Abstract
Obtaining full-length 16S rRNA gene sequences is important for generating accurate taxonomy assignments of bacteria, which normally is realized via clone library construction. However, the application of clone library has been hindered due to its limitations in sample throughput and in capturing minor populations (<1 % of total microorganisms). To overcome these limitations, a new strategy, two-step denaturing gradient gel electrophoresis (2S-DGGE), is developed to obtain full-length 16S rRNA gene sequences. 2S-DGGE can compare microbial communities based on its first-round DGGE profiles and generate partial 16S rRNA gene sequences (8–534 bp, Escherichia coli numbering). Then, strain-specific primers can be designed based on sequence information of bacteria of interest to PCR amplify their remaining 16S rRNA gene sequences (515–1541 bps, E. coli numbering). The second-round DGGE can confirm DNA sequence purity of these PCR products. Finally, the full-length 16S rRNA gene sequences can be obtained through combining the two partial DNA sequences. By employing 2S-DGGE, taxonomies of a group of dehalogenating bacteria have been assigned based on their full-length 16S rRNA gene sequences, several of which existed in dehalogenating microcosms as minor populations. In all, 2S-DGGE can be utilized as a medium throughput method for taxonomic identification of interested/minor populations from single or multiple microbial consortia.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The 16S rRNA gene-based molecular tools have revolutionized our understanding of the diversity and dynamics of bacterial community which normally involves microbial community profiling and taxonomic assignments of specific populations based on full-length 16S rRNA gene sequences. Among them, PCR-denaturing gradient gel electrophoresis (DGGE; Muyzer et al. 1993) has played a key role in fingerprinting of microbial communities. Compared with other methods (e.g., clone library), DGGE has advantages in loading multiple samples in a single run and in observing/comparing their microbial community profiles directly on a gel (Nocker et al. 2007). However, the commonly used DGGE method is limited by a maximum of ~500 nucleotides for good separation on DGGE gels (Nocker et al. 2007), therefore, only partial 16S rRNA gene sequences can be obtained from sequencing excised gel bands. Burr et al. (2006) reported a strategy to overcome this limitation by cloning full-length 16S rRNA PCR products, followed by PCR/DGGE to screen the clone library, and then by sequencing the DNA of clones. Nevertheless, the strategy offsets advantages of normal DGGE both in sample throughput and microbial community comparison. Moreover, this strategy has low genotyping capability due to the limitation step of PCR/DGGE which screens sequence differences only within one (i.e., the V3) out of nine highly variable regions of cloned full-length 16S rRNA gene sequences (Burr et al. 2006).
Clone library has been widely used to complement other methods to obtain full-length16S rRNA gene sequences for taxonomic identification (Nocker et al. 2007; Yoshida et al. 2009). However, the employment of clone library also brings several limitations, e.g., the gene sequences of minor populations (<1 % in total abundance) could be missed due to picking and sequencing limited clones (Nocker et al. 2007); clone library is a low throughput method, which makes it time- and labor-intensive to compare microbial communities through constructing multiple clone libraries.
In this study, we developed a new strategy, two-step denaturing gradient gel electrophoresis (2S-DGGE), to compare microbial communities through screening sequence differences across over five highly variable regions (V1–V5) of 16S rRNA genes, and to capture the full-length sequences of interested/minor populations from one or multiple samples for their taxonomic assignments without tedious clone library construction.
Materials and methods
Chemicals
Unless stated otherwise, chemicals were purchased from Sigma-Aldrich (SIGMA-ALDRICH, St. Louis, MO, USA) at the highest purity available. The DNA extraction kits and PCR reagents were obtained from Qiagen (QIAGEN, GmbH, Germany).
Culture and growth conditions
A 1,2-dichloroethane (1,2-DCA) dechlorinating microcosm was established with digester sludge, and its dechlorinating bacteria were characterized by 2S-DGGE. Dehalococcoides sp. strain MB (EU073964) that dechlorinates tetrachloroethylene (PCE) to dominant trans-1,2-dichloroethene (trans-DCE) was utilized to prepare an artificial DNA sample in this study and was maintained as described previously (Cheng and He 2009). A coculture containing a Dehalococcoides sp. and two Dehalobacter sp. was also selected to prepare the artificial DNA sample, which was enriched from a river sediment sample and could dechlorinate PCE completely to ethene within 30 days.
DNA extraction, PCR, clone library, and sequencing
Total genomic DNA was extracted from 1 ml of cell pellets collected from dechlorinating cultures and the controls according to a method described previously (Löffler et al. 1997). The concentration of the nucleic acid was determined by a Nanodrop-1000 (NanoDrop Technologies, Wilmington, DE, USA). PCR (Eppendorf, Hamburg, Germany) amplification of nearly complete 16S rRNA gene sequences were conducted under conditions described previously (Löffler et al. 1997). Clone libraries of 16S rRNA genes of dehalogenating cultures were established by using TOPO-TA cloning kit (Invitrogen, Carlsbad, CA, USA), and all further clone-based experiments were carried out as described previously (He et al. 2003). Purified plasmids or PCR products were sequenced and aligned by using MEGA4 (He et al. 2003; Tamura et al. 2007).
DGGE
PCR products amplified with a primer set of 8FGC and 518R or 519FGC and 926R were separated on an 8 % polyacrylamide gel with a gradient range of 30–60 % (100 % denaturant consisted of 7 M urea and 40 % deionized formamide) in 0.5× TAE buffer. Gradient gels were cast with Bio-Rad’s Model 475 gradient delivery system (Bio-Rad, Hercules, CA, USA). The electrophoresis was performed for 15 h at a constant electric current of 30 mA and a temperature of 60 °C with the DCode Mutation Detection System (Bio-Rad, Hercules, CA, USA). Gel images of SYBR® Gold (Invitrogen, Carlsbad, CA, USA) stained DNA were taken by using a Molecular Imager Gel Doc XR System (Bio-Rad, Hercules, CA, USA). Bands of interest were excised and DNA fragments were extracted by using the QIAEX II Gel Extraction Kit (QIAGEN, GmbH, Germany). The captured DNAs were then PCR re-amplified by using the primer set 8FGC and 518R or 519FGC and 926R, and re-analyzed by DGGE to confirm that single bands were obtained before sending the PCR re-amplified products for sequencing.
2S-DGGE procedures
The schematic diagram of 2S-DGGE was shown in Fig. 1, which included seven steps: (1) partial sequences (PCR amplified using primers 8FGC and 518R) of 16S rRNA genes were prepared; (2) DGGE was carried out to separate partial 16S rRNA gene fragments from Step 1, and bands of interest were excised from DGGE gels to elute the DNAs, followed by PCR re-amplification, purification, and then sequencing; (3) sequences from Step 2 were aligned with BioEdit assembly software (http://www.mbio.ncsu.edu/BioEdit/bioedit.html), and were analyzed with BLASTN (http://www.ncbi.nlm.nih.gov/) to design a strain-specific forward primer targeting sequences of highly variable regions in V1 (from base 69–100, Escherichia coli numbering), V2 (from base 137–226, E. coli numbering) or V3 (from base 440–496, E. coli numbering); (4) the remaining part of 16S rRNA gene sequences of interested species were PCR amplified with primer sets of the newly-designed strain-specific forward primers (BandX-F, X = 1, 2, 3……) and universal bacterial reverse primer (1392R/1541R); (5) diluted PCR products (50 times dilution) from Step 4 were used as templates for PCR amplification with another universal bacterial primer set (519FGC and 926R) for following DGGE analysis; (6) the second round of DGGE was conducted to separate DNA fragments from Step 5 to check their purity/diversity; (7) once DGGE gels showed single bands (e.g., Band 2 and Band 3 in Fig. 1), the corresponding purified PCR products from Step 4 would be sequenced and then nearly full-length 16S rRNA gene sequences were obtained by assembling sequences obtained at Step 2 and Step 7. However, if DGGE gel showed multiple bands (e.g., Band1-1 and Band1-2 in Fig. 1), bands could also be excised and sequenced. As a result, a ~920-bp sequence (from base 8–926, E. coli numbering) could be obtained by assembling both Step 7 and Step 2 sequences. Normally, single fragments could be obtained at Step 4 with well-designed strain-specific forward primers at Step 3.
Nucleotide sequence accession numbers
The nearly full-length 16S rRNA gene sequences of Dehalococcoides species are deposited in GenBank under accession numbers JF698641-JF698649.
Results
2S-DGGE development
To understand the phylogeny of microbes in a consortium and to obtain their full-length 16S rRNA gene sequences without referring to the tedious clone library construction, a 2S-DGGE approach was developed (Fig. 1). Selecting appropriate universal bacteria primer sets at Step 1 and Step 5 (Fig. 1) and designing bacterium strain-specific forward primers at Step 3 were crucial when developing the 2S-DGGE method. The primer sets selected should meet the following two criteria: firstly, to get nearly full-length 16S rRNA gene sequences, the forward universal bacteria primer used at Step 1 should be 8F (Reysenbach et al. 1994) or 11F (Kane et al. 1993), and the reverse universal bacteria primer used at Step 4 should be 1392R (Lane 1991) or 1541R (Paster et al. 2002); secondly, to screen sequence differences within continuous highly variable regions by 2S-DGGE, the reverse universal bacteria primer used at Step 1 and forward universal bacteria primer used at Step 5 should target the same positions in the 16S rRNA gene sequences. Several different universal bacteria primer sets (i.e., 8FGC-518R, 8F-518RGC, 519FGC-926R, 519F-926RGC, 519FGC-1115R, and 519F-1115RGC) for Step 1 and Step 5 in Fig. 1 were assessed through computer program MELT94 (Lerman et al. 1996). In the assessment, 16S rRNA gene sequences of known dehalogenators were used as model sequences to be amplified with those primer sets and then the melting behavior of simulative amplified products were predicted with MELT94. Two universal bacteria primer sets (8FGC and 518R, and 519FGC and 926R) were ultimately chosen to be used in the 2S-DGGE analysis, both of which exhibits good separation of dehalogenators’ 16S rRNA gene fragments based on their meltmaps calculated by MELT94. The predicted result was confirmed through conducting 2S-DGGE with 16S rRNA gene sequences from two Dehalococcoides species strains MB (EU073964) and BAV1 (CP000688) (data not shown). Additionally, the two primer sets were also able to effectively fingerprint complex microbial communities other than communities in dechlorinating cultures (Juck et al. 2003; Nakasaki et al. 2009).
The bacterium strain-specific forward primers used at Step 4 can be either the available genus-specific primers or primers designed at Step 3 based on the sequence information from Step 2, depending on the complexity of the microbial community in the studied samples. The available genus-specific forward primers can be utilized as forward primers at Step 4 only when single bacterium in this genus presented in DGGE results at Step 2, and the target positions of genus-specific forward primers should be located in V1, V2 or V3 (base 8–529, E. coli numbering). If multiple species from the same genus presented at Step 2, bacterium strain-specific forward primers should be designed at Step 3, and then the remaining 16S rRNA gene sequences of those species can be amplified with bacterium strain-specific forward primers combined with the reverse universal bacteria primer, 1392R (Lane 1991) or 1541R (Paster et al. 2002). 1392R was chosen in this study due to its comparatively broader matches compared to 1541R (Baker et al. 2003). The sequences (from base 8–529, E. coli numbering) from Step 2 covered three highly variable regions of 16S rRNA genes (Baker et al. 2003). The designed strain-specific forward primers for PCR amplifications at Step 4 should target positions within any of the three highly variable regions, and the particular variable base pairs specific for the targeted strain should be located at the 3′ end of the strain-specific forward primer, which usually ensures the primer specificity (Wu et al. 2009).
Phylogenetic characterization of Dehalococcoides in a 1,2-dichloroethane (1,2-DCA) dechlorinating culture by using 2S-DGGE
2S-DGGE can compare microbial community profiles directly on the gel and, at the same time, provide taxonomic assignments of bacteria with full-length 16S rRNA gene sequences. To demonstrate these capabilities, 2S-DGGE was employed to characterize dechlorinating bacteria in a 1,2-DCA dechlorinating consortium (Fig. 2). Through comparing DGGE profiles (with PCR products amplified with 8FGC and 518R at Step 1) of the 1,2-DCA dechlorinating culture (lane 1 in Fig. 2a) and its control (lane 2 in Fig. 2a) without 1,2-DCA amendment, band-1 was shown to represent the bacterium responsible for 1,2-DCA dechlorination (Step 2). DNA sequencing results show that band-1 represents a Dehalococcoides bacterium in Victoria-subgroup. Based on the sequence information, a forward primer, denoted as Dhc-Victoria, was designed to specifically target the V3 highly variable region of the Dehalococcoides 16S rRNA gene sequence (Step 3), of which the specificity was verified using RDP's ProbeMatch (Cole et al. 2009). The remaining 16S rRNA gene sequence of the Dehalococcoides bacterium could be amplified with primers Dhc-Victoria and 1392R (Step 4), of which diluted DNA samples (50 times dilution) were used as templates for PCR amplification with primers 519FGC and 926R (Step 5), and then subjected to DGGE separation (Step 6) to check whether the remaining 16S rRNA gene sequence was specifically amplified. The single band on DGGE gel (lane 3 in Fig. 2a) confirmed the specific PCR amplification of the remaining 16S rRNA gene sequences with primers Dhc-Victoria and 1392R, and thus, their sequence information could be obtained through sequencing purified PCR products from Step 4. Finally, near full-length 16S rRNA gene sequence of the Dehalococcoides bacterium was obtained by assembling sequences from Step 3 and Step 7.
a Application of 2S-DGGE to characterize dechlorinators in a 1,2-DCA dechlorinating culture (lane 1) and in its control culture without 1,2-DCA amendment (lane 2) (note: Step 2 using 8FGC/518R primers); lane 3, amplification specificity of Dehalococcoides bacteria with 519FGC/926R primers at Step 6. b Phylogenetic tree of dechlorinators in the 1,2-DCA dechlorinating culture (closed circle) and PBDE-debromination microcosms (open circles). Phylogenetic analyses were conducted in MEGA4 (Tamura et al. 2007)
Application of 2S-DGGE to probe minor populations
2S-DGGE can be utilized to phylogenetically characterize minor populations in a complex microbial community. To determine its applicability, an artificial DNA sample was prepared, in which 16S rRNA gene sequences of two Dehalococcoides (strain MB and AD14-1) and two Dehalobacter (strainTCP2 and AD14) were mixed with 16S rRNA gene sequences from a digester sludge sample. The amount of each of those four dehalogenators’ DNA was less than 1 % of total DNA (molar basis). A clone library was constructed as a control with the PCR products amplified with universal bacterial primer set (8F and 1392R) and the artificial DNA as the template. Before conducting 2S-DGGE, the mixed DNA templates were firstly PCR amplified with 8F forward primer and genus-specific reverse primers, DHC710R and DEB630R (Table 1), to specifically amplify 16S rRNA gene sequences of Dehalococcoides and Dehalobacter, respectively. The diluted PCR products were further used as templates for PCR amplifications with universal bacteria primer set, 8FGC and 518R, at Step 1, and their products were profiled on DGGE gels (lanes 1 and 2 in Fig. 3a). Figure 3a demonstrated that the two Dehalobacter (lane 1 in Fig. 3a) and two Dehalococcoides (lane 2 in Fig. 3a) were specifically amplified. Through serial procedures of band excising, DNA extraction, PCR re-amplification and subsequent sequencing, partial 16S rRNA gene sequences (from base 8-529, E. coli numbering) of those four dehalogenators were obtained, based on which four strain-specific forward primers (Deb-V2-F, DebTCP-V3-F, Dhc-V2-F, and DhcMB-V2-F in Table 1) were designed. Then, the remaining 16S rRNA gene sequences could be obtained through PCR amplifications with those strain-specific forward primers together with 1392R reverse primer, of which the specificities were confirmed by DGGE profiles (lanes 3, 4, 5, and 6 in Fig. 3a) on PCR products amplified with the primer set, 519FGC and 926R. The DNA fragments from Dehalococcoides sp. MB (lane 3 in Fig. 3b) and strain AD14-1 (lane 4 in Fig. 3a) stopped at positions with the same denaturant concentration for their identical sequences from base 519 to 1392 (E. coli numbering). Full-length 16S rRNA gene sequences of dehalogenators in this artificial sample were finally obtained through assembling sequences from Step 2 and Step 7, which were consistent with their previous sequencing results. In comparison, among 100 randomly picked clones in the control experiment, only one clone harbored the 16S rRNA gene sequence of Dehalococcoides sp. AD14-1.
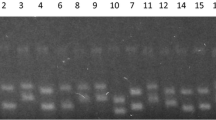
2S-DGGE analysis of minor populations: a charactering two Dehalococcoides and two Dehalobacter with less than 1 % in abundance over total microorganisms in an artificial complex microbial community. Lane 1 Dehalobacter bacteria (Step2); lane 2 Dehalococcoides bacteria (Step 2); lanes 3, 4, 5, and 6 amplification specificity of Dehalococcoides sp. MB, Dehalococcoides sp. unknown, Dehalobacter sp. unknown, and Dehalobacter sp. TCP2, respectively (Step 6). b Characterizing Dehalococcoides in PBDE-debromination microcosms established with soils and sediments from different geographic sites. Lane 1, S-T-1; lane 2, C-T-2; lane 3, C-T-3; lane 4, C-T-7; lane 5, C-T-10; lane 6, GY-T-11
To further demonstrate the advantage of 2S-DGGE over clone library in characterizing specific populations from actual samples, it was applied to polybrominated diphenyl ethers (PBDEs) debromination microcosms established with soils or sediments from different geographic sites (Lee and He 2010). Previous studies have identified that Dehalococcoides species presented in most PBDE-debromination microcosms as minor populations by using genus-specific primers (Lee and He 2010). However, the 16S rRNA gene sequence information of these Dehalococcoides bacteria was limited due to massive work in clone library construction. To obtain their nearly full-length 16S rRNA gene sequences without construction of multiple clone libraries, 2S-DGGE was applied to phylogenetically characterize Dehalococcoides populations in six randomly chosen PBDE-debromination microcosms (i.e., S-T-1, C-T-2, C-T-3, C-T-7, C-T-10, and GY-T-11) (Lee and He 2010). Figure 3b showed that three typical bands stopping at different gradient positions appeared on the DGGE gel, which was extracted and sequenced. Sequencing results demonstrated that they represented bacteria affiliated to three subgroups of Dehalococcoides, i.e., Cornell (195-like, Dehalococcoides sp. strain ST1, presented only in S-T-1), Victoria (VS-like, Dehalococcoides sp. strain CT3-2, CT10-2, and GY11-2 presented in C-T-3, C-T-10, and GY-T-11, respectively), and Pinellas (CBDB1-like, Dehalococcoides sp. strain CT2, CT3-1, CT7, CT10-1, and GYT11-1 presented in C-T-2, C-T-3, C-T-7, C-T-10, and GY-T-11, respectively). The remaining 16S rRNA gene sequences of 195-, VS- and CBDB1-like Dehalococcoides were obtained through PCR amplifications with their specific forward primers (Table 1) together with 1392R, and the amplification specificity was confirmed by DGGE at Step 6 (data not shown). The nearly full-length 16S rRNA gene sequences (1,353 bp) were identified in the microcosms, sharing 100 %, 99 % (1 bp difference over 1,353 bp), and 100 % similarities with that of strains 195, VS, and CBDB1, respectively (Fig. 2b).
Discussion
Microbial community analysis normally involves two steps: to screen and compare microbial communities by using community profiling techniques (e.g., Terminal Restriction Fragment Length Polymorphism or T-RFLP, DGGE, and next-generation sequencing platforms); to identify the taxonomy of the microbes based on full-length 16S rRNA gene sequences by clone library (Nocker et al. 2007). Though the clone library can generate high quality full-length 16S rRNA gene sequences, it has limitations in sample throughput and in capturing minor populations (<1 % of total microorganisms; Nocker et al. 2007; Ding and He 2012). In this study, the newly developed 2S-DGGE approach provides an alternative and convenient way to obtain full-length 16S rRNA gene sequences, which distinguishes from clone library by its advantages of running multiple samples in a single run and displaying microbial community profiles directly on the gel. Therefore, 2S-DGGE can be a complementary approach to clone library in studying specific populations in mixed cultures. 2S-DGGE also has high resolution capability in screening sequence difference. Out of nine highly variable regions in 16S rRNA genes (Baker et al. 2003), Liu et al. (Liu et al. 2008) reports that V2 and V4 give the lowest error rates when assigning taxonomy, and these two regions are also suitable for community clustering (Liu et al. 2008). Compared with normal DGGE by screening sequence difference within the sole V3 region, 2S-DGGE can screen sequence differences across over 5 highly variable regions (V1~V5) within 16S rRNA genes by using the two selected GC-clamped PCR primer sets (8FGC and 518R, and 519FGC and 926R), which thus could supply much better genotyping capability. For example, 2S-DGGE analysis demonstrates an insertion sequence in V1 region of the 16S rRNA gene of strain TCP2, which is not in the coverage of the widely used DGGE primer set 341FGC and 518R (Muyzer et al. 1993) targeting only the V3 region of the 16S rRNA genes.
2S-DGGE technique also shows advantage of probing minor dechlorinators (e.g., <1 % Dehalococcoides or Dehalobacter) in microbial reductive dechlorinating consortia or bioremediation sites, which are difficult to be captured by clone library unless picking more than 100 colonies. Furthermore, phylogenetic characterization of specific dechlorinators can be conducted by PCR analysis using genus-specific primers. To further differentiate their subgroups and make taxonomy assignments, multiple clone libraries are required for each genus-specific PCR product but only partial 16S rRNA gene sequences (depending on the amplified sequence length of genus-specific primers) can be obtained, e.g., ~830 bp sequences can be obtained by using Dehalobacter genus-specific primers Deb179F and 1007R (Holliger et al. 1998). The 2S-DGGE can differentiate distinct species in each genus and further pull out their nearly full-length 16S rRNA gene sequences of interested/minor populations within multiple samples simultaneously in a single run, e.g., phylogenetic characterization of Dehalococcoides populations present in PBDE-debromination microcosms by using a single run of 2S-DGGE instead of constructing 6 clone libraries (Fig. 3b). Additionally, 2S-DGGE might be a complementary strategy to next-generation sequencing (NGS) platforms to obtain full-length sequences of interested populations, e.g., dechlorinating bacteria, nitrification/denitrification bacteria, phosphate accumulation microorganisms, etc. Thus far, characterizing microbial community has been greatly facilitated by the development of NGS platforms (e.g., GS-FLX system, Illumina Solexa system, and SOLiD system) which supply unsurpassed detection capability. However, the application of NGS is still limited due to their relatively short read-lengths (e.g., average 330 bp for 454 GS-FLX system; Metzker 2010). Based on 16S rRNA gene sequence (V1–V3) information generated by NGS, bacterial strain-specific forward primers can be designed (Step 3 in Fig. 1) to probe bacteria of interest, and then their full-length 16S rRNA gene sequences can be obtained through following 2S-DGGE procedures.
Overall, the medium throughput 2S-DGGE approach is promising both in characterizing microbial communities and in taxonomy identification of interested/minor populations with full-length 16S rRNA gene sequence.
References
Baker GC, Smith JJ, Cowan DA (2003) Review and re-analysis of domain-specific 16S primers. J Microbiol Methods 55:541–555
Burr MD, Clark SJ, Spear CR, Camper AK (2006) Denaturing gradient gel electrophoresis can rapidly display the bacteria diversity contained in 16S rDNA clone libraries. Microbiol Ecol 51:479–486
Cheng D, He J (2009) Isolation and characterization of "Dehalococcoides" sp. strain MB, which dechlorinates tetrachloroethene to trans-1,2-dichloroethene. Appl Environ Microbiol 75:5910–5918
Cole JR, Wang Q, Cardenas E, Fish J, Chai B, Farris RJ, Kulam-Syed-Mohideen AS, McGarrell DM, Marsh T, Garrity GM, Tiedje JM (2009) The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res 37(Database issue):D141–D145. doi:10.1093/nar/gkn879
Ding C, He J (2012) Molecular techniques in the biotechnological fight against halogenated compounds in anoxic environments. Microb Biotechnol 5:347–367
He J, Ritalahti KM, Aiello MR, Löffler FE (2003) Complete detoxification of vinyl chloride by an anaerobic enrichment culture and identification of the reductively dechlorinating population as a Dehalococcoides species. Appl Environ Microbiol 69:996–1003
Holliger C, Hahn D, Harmsen H, Ludwig W, Schumacher W, Tindall B, Vazquez F, Weiss N, Zehnder AJ (1998) Dehalobacter restrictus gen. nov. and sp. nov., a strictly anaerobic bacterium that reductively dechlorinates tetra- and trichloroethene in an anaerobic respiration. Arch Microbiol 169:313–321
Juck D, Driscoll BT, Charles TC, Greer CW (2003) Effect of experimental contamination with the explosive hexahydro-1,3,5-trinitro-1,3,5-triazine on soil bacterial communities. FEMS Microbiol Ecol 43:255–262
Kane MD, Poulsen LK, Stahl DA (1993) Monitoring the enrichment and isolation of sulfate-reducing bacteria by using oligonucleotide hybridization probes designed from environmentally derived 16S rRNA sequences. Appl Environ Microbiol 59:682–686
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, Chichester, pp 115–175
Lee LK, He J (2010) Reductive debromination of polybrominated diphenyl ethers by anaerobic bacteria from soils and sediments. Appl Environ Microbiol 76:794–802
Lerman LS, Silverstein K, Fripp B, Sauer P, Dresselhaus C (1996) http://web.mit.edu.libproxy1.nus.edu.sg/osp/www/melt.html
Liu Z, DeSantis TZ, Andersen GL, Knight R (2008) Accurate taxonomy assignments from 16S rRNA sequences produced by highly parallel pyrosequencers. Nucleic Acids Res 36:e120. doi:10.1093/nar/gkn491
Löffler FE, Champine JE, Ritalahti KM, Sprague SJ, Tiedje JM (1997) Complete reductive dechlorination of 1,2-dichloropropane by anaerobic bacteria. Appl Environ Microbiol 63:2870–2875
Metzker ML (2010) Sequencing technologies—the next generation. Nat Rev Genet 11:31–46
Muyzer G, De Waal EC, Uitterlinden AG (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59:695–700
Nakasaki K, le Tran TH, Idemoto Y, Abe M, Rollon AP (2009) Comparison of organic matter degradation and microbial community during thermophilic composting of two different types of anaerobic sludge. Bioresour Technol 100:676–682
Nocker A, Burr M, Camper AK (2007) Genotypic microbial community profiling: a critical technical review. Microb Ecol 54:276–289
Paster BJ, Falkler WA Jr, Enwonwu CO, Idigbe EO, Savage KO, Levanos VA, Tamer MA, Ericson RL, Lau CN, Dewhirst FE (2002) Prevalent bacterial species and novel phylotypes in advanced noma lesions. J Clin Microbiol 40:2187–2191
Reysenbach AL, Wickham GS, Pace NR (1994) Phylogenetic analysis of the hyperthermophilic pink filament community in Octopus Spring, Yellowstone National Park. Appl Environ Microbiol 60:2113–2119
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Wu JH, Hong PY, Liu WT (2009) Quantitative effects of position and type of single mismatch on single base primer extension. J Microbiol Methods 77:267–275
Yoshida N, Ye L, Baba D, Katayama A (2009) A novel Dehalobacter species is involved in extensive 4,5,6,7-tetrachlorophthalide dechlorination. Appl Environ Microbiol 75:2400–2405
Acknowledgments
This study was supported by Singapore Agency for Science, Technology and Research (A*STAR) of the Science and Engineering Research Council under Project No: 102 101 0025.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, S., He, J. Two-step denaturing gradient gel electrophoresis (2S-DGGE), a gel-based strategy to capture full-length 16S rRNA gene sequences. Appl Microbiol Biotechnol 95, 1305–1312 (2012). https://doi.org/10.1007/s00253-012-4251-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-012-4251-5