Abstract
This study examined the effects of three lactic acid bacteria (LAB) strains on the nematode Caenorhabditis elegans. Lactobacillus salivarius, Lactobacillus reuteri, and Pediococcus acidilactici were found to inhibit the development and growth of the worm. Compared to Escherichia coli used as the control, L. reuteri and P. acidilactici reduced the lifespan of wild-type and short-lived daf-16 worms. On the contrary, L. salivarius extended the lifespan of daf-16 worms when used live, but reduced it as UV-killed bacteria. The three LAB induced the expression of genes involved in pathogen response and inhibited the growth of tumor-like germ cells, without affecting DAF16 localization or increasing corpse cells. Our results suggest the possible use of C. elegans as a model for studying the antitumor attributes of LAB. The negative effects of these LAB strains on the nematode also indicate their potential use against parasitic nematodes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Many strains of lactic acid bacteria (LAB) have been described and exploited commercially as probiotics (Gerritsen et al. 2011). The positive effects of these strains on animal and human health, that include improvement of digestion and nutrient uptake, increased resistance to pathogenic bacteria, antitumor effects, and increased longevity, have been the subject of many reviews (Gerritsen et al. 2011; Taverniti and Guglielmetti 2011; Ottaviani et al. 2011).
The mechanisms behind the probiotic effects of bacteria are not entirely understood. Probiotic bacteria have been shown to compete against other microorganisms that are found within the intestine and may be pathogenic. This is most likely achieved through the secretion of bacteriocins (Gerritsen et al. 2011). There is now evidence to suggest that the beneficial modulation of the intestinal microbiota could also protect against colorectal cancer (Candela et al. 2011). Production of hydrogen peroxide by LAB has been shown to be effective against food-borne pathogens (Nakajima et al. 2003). LAB have also been shown to stimulate local epithelial innate immune responses that prevent intestinal inflammation in a Crohn’s disease mouse model, as well as production of TNF-α and activation of NF-κB in vitro (Pagnini et al. 2010). In BALB/c mice fed LAB, a dose-dependent increase of the Bcl2 protein was observed, and an analysis of cytokine-producing cells in the lamina propria of the gut showed increased TNF-α and INF-γ values (Perdigón et al. 2002). Strain-specific transcriptional responses to LAB, involving the Toll-like receptor pathway, have also been reported using human intestinal epithelial cell cultures (Putaala et al. 2010).
Caenorhabditis elegans is susceptible to a wide range of pathogens and has been proven to be a very useful model for studying host–microbe interactions and virulence, providing many genetic and molecular tools for this purpose (Aballay et al. 2000; Labrousse et al. 2000; Garsin et al. 2001; Couillault and Ewbank 2002; Zhang et al. 2005; Tan and Shapira 2011). Virulence mechanisms have been shown to be highly conserved among divergent hosts (Mahajan-Miklos et al. 2000) and the worm has been used to study a wide range of pathogens such as Salmonella enterica, Pseudomonas aeruginosa (Alegado et al. 2003), and Leigonella pneumophila (Komura et al. 2010).
It is reasonable to consider that C. elegans should be useful in providing insights regarding the mechanisms by which probiotic bacteria exert beneficial effects. Little work has been done in this area, and so far, it has been shown that Bifidobacterium infantis, Bifidobacterium longum, Lactobacillus helveticus, Lactobacillus plantarum, and Lactobacillus rhamnosus were able to extend the worm’s lifespan as well as increase resistance to Salmonella enterica (Ikeda et al. 2007). Similarly, Komura et al. (2010) demonstrated the protective effects of B. infantis against L. pneumophila. Moreover, Wang et al. (2011) have suggested the use of C. elegans for preselecting Lactobacillus isolates to control Salmonella infections.
The aim of this study was to examine the effect of LAB on growth, aging, immune response, and tumor-like growth in C. elegans. For this purpose, the following three LAB strains were used: Lactobacillus reuteri (LR), a common LAB strain found in the intestine of humans that produces reuterin, a broad spectrum microbial inhibitor (Talarico et al. 1988; Casas and Dobrogosz 2000); Lactobacillus salivarius (LS) that has been shown to have immunomodulatory effects in mice and can also stimulate the maturation of immature mouse spleen dendritic cells and inhibit the in vitro infectivity of HIV-1 (Galvez et al. 2008; Langa et al. 2012); and Pediococcus acidilactici (PA) that has been shown to modulate intestinal bacterial communities in on-growing red tilapia and stimulate some aspects of the nonspecific immune response (Ferguson et al. 2010) as well as suppress autoimmune encephalomyelitis by inducing IL-10-producing regulatory T cells (Takata et al. 2011). LAB were compared to the standard food source of C. elegans, Escherichia coli strain OP50 (OP).
Materials and methods
Worm and bacterial strains used
L. salivarius DSM 20555, L. reuteri DSM 20016, and P. acidilactici DSM 20284 were obtained as lyophilized stock cultures (DSMZ GmbH, Braunschweig, Germany). C. elegans strains used were N2 (wild-type), CF1038 daf-16(mu86), JK1466 gld-1(q485)/dpy-5(e61);unc-13(e51), MT2547 ced-4(n1162), and CF1407 daf-16(mu86);muIs71[pKL99(daf-16a::GFP/bKO) + pRF4(rol-6)] purchased from the Caenorhabditis Genetics Center (CGC, St Paul, MN, USA). E. coli OP50 was used as the standard food source for growing and maintaining C. elegans.
LAB cultures and worm plates
LS and LR were grown on MRS (Sigma-Aldrich, St. Louis, MO, USA) agar plates supplemented with Tween 20 at 37 °C; PA was grown on MRS agar plates supplemented with Tween 20 at 30 °C; and OP was grown on LB plates at 37 °C. Because the medium used to grow bacteria can influence their effect on C. elegans (Garsin et al. 2001), all cultures used to seed nematode growth medium (NGM) plates (Stiernagle 2006) were grown overnight in MRS broth supplemented with Tween 20, at 30 or 37 °C accordingly. NGM agar plates were then seeded with 0.25–0.5 ml of bacterial culture and incubated overnight at 30 or 37 °C. For experiments using killed bacteria, seeded plates were UV-irradiated.
Longevity assays
Worms grown at 20 °C on NGM plates seeded with OP were hypochloride-treated (Stiernagle 2006) to remove any live bacteria and moved to plates with UV-killed OP. Worms from these plates were used to create synchronized cultures (Sutphin and Kaeberlein 2009). One-day-old adult worms (30 per plate) were transferred to NGM plates seeded with live or UV-killed cultures of OP, LS, LR, or PA. The plates were kept at 20 °C and the worms were transferred every 2 days to fresh plates. During transfer, worms were scored as dead if they failed to respond when touched with a pick. Worms that crawled off the agar or died as a result of internal hatching were removed from the experiment.
Body size
N2 worms grown on plates seeded with UV-killed OP were moved to live OP or LAB plates as 1-day-old adults. After 3, 7, and 11 days, the worms were immobilized in M9 buffer (Stiernagle 2006) containing 25 mM levamisole on agar pads and photographed using a Leica M205 FA stereoscope. The body sizes were calculated using ImageJ software (Abramoff et al. 2004). Each data point represents the mean size of at least ten worms with standard errors.
Intestinal colonization
The measurement of bacteria within the worm’s intestinal tract was performed according to Garsin et al. (2001). Briefly, N2 worms grown on plates seeded with UV-killed OP were moved to live OP or LAB plates as 1-day-old adults. After 2 days, worms were collected and washed four times in M9 buffer. Three worms (per sample) were placed in a microcentrifuge tube containing 20 μl M9 containing 1 % Triton X-100 and mechanically disrupted using a pestle. A series of dilutions were plated on LB agar (OP) or MRS agar (LAB). Each data point represents the mean cfu from triplicate samples with standard errors.
Confocal microscopy
Synchronized 1-day-old adult N2 and ced-4 and gld-1 worms were moved from UV-killed OP plates to live OP and LAB plates. After 1–5 days, nuclei visualization was achieved by staining with 4′,6-diamidino-2-phenylindole (DAPI, Sigma-Aldrich). Briefly, worms were washed in M9 and fixed in methanol for 20 min at −20 °C. Samples were washed again in M9, incubated for 15 min in 1 mg ml−1 DAPI in M9 at RT, washed in M9, and mounted on 3 % agarose pads for microscopy. For visualizing corpse cells, the worms were stained with SYTO 12 (Molecular Probes, Eugene, OR, USA) as previously described (Gumienny et al. 1999). Briefly, worms were washed in M9 and incubated in 33 μM SYTO 12 in M9 for 4–5 h at RT. Worms were then washed in M9 and transferred to fresh OP plates. After 30 min, worms were anesthetized in levamisole and mounted on 3 % agarose pads for microscopy.
Synchronized 1-day-old adult transgenic worms expressing DAF16::GFP were moved from UV-killed OP plates to live OP and LAB plates. After 24 h, worms were anesthetized in levamisole and mounted on 3 % agarose pads. All samples were visualized using a Leica TCS SP5 confocal microscope (Leica Microsystems GmbH, Wetzlar, Germany).
Transmission electron microscopy
Wild-type N2 worms were grown to L4 stage on UV-killed OP and then moved to live OP and LAB plates. Five days later, samples for transmission electron microscopy (TEM) were prepared by fixing the worms in 0.1 M KPO4 buffer containing 2.5 % glutaraldehyde, 1 % paraformaldehyde, and 0.1 M sucrose for 2 h at 4 °C. Samples were washed in 0.2 M KPO4 buffer and incubated for 90 min at 4 °C in 0.5 % OsO4 in 0.1 M KPO4 buffer. Samples were washed again in 0.2 M KPO4 buffer and enclosed in 3 % agarose cubes. Dehydration and UAc staining took place by incubating for 10 min in 30 and 50 % acetone, 30 min in 70 % acetone containing 1 % UAc, and 10 min in 95 and 100 % acetone. Samples were then placed in freshly made resin and cured for 3 days. Sections were cut using a Reichert OMU3 ultramicrotome and visualized using a JEOL JEM-100S TEM (JEOL Ltd, Tokyo, Japan).
Quantitative real-time PCR
One-day-old adult N2 worms were moved from UV-killed OP plates to live OP and LAB plates. After 24 h at 20 °C, worms were washed in M9 and total mRNA was isolated using RNAzol (Sigma-Aldrich), according to the manufacturer’s instructions. RNA was transcribed to cDNA using iSctipt cDNA synthesis kit (Bio-Rad Laboratories, Hercules, CA, USA) with random primers. Quantitative real-time PCR was performed using a Bio-Rad MJ Mini PCR unit equipped with a Mini Opticon Detection System using SsoFast™ EvaGreen Supermix (Bio-Rad Laboratories). Calculation of reaction efficiencies and analysis of expression data were performed using CFX manager™ software (Bio-Rad Laboratories). Primers were designed using Primer3 (http://primer3.sourceforge.net/). Primer sequences are available upon request.
Statistical analysis
Calculation and comparison of mean lifespans was done using GraphPad Prism 5.00 software (GraphPad Software, San Diego, CA, USA, www.graphpad.com) and the log-rank (Mantel–Cox) test. Body size and gene expression results were compared by one-way analysis of variance using GraphPad Prism 5.00.
Results
Effect on fertility and growth
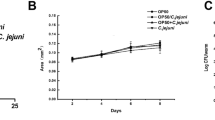
When adult worms were moved to plates seeded with LAB, the first eggs they laid hatched and arrested as L1s. Eggs laid after a few hours did not hatch at all. In fact, all larval stages arrested when they were introduced to LAB plates except for L4s that developed into adults but with a significantly (P ≤ 0.05) smaller body size compared to those fed OP (Fig. 1).
Body size of worms moved to LAB plates as L4 larvae and measured after 3, 7, and 11 days. N2 worms grown on plates seeded with UV-killed OP were moved to live OP or LAB plates as 1-day-old adults. Worms photographed and the body size was calculated using ImageJ. Each data point represents the mean size of at least ten worms ± standard errors. Points next to different superscripts (a–d) differ significantly (P < 0.05)
Intestinal colonization and TEM
In order to verify that LAB were consumed by the worms, intestinal colonization experiments were performed and showed that LAB cells were indeed present within the intestinal track (Fig. 2). Bacterial cells being digested were also visible in images of the intestine produced by TEM (Fig. 3). TEM images of the body wall muscle of 6-day-old adult worms fed OP or LAB did not reveal any morphological differences between treatments (data not shown).
Numbers of live bacterial cells recovered from worms. N2 worms grown on plates seeded with UV-killed OP were moved to live OP or LAB plates as 1-day-old adults. After 2 days, worms were washed and mechanically disrupted in M9 containing 1 % Triton X-100 and a series of dilutions were plated on LB (OP) or MRS agar (LAB). Each data point represents the mean cfu from triplicate samples ± standard errors. Bars with different superscripts (a, b) differ significantly (P < 0.05)
TEM images of N2 worms showing a OP or b LS cells present within the intestinal track. Synchronized N2 L4 worms were moved from UV-killed OP plates to live OP and LS plates. After 7 days, worms were fixed and prepared for TEM. Images of the intestine show OP and LS being digested; black arrow: bacterial cells; white arrow: microvilli; hollow arrow: bacterial cell being digested (scale bar equals 1 μm)
Effect on longevity
As the nematodes did not develop on the LAB plates, the longevity experiments began using 1-day-old adults. The results showed that the LAB significantly (P < 0.01) reduced the lifespan of WT (N2) worms (Fig. 4a). UV-killed LS and LR further reduced the worm’s lifespan while UV-killed PA did not significantly (P > 0.05) affect it (Fig. 4b).
Lifespan of N2 worms fed a live and b UV-killed LAB. One-day-old adult worms were transferred to NGM plates seeded with live or UV-killed cultures of OP or LAB. The plates were kept at 20 °C and the worms were transferred every 2 days to fresh plates. During transfer, worms were scored as dead if they failed to respond when touched with a pick. Worms that crawled off the agar or died as a result of internal hatching were removed from the experiment. Graph represents data from two independent experiments for each treatment. Mean lifespans of worms fed live OP, LS, LR, and PA were 19, 17, 15, and 15, respectively. Mean lifespans of worms fed UV-killed OP, LS, LR, and PA were 20, 14, 14, and 18, respectively. Different superscripts (a–c) indicate significant difference in mean lifespan (P < 0.05)
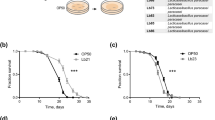
For LR and PA, the use of short-lived daf-16 mutants revealed an even greater sensitivity to the LAB indicating that the insulin/insulin-like signaling cascade may play a role in this effect (Fig. 5a). However, LS significantly (P < 0.001) increased the mean lifespan, suggesting a different mechanism of action (Fig. 5a). With UV-killed bacteria, LR and PA again reduced the lifespan (Fig. 5b), while the increased lifespan observed with the live LS bacteria was reversed (Fig. 5b).
Lifespan of daf-16 worms fed a live and b UV-killed LAB. One-day-old adult worms were transferred to NGM plates seeded with live or UV-killed cultures of OP or LAB. The plates were kept at 20 °C, and the worms were transferred every 2 days to fresh plates. During transfer, worms were scored as dead if they failed to respond when touched with a pick. Worms that crawled off the agar or died as a result of internal hatching were removed from the experiment. Graph represents data from two independent experiments for each treatment. Mean lifespans of daf-16 worms fed live OP, LS, LR, and PA were 14, 17, 11, and 9, respectively. Mean lifespans of worms fed UV-killed OP, LS, LR, and PA were 17, 15, 13, and 15, respectively. Different superscripts (a–c) indicate significant difference in mean lifespan (P < 0.05)
Tumor-like germ cell growth, DAF16 localization, and programmed cell death
When gld-1 −/− worms were moved to LAB plates as 1-day-old adults, the growth of tumor-like germ cells was significantly inhibited (Fig. 6). The results were similar when UV-killed plates were used (data not shown). The result of tumor-like growth inhibition was also demonstrated by a survival experiment where gld-1 −/− worms lived significantly longer on LAB compared to OP (Fig. 7).
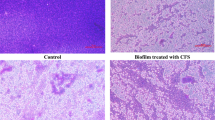
DAPI stained gld-1 worms fed a OP, b LS, c LR, or d PA. One-day-old adult gld-1 −/− worms were transferred to NGM plates seeded with live lawns of OP or LAB. Three days later, worms were methanol-fixed and stained with DAPI. Confocal images show whole worms with stained nuclei of germ cells visible within the gonads (densely stained areas that fill the body). A reduced number of stained nuclei in the LAB samples compared to the OP sample where the images become saturated. Representative images of four independent experiments are shown (scale bar equals 50 μm)
Lifespan of gld-1 worms fed OP or LAB. One-day-old adult gld-1 −/− worms were transferred to NGM plates seeded with live lawns of OP or LAB. The plates were kept at 20 °C, and the worms were transferred daily to fresh plates. Worms were scored as dead when they burst and failed to move when touched with a pick. Mean lifespans of worms fed OP, LS, LR, and PA were 5, 10, 12, and 7, respectively. Different superscripts (a–d) indicate significant difference in mean lifespan (P < 0.05)
We used transgenic worms expressing DAF16 fused to GFP to examine whether the presence of LAB could have an effect on DAF16 localization. Under normal conditions, GFP fluorescence is visible in the intestinal nuclei and cytoplasm. Stress (such as heat) causes DAF16 to localize in the nuclei that become much more visible. It has also been shown that P. aeruginosa can inhibit nuclear localization of DAF16 (Evans et al. 2008). This can be seen following a heat shock. The results showed no difference in DAF16 localization between worms fed LAB and OP both under normal conditions and after heat stress (2 h at 35 °C and 16 h recovery) (Fig. 8).
Transgenic worms expressing DAF16:GFP fed a OP or b LS under normal conditions and c OP or d LS 16 h after heat stress (2 h at 35 °C). Fluorescence is seen in both nuclei (discs) and cytoplasm (background) under normal conditions for OP and LS (small dots are a result of intestinal autofluorescence). After heat shock, nuclear localization of DAF16 is visible in OP- and LS-fed worms (only nuclei and intestinal autofluorescence visible). Images are representative of at least three independent experiments. Results were the same for all LAB strains (scale bar equals 50 μm)
We then examined whether LAB can increase corpse cells in N2 worms, ced-4 mutants that lack programmed cell death (PCD) activity (Aballay and Ausubel 2001), and gld-1 worms. The results showed no visible difference between the LAB plates and the control (OP) for all three worm strains (data not shown).
Immune response
To investigate whether LAB induce an immune response in C. elegans, we examined the expression of genes lys-7, lys-8, thn-2, and sod-3. When 1-day-old adult N2 worms were left for 24 h on live LS plates, the expression levels of lys-7 significantly (P < 0.05) dropped (compared to live OP control) and the expression of thn-2 increased approximately sevenfold (P < 0.05) (Fig. 9a). On LR plates, expression of both lys-7 and thn-2 significantly increased (P < 0.05 and P < 0.01, respectively) with thn-2 showing an 11-fold increase compared to the control (Fig. 9b). On PA plates, lys-7 expression did not significantly (P > 0.05) change, while thn-2 transcript levels were nearly seven times higher (P < 0.05) (Fig. 9c). No significant (P > 0.05) change in expression levels of lys-8 or sod-3 were seen for any of the LAB plates (Fig. 9a–c). These results indicate that despite the fact that exposure to the LAB did not appear to cause nuclear localization of DAF16, it did induce an immune response involving thn-2 and/or lys-7. As all four genes are targets of DAF16, the upregulation of only lys-7 and thn-2, without nuclear localization of DAF16, suggests that pathways other than the insulin/insulin-like signaling are involved.
Results from real-time RT-PCR showing transcriptional response of lys-7, lys-8, thn-2, and sod-3 to LAB. One-day-old adult N2 worms were left for 24 h on live OP or a LS, b LR, and c PA plates, and total RNA was isolated and reverse-transcribed to cDNA. Expression levels are relative to ama-1 and the control treatment (OP) ± standard errors (*P < 0.05; **P < 0.01)
Discussion
The first evidence that LAB can be harmful to nematodes
Our results showed that the LAB inhibited the reproduction, growth, and development of C. elegans. For this reason, the longevity experiments began using 1-day-old adults. Wild-type N2 worms, fed LAB, showed a significantly reduced lifespan (Fig. 4). This result was unexpected given that other LAB tested so far have been shown to have a beneficial effect on longevity (Ikeda et al. 2007). Generally, in C. elegans, dietary restriction or starvation extends lifespan (Houthoofd et al. 2007; Kaeberlein et al. 2006; Sutphin and Kaeberlein 2008). Therefore, a reduced lifespan should not reflect a lack in nutritional value. One factor that reduces longevity is the packing of proliferating bacteria within the intestine; feeding UV-killed OP reduces this effect (Garigan et al. 2002). For this reason, experiments using UV-killed bacteria were also conducted in this study. The results showed that UV-killed LS and LR further reduced the worm’s lifespan while UV-killed PA did not significantly (P > 0.05) affect it (Fig. 4b). This shows that live bacterial activity was not the cause of death, at least for LS and LR. Instead, the presence of live bacteria had a protective effect against the factor(s) that reduced the worm’s lifespan.
To our knowledge, this is the first time LAB have been shown to have a negative effect on nematodes. Although this was not expected, given the results of other researchers, it could be suggested that the effect of LAB on C. elegans may differ significantly from strain to strain.
Ikeda et al. (2007) also observed that adult worms moved to LAB plates lagged slightly in growth for the first 3 days; however, after 4 days, they achieved the same body size as those fed OP. The LAB strains in this study did not allow the worms to grow as much as OP did even after 11 days (Fig. 1). As LAB produce bacteriocins effective against other microbes, the possibility that such a compound, produced by the specific strains used, may be responsible for this effect on C. elegans, cannot be excluded.
Effect of LAB may be the result of competing mechanisms
To further examine the effect of the LAB strains on the worm’s lifespan, we also used the daf-16(mu86) mutant strain. DAF16 is a FOXO family transcription factor that is part of the insulin/insulin-like growth factor 1 signaling mechanism. The lack of DAF16 localization in the nucleus results in downregulation of a number of target genes involved in stress and immune responses. As such, daf-16 mutants are short-lived and sensitive to pathogens or stress (Hsu et al. 2003). The fact that live LS counteracts the negative effect of UV-killed LS and that this is much more obvious in daf-16 worms is interesting. In fact, daf-16 worms fed LS lived as long as N2 worms fed LS (Figs. 4 and 5). One possible explanation for this may be that the reduced growth of worms fed LS results in less oxidative damage. This in turn has a greater effect on daf-16 mutants that are more sensitive to such stress. However, this does not explain why the same effect is not observed with LR and PA. The presence of certain compounds may also be the cause of this effect. Such compounds have been found in royal jelly (Honda et al. 2011). Their presence extended the lifespan of both N2 and daf-16 worms though the exact mechanism behind this remains unclear. It is possible that intermediate metabolites or rapidly degradable compounds found in LS are only present in sufficient concentrations in live cells. Overall, it seems likely that the effect of LAB is not simply due to bacteriocins and may be the result of competing mechanisms.
LAB inhibited tumor-like germ cell growth but did not affect DAF16 localization or increase corpse cells
As LAB are presumed to have antitumor effects, we used worm strain JK1466 to isolate gld-1 −/− individuals and examine the effect of LAB on tumor-like growth. gld-1 is a tumor suppressor the lack of which causes germ cells that enter the meiotic pathway normally to then exit pachytene and return to the mitotic cycle, leading to continuous proliferation of cells that fill the body of the worm and finally kill it (Francis et al. 1995). This strain has been used to study the effectiveness of antitumor drugs (Wilson et al. 2008). When gld-1 −/− worms were moved to LAB plates as 1-day-old adults the growth of tumor-like germ cells was significantly inhibited (Fig. 6). This effect was expected given that the worm’s growth was also inhibited. The result of tumor-like growth inhibition was also demonstrated by a survival experiment where gld-1 −/− worms lived significantly longer on LAB compared to OP (Fig. 7).
Localization of DAF16 in the nucleus, as part of an immune response, can be induced or inhibited by pathogens (Evans et al. 2008; Singh and Aballay 2009). We therefore used transgenic worms expressing DAF16 fused to GFP to examine whether the presence of LAB could have such an effect. The results showed no difference between worms fed LAB and OP both under normal conditions and after heat stress (DAF16 localizes in the nucleus) (data not shown).
In C. elegans, another response to pathogens or stress can be the activation of PCD (Aballay and Ausubel 2001). LR has been shown to induce apoptosis in myeloid leukemia-derived cells induced by tumor necrosis factor (Iyer et al. 2008). We examined whether LAB can increase corpse cells in N2 worms, ced-4 mutants that lack PCD activity (Aballay and Ausubel 2001), and gld-1 worms where we previously demonstrated a significant reduction in tumor-like germ cell proliferation (Fig. 6). The results showed no visible difference between the LAB plates and the control (OP) for all three worm strains (data not shown).
LAB strains induce a DAF16-independent immune response
To investigate whether LAB induce an immune response in C. elegans, we examined the expression of genes lys-7, lys-8, thn-2, and sod-3. lys-7 and lys-8 are expressed downstream of daf-16 and encode lysozymes. Expression levels of lys-7 and lys-8 have been shown to be upregulated in response to infection by the Gram-negative bacterium Serratia marcescens (Mallo et al. 2002). thn-2 encodes a member of the thaumatin family, an immune effector the downregulation of which results in increased sensitivity to P. aeruginosa infection (Evans et al. 2008). thn-2 expression is upregulated in daf-2 mutants suggesting that it is also a target of DAF16 (Murphy et al. 2003). sod-3, another target of DAF16, encodes an iron/manganese superoxide dismutase that protects against oxidative stress and whose expression levels are correlated with the worm’s remaining lifespan (Sánchez-Blanco and Kim 2011).
The response to the three LAB strains seen here shows that despite the fact that exposure to the LAB did not appear to cause nuclear localization of DAF16, it did induce an immune response involving thn-2 and/or lys-7. As all four genes are targets of DAF16, the upregulation of only lys-7 and thn-2, without nuclear localization of DAF16, suggests that pathways other than the insulin/insulin-like signaling are involved. This response also conforms with the strain-specific immune responses described by Alper et al. (2007).
Inhibitory effect on tumor-like growth is confirmed, possible use as model system
The inhibition of tumor-like germ cell growth demonstrated here suggests that the worm could be useful for examining the mechanism by which LAB can exert anticancer activity. Though this has been the subject of many studies, there is mostly indirect evidence so far that LAB can protect against or suppress cancer (Kumar et al. 2010). While the overall effect of the examined LAB strains on C. elegans appeared to be negative, there is evidence here that they can also have a positive effect (such as the extended lifespan of gld-1 and daf-16 worms fed LS). Further investigation of such mechanisms in C. elegans may lead to the identification and isolation of compounds produced by LAB with therapeutic or antitumor activity.
LAB may be effective against parasitic nematodes
Our work showed that not only did the LS, LR, and PA cultures fail to extend the worm’s lifespan, but they also significantly reduced it while also inhibiting egg hatching and larval development (Figs. 1 and 4). C. elegans has also been used as a model for parasitic nematodes (Gilleard 2004). Even though there is a significant genetic variation among these species, the evidence here should justify examining the possibility of using LAB as anthelminthics. There are numerous studies concerning the use of such cultures in farm animals to fight bacterial and viral infections though little attention has been given to their effectiveness against parasites (Gill 2003; Glass et al. 2004; Tierney et al. 2004). A recent review by Travers et al. (2011) describes how this subject is gaining interest along with the need to better describe the molecular mechanisms involved.
References
Aballay A, Ausubel FM (2001) Programmed cell death mediated by ced-3 and ced-4 protects Caenorhabditis elegans from Salmonella typhimurium-mediated killing. PNAS 98:2735–2739. doi:10.1073/pnas.041613098
Aballay A, Yorgey P, Ausubel FM (2000) Salmonella typhimurium proliferates and establishes a persistent infection in the intestine of Caenorhabditis elegans. Curr Biol: CB 10:1539–1542
Abramoff MD, Magalhães Paulo J, Ram SJ (2004) Image processing with ImageJ. Biophoton Int 11:36–42
Alegado RA, Campbell MC, Chen WC, Slutz SS, Tan M-W (2003) Characterization of mediators of microbial virulence and innate immunity using the Caenorhabditis elegans host–pathogen model. Cell Microbiol 5:435–444. doi:10.1046/j.1462-5822.2003.00287.x
Alper S, McBride SJ, Lackford B, Freedman JH, Schwartz D (2007) Specificity and complexity of the Caenorhabditis elegans innate immune response. Mol Cell Biol 27:5544–5553. doi:10.1128/MCB.02070-06
Candela M, Guidotti M, Fabbri A, Brigidi P, Franceschi C, Fiorentini C (2011) Human intestinal microbiota: cross-talk with the host and its potential role in colorectal cancer. Crit Rev Microbiol 37:1–14. doi:10.3109/1040841X.2010.501760
Casas IA, Dobrogosz WJ (2000) Validation of the probiotic concept: Lactobacillus reuteri confers broad-spectrum protection against disease in humans and animals. Microb Ecol Health D 12:247–285
Couillault C, Ewbank JJ (2002) Diverse bacteria are pathogens of Caenorhabditis elegans. Infect Immun 70:4705. doi:10.1128/IAI.70.8.4705
Evans EA, Kawli T, Tan M-W (2008) Pseudomonas aeruginosa suppresses host immunity by activating the DAF-2 insulin-like signaling pathway in Caenorhabditis elegans. PLoS Pathog 4:e1000175. doi:10.1371/journal.ppat.1000175
Ferguson RMW, Merrifield DL, Harper GM, Rawling MD, Mustafa S, Picchietti S, Balcázar JL, Davies SJ (2010) The effect of Pediococcus acidilactici on the gut microbiota and immune status of on-growing red tilapia (Oreochromis niloticus). J Appl Microbiol 109:851–862. doi:10.1111/j.1365-2672.2010.04713.x
Francis R, Barton MK, Kimble J, Schedl T (1995) gld-1, a tumor suppressor gene required for oocyte development in Caenorhabditis elegans. Genetics 139:579–606
Galvez J, Arribas B, Rodríguez-Cabezas ME, Bailon E, Comalada M, Camuesco D, Xaus J, Zarzuelo A (2008) Immunomodulatory properties of Lactobacillus salivarius are not limited to the intestine. P Nutr Soc 67:E40. doi:10.1017/S0029665108006496
Garigan D, Hsu A, Fraser A, Kamath R (2002) Genetic analysis of tissue aging in Caenorhabditis elegans: a role for heat-shock factor and bacterial proliferation. Genetics 16:1101–1112
Garsin DA, Sifri CD, Mylonakis E, Qin X, Singh KV, Murray BE, Calderwood SB, Ausubel FM (2001) A simple model host for identifying Gram-positive virulence factors. PNAS 98:10892–10897. doi:10.1073/pnas.191378698
Gerritsen J, Smidt H, Rijkers GT, de Vos WM (2011) Intestinal microbiota in human health and disease: the impact of probiotics. Genes Nutr 6(3):209–240. doi:10.1007/s12263-011-0229-7
Gill H (2003) Probiotics to enhance anti-infective defences in the gastrointestinal tract. Best Pract Res Cl Ga 17:755–773. doi:10.1016/S1521-6918(03)00074-X
Gilleard JS (2004) The use of Caenorhabditis elegans in parasitic nematode research. Parasitology 128(Suppl 1):S49–S70. doi:10.1017/S003118200400647X
Glass MD, Courtney PD, LeJeune JT, Ward LA (2004) Effects of Lactobacillus acidophilus and Lactobacillus reuteri cell-free supernatants on Cryptosporidium viability and infectivity in vitro. Food Microbiol 21:423–429. doi:10.1016/j.fm.2003.11.001
Gumienny TL, Lambie E, Hartwieg E, Horvitz HR, Hengartner MO (1999) Genetic control of programmed cell death in the Caenorhabditis elegans hermaphrodite germline. Development 126:1011–1022
Honda Y, Fujita Y, Maruyama H, Araki Y, Ichihara K, Sato A, Kojima T, Tanaka M, Nozawa Y, Ito M, Honda S (2011) Lifespan-extending effects of royal jelly and its related substances on the nematode Caenorhabditis elegans. PLoS One 6:e23527. doi:10.1371/journal.pone.0023527
Houthoofd K, Gems D, Johnson TE, Vanfleteren JR (2007) Dietary restriction in the nematode Caenorhabditis elegans. Interdisc Topics Ger 35:98–114. doi:10.1159/000096558
Hsu A-L, Murphy CT, Kenyon C (2003) Regulation of aging and age-related disease by DAF-16 and heat-shock factor. Science (New York, NY) 300:1142–1145. doi:10.1126/science.1083701
Ikeda T, Yasui C, Hoshino K, Arikawa K, Nishikawa Y (2007) Influence of lactic acid bacteria on longevity of Caenorhabditis elegans and host defense against Salmonella enterica serovar enteritidis. Appl Environ Microb 73:6404–6409. doi:10.1128/AEM.00704-07
Iyer C, Kosters A, Sethi G, Kunnumakkara AB, Aggarwal BB, Versalovic J (2008) Probiotic Lactobacillus reuteri promotes TNF-induced apoptosis in human myeloid leukemia-derived cells by modulation of NF-kappaB and MAPK signalling. Cell Microbiol 10:1442–1452. doi:10.1111/j.1462-5822.2008.01137.x
Kaeberlein TL, Smith ED, Tsuchiya M, Welton KL, Thomas JH, Fields S, Kennedy BK, Kaeberlein M (2006) Lifespan extension in Caenorhabditis elegans by complete removal of food. Aging cell 5:487–494. doi:10.1111/j.1474-9726.2006.00238.x
Komura T, Yasui C, Miyamoto H, Nishikawa Y (2010) Caenorhabditis elegans as an alternative model host for Legionella pneumophila, and protective effects of Bifidobacterium infantis. Appl Environ Microb 76:4105–4108. doi:10.1128/AEM.03021-09
Kumar M, Kumar A, Nagpal R, Mohania D, Behare P, Verma V, Kumar P, Poddar D, Aggarwal PK, Henry CJK, Jain S, Yadav H (2010) Cancer-preventing attributes of probiotics: an update. Int J Food Sci Nutr 61:473–496. doi:10.3109/09637480903455971
Labrousse A, Chauvet S, Couillault C, Kurz CL, Ewbank JJ (2000) Caenorhabditis elegans is a model host for Salmonella typhimurium. Curr Biol: CB 10:1543–1545
Langa S, Maldonado-Barragán A, Delgado S, Martín R, Martín V, Jiménez E, Ruíz-Barba JL, Mayo B, Connor RI, Suárez JE, Rodríguez JM (2012) Characterization of Lactobacillus salivarius CECT 5713, a strain isolated from human milk: from genotype to phenotype. Appl Microbiol Biot 94(5):1279–1287. doi:10.1007/s00253-012-4032-1
Mahajan-Miklos S, Rahme LG, Ausubel FM (2000) Elucidating the molecular mechanisms of bacterial virulence using non-mammalian hosts. Mol Microbiol 37:981–988
Mallo GV, Kurz CL, Couillault C, Pujol N, Granjeaud S, Kohara Y, Ewbank JJ (2002) Inducible antibacterial defense system in C. elegans. Curr Biol: CB 12:1209–1214
Murphy C, McCarroll S, Bargmann C, Fraser A, Kamath RS, Ahringer J, Li H, Kenyon C (2003) Genes that act downstream of DAF-16 to influence the lifespan of Caenorhabditis elegans. Nature 424(6946):277–283
Nakajima H, Toba T, Ito A, Kudo S, Sato S, Sato Y (2003) The screening of hydrogen peroxide-producing lactic acid bacteria and their application to inactivating psychrotrophic food-borne pathogens. Curr Microbiol 47:231–236. doi:10.1007/s00284-002-3993-1
Ottaviani E, Ventura N, Mandrioli M, Candela M, Franchini A, Franceschi C (2011) Gut microbiota as a candidate for lifespan extension: an ecological/evolutionary perspective targeted on living organisms as metaorganisms. Biogerontology 12(6):599–609. doi:10.1007/s10522-011-9352-5
Pagnini C, Saeed R, Bamias G, Arseneau KO, Pizarro TT, Cominelli F (2010) Probiotics promote gut health through stimulation of epithelial innate immunity. PNAS 107:454–459. doi:10.1073/pnas.0910307107
Perdigón G, Maldonado Galdeano C, Valdez JC, Medici M (2002) Interaction of lactic acid bacteria with the gut immune system. Eur J Clin Nutr 56(Suppl 4):S21–S26. doi:10.1038/sj.ejcn.1601658
Putaala H, Barrangou R, Leyer GJ, Ouwehand AC, Hansen EB, Romero DA, Rautonen N (2010) Analysis of the human intestinal epithelial cell transcriptional response to Lactobacillus acidophilus, Lactobacillus salivarius, Bifidobacterium lactis and Escherichia coli. Beneficial microbes 1:283–295. doi:10.3920/BM2010.0003
Singh V, Aballay A (2009) Regulation of DAF-16-mediated innate immunity in Caenorhabditis elegans. J Biol Chem 284:35580–35587. doi:10.1074/jbc.M109.060905
Stiernagle T (2006) Maintenance of C. elegans. WormBook: Online Rev C elegans Biol 11:1–11. doi:10.1895/wormbook.1.101.1
Sutphin GL, Kaeberlein M (2009) Measuring Caenorhabditis elegans life span on solid media. J Vis Exp. doi:10.3791/1152
Sutphin GL, Kaeberlein M (2008) Dietary restriction by bacterial deprivation increases life span in wild-derived nematodes. Exp Gerontol 43:130–135. doi:10.1016/j.exger.2007.10.019
Sánchez-Blanco A, Kim SK (2011) Variable pathogenicity determines individual lifespan in Caenorhabditis elegans. PLoS genetics 7:e1002047. doi:10.1371/journal.pgen.1002047
Takata K, Kinoshita M, Okuno T, Moriya M, Kohda T, Honorat JA, Sugimoto T, Kumanogoh A, Kayama H, Takeda K, Sakoda S, Nakatsuji Y (2011) The lactic acid bacterium Pediococcus acidilactici suppresses autoimmune encephalomyelitis by inducing IL-10-producing regulatory T cells. PLoS One 6:e27644. doi:10.1371/journal.pone.0027644
Talarico T, Casas I, Chung TC, Dobrogosz W (1988) Production and isolation of reuterin, a growth inhibitor produced by Lactobacillus reuteri. Antimicrob Agents Ch 32:1854. doi:10.1128/AAC.32.12.1854.Updated
Tan M-W, Shapira M (2011) Genetic and molecular analysis of nematode–microbe interactions. Cell Microbiol 13:497–507. doi:10.1111/j.1462-5822.2011.01570.x
Taverniti V, Guglielmetti S (2011) The immunomodulatory properties of probiotic microorganisms beyond their viability (ghost probiotics: proposal of paraprobiotic concept). Genes Nutr 6:261–274. doi:10.1007/s12263-011-0218-x
Tierney J, Gowing H, Van Sinderen D, Flynn S, Stanley L, McHardy N, Hallahan S, Mulcahy G (2004) In vitro inhibition of Eimeria tenella invasion by indigenous chicken Lactobacillus species. Vet Parasitol 122:171–182. doi:10.1016/j.vetpar.2004.05.001
Travers M-A, Florent I, Kohl L, Grellier P (2011) Probiotics for the control of parasites: an overview. J Parasitol Res 2011:610769. doi:10.1155/2011/610769
Wang C, Wang J, Gong J, Yu H, Pacan JC, Niu Z, Si W, Sabour PM (2011) Use of Caenorhabditis elegans for preselecting Lactobacillus isolates to control Salmonella typhimurium. J Food Protect 74:86–93. doi:10.4315/0362-028X.JFP-10-155
Wilson MA, Rimando AM, Wolkow CA (2008) Methoxylation enhances stilbene bioactivity in Caenorhabditis elegans. BMC Pharmacol 8:15. doi:10.1186/1471-2210-8-15
Zhang Y, Lu H, Bargmann CI (2005) Pathogenic bacteria induce aversive olfactory learning in Caenorhabditis elegans. Nature 438:179–184. doi:10.1038/nature04216
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Fasseas, M.K., Fasseas, C., Mountzouris, K.C. et al. Effects of Lactobacillus salivarius, Lactobacillus reuteri, and Pediococcus acidilactici on the nematode Caenorhabditis elegans include possible antitumor activity. Appl Microbiol Biotechnol 97, 2109–2118 (2013). https://doi.org/10.1007/s00253-012-4357-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-012-4357-9