinbrecht RA (1998) Odorant-binding proteins: expression and function. Ann N Y Acad Sci 855:323–332
Sun YL, Huang LQ, Pelosi P, Wang CZ (2012a) Expression in antennae and reproductive organs suggests a dual role of an odorant-binding protein in two sibling Helicoverpa species. PLoS One 7:e30040
Sun YF, De Biasio F, Qiao HL, Iovinella I, Yang SX, Ling Y, Riviello L, Battaglia D, Falabella P, Yang XL, Pelosi P (2012b) Two odorant-binding proteins mediate the behavioural response of aphids to the alarm pheromone (E)-β-farnesene and structural analogues. PLoS One 7:e32759
Swarup S, Williams TI, Anholt RR (2011) Functional dissection of odorant binding protein genes in Drosophila melanogaster. Genes Brain Behav 10:648–657
Tcatchoff L, Nespoulous C, Pernollet JC, Briand L (2006) A single lysyl residue defines the binding specificity of a human odorant-binding protein for aldehydes. FEBS Lett 580(8):2102–2108
Tegoni M, Ramoni R, Bignetti E, Spinelli S, Cambillau C (1996) Domain swapping creates a third putative combining site in bovine odorant binding protein dimer. Nat Struct Biol 3:863–867
Tegoni M, Pelosi P, Vincent F, Spinelli S, Campanacci V, Grolli S, Ramoni R, Cambillau C (2000) Mammalian odorant binding proteins. Biochim Biophys Acta 1482:229–240
Tegoni M, Campanacci V, Cambillau C (2004) Structural aspects of sexual attraction and chemical communication in insects. Trends Biochem Sci 29:257–264
Topazzini A, Pelosi P, Pasqualetto PL, Baldaccini NE (1985) Specificity of a pyrazine binding protein from cow olfactory mucosa. Chem Senses 10:45–49
Vincent F, Spinelli S, Ramoni R, Grolli S, Pelosi P, Cambillau C, Tegoni M (2000) Complexes of porcine odorant binding protein with odorant molecules belonging to different chemical classes. J Mol Biol 300:127–139
Vincent F, Löbel D, Brown K, Spinelli S, Grote P, Breer H, Cambillau C, Tegoni M (2001) Crystal structure of aphrodisin, a sex pheromone from female hamster. J Mol Biol 305:459–469
Virtanen T, Kinnunen T, Rytkönen-Nissinen M (2012) Mammalian lipocalin allergens-insights into their enigmatic allergenicity. Clin Exp Allergy 42:494–504
Vogt RG (2003) Biochemical diversity of odor detection: OBPs, ODEs and SNMPs. In: Blomquist GJ, Vogt RG (eds) Insect pheromone biochemistry and molecular biology. Elsevier Academic, London, pp 391–446
Vogt RG (2005) Molecular basis of pheromone detection in insects. In: Gilbert LI, Iatrou K, Gill S (eds) Comprehensive insect physiology, biochemistry, pharmacology and molecular biology, vol 3, Endocrinology. Elsevier, London, pp 753–804
Vogt RG, Riddiford LM (1981) Pheromone binding and inactivation by moth antennae. Nature 293:161–163
Wanner KW, Willis LG, Theilmann DA, Isman MB, Feng Q, Plettner E (2004) Analysis of the insect OS-D-like gene family. J Chem Ecol 30:889–911
Wei Y, Brandazza A, Pelosi P (2008) Binding of polyclycic aromatic hydrocarbons to mutants of odorant-binding protein: a first step towards biosensors for environmental monitoring. Biochim Biophys Acta 1784:666–671
Wyszynski B, Nakamoto T (2009) Linking biological and artificial olfaction: biomimetic quartz crystal microbalance odor sensors. IEEJ Trans Electr Electron Eng 4:334–338
Xu P, Atkinson R, Jones DN, Smith DP (2005) Drosophila OBP LUSH is required for activity of pheromone-sensitive neurons. Neuron 45:193–200
Yabuki M, Scott DJ, Briand L, Taylor AJ (2011) Dynamics of odorant binding to thin aqueous films of rat-OBP3. Chem Senses 36:659–671
Yoon H, Lee SH, Kwon OS, Song HS, Oh EH, Park TH, Jang J (2009) Polypyrrole nanotubes conjugated with human olfactory receptors: high-performance transducers for FET-type bioelectronic noses. Angew Chem Int Ed 48:2755–2758
Zhou J-J, Zhang G-A, Huang W, Birkett MA, Field LM, Pickett JA, Pelosi P (2004) Revisiting the odorant-binding protein LUSH of Drosophila melanogaster: evidence for odour recognition and discrimination. FEBS Lett 558:23–26
Zhou XH, Ban LP, Iovinella I, Zhao LJ, Gao Q, Felicioli A, Sagona S, Pieraccini G, Pelosi P, Zhang L, Dani FR (2013) Diversity, abundance and sex-specific expression of chemosensory proteins in the reproductive organs of the locust Locusta migratoria manilensis. Biol Chem 394:43–54
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pelosi, P., Mastrogiacomo, R., Iovinella, I. et al. Structure and biotechnological applications of odorant-binding proteins. Appl Microbiol Biotechnol 98, 61–70 (2014). https://doi.org/10.1007/s00253-013-5383-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-013-5383-y