Abstract
The budC gene coding for a new meso-2,3-butanediol dehydrogenase (BDH) from Serratia marcescens H30 was cloned and expressed in Escherichia coli BL21(DE3), purified, and characterized for its properties. The recombinant BDH with a molecular weight of 27.4 kDa exhibited a reversible transformation between acetoin and 2,3-butanediol. In the presence of NADH, BDH could catalyze the reduction of diacetyl and (3R)-acetoin to (3S)-acetoin and meso-2,3-butanediol, respectively, while (3S)-acetoin as a substrate could be further transformed into (2S, 3S)-2,3-butanediol at pH 9.0. For diol oxidation reactions, (3R)-acetoin and (3S)-acetoin were obtained when meso-2,3-butanediol and (2S,3S)-2,3-butanediol were used as the substrates with BDH and NAD+. (2R,3R)-2,3-butanediol was not a substrate for the BDH at all. The low K m value (4.1 mM) in meso-2,3-butanediol oxidation reaction and no activity for diacetyl, acetoin, and 2,3-butanediol as the substrates with NADP+/NADPH suggested that the budC gene product belongs to a NAD(H)-dependent meso-2,3-BDH. Maximum activities for diacetyl and (3S/3R)-acetoin reduction were observed at pH 8.0 and pH 5.0 while for meso-2,3-butanediol oxidation it was pH 8.0. However, the optimum temperature for oxidation and reduction reactions was about 40 °C. In addition, the BDH activity for meso-2,3-butanediol oxidation was enhanced in the presence of Fe2+ and for diacetyl and (3S/3R)-acetoin reduction in the presence of Mg2+ and Mn2+, while several metal ions inhibited its activity, particularly Fe3+ for reduction of diacetyl and acetoin. Sequence analysis showed that the BDH from S. marcescens H30 possessed two conserved sequences including the coenzyme binding motif (GxxxGxG) and the active-site motif (YxxxK), which are present in the short-chain dehydrogenase/reductase superfamily.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
2,3-Butanediol (2,3-BD) and its dehydrogenation product, acetoin (AC), are important platform compounds due to its potential industrial applications (Ji et al. 2009; Liu et al. 2011). 2,3-BD contains two stereo centers and has three stereo isomers including (2R,3R)-, meso- and (2S,3S)-forms, and AC exists in two stereoisomeric forms: (3R)-acetoin and (3S)-acetoin, which are important potential pharmaceutical intermediates (Yan et al. 2009; Ji et al. 2011). All of the isomers of 2,3-BD and AC could be produced by the homologous pathways involving pyruvate in a mixed acid fermentation process in several native strains such as Klebsiella pneumoniae (Borim et al. 2012), Klebsiella oxytoca (Yu and Saddler 1983; Han et al. 2013), Enterobacter aerogenes (Zeng et al. 1990), Enterobacter cloacae (Li et al. 2011), Bacillus polymyxa (De Mas et al. 1987), and Serratia marcescens (Zhang et al. 2010; Sun et al. 2012; Rao et al. 2012). α-Acetolactate, diacetyl, and acetoin are the main three intermediate compounds of 2,3-BD fermentation in bacteria (Celinska and Grajek 2009; Zhang et al. 2012). In general, the stereoisomeric composition of 2,3-BD and AC formed by bacteria differs among strains (Table 1), a phenomenon that is related to the existence of various 2,3-butanediol dehydrogenases (BDHs) differing in their stereospecificities (Ui et al. 1998).
Several BDHs from different strains have been characterized in previous studies (Table 2). In summary, BDHs can be divided into three classes: meso-BDH, (2S,3S)-BDH and (2R,3R)-BDH. According to the configuration of the AC product from 2,3-BD as a substrate, BDHs also were classified into two types: D-(−)-Acetoin forming and L-(+)-Acetoin forming. meso-BDHs in K. pneumoniae CICC10011 and Enterobacter sp. 638 could catalyze the conversion of (3R)-AC to meso-2,3-BD and (3S)-AC to (2S,3S)-2,3-BD in the presence of NADH. Meanwhile, meso- and (2S,3S)-2,3-BD as the substrates could also be transformed into (3R)- and (3S)-AC, respectively, with the BDHs and NAD+ (Liu et al. 2011; Li et al. 2011). While (2S,3S)-BDH from Brevibacterium saccharolyticum C-1012 was found to possess substrate specificity in the interconversion between (3S)-AC and (2S,3S)-2,3-BD (Ui et al. 1998; Takusagawa et al. 2001), which has not yet been reported in other strains. Recently, three (2R,3R)-BDHs from Paenibacillus polymyxa ATCC12321, Bacillus subtilis 168, and Saccharomyces cerevisiae were identified, and analysis of their sequences revealed that the three enzymes belonged to the medium-chain dehydrogenase/reductase superfamily. The three BDHs showed the abilities in the interconversion of (3S)-AC/meso-2,3-BD and (3R)-AC/(2R,3R)-2,3-BD (Gonzalez et al. 2000; Xiao et al. 2010; Yu et al. 2011; Liu et al. 2011). During the 2,3-BD fermentation process, (3R)-AC is the decarboxylation product of α-acetolactate catalyzed by α-acetolactate decarboxylase (Xiao and Xu 2007). The precursor of (3S)-AC is diacetyl (DA), a byproduct of 2,3-BD fermentation produced by the non-enzymatic oxidation of α-acetolactate (Nicholson 2008). Due to the low concentration of DA in microbial fermentation, (3S)-AC can only be limited at low concentrations (Liu et al. 2011).
In our previous study, we reported a S. marcescens strain H30 possessing high productivity and yield for AC and 2,3-BD production (Zhang et al. 2010; Sun et al. 2012). During the fermentation process, the strain H30 exhibited broad substrate spectrum, cultural adaptability, and better resistance to bacteria contamination due to prodigiosin production. Therefore, it is considered as a promising producer with high industrial potential (Zhang et al. 2010; Ji et al. 2011). Interestingly, unlike other 2,3-BD producing Gram-negative strains, this strain mainly produced meso-2,3-BD (a weight fraction of over 98 %) with a little (2S,3S)-2,3-BD (Zhang et al. 2010). To understand the characteristics of BDH from S. marcescens H30 and conduct a study on fermentation, the BDH gene (budC) was cloned, and the encoded amino acid sequence was compared with those of other BDHs reported previously. The enzyme was characterized according to the optimal pH, temperature, and substrate stereospecificity after the expression and purification.
Materials and methods
Strains and bacterial growth condition
S. marcescens H30 (deposited in the China Center for Industrial Culture Collection, accession number: CICC 20066) used as the source of budC gene was grown at 30 °C. Escherichia coli DH5α and BL21(DE3) as the cloning and expression hosts were grown at 37 °C. Plasmid pET-28a (+) (Novagen) was used for the BDH expression. All strains were cultured in Luria–Bertani (LB) medium. Antibiotics were added in the following amounts (per milliliter) if necessary: 50 μg kanamycin.
Reagents, primers, and genomic isolation
Restriction enzyme, T4 DNA ligase, and Taq DNA polymerase high fidelity were from TaKaRa Biotech (Dalian, China). DNA and protein marker were purchased from Tiangen Biotech (Shanghai, China). Oligonucleotied primers were systhesized in SBSbio (Shanghai, China). Genomic DNA from S. marcescens H30, which were used as template for PCR amplification, was prepared with the Bacterial Genomic DNA Mini-prep Kit (BIODEV Corp., Beijing, China). (3S/3R)-AC, (2S,3S)-2,3-BD (97.0 %), (2R,3R)-2,3-BD (97.0 %), meso-2,3-BD (99.0 %), and other chemicals, unless otherwise indicated, were obtained from Sigma-Aldrich (Shanghai, China).
Development of the budC recombinant strain
The encoding sequence of budC gene was amplified by PCR with the genomic DNA of S. marcescens H30 as template using the following primers: budC-F (5′-TCC GAA TTC ATG CGT TTT GAC AAT AAA G-3′) and budC-R (5′-GAC AAG CTT TTA GAC GAT CTT CGG TTG G-3′). Amplification was carried out in a TaKaRa PCR thermal cycler with the parameter settings as follows: 5 min at 94 °C for predenaturation, 30 s at 94 °C for denaturation, 30 s at 55 °C for annealing and 50 s at 72 °C for elongation. After 30 cycles of amplification, an additional elongation period for 10 min at 72 °C was used to ensure the completeness of the products. The amplified product was ligated into the vector pET-28a (+) at EcoRI and HindIII sites, resulting in the recombinant plasmid designated as pET28a-budC. The recombinant E. coli BL21(DE3)/pET28a-budC was obtained using heat shock transformation.
Preparation and purification of recombinant BDH
The recombinant strain was cultured at 37 °C in a 250-ml flask containing 50 ml LB medium (pH 7.0) with kanamycin (50 μg/ml). The cells were induced at about 0.6 OD600 with 0.5 mM isopropyl-beta-d-thiogalactopyranoside (IPTG) and harvested by centrifugation after 6 h. The precipitate was resuspended in binding buffer (20 mM phosphate, 500 mM NaCl, and 20 mM imidazole, pH 7.4) and disrupted by sonication in an ice bath. The homogenate was centrifuged at 13,000×g for 10 min to remove the debris. The soluble fraction was subjected to purification under nondenaturing conditions with Ni-affinity chromatography using a Histrap HP column according the purification protocol (GE Healthcare, USA). The eluate from the column was pooled and desalted by a Hitrap desalting column (GE Healthcare, USA). To avoid the effect of His6-tag on the properties of BDH enzyme, the His6-tag of the purified enzyme was cut off using the THROMBIN Kit (Jianglaibio, Shanghai). The obtained enzyme was analyzed via sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE).
Enzyme assays
Enzyme activity was determined spectrophotometrically by measuring the changes in absorbance at 340 nm and 40 °C corresponding to the oxidation of NADH or the reduction of NAD+. The reaction mixtures containing 50 mM potassium phosphate buffer (pH 8.0), 4 mM NAD+ for the oxidation reactions or 50 mM sodium acetate buffer (pH 5.0), 0.2 mM NADH for the reduction reactions were incubated at 40 °C for 5 min. After adding 10 μl of enzyme solution, the reaction was started by the addition of the substrates. One unit of BDH activity was defined as the amount of enzyme required to reduce 1 μmol of NAD(H) in 1 min. All enzyme activities were determined in triplicate.
Stereospecificity of BDH
Stereospecificity of the purified BDH enzyme was investigated in the oxidation–reduction processes of the 2,3-BD/AC/DA interconversion. For the oxidation processes, a mixture containing 100 mM meso-2,3-BD/(2S,3S)-2,3-BD, 4 mM NAD+, 50 mM of potassium phosphate buffer (pH 8.0) and 10 μl of purified BDH enzyme in a final volume of 1 ml was incubated at 40 °C for 2 h. The reduction processes was carried out in 1-ml reaction system containing 100 mM DA /(3S/3R)-AC, 0.2 mM NADH, 50 mM of sodium acetate buffer (pH 5.0) and 10 μl of purified BDH enzyme at 40 °C for 2 h. The products in these reaction systems were extracted by ethyl acetate and then used to check the enzyme stereospecificity using a GC system (Agilent GC9860) equipped with a chiral column (Supelco β-DEX™ 120, 30 m length, 0.25 mm inner diameter). The operation conditions were as follows: N2 was used as the carrier gas at flow rate of 1.2 ml/min; the injector temperature and the detector temperature were 215 and 245 °C, respectively; and the column temperature was maintained at 50 °C for 1.5 min, then raised to 180 °C at a rate of 15 °C/min.
Results
Cloning and sequence analysis of the budC gene
The budC gene was obtained by PCR method using the primers (budC-F/budC-R) designed according to our previous submitted sequence (Genbank accession number AFH00999). The amino acid sequence as deduced from the nucleotide sequence of 756 bp was compared with other reported BDHs by multiple alignment (Table 3). The BDH showed low identity with other known functional BDHs except that from S. marcescens MG1 with 98 % identity. Using InterProScan web server (http://www.ebi.ac.uk/Tools/pfa/iprscan/) to search the conserved domain of the BDH enzyme, two conserved sequences for the coenzyme binding motif (GxxxGxG) in the N-terminal and the active site motif (YxxxK) in the C-terminal were found (Fig. 1), which are present in the short-chain dehydrogenase/reductase (SDR) superfamily (Jornvall et al. 1995; Kallberg et al. 2002). So far, almost all the meso-BDHs and (2S,3S)-BDH characterized belong to the SDR superfamily (Yu et al. 2011). Such observations indicated that the budC gene product from S. marcescens H30 may be meso-BDH or (2S,3S)-BDH belonging to the SDR superfamily.
Expression and purification of BDH enzyme
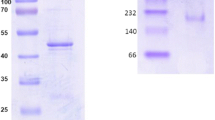
The recombinant E. coli BL21(DE3)/pET28a-budC was induced with 0.5 mM IPTG at 30 °C when OD600 reached 0.6. SDS-PAGE analysis on the soluble fractions from the cell lysate revealed the presence of over-expressed protein (Fig. 2), implying that the target protein was successfully expressed.
Expression and purification of the BDH protein: M protein marker (200, 116, 97.2, 66.4, 44.3, 29, 20.1, 14.3, 6.5 kDa); 1 purified His6-tag-free BDH protein; 2 purified His6-tag BDH protein; 3 soluble protein from the cell lysate, were electrophoresed on a 12 % polyacrylamide gel under denaturing conditions
The purified His6-tag BDH and His6-tag-free BDH were given in Fig. 2. The BDH activity was determined using two buffer systems for the oxidation-reduction reactions. For the oxidation reaction, the reaction mixtures contained 50 mM potassium phosphate buffer (pH 8.0), 4 mM NAD+ and 100 mM meso-2,3-BD, while in the reduction reactions 50 mM sodium acetate buffer (pH 5.0), 0.2 mM NADH and 100 mM (3S/3R)-AC were employed. The results showed that the highest activities were determined as 215 U/mg for reduction of (3S/3R)-AC and 67 U/mg for oxidation of meso-2,3-BD. After purification, the BDH was purified 1.9-fold during the purification process.
Specificity of BDH for substrates and coenzymes
The substrate specificity of BDH enzyme in the oxidation reactions was studied using 100 mM diols with 2 different coenzymes, NAD+ and NADP+. Reduction activities were measured with 100 mM ketones in the presence of NADH and NADPH as coenzymes. As shown in Table 4, the recombinant BDH could oxidize meso-2,3-BD and (2S,3S)-2,3-BD, while (2R,3R)-2,3-BD was not a substrate for the recombinant BDH at all. In addition, several primary alcohols (such as glycerol, 1,2-pentanediol and 1,2-propanediol) could also be oxidized, though showing less activity than meso-2,3-BD. (3S/3R)-AC was the best substrate in the ketone reduction reaction, followed by diacetyl which had 75 % of the specific activity of (3S/3R)-AC. No activity (data not shown) was detected for DA, AC and 2,3-BD as the substrates with NADPH/NADP+, indicating that the budC gene product belongs to a NAD(H)-dependent 2,3-BDH.
The comparative data of apparent K m and K cat values for BDH from S. marcescens H30 were given in Table 4. The K m and K cat values of BDH were 4.1 mM and 6.2 s−1 for meso-2,3-BD, 31.2 mM and 1.02 s−1 for (2S,3S)-2,3-BD, 0.97 mM, and 19.7 s−1 for (3S/3R)-AC, and 3.3 mM and 11.5 s−1 for diacetyl, respectively. The K cat/K m ratio for meso-2,3-BD is much higher than that for (2S,3S)-2,3-BD, and combined with the results from substrate specificity test, BDH from S. marcescens H30, therefore, could be categorized as a NAD(H)-dependent meso-2,3-BDH.
Effects of temperature and pH on BDH activity
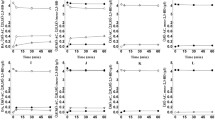
The effects of temperature and pH on BDH activity were determined using 100 mM DA, 100 mM (3S/3R)-AC, and 100 mM meso-2,3-BD as substrates and NAD(H) as coenzymes. The pH effects were determined in the range of pH 4–10 at 40 °C using 50 mM sodium acetate (pH 4–5), potassium phosphate (pH 6–8), and glycine–NaOH buffers (pH 9–10). As shown in Fig. 3a, maximun activity for (3S/3R)-AC and DA reduction was observed at pH 5.0 and pH 8.0, while it was pH 8.0 for the meso-2,3-BD oxidation reaction.
Temperature effects were studied in the range of 20–70 °C for meso-2,3-BD oxidation and DA reduction at pH 8.0, and for (3S/3R)-AC reduction at pH 5.0 (Fig. 3b). The effects of temperature on the BDH activity appeared to be similar in both reduction and oxidation reactions, and the optimal temperature of 40 °C for these three substrates was determined. More than 50 % of maximum activity was retained between 20 and 50 °C, but the recombinant BDH became unstable at 60 and 70 °C, most likely due to thermal denaturation.
Effects of metal ions on BDH activity
The effects of different metal ions (NH4 +, Na+, Mn2+, Mg2+, Zn2+, Ca2+, Fe2+, and Fe3+), all in the form of chloride/sulfate salts, on the BDH activity were evaluate at 1 mM. As shown in Table 5, the BDH activity in the oxidation and reduction processes could be inhibited by most of metal ions such as NH4 +, Na+, Zn2+, Ca2+, and Fe3+. Especially Fe3+ exhibited strong inhibition with less than 20 % of maximum activity for the DA and (3S/3R)-AC reduction. In contrast, Mg2+ and Mn2+ could slightly activate and increase its activity (<10 %). For meso-2,3-BD oxidation, the activity of BDH could obviously be enhanced by 27 % in the presence of Fe2+, which resulted in the loss of 64 and 52 % with DA and (3S/3R)-AC as substrates and NADH as coenzyme.
Stereospecificity of the BDH
Figure 4 demonstrated the BDH stereospecificity in the oxidation-reduction processes of the 2,3-BD/AC/DA interconversion. With respect to 2,3-BD oxidation reactions, when meso-2,3-BD was used as the substrate with NAD+ and BDH, (3R)-AC was the only product detected (Fig. 4b). Accordingly, (3S)-AC was, as expected, the only product obtained from (2S,3S)-2,3-BD (Fig. 4c). And more, (3S)-AC could be obtained from DA by BDH (Fig. 4d). Unexpectedly, (3S)-AC could not further be transformed into any form of 2,3-BD at pH 5.0 (Fig. 4d), which is different from other meso-2,3-BDHs reported previously. To investigate the reason, the DA reduction reactions were carried out with different buffer systems from pH 5.0 to pH 9.0. The results (Fig. 5) showed that only (3S)-AC was obtained from DA when the pH value was below 8.0, while the (3S)-AC product from DA could be further transformed into (2S,3S)-2,3-BD at pH 9.0. When a racemic mixture of (3S/3R)-AC was incubated with BDH and NADH, only meso-2,3-BD was formed (Fig. 4e). Considering that (2S,3S)-2,3-BD is the only product from (3S)-AC, therefore, meso-2,3-BD was produced from (3R)-AC as a substrate.
Chiral-column GC analysis of the substrates and products in the oxidation and reduction reaction catalyzed by meso-2,3-butanediol dehydrogenase from S. marcescens H30. a Profile of mixture of standard chemicals. b The product from meso-2,3-butanediol. c The product from (2S,3S)-2,3-butanediol. d The product from diacetyl. e The product from (3S/3R)-acetoin (the peak of diacetyl was in front of solvent peak and not shown in the picture)
Chiral-column GC analysis of the substrate and products in the DA reduction reaction catalyzed by meso-2,3-butanediol dehydrogenase from S. marcescens H30 with different pH system. a Profile of mixture of standard chemicals. b pH 5.0, c pH 6.0, d pH 7.0, e pH 8.0, f pH 9.0. IA was the abbreviation of isoamyl alcohol, which was used as internal standard for GC analysis
Discussion
Single configuration production of AC and 2,3-BD was recently paid increasing attention due to their potential industrial and pharmaceutical applications (Ji et al. 2011). However, natural microorganism usually produced a mixture of (2S,3S)-2,3-BD/meso-2,3-BD or (2R,3R)-2,3-BD/meso-2,3-BD with (3S/3R)-AC, such as (2S,3S)-2,3-BD/meso-2,3-BD with (3S/3R)-AC were formed by K. pneumoniae (Liu et al. 2011), and P. polymyxa could produce (2R,3R)-2,3-BD and (3S/3R)-AC with a little meso-2,3-BD (Ji et al. 2011; Yu et al. 2011). Hence, it is very difficult that single configuration of AC and 2,3-BD in the direct fermentative process was achieved by natural strains. There are several possible explanations for the mixed formation of AC and 2,3-BD isomers, including aeration conditions (redox balance), non-stereospecific dehydrogenases, multiple pathways, and multiple stereospecific dehydrogenases (Syu 2011; Yan et al. 2009). In previous studies, several 2,3-butanediol dehydrogenases from different strains were characterized, exhibiting the variation of their stereospecificities and potential applications. Ui et al. introduced gene fragments containing genes encoding acetolactate synthase (ALS), acetolactate decarboxylase (ALDC), and a single meso-2,3-BDH from K. pneumoniae IAM 1063 into E. coli, and obtained production of pure meso-2,3-BD (Ui et al. 1997). While whole cells of engineered E. coli with overexpression of (2S,3S)-2,3-BDH from B. saccharolyticum C-1012 were developed for biocatalytic production of pure (3S)-AC and (3S,3S)-2,3-BD from DA (Ui et al. 2004). Pure (2R,3R)-2,3-BD has been achieved by introducing three genes encoding ALS, ALDC, and a (2R,3R)-2,3-BDH from B. subtilis into E. coli JCL16 and JCL 260 (Yan et al. 2009). These studies indicated that mixed formation of AC and 2,3-BD isomers is mainly due to the existence of multiple pathways or dehydrogenases.
In this study, the budC gene encoding the BDH from S. marcescens H30 was cloned and expressed in E. coli BL21(DE3), purified and characterized for its properties. Amino acid sequence alignment showed the BDH shared low identity (<35 %) with other reported BDHs except that with a 98 % identity from S. marcescens MG1, implying that the enzyme from S. marcescens is a new BDH. However, the BDH similar to that from K. pneumoniae and B. saccharolyticum possessed two conserved residues: coenzyme binding motif (GxxxGxG) in the N-terminal part and substrate binding region (YxxxK) in the C-terminal. So the BDH from this study should also belong to the SDR superfamily. In the oxidation and reduction reactions, no activity could be detected with NADP(H) as coenzyme, NAD(H) was the only coenzyme for the BDH, indicating that the budC gene product is a NAD(H)-specific BDH. The maximum activity and low K m value for meso-2,3-BD in the oxidation process were determined, and combined with the results from substrate specificity test, therefore, the BDH from S. marcescens H30 could be categorized as a NAD(H)-dependent meso-2,3-BDH.
The BDH optimal values of pH 5 and pH 8 determined with (3S/3R)-AC and meso-2,3-BD as substrates demonstrated that pH value played an important role between oxidation and reduction reactions. The BDH activity could also be influenced by several metal ions. Fe2+ could increase its activity by 27 % with meso-2,3-BD as a substrate. In contrast, Fe2+ exhibited inhibition activity in DA and (3S/3R)-AC reduction process and resulted in the loss of 64 and 52 %, respectively. In addition, Mg2+ and Mn2+ could slightly improve the 2,3-BDH activity in the DA and (3S/3R)-AC reduction, while its activity could be strongly inhibited (>90 %) by Fe3+ with (3S/3R)-AC as a substrate. Hence, the ratio between AC and 2,3-BD by S. marcescens H30 during the fermentation process could be controlled by the fermentative pH value adjusted and metal ions added. These results would provide further useful hints for fermentation strategy development of AC or 2,3-BD production.
The isomers formation mechanism of AC and 2,3-BD by native strains during the fermentation process was still elucidated unclearly as far. But stereospecificity of the BDH have been accepted as a key factor. Similar to the genus of Klebsiella and Enterobacter, the BDH from S. marcescens H30 exhibited the abilities in the interconversion of (3R)-AC/meso-2,3-BD and (3S)-AC/(2S,3S)-2,3-BD, and (3S)-AC was obtained from DA as a substrate by the BDH. The major difference was that (2S,3S)-2,3-BD produced from (3S)-AC was limited at the pH value of 9.0. When the pH value was below 8.0, (3S)-AC with the BDH enzyme and NADH as a coenzyme could not be further oxidized and transformed into (2S,3S)-2,3-BD. From another perspective, this characteristics of the BDH from S. marcescens H30 is very helpful for chiral (3S)-AC production from DA by whole cells catalysis with over-expression, since the pH adjustment could effectively prevent (3S)-AC from forming (2S,3S)-2,3-BD.
In conclusion, we reported a new NAD(H)-dependent meso-2,3-BDH from S. marcescens H30. And the meso-2,3-BDH was purified and characterized, exhibiting some different characteristics compared with others reported meso-2,3-BDH. These properties should lead to its potential application for chiral (3S)-AC production, and more suitable fermentation strategy for AC or 2,3-BD production by S. marcescens H30 in industry.
References
Borim K, Lee S, Park J, Lu M, Oh M, Kim Y, Lee J (2012) Enhanced 2,3-butanediol production in recombinant Klebsiella pneumoniae via overexpression of synthesis-related genes. J Microbiol Biotechnol 22(9):1258–1263
Celinska E, Grajek W (2009) Biotechnological production of 2,3-butanediol—current state and prospects. Biotechnol Adv 27:715–725
De Mas C, Jansen NB, Tsao GT (1987) Production of optically active 2,3-butanediol by Bacillus polymyxa. Biotechnol Bioeng 31:366–377
Gonzalez E, Rosario Fernanzdez M, Larroy C, Lluis S, Pericas MA (2000) Characterization of a (2R,3R)-2,3-butanediol dehydrogenase as the Saccharomyces cerevisiae YAL060W gene product. J Biol Chem 275(46):35876–35885
Han SH, Lee JE, Park K, Park YC (2013) Production of 2,3-butanediol by a low-acid producing Klebsiella oxytoca NBRF4. New Biotechnol 30(2):166–172
Ji XJ, Huang H, Du J, Zhu JG, Ren LJ, Hu N, Li S (2009) Enhanced 2,3-butanediol production by Klebsiella oxytoca using a two-stage agitation speed control strategy. Bioresoure Technol 100:3410–3414
Ji XJ, Huang H, Ouyang PK (2011) Microbial 2,3-butanediol production: a state-of-the-art review. Biotechnol Adv 29:351–364
Jornvall H, Persson B, Krook M, Atrian S, Gonzalez-Duarte R, Jeffery J, Ghosh D (1995) Short-chain dehydrogenase/reductases (SDR). Biochemistry 34:6003–6013
Kallberg Y, Oppermann U, Jornvall H, Persson B (2002) Short-chain dehydrogenase/reductases (SDRs): coenzyme-based functional assignments in completed genomes. Eur J Biochem 269:4409–4417
Li LX, Wang Y, Zhang LJ, Ma CQ, Wang AL, Tao F, Xu P (2011) Biocatalytic production of (2S,3S)-2,3-butanediol from diacetyl using whole cells of engineered Escherichia coli. Bioresource Technol 115:111–116
Liu Z, Qin JY, Gao C, Hua DL, Ma CQ, Li LX, Wang Y, Xu P (2011) Production of (2S,3S)-2,3-butanediol and (3S)-acetoin from glucose using resting cells of Klebsiella pneumonia and Bacillus subtilis. Bioresource Technol 102:10741–10744
Nicholson WL (2008) The Bacillus subtilis ydjL (bdhA) gene encodes acetoin reductase/2,3-butanediol dehydrogenase. Appl Environ Microbiol 74(22):6832–6838
Rao B, Zhang LY, Sun JA, Su G, Wei DZ, Chu J, Zhu JW, Shen YL (2012) Characterization and regulation of the 2,3-butanediol pathway in Serratia marcescens. Appl Microbiol Biotechnol 93(5):2147–2159
Sun JA, Zhang LY, Rao B, Shen YL, Wei DZ (2012) Enhanced acetoin production by Serratia marcescens H32 with expression of a water-forming NADH oxidase. Bioresource Technol 119:94–98
Syu MJ (2011) Biological production of 2,3-butanediol. Appl Microbiol Biotechnol 55:10–18
Takusagawa Y, Otagiri M, Ui S, Ohtsuki T, Mimura A, Ohkuma M, Kudo T (2001) Purification and characterization of l-2,3-butanediol dehydrogenase of Brevibacterium saccharolyticum C-1012 expressed in Escherichia coli. Biosci Biotechnol Biochem 65(8):1876–1878
Ui S, Masuda H, Muraki H (1983) Laboratory-scale production of 2,3-butanediol isomers (D(−)), L(+), and (meso) by bacterial fermentations. J Ferment Technol 61:253–259
Ui S, Okajima Y, Mimura A, Kanai H, Kudo T (1997) Molecular generation of an Escherichia coli strain producing only the meso-isomer of 2,3-butanediol. J Ferment Bioeng 84:185–189
Ui S, Otagiri M, Mimura A, Dohmae N, Takio K, Ohkuma M, Kudo T (1998) Cloning, expression and nucleotide sequence of the l-2,3-butanediol dehydrogenase gene from Brevibacterium saccharolyticum C-1012. J Ferment Bioeng 86(3):290–295
Ui S, Takusagawa Y, Sato T, Ohtsuki T, Mumura A, Ohkuma M, Kudo T (2004) Production of l-2,3-butanediol by a new pathway constructed in Escherichia coli. Lett Appl Microbiol 39:533–537
Xiao ZJ, Xu P (2007) Acetoin metabolism in bacteria. Crit Rev Microbiol 33:127–140
Xiao ZJ, Lv CJ, Gao C, Qin JY, Ma CQ, Liu Z, Liu PH, Li LX, Xu P (2010) A novel whole-cell biocatalyst with NAD+ regeneration for production of chiral chemicals. PLoS One 5(1):e8860
Yan YJ, Lee CC, Liao JC (2009) Enantioselective synthesis of pure (R, R)-2,3-butanediol in Escherichia coli with stereospecific secondary alcohol dehydrogenases. Org Biomol Chem 7:3914–3917
Yu B, Sun JB, Bommareddy RR, Song LF, Zeng AP (2011) Novel (2R,3R)-2,3-butanediol dehydrogenase from potential industrial strain Paenibacillus polymyxa ATCC 12321. Appl Environ Microbiol 77(12):4230–4233
Yu EKC, Saddler JN (1983) Fed-batch approach to production of 2,3-butanediol by Klebsiella pneumoniae grown on high substrate concentrations. Appl Environ Microbiol 46:630–635
Zeng AP, Biebl H, Deckwer WD (1990) Effect of pH and acetic acid on the growth and 2,3-butanediol of Enterobacter aerogenes in continuous culture. Appl Microbiol Biotechnol 33(5):485–489
Zhang GL, Wang CW, Li C (2012) Cloning, expression and characterization of meso-2,3-butanediol dehydrogenase from Klebsiella pneumoniae. Biotechnol Lett 34:1519–1523
Zhang LY, Yang YL, Sun JA, Shen YL, Wei DZ, Zhu JW, Chu J (2010) Microbial production of 2,3-butanediol by a mutagenized strain of Serratia marcescens H30. Bioresource Technol 101:1961–1967
Acknowledgments
This work was supported by the Open Funding Project of the State Key Laboratory of Bioreactor Engineering, the Natural Science Foundation of Fujian Province of China (no. 2011J05048) and Science Fund of the Provincial Education Department of Fujian Province of China (no. JA11089).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Zhang, L., Xu, Q., Zhan, S. et al. A new NAD(H)-dependent meso-2,3-butanediol dehydrogenase from an industrially potential strain Serratia marcescens H30. Appl Microbiol Biotechnol 98, 1175–1184 (2014). https://doi.org/10.1007/s00253-013-4959-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-013-4959-x