Abstract
Lactic acid bacteria populations of red wine samples from industrial fermentations, including two different vinification methods were studied. For this investigation, polymerase chain reaction–denaturing gradient gel electrophoresis (PCR-DGGE) analysis was employed to supplement previous results that were obtained by culture-dependent methods. PCR-DGGE was aimed to study two targeted genes, 16S ribosomal DNA (rDNA) and rpoB, and the results were useful to evaluate the microbial populations in wine samples. Moreover, an improvement of a detection limit determined so far for DGGE analysis was obtained with the method described in this study, what made possible to identify lactic acid bacteria populations below 101 colony-forming unit/mL. The species Oenococcus oeni was the most frequently detected bacterium, but identifications close to species Oenococcus kitaharae and Lactococcus lactis that are not often found in wine were firstly identified in samples of this research. PCR-DGGE allowed to detect 9 out of 11 lactic acid bacteria species identified in this study (nine by PCR-16S rDNA/DGGE and four by PCR-rpoB/DGGE), while five species were detected using the modified de Man, Rogosa and Sharpe agar. Therefore, the two methods were demonstrated to be complementary. This finding suggests that analysis of the lactic acid bacteria population structure in wine should be carried out using both culture-dependent and culture-independent techniques with more than one primer pair.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Winemaking is a complex process that involves numerous microorganisms from which the main microorganisms are yeast and bacteria. The yeasts lead the alcoholic fermentation (AF), wherein glucose is mainly converted to ethanol, and the lactic acid bacteria (LAB) perform the malolactic fermentation (MLF). MLF is a part of LAB metabolism and mainly consists on the transformation of l-malic acid into l-lactic acid and carbon dioxide, producing several intermediate and final compounds (Vanvuuren and Dicks 1993). This biological transformation is a recommended oenological practice for certain types of white wines and for almost all red wines because of its significant contribution to increasing the microbiological stability of wines (Alexandre et al. 2004; Lonvaud-Funel 1999, 2008) and to improving its sensorial characteristics (Andorrà et al. 2010; de Revel et al. 1999; Torriani et al. 2011).
The LAB ecology in wines represents a microbiological factor that should be monitored to achieve high-quality wines (Andorrà et al. 2010; Kim et al. 2011; Reguant et al. 2005). To understand the ecosystem structure of LAB in wines, many studies about LAB species in red wine have been conducted by using culture-dependent methods (González-Arenzana et al. 2012a, 2013), but results have always been limited by the ability of the microorganisms to develop their metabolisms in culture media (Cocolin et al. 2011a; Renouf et al. 2005a, b). Moreover, the presence of viable but non-culturable microorganisms in wine (Divol and Lonvaud-Funel 2005; Millet and Lonvaud-Funel 2000), which are not able to form colonies on microbiological media, has made necessary the use of different methods based on culture-independent techniques such as polymerase chain reaction (PCR) and denaturing gradient gel electrophoresis (DGGE) to resolve this difficulty (Andorrà et al. 2008; Renouf et al. 2006a, b).
This qualitative technique makes possible separation of PCR amplicons of the same size but of different sequences in denaturing gradient gels (Cocolin and Mills 2003; Cocolin et al. 2011b). This technique is a common method employed to characterise microbial communities from specific environmental niches (Muyzer and Smalla 1998) because it enables detection of individual species as well as overall profiling of community structure changes with time. The 16S ribosomal DNA (rDNA) gene has been the most employed one among all the universal genes to detect LAB species by culture-independent techniques (López et al. 2003). However, several authors have proposed that PCR-DGGE technique should be focused on the study of rpoB gene in order to minimise the biases caused by intraspecies heterogeneity that 16S rDNA genes can produce (Renouf et al. 2006a, b).
In this research, LAB community structure and its evolution along the fermenting process of Rioja red wines was analysed with 16S rDNA and rpoB/PCR-DGGE. Rioja wine comes from the Rioja region of northern Spain. Rioja is a region with a long, glorious viniculture history and was the first Qualified Appellation of Origin region in Spain. Rioja wine, especially the red, is the most well-known Rioja style. Classic, bold, these wines taste mostly of their Tempranillo roots and have a bright, fresh flavour to them. The study described in this paper was conducted in these fermenting wines to expand on previous ecological research that used culture-dependent methods (González-Arenzana et al. 2012a).
Materials and methods
Wine fermentations and sampling
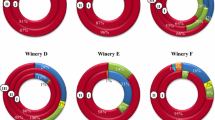
The samples of Tempranillo musts and red wines were taken from four wineries located in the Rioja Appellation in one vintage. The winemaking process involves manual harvesting of grapes followed by two different vinification methods that are representative of the most important types of red winemaking: wineries A, B and C, where AF was performed with the destemming and crushing method in stainless steel tanks, and winery D, where AF was conducted by the traditional semi-carbonic maceration method (Ribéreau-Gayon et al. 2007) in open cement tanks (Fig. 1). When AF was completed, wines were racked and placed in stainless steel tanks for the case of wineries A, B and C and in an open cement tank for winery D. The wines underwent spontaneous MLF with the endogenous microbiota (no starter inocula were used).
One fermentation tank was sampled in each winery. Must and wine samples were taken aseptically at different moments: must (stage 1), middle AF (density around 1,025; stage 2), at the end of AF (<2 g/L glucose + fructose; stage 3), initial MLF (consumption of 10 % of the initial malic acid; stage 4), middle MLF (consumption of 60 % of the initial malic acid; stage 5) and at the end of MLF (l-malic acid concentration of <0.5 g/L; stage 6). Wineries A and B were not sampled during AF.
Bacterial identification by culture-independent methods
Direct DNA extraction from wine samples for culture-independent methods
A previous study was performed to optimise direct DNA extraction from red wine (González-Arenzana et al. 2010). In this study, different commercial DNA extraction kits (Plant DNeasy kit from Qiagen, PowerSoil® and PowerFood® DNA isolation kit from MO BIO) were combined with the addition of zirconium hydroxide. The best result was obtained using the procedure described below. A volume of 10 mL of each sample was centrifuged (20 min, 4,000×g, 4 °C). The supernatant was discarded, and 1.2 mL of saline solution (NaCl 0.9 %) and 2.4 mL of zirconium hydroxide (7 g/L) were added to the pellet to facilitate pelleting of the bacteria in wine (Lucore et al. 2000). After 10 min of horizontal shaking at room temperature, the suspension was again centrifuged (10 min, 500×g, 7 °C). The DNA was subsequently extracted and purified from the cell pellet using the PowerSoil® DNA isolation kit (MO BIO Laboratories, Inc., Carlsbad, CA USA) and Fast Prep™ (FP120, BIO 101, Thermo Electron Corporation, USA) bead beater instrument (twice for 45 s at speed setting of 6) as per the manufacturer's instructions.
PCR conditions
PCR was performed using the Applied Biosystem, GeneAmp® PCR System 2700 thermocycler at a final volume of 50 μL. To amplify the region V4 to V5 of 16S rDNA gene, primers WLAB1 and WLAB2GC were used as described by López et al. (2003). Moreover, the rpoB1, rpoB1o and rpoB2GC primers were employed to amplify the region of the rpoB gene as described by Renouf et al. (2006a, b) with the following modifications: 0.5 μM of each primer, 1 mM dNTP mix and 0.5 μL of PfuUltra II Fusion HS DNA Polymerase (Stratagene).
DGGE analysis
The separation of the respective PCR products was performed with the DCode™ Universal Mutation Detection System (Bio-Rad, Hercules, CA). PCR products were run on 8 % (w/v) polyacrylamide gels in TAE buffer (2 M Tris, 1 M glacial acetic acid and 50 mM EDTA pH 8) at a constant temperature of 60 °C. WLAB1–WLAB2GC amplicons were separated with gels containing 35 to 55 % urea–formamide gradient, and electrophoresis was performed for 10 min at 20 V and for 18 h at 80 V. rpoB amplicons were separated with gels containing 32 to 50 % urea–formamide gradient, and the electrophoresis was performed for 10 min at 20 V and for 15 h at 60 V. Gels were stained in ethidium bromide after the electrophoresis and then were visualised with UV transillumination (Gel Doc, Bio-Rad). Blocks of the polyacrylamide gels which contained the selected DGGE bands were excised and subsequently incubated overnight in 20 μL of sterile and pure water at 4 °C to make DNA bands diffuse to the liquid. One microlitre of this elution was reamplified using the PCR conditions described above to DNA sequencing.
DNA sequencing and phylogenetic analysis
PCR products were sequenced by Macrogen, Inc. (Seoul, South Korea). The quality and characteristic of the obtained sequences were analysed with the software InfoQuest™ 5.1, and only those products considered as appropriate were used for comparison to the GenBank nucleotide database with the Basic Local Alignment Search Tool (BLAST) (Altschul et al. 1990). The 16S rDNA and rpoB sequences were deposited in the GenBank nucleotide database under the accession numbers JQ838834–JQ838875 and JQ838876–JQ838890, respectively. After this preliminary study, our sequences were assembled and submitted to phylogenetic and evolutionary analysis with the Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0.1 (Tamura et al. 2007). The neighbour-joining method (Saitou and Nei 1987) identified the relationships between the obtained sequences and the reliability of the identifications that were provided by the GenBank databases. The bootstrap test was based on 1,000 random revamping (Felsenstein 1985). The evolutionary distances were computed using the maximum composite likelihood method (Tamura et al. 2007) that made it possible to calculate the units that were equivalent to the base substitutions per site.
Bacterial identification by culture-dependent methods
The bacterial isolation was performed in de Man, Rogosa and Sharpe (MRS) agar (Scharlau Chemie S.A., Barcelona. Spain) plates supplemented with tomato juice (10 % v/v), fructose (6 g/L), cysteine HCl (0.5 g/L), dl-malic acid (5 g/L) and 50 mg/L of pymaricine to avoid yeast growth (Acofarma, S. Coop., Spain), and species identification was carried out as described by González-Arenzana et al. (2012a).
Results
LAB species detected by PCR-16S rDNA/DGGE
The DNA was extracted directly from must and wine samples and was submitted to PCR targeted to the 16S rDNA gene and later to DGGE. One hundred thirty-seven bands were retrieved from different DGGE gels (Fig. 2) and were sequenced by Macrogen, Inc.
The obtained sequences were carefully analysed with the bioinformatic software InfoQuest™ 5.1. Only those with reliable sequencing were included in the GenBank Data Library, referred each one to an accession number and then submitted to BLAST searches at the National Centre of Biotechnology Information GenBank. Some sequences were clearly identified, and they corresponded to sequences available in the databank, and others were compared to the most closely related species (Fig. 3). Finally, 38 sequences were processed with the MEGA software 4.0.2 and shaped the phylogenetic neighbour-joining tree displayed in Fig. 3. This tree was established by two main ramifications or branches. The “branch a” was composed by the LAB families Leuconostococaceae and Lactobacillaceae. The family Leuconostococaceae was formed by the genera Oenococcus and Leuconostoc. Eighteen bands were identified as Oenococcus oeni; three, close to Oenococcus kitaharae; four, as or similar to Leuconostoc mesenteroides; and finally, one, close to Leuconostoc fallax. The family Lactobacillaceae was integrated by the genera Lactobacillus and Pediococcus. Five sequences were identified as or close to Lactobacillus plantarum; two, near to Lactobacillus buchneri; one, as Pediococcus pentosaceus; and two, close to Pediococcus parvulus. Otherwise, the “branch b” included two sequences belonging to the family Streptococcaceae and the genus Lactococcus, which were identified as close to the species Lactococcus lactis.
Phylogenetic relationship of DNA sequences obtained from 16S rDNA PCR/DGGE. The phylogenetic tree was inferred using the neighbour-joining method (Saitou and Nei 1987) (1,000 bootstrap analysis). The evolutionary distances were computed using the maximum composite likelihood method (Tamura et al. 2007) and are in the units of the number of base substitutions per site. Each sequence is defined with the most accurate identification and the identity percentage (%), with the given accession number and with a letter and a number that mean the winery and isolation stage, respectively
LAB species detected by PCR-rpoB/DGGE
The LAB species detected in the fermenting wine samples with PCR-rpoB/DGGE are displayed in Fig. 4. In this case, 29 bands were sequenced, and 15 were considered qualified enough to be submitted in the GenBank nucleotide database. These sequences were linked to a different accession number and were included in the phylogenetic study. Two branches could be observed in the tree (Fig. 4). The branch a was made up by sequences belonging to the families Leuconostococaceae and Lactobacillaceae. Eight sequences were identified as O. oeni; one, as L. mesenteroides; and two, similar to L. buchneri. The family Lactobacillaceae was also represented in the branch b where four sequences were confirmed near to P. parvulus.
Phylogenetic relationship of DNA sequences obtained from rpoB PCR/DGGE. The phylogenetic tree was inferred using the neighbour-joining method (Saitou and Nei 1987) (1,000 bootstrap analysis). The evolutionary distances were computed using the maximum composite likelihood method (Tamura et al. 2007) and are in the units of the number of base substitutions per site. Each sequence is defined with the most accurate identification and the identity percentage (%), with the given accession number and with a letter and a number that mean the winery and isolation stage, respectively
Distribution of LAB in each winery and stage of vinification
The LAB species identified at each fermentative stage in every sampled winery are shown in Table 1. In this table, both results from culture-independent (PCR-16S rDNA/DGGE and PCR-rpoB/DGGE) and culture-dependent techniques of a previous study (González-Arenzana et al. 2012a) of the same wine samples are included.
Cellars A and B were not sampled during AF, while C and D wineries were studied during AF and MLF. In relation to the LAB species distribution during the process of winemaking, it was observed by culture-independent methods that six species were detected at final AF (stage 3) in winery B and between one and three species in wineries C and D during the whole AF process. In relation to the LAB diversity during MLF, it was observed that O. oeni was the only species found at this period in fermenting wines from cellars A and C. In contrast, five different identifications besides O. oeni were found in the wines from winery B at stage 4. In addition, identifications close to P. parvulus were also detected, apart from O. oeni, at all the sampled MLF stages in winery D, and in the middle period (stage 5), another LAB species was found and was later identified near to O. kitaharae.
Discussion
In this study, the LAB community structure and its evolution along the fermenting process of Rioja wines were analysed using 16S rDNA (Fig. 3) and rpoB/PCR-DGGE (Fig. 4).
The PCR-16S rDNA/DGGE enabled us to detect the highest number species of this study (nine) belonging to three families such as Leuconostococaceae and Lactobacillaceae that are widely found in winemaking according to the literature (Dicks and Endo 2009; Zhang et al. 2011) and Streptococcaceae that is rarely associated to wine. These three families were represented by five genera. The genus Oenococcus was the most representative inside the family Leuconostococaceae, and O. oeni was the most detected LAB species due to its great adaptation to the wine conditions (López et al. 2008; Renouf and Favier 2010). Surprisingly, in the genus Oenococcus, several sequences were identified with BLAST close to O. kitaharae (JQ838839, JQ838856 and JQ838866), despite having never been detected in wine because of its poor adaptation to the stressful wine environment (Endo and Okada 2006). O. kitaharae could have been present at the wine samples, but not necessarily with an active metabolism during fermentation, as the PCR-DGGE was based on DNA/PCR-DGGE (Cocolin et al. 2011a). To the best of our knowledge, the proximity of those sequences placed O. kitaharae in the group of the genus Oenococcus in the phylogenetic tree (Fig. 3), which meant that the identification in GenBank nucleotide database may have been adequate. However, the unusual detection of O. kitaharae in fermenting wines could be due to the Library of GenBank nucleotide database not being complete enough to achieve a more accurate identification. These sequences could be considered to a different group inside the genus Oenococcus as other authors have already reported in their studies (Lucena et al. 2010; Renouf et al. 2009). In relation to the genus Leuconostoc, species such as L. mesenteroides and L. fallax that were previously reported by many authors in wines have been also detected in this research (Dicks and Endo 2009; Ogier et al. 2008). Relating to the genus Lactobacillus (Fig. 3), some sequences were identified as similar to L. plantarum and L. buchneri. The first one has been considered as habitual species in stages previous to the beginning of MLF because they have malolactic activities, but they are not usually adapted to the pH and ethanol of wine (Dicks and Endo 2009; du Toit et al. 2011). The species L. buchneri that was isolated from must and wines has been deeply analysed along with O. oeni in relation to arginine metabolism (Liu et al. 1994; Mira de Orduña et al. 2001). Inside this same family and very close to the genus Lactobacillus, two species of the genus Pediococcus were included. These species are sometimes related to spoilage wines or to difficult MLF situations (Dobson et al. 2002). P. parvulus is a species usually present in must and wine samples that can produce extracellular polysaccharides (Dols-Lafargue et al. 2008), and regarding to P. pentosaceus, it is known that under glucose limitation, it is able to use glycerol-producing pyruvate (Ribéreau-Gayon et al. 2007). Finally, the other genus found in this study with 16S rDNA/PCR-DGGE was Lactococcus, specifically the species L. lactis which has been usually attached to dairy products. This species is not very frequently reported in wine samples, although it has been recently detected in grapes samples from Australian vineyards (Bae et al. 2006), in musts from Ribera Sacra (Mesas et al. 2011) and from Rioja (López-Alfaro 2004).
Respect to the analysis of DGGE based on the PCR of the rpoB gene was rather a lack of results although allowed to detect four different species such as O. oeni, L. mesenteroides, L. buchneri and P. parvulus. The main reason to use PCR-rpoB/DGGE was the deviation caused by the presence of several copies of the 16S rDNA gene in the genome of some microorganisms (Renouf et al. 2006a, b). However, in certain wine samples that were submitted to PCR-rpoB/DGGE, more than five bands proceeding from the same direct DNA extraction were later identified as the same species (data not shown). Thus, we were not able to avoid the intraspecies heterogeneity associated to 16S rDNA gene (Case et al. 2007). In addition to these considerations, it was important to take into account that GenBank nucleotide database includes few sequences of rpoB gene from different LAB species, and this lack of sequences made the identification rather limited, as other authors have already described (Ruiz et al. 2010). Nevertheless, the results that were obtained with PCR-rpoB/DGGE could be complementary to PCR-16S rDNA/DGGE results when the study was focused on each fermentation stage.
A comparison of the PCR-16S rDNA and rpoB/DGGE results with those obtained from the identification of the colonies isolated from modified MRS plates in a previous study (González-Arenzana et al. 2012a) revealed coincidental results in certain wine samples, but some differences in others. A total of 11 different LAB species were identified in the four studied wineries by traditional and culture-independent methods. PCR-DGGE of the 16S rDNA-targeted gene allowed identifying nine species in comparison with the four species that were detected by PCR-rpoB/DGGE and the five species detected by culture in plates of the samples. Thus, L. buchneri, L. lactis, L. fallax, O. kitaharae, P. parvulus and P. pentosaceus were not detected in the employed culture medium, while Lactobacillus mali was not detected by PCR-DGGE.
The LAB species distribution at each winery during the winemaking process revealed different situations. In winery A, only O. oeni was detected by PCR-DGGE during MLF, and this finding was concordant with the plating results in which the 100 % of the isolates were identified as O. oeni.
The winery B showed the highest LAB species richness, detecting 9 out of 11 LAB species that were identified in this study. At the end of AF (no colony isolation performed, stage 3), six LAB species were found; L. mesenteroides was the only species detected by DGGE with the two targeted genes and the only one detected by PCR-rpoB/DGGE. This coincidental result could be due to the presence of an important population of this species in the analysed samples (Cocolin et al. 2011a). A great LAB species diversity (six species, between them L. lactis, L. fallax, L. mesenteroides, O. kitaharae, O. oeni and L. plantarum) was also observed at the initial MLF (stage 4) and was in accordance with the results that were obtained from culture plates (four identified species) in terms of the highest detected species diversity, but not in terms of microbial composition. This observation has been also reported in other fermented foods (Meroth et al. 2003; Pérez-Pulido et al. 2006). Moreover, L. mali was the 50 % of the species isolated in plate, but it was not detected by PCR-DGGE. This lack of concordance between techniques could be related first to the effect of the enrichment medium that would favour growth of some species over the others. In this sense, other authors have already reported that high populations on plating media were not found by PCR-DGGE (Rantsiou et al. 2008). Secondly, this disagreement could also be in relation to other technique biases such as co-migrations, primer competition and limit of detection (Cocolin et al. 2011a). In this winery B, O. oeni was not able to grow on plating media despite being detected by PCR-DGGE at stage 4, most likely due to the competition with other LAB species that were present in the media, which could suggest that MLF did not actually begin as was indicated by the viable LAB count and the malic acid consumption at this stage (González-Arenzana et al. 2012a). Moreover, the O. oeni detection with the two targeted genes during stages 5 and 6 was in agreement with the results derived from the previous plating study and supported viability DGGE in analysing large populations (106–107 colony-forming unit (CFU)/mL) from winemaking samples (Bester et al. 2010).
The whole process of winemaking (AF and MLF) was studied in wineries C and D. Curiously, in winery C, O. oeni was identified by PCR-DGGE and also in the MRS agar during every stage of AF and MLF, despite the fact that the LAB viable populations were low at some stages (101 CFU/mL at stage 1 and 102 CFU/mL at stage 2). This fact meant an increase in the detection limit for PCR-DGGE previously established by Cocolin et al. (2011b). During the early AF (stages 1 and 2), four LAB species were detected in addition to O. oeni, being L. plantarum noticed by PCR-16S rDNA/DGGE and by plating. L. mali was not detected by culture-independent techniques similar to the results from winery B. L. buchneri and P. parvulus were detected in AF by PCR-DGGE which was in agreement with previous studies (Renouf et al. 2005a, b). During MLF, O. oeni was the only species detected by all the employed techniques, supporting previous studies (González-Arenzana et al. 2012a, b), which highlighted the enormous adaptation of this species to the strict wine conditions.
Surprisingly, in winery D, L. mesenteroides and L. plantarum were detected at stage 1 by PCR-16S DNA/DGGE, despite any LAB species that was able to grow in plating media at this moment. This result improved the detection limit for PCR-DGGE which is described above. The immobilisation with metal hydroxides to concentrate food-borne bacteria was developed by Lucore for bacterial detection by cultural and molecular methods in dairy products by Lucore et al. (2000) and was applied to bacterial DNA extraction for RT-PCR in wine by Phister et al. (2007). To our knowledge, this study details for the first time that direct DNA extraction with a soil kit has been combined with the application of zirconium hydroxide, and the present study demonstrated that this method was a successful strategy to study the dynamics of LAB species in red wine samples by PCR-DGGE.
In this winery D, four LAB species that were different to O. oeni were detected using the only one of the methods at middle and final AF (LAB populations around 101 CFU/mL). L. mesenteroides was an exception because it was detected by PCR-16S DNA/DGGE and by plating. During MLF (LAB populations of >106 CFU/mL), identifying two different LAB species, in addition to O. oeni, was also surprising. This finding represents for the first time that O. kitaharae and P. parvulus were detected together with O. oeni in a correct (without quality deviation) wine fermentation that was conducted in open cement tanks with whole grapes. This peculiar result was in accordance with the suggestion that the employed winemaking method might create a special ecosystem with its own microbiota, as reported in a previous study (González-Arenzana et al. 2012a).
In this study, results obtained by culture-independent methods were in agreement with the ones in culture in plating media. Both methods were complementary during AF (with low LAB viable populations) and during MLF (with high populations), but in most of the studied fermentation stages, PCR-DGGE allowed to detect a greater number of species that were observed growing on MRS agar. The highest LAB species diversity was detected during AF, and an alternation of species was observed at a previous stage of the beginning of MLF as it has been reported in other wine fermentations (López-Alfaro 2004; Renouf et al. 2005a, b; Ruiz-Larrea et al. 2001).
In summary, this study contributed to increasing the knowledge of LAB diversity in red fermenting wines from La Rioja (Spain). The method developed in this study allowed to improve the detection limit that has been established to date for the T/DGGE direct analysis below 101 CFU/mL in some wine samples. It was firstly showed in the presence of the species O. kitaharae-like and L. lactis in fermenting wine samples. PCR/DGGE allowed us to increase the number of LAB species usually detected during MLF in a wine. Further research will be necessary to determine if this result was in relation to the different types of winemaking carried out in winery D (semi-carbonic maceration). The results from PCR/DGGE with two targeted genes and plating were complementary throughout the entire winemaking process; therefore, a parallel analysis based on culture-dependent and culture-independent methods should be developed to achieve a complete knowledge of LAB ecology in wine.
References
Alexandre H, Costello PJ, Remize F, Guzzo J, Guilloux-Benatier M (2004) Saccharomyces cerevisiae–Oenococcus oeni interactions in wine: current knowledge and perspectives. Int J Food Microbiol 93:141–154. doi:10.1016/j.ijfoodmicro.2003.10.103
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic Local Alignment Search Tool. J Mol Biol 215:403–410
Andorrà I, Landi S, Mas A, Guillamón JM, Esteve-Zarzoso B (2008) Effect of oenological practices on microbial populations using culture-independent techniques. Food Microbiol 25:849–856. doi:10.1016/j.fm.2008.05.005
Andorrà I, Landi S, Mas A, Esteve-Zarzoso B, Guillamón JM (2010) Effect of fermentation temperature on microbial population evolution using culture-independent and dependent techniques. Food Res Int 43:773–779. doi:10.1016/j.foodres.2009.11.014
Bae S, Fleet GH, Heard GM (2006) Lactic acid bacteria associated with wine grapes from several Australian vineyards. J Appl Microbiol 100:712–727
Bester L, Cameron M, du Toit M, Witthuhn RC (2010) PCR and DGGE detection limits for wine spoilage microbes. S Afr J Enol Vitic 31:26–33
Case RJ, Boucher Y, Dahllöf I, Holmström C, Doolittle WF, Kjelleberg S (2007) Use of 16S rRNA and rpoB genes as molecular markers for microbial ecology studies. Appl Environ Microbiol 73:278–288. doi:10.1128/AEM.01177-06
Cocolin L, Mills D (2003) Wine yeast inhibition by sulphur dioxide: a comparison of culture-dependent and independent methods. Am J Enol Vitic 54:125–130
Cocolin L, Campolongo S, Alessandria V, Dolci P, Rantsiou K (2011a) Culture independent analyses and wine fermentation: an overview of achievements 10 years after first application. Ann Microbiol 61:17–23. doi:10.1007/s13213-010-0076-6
Cocolin L, Dolci P, Rantsiou K (2011b) Biodiversity and dynamics of meat fermentations: the contribution of molecular methods for a better comprehension of a complex ecosystem. Meat Sci 89:296–302. doi:10.1016/j.meatsci.2011.04.011
de Revel G, Martin N, Pripis-Nicolau L, Lonvaud-Funel A, Bertrand A (1999) Contribution to the knowledge of malolactic fermentation influence on wine aroma. J Agric Food Chem 47:4003–4008
Dicks LMT, Endo A (2009) Taxonomic status of lactic acid bacteria in wine and key characteristics to differentiate species. S Afr J Enol Vitic 30:72–90
Divol B, Lonvaud-Funel A (2005) Evidence for viable but nonculturable yeasts in Botrytis-affected wine. J Appl Microbiol 99:85–93. doi:10.1111/j.1365-2672.2005.02578.x
Dobson CM, Deneer H, Lee S, Hemmingsen S, Glaze S, Ziola B (2002) Phylogenetic analysis of the genus Pediococcus, including Pediococcus claussenii sp. nov., a novel lactic acid bacterium isolated from beer. Int J Syst Evol Microbiol 52:2003–2010. doi:10.1099/ijs.0.02191-0
Dols-Lafargue M, Lee HY, Le Marrec C, Heyraud A, Chambat G, Lonvaud-Funel A (2008) Characterization of gtf, a glucosyltransferase gene in the genomes of Pediococcus parvulus and Oenococcus oeni, two bacterial species commonly found in wine. Appl Environ Microbiol 74:4079–4090. doi:10.1128/AEM.00673-08
du Toit M, Engelbrecht L, Lerm E, Krieger-Weber S (2011) Lactobacillus: the next generation of malolactic fermentation starter cultures—an overview. Food Bioproc Technol 4:876–906. doi:10.1007/s11947-010-0448-8
Endo A, Okada S (2006) Oenococcus kitaharae sp. nov., a non-acidophilic and non-malolactic-fermenting Oenococcus isolated from a composting distilled shochu residue. Int J Syst Evol Microbiol 56:2345–2348. doi:10.1099/ijs.0.64288-0
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791. doi:10.2307/2408678
González-Arenzana L, López R, Santamaría P, Garijo P, Gutiérrez AR, López-Alfaro I, Tenorio C (2010) Comparación de distintos métodos de extracción directa de ADN de vino tinto para el estudio de bacterias lácticas. VI Foro Mundial del Vino, Logroño
González-Arenzana L, Santamaría P, López R, Tenorio C, López-Alfaro I (2012a) Ecology of indigenous lactic acid bacteria along different winemaking processes of Tempranillo red wine from La Rioja (Spain). Sci World J. doi:10.1100/2012/796327
González-Arenzana L, López R, Santamaría P, Tenorio C, López-Alfaro I (2012b) Dynamics of indigenous lactic acid bacteria populations in wine fermentations from La Rioja (Spain) during three vintages. Microb Ecol 62:1–8. doi:10.1007/s00248-011-9911-y
González-Arenzana L, Santamaría P, López R, López-Alfaro I (2013) Indigenous lactic acid bacteria communities in alcoholic and malolactic fermentations of Tempranillo wines elaborated in ten wineries of La Rioja (Spain). Food Res Int 50:538–545. doi:10.1016/j.foodres.2012.11.008
Kim JY, Kim D, Park P, Kang H, Ryu EK, Kim SM (2011) Effects of storage temperature and time on the biogenic amine content and microflora in Korean turbid rice wine, Makgeolli. Food Chem 128:87–92. doi:10.1016/j.foodchem.2011.02.081
Liu SQ, Pritchard GG, Hardman MJ, Pilone GJ (1994) Citrulline production and ethyl carbamate (urethane) precursor formation from arginine degradation by wine lactic acid bacteria Leuconostoc oenos and Lactobacillus buchneri. Am J Enol Vitic 45:235–242
Lonvaud-Funel A (1999) Lactic acid bacteria in the quality improvement and depreciation of wine. Anton Leeuw Int J Gen Mol Microbiol 76:317–331
Lonvaud-Funel A (2008) From raisin to wine: activity of a dynamic microbial system. Biofutur 26–29
López I, Ruiz-Larrea F, Cocolin L, Orr E, Phister T, Marshall M, VanderGheynst J, Mills DA (2003) Design and evaluation of PCR primers for analysis of bacterial populations in wine by denaturing gradient gel electrophoresis. Appl Environ Microbiol 69:6801–6807. doi:10.1128/AEM.69.11.6801-6807.2003
López I, López R, Santamaría P, Torres C, Ruiz-Larrea F (2008) Performance of malolactic fermentation by inoculation of selected Lactobacillus plantarum and Oenococcus oeni strains isolated from Rioja red wines. Vitis 47:123–129
López-Alfaro I (2004) Detección y Control por Técnicas de la Biología Molecular de Bacterias Lácticas Autóctonas Responsables de la Fermentación maloláctica en Vinos de D.O.Ca. Rioja. Universidad de La Rioja, Logroño
Lucena BTL, dos Santos BM, Moreira JLS, Moreira APB, Nunes AC, Azevedo V, Miyoshi A, Thompson FL, de Morais Junior MA (2010) Diversity of lactic acid bacteria of the bioethanol process. BMC Microbiol 10:298. doi:10.1186/1471-2180-10-298
Lucore L, Cullison M, Jaykus L (2000) Immobilization with metal hydroxides as a means to concentrate food-borne bacteria for detection by cultural and molecular methods. Appl Environ Microbiol 66:1769–1776. doi:10.1128/AEM.66.5.1769-1776.2000
Meroth CB, Walter J, Hertel C, Brandt MJ, Hammes WP (2003) Monitoring the bacterial population dynamics in sourdough fermentation processes by using PCR-denaturing gradient gel electrophoresis. Appl Environ Microbiol 69:475–482. doi:10.1128/AEM.69.1.475-482.2003
Mesas JM, Rodríguez MC, Alegre MT (2011) Characterization of lactic acid bacteria from musts and wines of three consecutive vintages of Ribeira Sacra. Lett Appl Microbiol 52:258–268. doi:10.1111/j.1472-765X.2010.02991.x
Millet V, Lonvaud-Funel A (2000) The viable but non-culturable state of wine micro-organisms during storage. Lett Appl Microbiol 30:136–141
Mira de Orduña R, Patchett M, Liu S, Pilone G (2001) Growth and arginine metabolism of the wine lactic acid bacteria Lactobacillus buchneri and Oenococcus oeni at different pH values and arginine concentrations RID B-9010-2009. Appl Environ Microbiol 67:1657–1662. doi:10.1128/AEM.67.4.1657-1662.2001
Muyzer G, Smalla K (1998) Application of denaturing gradient gel electrophoresis (DGGE) and temperature gradient gel electrophoresis (TGGE) in microbial ecology RID H-4002-2011. Anton Leeuw Int J Gen Mol Microbiol 73:127–141. doi:10.1023/A:1000669317571
Ogier J, Casalta E, Farrokh C, Saihi A (2008) Safety assessment of dairy microorganisms: the Leuconostoc genus. Int J Food Microbiol 126:286–290. doi:10.1016/j.ijfoodmicro.2007.08.012
Pérez-Pulido R, Abriouel H, Ben Omar N, Lucas R, Martínez-Canamero M, Galvez A (2006) Safety and potential risks of enterococci isolated from traditional fermented capers. Food Chem Toxicol 44:2070–2077. doi:10.1016/j.fct.2006.07.008
Phister TG, Rawsthorne H, Joseph CML, Mills DA (2007) Real-time PCR assay for detection and enumeration of Hanseniaspora species from wine and juice RID G-2282-2011. Am J Enol Vitic 58:229–233
Rantsiou K, Urso R, Dolci P, Comi G, Cocolin L (2008) Microflora of Feta cheese from four Greek manufacturers. Int J Food Microbiol 126:36–42. doi:10.1016/j.ijfoodmicro.2008.04.031
Reguant C, Carreté R, Constanti M, Bordons A (2005) Population dynamics of Oenococcus oeni strains in a new winery and the effect of SO2 and yeast strain. FEMS Microbiol Lett 246:111–117. doi:10.1016/j.femsle.2005.03.045
Renouf V, Favier M (2010) Genetic and physiological characterisation of Oenococcus oeni strains to perform malolactic fermentation in wines. S Afr J Enol Vitic 31:75–81
Renouf V, Claisse O, Lonvaud-Funel A (2005a) Understanding the microbial ecosystem on the grape berry surface through numeration and identification of yeast and bacteria. Aust J Grape Wine Res 11:316–327
Renouf V, Gindreau E, Claisse O, Lonvaud-Funel A (2005b) Microbial changes during malolactic fermentation in red wine elaboration. J Int Des Sci De La Vigne Et Du Vin 39:179–190
Renouf V, Claisse O, Lonvaud-Funel A (2006a) RpoB gene: a target for identification of LAB cocci by PCR-DGGE and melting curves analyses in real time PCR. J Microbiol Methods 67:162–170. doi:10.1016/j.mimet.2006.03.008
Renouf V, Claisse O, Miot-Sertier C, Lonvaud-Funel A (2006b) Lactic acid bacteria evolution during winemaking: use of rpoB gene as a target for PCR-DGGE analysis. Food Microbiol 23:136–145. doi:10.1016/j.fm.2005.01.019
Renouf V, Vayssieres LC, Claisse O, Lonvaud-Funel A (2009) Genetic and phenotypic evidence for two groups of Oenococcus oeni strains and their prevalence during winemaking. Appl Microbiol Biotechnol 83:85–97. doi:10.1007/s00253-008-1843-1
Ribéreau-Gayon P, Dubourdieu D, Donèche B, Lonvaud A (2007) Handbook of enology. Wiley, England
Ruiz P, Izquierdo PM, Seseña S, Palop ML (2010) Analysis of lactic acid bacteria populations during spontaneous malolactic fermentation of Tempranillo wines at five wineries during two consecutive vintages. Food Control 21:70–75. doi:10.1016/j.foodcont.2009.04.002
Ruiz-Larrea F, López-Alfaro I, Alegría E, Zarazaga M, Torres C (2001) Aspectos prácticos de la fermentación malolácica. Gobierno de La Rioja, Logroño
Saitou N, Nei M (1987) The neighbor-joining method—a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics (MEGA) software version 4.0 RID E-9283-2010. Mol Biol Evol 24:1596–1599. doi:10.1093/molbev/msm092
Torriani S, Felis GE, Fracchetti F (2011) Selection criteria and tools for malolactic starters development: an update. Ann Microbiol 61:33–39. doi:10.1007/s13213-010-0072-x
Vanvuuren HJJ, Dicks LMT (1993) Leuconostoc oenos—a review. Am J Enol Vitic 44:99–112
Zhang Z, Ye Z, Yu L, Shi P (2011) Phylogenomic reconstruction of lactic acid bacteria: an update. BMC Evol Biol 11:1. doi:10.1186/1471-2148-11-1
Acknowledgments
This work was supported by funding and predoctoral grant (BOR 6 March 2009) from the Government of La Rioja, the INIA project RTA2007-00104-00-00 and the FEDER of the European Community and was made possible by the collaborating wineries.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
González-Arenzana, L., López, R., Santamaría, P. et al. Dynamics of lactic acid bacteria populations in Rioja wines by PCR-DGGE, comparison with culture-dependent methods. Appl Microbiol Biotechnol 97, 6931–6941 (2013). https://doi.org/10.1007/s00253-013-4974-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-013-4974-y