Abstract
Native proinsulin (PI) belongs to the class of the difficult-to-express proteins in Escherichia coli. Problems mainly arise due to its high proteolytic decay and troubles to reproduce the native disulphide pattern. In the present study, human PI was produced in E. coli as a fusion thioredoxin protein (Trx-PI). Such chimeric protein was obtained from the intracellular soluble fraction, and it was purified in one step by affinity chromatography on immobilized phenylarsine oxide. Trx-PI was also recovered from inclusion bodies and purified by anion exchange chromatography. The product identity and integrity were verified by mass analysis (22,173.5 Da) and mapping with Staphylococcus aureus V8 protease. Native PI folding was evaluated by biochemical and also by immunochemical analysis using specific sera from PI antibody-positive diabetic patients that recognise conformational discontinue epitopes. Dose–response curves showed identity between standard PI and Trx-PI. Moreover, surface plasmon resonance technique verified the correct conformation of the recombinant protein. The biochemical and immunochemical assays demonstrated the integrity of the chimera and the epitopes involved in the interaction with antibodies. In conclusion, it was possible to obtain with high-yield purified human PI as a fusion protein in E. coli and useful for analytical purposes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nearly 0.7% of the world population suffers from insulin-dependent diabetes with a continuously increasing number of patients. Consequently, the requirement for recombinant insulin for treatment is also increasing. On the other hand, relatively large amounts of properly folded insulin precursor, proinsulin (PI), are needed for various reasons. One of them is to produce reliable non-radiometric immunochemical tests for the large-scale proinsulin/insulin autoantibody (PAA/IAA) screening of populations deemed at risk for type 1 diabetes. Another reason is its use for basic studies focused on PI as an autoantigen in early prodromic phase of autoimmune aggression and on its role in humoural specific response eliciting high-affinity PAA/IAA. In this field, it was demonstrated that high-affinity insulin autoantibodies in children at risk for type 1 diabetes were mostly associated with HLA DRB1*0302/*0201, young age of IAA appearance and subsequent progression to multiple islet autoantibodies and insulin dependence (Achenbach et al. 2004).
In the human pancreas, insulin is produced as a single polypeptide chain, PI, with the A-chain and B-chain joined by the connecting peptide (C-peptide) (Mackin 1998). During in vivo synthesis, the native folding and concomitant disulphide bond formation is achieved, and then the PI is converted to insulin by enzymatic processing whereby the C-peptide is cleaved off. In addition, PI cleavage can be performed in vitro with trypsin and carboxypeptidase B (Kemmler et al. 1971).
Mature human insulin cannot be effectively produced in its native conformation in prokaryotic host cells using recombinant techniques, essentially because correct disulphide bond formation is only favoured in islet beta cells context during PI biosynthesis. Despite this, different strategies have been described in order to produce PI as soluble form or as inclusion bodies in Escherichia coli (Sung et al. 1986; Kang and Yoon 1991; Tang and Hu 1993; Winter et al. 2000; Mergulhão et al. 2004). Although the PI molecules as inclusion bodies were produced in reasonable high yield, the complex recovery procedures needed for the correct formation of the disulphide bonds during the folding process and the consecutive purification steps applied were considered as difficult and critical cost factors. Alternatively, PI has been produced as a secreted protein in E. coli by routing the recombinant protein to the periplasmic space using appropriate signal sequences. In this case, however, the yield of correctly folded PI was very low compared to that obtained by intracellular production (Talmadge et al. 1981; Chan et al. 1981; Winter et al. 2000; Malik et al. 2007). The first report of an efficient secretory expression of a modified PI in E. coli was published by Kang and Yoon (1994). The authors generated a so-called ZZ-proinsulin fusion construct in which the C-peptide was either totally deleted or drastically shortened (only 1–11 residues remained), which resulted in significantly increased expression yields. A similar approach was recently reported using unmodified PI fused to zero, one or two Z domains, to evaluate potential bottlenecks in PI secretion (Mergulhão et al. 2004). Other strategy used by Winter et al. (2000) was to fuse PI to full-length DsbA.
On the other hand, alternative expression systems such as Bacillus subtilis, Streptomyces lividans and Saccharomyces cerevisiae were used to secret PI to the culture medium (Thim et al. 1986; Koller et al. 1989; Novikov et al. 1990; Kjeldsen 2000; Olmos-Soto and Contreras-Flores 2003), but yields reached were not significantly higher than those obtained in E. coli. So, production of native PI in E. coli is still a challenging task. Beside the above-mentioned limitations, native PI is a quite difficult to express protein, mainly due to severe proteolysis after production in the cytoplasm (Talmadge and Gilbert 1982). The efficient folding in vivo of a recombinant protein produced in E. coli affects directly on stability against its degradation. Upon secretion into the periplasm, the half-life of PI increased from 2 to 20 min (Talmadge and Gilbert 1982). The oxidizing environment of the periplasm also leads to the formation of proper disulphide bonds in PI. However, the yield is very low as compared to production in the cytoplasm (Talmadge et al. 1981; Chan et al. 1981).
In this study, we have investigated the intracellular production of PI in E. coli as an N-terminal chimera with thioredoxin. Such thioredoxin fusion protein (Trx-PI) was designed to accomplish several objectives: (a) to efficiently protect PI from cytoplasmic proteases, (b) to achieve the correct folding of the protein and (c) to easily purify the final product. In addition, it is accepted that the higher solubility of fusion proteins can also induce higher refolding yield (Samuelsson et al. 1994; Tikhonov et al. 2001). Accordingly and as it is detailed in this paper, Trx-PI was produced as a soluble and properly folded protein, checked by biochemical and physical procedures. Moreover, the native and fully immunoreactive conformation of the PI moiety in the chimera was confirmed by using a rabbit anti-PI serum and sera from 30 autoimmune PAA-positive diabetic patients, which mostly recognise native conformational discontinuous epitopes. On the other hand, Trx-PI was recovered from inclusion bodies in order to improve its yielding. Further significant improvements in the production of the fusion protein could be obtained by optimisation of the bacterial growth conditions. Our findings provide a new example of a protein that is successfully expressed as a thioredoxin fusion protein presenting the immunochemical behaviour of the native protein (Georgiou and Valax 1996; Papouchado et al. 1997).
Materials and methods
Reagents
PI and insulin standards were a gift from Eli Lilly (Indianapolis, IN, USA). Restriction enzymes, Pfu polymerase, S1 nuclease and T4 ligase were from Promega (Madison, WI, USA). Dialysis membrane, with a cutoff of 3.5 kDa, was from Spectrum Medical Industries, Inc. (Houston, TX, USA). Acetonitrile and trifluoroacetic acid were high-performance liquid chromatography (HPLC) grade. All other reagents were analytical grade.
Human sera collection
Sera were obtained from 30 selected type 1 diabetic patients spanning a wide range of PAA reactivity. These sera corresponded to children and adolescents admitted to the Nutrition Service at the J. P. Garrahan Pediatric Hospital (Buenos Aires, Argentina), with a mean age of 8.31 ± 4.20 at diagnosis. Serum samples were collected before or within 72 h of insulin treatment initiation. Type 1 diabetes was diagnosed according to WHO criteria (Anonymous 1985). Control sera were from 30 healthy subjects without personal or family history of autoimmune disease. The reported investigation has been performed with the approval of the Ethical Committees of J. P. Garrahan Pediatric Hospital and José de San Martín Clinical Hospital, Buenos Aires, Argentina.
Rabbit PI antibodies
PI antibodies were obtained by immunizing two New Zealand White rabbits with 0.1 mg of standard PI emulsified in complete Freund’s adjuvant. The initial injection was followed by booster injections with 0.1 mg of PI in incomplete Freund’s adjuvant at 3-week intervals. The rabbits were bled 15 days after booster dose. The immunoreactivity of the polyclonal antibodies was tested by ELISA, using PI-coated polystyrene plates, and also by western blot. The antibody titre of both sera (HPI-1 and HPI-2) was higher than 1/10,000 by ELISA and higher than 1/1,000 by western blot. When ELISA plates were coated with insulin, titres of HPI-1 and HPI-2 were lower than 1/100, indicating that both sera were specific for PI. That result was in agreement with western blot analysis when using insulin as antigen.
Expression vector
Unless otherwise indicated, standard molecular biology protocols were used according to Sambrook et al. (1989). pBR328, harbouring full-length preproinsulin (PPI) cDNA (kindly provided by Graeme Bell, University of Chicago, Chicago, IL, USA), was digested with EcoRI. The restriction fragment encoding PPI was cloned into pGem 3Zf (Promega, Madison, WI, USA), yielding pGem3Zf-PPI. The human PI gene was amplified by PCR from pGem3Zf-PPI using 5′ CCCAGCCATGGCCTTTGTGAACCAACACCTGT 3′ and 5′ TTTATTCGAGCTCTCTCGGTGCAGGAGGCG 3′ primers. The amplification product incorporated NcoI and SacI sites at the 3′ and 5′ ends, respectively. The PCR product was ligated into the SmaI site of pGem3Zf to yield pGem3Zf-PI. The identity of the new DNA molecule encoding for PI was corroborated by sequencing.
pGem3Zf-PI was digested with NcoI and EcoRI and treated with S1 nuclease. The fragment containing the PI gene, with blunt 5′ and 3′ ends, was isolated and ligated into linearized pTrxFus (Invitrogen, Carlsbad, CA, USA). The plasmid was linearized with SmaI. The resulting construct pTrx-PI codes for the fusion protein thioredoxin PI (Fig. 1). The vector pTrxFus is used to create C-terminal fusions to E. coli thioredoxin. Foreign genes inserted into the multiple cloning site of the expression vector are expressed as amino terminal fusions to the E. coli protein thioredoxin (trxA). This vector includes an enterokinase cleave site that allows release of the native protein from Trx if wanted. To drive expression of thioredoxin fusions, pTrxFus uses the P L promoter from the bacteriophage λ and aspA transcription terminator.
Transformed E. coli cultures and protein expression
E. coli GI 724 was transformed with pTrx-PI. Bacteria were cultured at 30 °C in 0.2% casein amino acids, 0.5% glucose, 1 mM MgCl2 and 100 μg/ml ampicillin, and protein expression was induced with 100 μg/ml tryptophan for 3 h at 37 °C.
E. coli disruption and intracellular protein isolation
Total cell lysate
Bacteria from a 0.5-ml culture were collected by centrifugation and suspended in 0.2 ml of sodium dodecyl sulphate–polyacrylamide gel electrophoresis (SDS-PAGE) sample buffer (0.05 M Tris/HCI, 12% glycerol, 0.05% bromophenol blue, 4% SDS, 4% 2-mercaptoethanol, pH 6.8).
Intracellular soluble fraction
Bacteria from 200 ml culture were collected by centrifugation, suspended in 4.0 ml of lysis buffer I (50 mM sodium phosphate, 100 mM NaCl, pH 7) and sonicated in the presence of protease inhibitors (0.1% aprotinin and 2 mM phenylmethylsulphonyl fluoride). After sonication, Triton X-100 was added to a final concentration of 0.1%, and the mixture was incubated for 10 min at 0 °C. The soluble intracellular fraction was separated by centrifugation at 12,000 × g for 15 min at 4 °C.
Inclusion bodies
Inclusion bodies from 200 ml of culture were washed with 2 M urea in Tris 0.1 M, pH 8.5, and solubilized with 5 M urea in Tris 0.1 M, pH 8.5. Oxidative refolding was initiated by dialysis at 4 °C against 0.5 M l-arginine, 50 mM Tris–HCl, 5 mM EDTA, 5 mM reduced glutathione and 0.5 mM oxidized glutathione, pH 9.5 (Valdez et al. 2003).
Western blotting for recombinant Trx-PI
Total E. coli lysates, intracellular soluble proteins and the inclusion bodies fraction were analysed by SDS-PAGE (Shägger and von Jagow 1987) and western blotting. For comparison, all SDS-PAGE lanes in each gel contained proteins recovered from the same amount of cells. Protein bands were transferred to nitrocellulose membranes. Unoccupied binding sites in the membranes were blocked by overnight incubation at 4 °C with 3% skim milk in 0.05 M Tris/HCl, pH 7.5, and 0.15 M NaCl (Tris/NaCl). After three washing steps with 0.05 M Tris/NaCl, 0.05% Tween 20, membranes were incubated for 1.5 h at room temperature with polyclonal sera to PI (HPI-2) diluted 1:100 in Tris/NaCl, 0.05% Tween 20 and then washed five times with 0.05 M Tris/NaCl, 0.05% Tween 20. Bound antibodies were visualized by incubation with peroxidase-conjugated goat antibodies to rabbit IgG (Jackson ImmunoResearch Laboratories, Inc., West Grove, PA, USA) followed by the addition of a-chloronaphthol (Sigma-Aldrich, Inc., St. Louis, MO, USA).
Purification of Trx-PI
Trx-PI from intracellular soluble fraction was purified by means of affinity chromatography. Briefly, the resin was based on agarose support covalently modified with phenylarsine oxide, designed for affinity purification of proteins containing vicinal dithiols (Hoffman and Lane 1992). The resin equilibrated in lysis buffer (2 ml) was added to the lysate (4 ml), and the suspension was incubated for 1.5 h at 4 °C. The resin was poured into a column (0.7 × 9.0 cm) and washed sequentially with six-column volume (CV) lysis buffer containing 1 mM 2-mercaptoethanol and three-CV lysis buffer containing 5 mM 2-mercaptoethanol. Bound proteins were eluted with two-CV lysis buffer containing 100 mM 2-mercaptoethanol.
After overnight refolding of solubilized inclusion bodies, Trx-PI was isolated by FPLC (GE Healthcare, Sweden) on a 1.5 × 5-cm Q-Sepharose column (GE Healthcare, Sweden). Buffer A was 20 mM Tris–HCl pH 8.5, and buffer B was 20 mM Tris–HCl and 1 M NaCl, pH 8.5. A flow rate of 1 ml/min was employed, and detection was performed by measuring absorbance at 280/215 nm. Trx-PI was eluted at 30% buffer B.
Analysis and biochemical characterization of Trx-PI
The purified Trx-PI was subjected to ion-spray ionization mass spectrometry analysis (Laboratorio Nacional de Investigación y Servicios en Péptidos y Proteínas (LANAIS)-CONICET-UBA). Briefly, the protein was analysed by reversed-phase (RP)-HPLC mass spectrometry using a 1.00 × 30-mm Vydac C8 column, operating at 40 μl/min, and connected to Surveyor HPLC System online with an LCQ Duo (ESlion trap) mass spectrometer (Thermo Fisher, San José, CA, USA). Then, it was eluted using a 15-min gradient from 10% to 100% solvent B (solvent A—2% acetic acid, 2% acetonitrile; solvent B—2% acetic acid, 96% acetonitrile). Protein molecular mass assay was performed by Full Scan 300–2000 amu and ProMass Deconvolution program. The needle voltage was 4.5 V and the capillary temperature was 190 °C, approximate error 0.06%. To determine the proportion of disulphide bridges correctly formed, a proteolytic digestion was performed using Staphylococcus aureus protease V8. Thirty micrograms of both standard PI and Trx-PI were incubated overnight with 5 μg of protease, whereas no protein was added to the control. Another 30 μg of Trx-PI were treated with 0.2 M DTT after V8 digestion. The reactions were stopped by freezing at −20 °C. The samples were then analysed by RP-HPLC using a Vydac C18 column equilibrated in 0.07% TFA and 90% acetonitrile. Bound material was eluted using a linear gradient of increasing acetonitrile concentration (up to a maximum of 90%). Chromatograms from samples reduced with DTT and those non-reduced were compared, and relevant peaks were selected and examined by matrix-assisted laser desorption/ionization time-of-flight/time-of-flight analysis (MALDI-TOF/TOF spectrometer; UltraflexII, Bruker; Centro de Estudios Químicos y Biológicos por Espectrometría de Masa (CEQUIBIEM)-CONICET-UBA, Argentina).
To further characterize the peptide structure involving the disulphide bonds of Trx-PI, the reduced and non-reduced protein was subjected to chymotrypsin digestion. A proportion of Trx-PI was first reduced with 0.2 M DTT in urea 6 M, pH 8.2. After 1 h of incubation at room temperature, 5 μl of iodoacetamide 1 M was added in the dark at room temperature for 1 h. The reaction was stopped with 5 μl TFA. The reduced and non-reduced protein was then digested with a 1/25 chymotrypsin/Trx-PI ratio at room temperature overnight. The products of the digestion were analysed by MALDI-TOF-TOF spectrometer (CEQUIBIEM-CONICET-UBA, Argentina).
Immunochemical characterization of Trx-PI
PAA in human sera were detected by radioligand binding assay (RBA) using [35S]-PI as tracer, as previously described by Valdez et al. (2003). Results were calculated as B% = 100 × bound cpm/total cpm and expressed as SD score = (B% – B C%)/SDC, where B C% was the mean B% of 30 normal control sera and SDC its standard deviation. An assay was considered positive if SD score >3. The intra-assay CVs in triplicates were 10.34% and 3.78% for a SD score of 17.2 and 33.3, respectively. The ability of Trx-PI to compete with human [35S]-PI was assessed qualitatively by incubating sera from 30 type 1 diabetic patients with the tracer in the presence of 1 μM purified fusion protein, either from intracellular soluble fraction or recover from inclusion bodies.
Quantitative competition assays were performed by a standard radioimmunoassay (RIA). This method was carried out by incubating 30 μl of 1/100 final dilution of the rabbit polyclonal serum against PI or 30 μl of a pool of diabetic patients sera for 7 days at 4 °C with 1,000 cpm of [35S]-PI in the presence of 90 μl of serial concentrations (10 pM to 1 μM) of standard PI or purified Trx-PI from inclusion bodies. Immunocomplexes were isolated with Protein G Sepharose 4B FF, the pellets were washed and suspended in 1% SDS. Radioactivity of supernatants was counted in an automatic beta counter.
Dose–response curves were fitted to the following mathematical function:
where B%min and B%max are the minimal and maximal response, respectively, and the parameter EC50 represents the doses at the 50% of B%max. The identity between curves obtained with standard PI and Trx-PI was analysed by comparing slopes and EC50 values.
Aiming to further evaluate the proper folding of epitopes in Trx-PI, specific antigen-antibody interaction was analysed using surface plasmon resonance (SPR) technique (BIAcore, GE Healthcare, Uppsala, Sweden). For this analysis, flow cells from the sensor chip were immobilized with standard PI or Trx-PI. Each patient’s serum was used pure or diluted 1/2, 1/4 and 1/8 in running buffer (0.14 M NaCl, 2.7 mM KCl, 1.5 mM KPO4H2, 8.1 mM Na2PO4H, pH 7.4, 0.05% Tween 20). In either case, these starting samples were diluted 1/2 with carboxy-methyl dextran and NaCl to a final concentration of 1 mg/ml and 0.35 M, respectively, in order to eliminate the nonspecific reaction. Each sample was injected over 300 s, and the binding rate in running buffer was measured after 300 s. Assays were carried out at 20 °C using a flow rate of 10 μl/min. The association rate constant (k 1), the dissociation rate constant (k −1) and the equilibrium affinity constant (K a) were calculated from the sensorgram analysis using the BIAevaluation software.
Statistical analysis
ANOVA tests were used to analyse parallelism between slopes of two regression lines corresponding to dose–response curves. When the P value obtained was greater than 0.05, the test concluded that there were no significant differences between slopes (Real Farmacopea Española 1997).
Differences between EC50 were evaluated by the Mann–Whitney U test for comparison of the results derived from dose–response curves. P values less than 0.1 were considered as significant.
Inter-assay correlation was assessed by standard linear regression.
Results
Proinsulin expression as a fusion protein in E. coli
Recombinant fusion protein Trx-PI was expressed in E. coli strain GI724. Efficient Trx-PI production was achieved with 3-h induction. A western blot analysis of total cell lysates with rabbit specific antisera to PI (HPI-2) (Fig. 2b) showed one band with the expected mass (~22 kDa) for the full-length engineered protein.
Analysis of recombinant Trx-PI fractions. SDS-PAGE (a) and western blot (b) at different stages of purification of Trx-PI from intracellular soluble fraction: lane 1 molecular weight markers, lane 2 total cell lysates of bacteria before induction, lane 3 total cell lysates of Trp-induced bacteria, lane 4 intracellular soluble fraction of Trp-induced bacteria, lane 5 unbound material, lane 6 eluate obtained after 5 mM 2-mercaptoethanol treatment, lane 7 affinity-purified Trx-PI. SDS-PAGE (c) and UV scanner (d) of Trx-PI recovered and purified from inclusion bodies; dotted line represents the raw data, and dashed line represents the data corrected by scattering subtraction
A rough estimation based on the intensity of the PI bands quantified using Scion Image for Windows (Scion Corporation; Beta 4.0.2) indicates that approximately 5–10% of the Trx-PI produced was soluble. No bands were detected in the western blot analysis of the periplasmic fraction or the culture medium, indicating that Trx-PI was not exported significantly (data not shown).
Purification of Trx-PI
Recombinant Trx-PI was purified from the soluble intracellular fraction of bacterial lysates by affinity chromatography on a matrix that binds specifically the active site of thioredoxin (see “Materials and methods”). The single-step purification yielded 1.5 mg 90–95% pure Trx-PI/L culture (Fig. 2a, b).
On the other hand, Trx-PI isolated from inclusion bodies was refolded and purified by anion exchange chromatography (Fig. 2c). Final yield of properly refolded 90–95% pure Trx-PI was 10 mg/L bacterial culture, as determined by absorbance at 280 nm (Fig. 2d). Since the majority of fusion protein was present as inclusion bodies, these were selected as a more viable option for the biochemical and immunochemical characterization.
Physical and biochemical characterization of Trx-PI
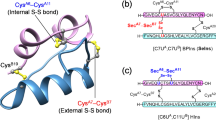
Mass spectrometry analysis of Trx-PI verified the agreement of the molecular mass of our expressed chimera with the expected value (actual mass 22,173.5 Da; theoretical mass 22,174.2 Da), indicating that the product is in fact Trx-PI and that all disulphide bonds were formed. To further test whether the recombinant Trx-PI had properly refolded, a proteolytic digestion was performed using S. aureus protease V8 (Fig. 3). Standard PI was used as a reference product, and the digested peptides were subjected to RP-HPLC. The analysis demonstrated that V8 digestion of both the standard and recombinant human PI produced comparable chromatograms (Fig. 4). The two peaks suspected of having peptides from the PI moiety in Trx-PI with hemicystine residues (peptides 6 and 7) which disappeared when injecting the reduced products were collected and analysed by MALDI-TOF. One of them (peptide 7, theoretical mass 1,378 Da) was successfully characterized being consistent with the formation of the expected disulphide bond between Cys140 and Cys206 (Table 1). No material with a mass corresponding to the other peptide of interest (peptide 6; theoretical mass 4,845 Da, Cys128–Cys193 and Cys192–Cys197) could be detected, not even using sinapinic acid matrix, probably due to its high molecular size.
To extra characterize the formation of correct disulphide bonds, additional analysis was done by subjecting Trx-PI to chymotrypsin digestion with or without reduction and blocking and analysing the peptides by MALDI-TOF. This procedure confirmed once again the formation of disulphide bond between Cys140 and Cys206 (Table 1). In addition, the presence of a peak of 3,869 Da (Table 1) which disappeared when Trx-PI was reduced and carbamidomethylated supported the peptide structure involving the disulphide bonds between Cys128–Cys193 and Cys192–Cys197. Moreover, when reduced and blocked, the identity of peptides 123–137 containing Cys128 was verified by fragmentation.
Immunochemical characterization of Trx-PI
To test the ability of Trx-PI to react with PAA, radio competition assays were performed. Sera from 30 type 1 diabetic patients that scored positive in the RBA (Valdez et al. 2003) were used. All sera tested were PAA-positive with a SD score 11.64 ± 8.35 (mean ± SD, range 3.58 to 35.85, cutoff value for positivity SD score = 3.00). The assay was carried out in parallel in the presence of 1 μM Trx-PI from intracellular soluble fraction or Trx-PI refolded from inclusion bodies. All 30 positive sera became virtually negative under this condition of cold antigen excess, with a SD score of 1.01 ± 1.22 (mean ± SD, range 0.77 to 3.52) (Fig. 5).
Inhibition capacity of Trx-PI in the RBA of 30 PAA-positive sera: a binding of [35S]PI to control human sera, b binding to PAA-positive sera, c binding to PAA-positive sera in the presence of 1 μM unlabeled Trx-PI from intracellular soluble fraction and d binding to PAA-positive sera in the presence of 1 μM unlabeled Trx-PI from inclusion bodies. Binding is expressed as SD scores; the cutoff value is shown as a dotted line
In order to study the immunochemical behaviour of Trx-PI compared to standard PI, dose–response curves were performed with high titred anti-PI rabbit HPI-2 serum or with a pool of sera from PAA-positive diabetic patients (Fig. 6). The immunochemical identities between both proteins were analysed and expressed in terms of parallelism and the similarity of EC50 values, as described in “Materials and methods”. Variance analyses of the data were carried out by parallel line model and completely randomized design. When using either rabbit HPI-2 serum or PAA + human sera, parallelism between curves was achieved (p<0.05 and p<0.1, respectively), and the EC50 showed no significant differences (Table 2).
To further assess the proper folding of the recombinant PI chimera, SPR technology was used. When injecting PAA-positive sera from diabetic patients to the flow cell on the chip with immobilized Trx-PI, the K a values calculated did not show significant difference with those obtained with standard PI (mean 1.2 ± 1.6 × 108 vs 1.1 ± 2.2 × 108 M−1, respectively; p < 0.05). Moreover, there was a good correlation (r 2 0.80) between results achieved with both immobilized antigens (Fig. 7).
Discussion
E. coli is the prokaryotic host for the production of recombinant proteins most widely used by diverse research groups and also in private industries (Baneyx 1999; Pines and Inouye 1999). This expression system is simple, versatile due to the high amount of cloning vectors and host strains available, and implies a rapid generation of biomass at low cost. Disulphide bonds are essential for the correct folding and stability of secreted proteins, most of which are frequently used in therapy and diagnosis. However, most proteins form inclusion bodies when they are expressed in the cytoplasm of E. coli. This could be an advantage for ease purification when in vitro refolding leads to the formation of the correct disulphide bonds.
Alternatively, the protein can be directed to the periplasm and will be natively folded due to the oxidizing conditions in this compartment. A major drawback of this later approach is the limited secretion capacity of the E. coli and the limited space of the periplasmic compartment.
It is not in the nature of E. coli to secret high amounts of proteins (Francetic et al. 2000), while the transport to the periplasm or to the culture medium implies very complex processes (Economou 1999; Pugsley et al. 2004). Thus, the presence of a signal peptide is essential but does not guarantee translocation of proteins to the periplasmic compartment or to the extracellular culture medium (Kajava et al. 2000; Khokhlova and Nesmeianova 2003). Previous attempts by other investigators had suggested that bacterial expression of PI resulted in very poor yields due to N-terminal degradation (Shen 1984). An alternative approach is the use of a larger fusion partner which may lead to efficient translocation and increases the chances of correct folding. Our approach involved the production of soluble cytoplasmic PI by means of the apparently general solubilizing effect of the thioredoxin moiety (LaVallie et al. 1993). Although the exact mechanism by which this happens is not known, thioredoxin has been shown to facilitate the soluble expression of several cytokines, peptides and other proteins (Chopra et al. 1994; Wilkinson et al. 1995; Papouchado et al. 1997).
In this work, Trx-PI was successfully expressed in the intracellular soluble fraction of E. coli GI724 at 37 °C. The thioredoxin moiety allowed the one-step purification by an arsine derivative affinity chromatography yielding 1.5 mg of protein/L of culture medium with 90–95% of purity. In addition, Trx-PI was recovered from inclusion bodies, subjected to in vitro refolding and purified by anion exchange chromatography, yielding 10 mg of protein/L of culture medium, 90–95% pure. Because the majority of the chimera was present as inclusion bodies, these were selected as a more viable option for biochemical and immunochemical characterization.
Our first quality control step was to verify if Trx-PI had the theoretical expected molecular mass by means of mass spectrometry. This analysis indicated that the fusion protein was expressed with all its disulphide bonds formed. To assure that proper refolding and the correct disulphide bond formation of our recombinant protein had occurred, S. aureus V8 protease was then used for peptide mapping. After V8 digestion of Trx-PI, there should be two peptides suspected to have hemicystine residues (Fig. 3). Combined RP-HPLC and MALDI-TOF-TOF analysis identified one of these peptides (peptide 7, theoretical mass 1,378 Da, actual mass 1,377 Da). The presence of the disulphide bond was confirmed by reduction and modification with iodoacetamide (data not shown). The other peptide of interest (peptide 6; theoretical mass 4,845 Da) could not be detected, not even using sinapinic acid matrix probably because of its high molecular weight that hindered the analysis. The peptide structure involving the disulphide bonds between Cys128–Cys193 and Cys192–Cys197 was characterized by digesting the chimera with chymotrypsin. With this approach, a peptide of 3,869 Da (aa 123–137 + 184–202) was identified in non-reduced protein, which disappeared when evaluating the reduced form. This was further confirmed with fragmentation of the peptide formed by the aa 123–137.
Taking into account the results obtained from mass spectrometry of the entire protein together with the successful characterization of the peptides from the V8 and chymotrypsin digestions, we assume that we have expressed genuine human PI as a fusion protein with thioredoxin. These results were further confirmed by the immunochemical behaviour of Trx-PI compared with that of standard PI.
Considering that most sera from diabetic patients only target conformational discontinue epitopes of PI, native conformation of Trx-PI was evaluated by means of 30 PAA-positive sera from type 1 diabetic patients. All 30 positive sera became virtually negative when incubated in the presence of an excess of Trx-PI either from intracellular soluble fraction or Trx-PI refolded from inclusion bodies.
The dose–response curves generated by RIA with the polyclonal rabbit HPI-2 serum and with the pool of diabetic patient sera showed identity between the standard PI and Trx-PI (Table 2 and Fig. 6) according to the general principles of immunochemical cross-reactivity (Berzofsky and Schechter 1981). In order to establish the conformation of native PI, the use of a RIA with a pool of polyclonal antibodies and the analysis of identity between standard PI and Trx-PI are a more adequate and complete alternative than the use of a sandwich ELISA using two monoclonal antibodies (Malik et al. 2007). By means of a pool of polyclonal antibodies, virtually all immunodominant epitopes existing in the antigen are mapped, while monoclonal antibodies only map one specific epitope.
Another way to evaluate the proper folding of the chimera was using SPR technology. By this methodology, it is possible to determine kinetic parameters of antigen–antibody interaction. Herein we used PAA-positive sera from diabetic patients, which were injected to the flow cell immobilized with standard PI or Trx-PI, and results were compared between them. As shown in Fig. 7, there is a good correlation (r 2 0.80) between both immobilized antigens. This is another evidence that demonstrates the correct conformation of the recombinant protein.
We have previously assayed cleaving Trx, but the yield of protein recovery was too low. Besides, taking into account that our final aim is to use the chimera as antigen in SPR technique where the immobilization of proteins on CM5 sensor chip is based on covalent binding between carboxyl groups of the chip surface and amino groups of the proteins, we decided not to cleave Trx from PI. The goal of this approach was to improve the orientation of PI assuring that all epitopes were better exposed to antibodies.
In conclusion, human PI was expressed in E. coli as a soluble and properly folded fusion protein with thioredoxin. Additionally, it was possible to improve its yielding by recovering the chimera from inclusion bodies. Its structure and proper refolding and native conformation were demonstrated by physical, biochemical and immunochemical studies. Our approach provides a solution to the main problems that had hindered the expression of recombinant human PI in E. coli. By fusing the PI to thioredoxin, the solubility of the product was greatly increased, and a significant fraction of the protein entered a native folding pathway. In addition, Trx-PI was far less prone than PI to proteolysis in the cytoplasm. Also, the refolding in vitro of the chimera was assisted by the presence of thioredoxin. The strategy reported herein should be accessible to many laboratories and should allow the easy production of Trx-PI. In turn, the availability of properly folded PI would encourage researchers to improve current and develop new immunochemical tests for PAA/IAA assessment for routine screening of individuals at risk of type 1 diabetes mellitus.
References
Achenbach P, Koczwara K, Knopff A, Naserke H, Ziegler A-G, Bonifacio E (2004) Mature high-affinity immune responses to (pro)insulin anticipate the autoimmune cascade that leads to type 1 diabetes. J Clin Invest 114(4):589–597
Anonymous (1985) Diabetes mellitus. Report of a WHO Study Group. World Health Organ Tech Rep Ser 727:1–113
Baneyx F (1999) Recombinant protein expression in Escherichia coli. Curr Opin Biotechnol 10(5):411–421
Berzofsky JA, Schechter AN (1981) The concepts of crossreactivity and specificity in immunology. Mol Immunol 18(8):751–763
Chan SJ, Weiss J, Konrad M, White T, Bahl C, Yu SD, Marks D, Steiner DF (1981) Biosynthesis and periplasmic segregation of human proinsulin in Escherichia coli. Proc Natl Acad Sci U S A 78(9):5401–5405
Chopra AK, Brasier AR, Das M, Xu XJ, Peterson JW (1994) Improved synthesis of Salmonella typhimurium enterotoxin using gene fusion expression systems. Gene 144(1):81–85
Economou A (1999) Following the leader: bacterial protein export through the Sec pathway. Trends Microbiol 7(8):315–320
Francetic O, Belin D, Badaut C, Pugsley AP (2000) Expression of the endogenous type II secretion pathway in Escherichia coli leads to chitinase secretion. EMBO J 19(24):6697–6703
Georgiou G, Valax P (1996) Expression of correctly folded proteins in Escherichia coli. Curr Opin Biotechnol 7(2):190–197
Hoffman RD, Lane MD (1992) Iodophenylarsine oxide and arsenical affinity chromatography: new probes for dithiol proteins. Application to tubulins and to components of the insulin receptor–glucose transporter signal transduction pathway. J Biol Chem 267(20):14005–14011
Kajava AV, Zolov SN, Kalinin AE, Nesmeyanova MA (2000) The net charge of the first 18 residues of the mature sequence affects protein translocation across the cytoplasmic membrane of gram-negative bacteria. J Bacteriol 182(8):2163–2169
Kang Y, Yoon JW (1991) Development of a high-expression vector (Pyk 10–9) of human proinsulin gene. Biotechnol Lett 13:755–760
Kang Y, Yoon JW (1994) Effect of modification of connecting peptide of proinsulin on its export. J Biotechnol 36(1):45–54
Kemmler W, Peterson JD, Steiner DF (1971) Studies on the conversion of proinsulin to insulin. I. Conversion in vitro with trypsin and carboxypeptidase. B J Biol Chem 246(22):6786–6791
Khokhlova OV, Nesmeianova MA (2003) Interaction of SecB and SecA with the N-terminal region of mature alkaline phosphatase on its secretion in Escherichia coli. Mol Biol (Mosk) 37(4):712–718
Kjeldsen T (2000) Yeast secretory expression of insulin precursors. Appl Microbiol Biotechnol 54(3):277–286
Koller DP, Rieb G, Sauber K, Uhlmann E, Wallmeier H (1989) Recombinant Streptomyces lividans secretes a fusion protein of tendamistat and proinsulin. Biotechnology (NY) 7:1055–1059
LaVallie ER, DiBlasio EA, Kovacic S, Grant KL, Schendel PF, McCoy JM (1993) A thioredoxin gene fusion expression system that circumvents inclusion body formation in the E. coli cytoplasm. Biotechnology 11(2):187–193
Mackin RB (1998) Proinsulin: recent observations and controversies. Cell Mol Life Sci 54(7):696–702
Malik A, Jenzsch M, Lübbert A, Rudolph R, Söhling B (2007) Periplasmic production of native human proinsulin as a fusion to E. coli ecotin. Protein Expr Purif 55(1):100–111
Mergulhão FJ, Taipa MA, Cabral JM, Monteiro GA (2004) Evaluation of bottlenecks in proinsulin secretion by Escherichia coli. J Biotechnol 109(1–2):31–43
Novikov AA, Borukhov SI, Strongin AY (1990) Bacillus amyloliquefaciens alpha-amylase signal sequence fused in frame with human proinsulin is properly processed by Bacillus subtilis cells. Biochem Biophys Res Commun 169(1):297–301
Olmos-Soto J, Contreras-Flores R (2003) Genetic system constructed to overproduce and secrete proinsulin in Bacillus subtilis. Appl Microbiol Biotechnol 62(4):369–373
Papouchado ML, Valdez SN, Ghiringhelli D, Poskus E, Ermacora MR (1997) Expression of properly folded human glutamate decarboxylase 65 as a fusion protein in Escherichia coli. Eur J Biochem 246(2):350–359
Pines O, Inouye M (1999) Expression and secretion of proteins in E. coli. Mol Biotechnol 12(1):25–34
Pugsley AP, Francetic O, Driessen AJ, de Lorenzo V (2004) Getting out: protein traffic in prokaryotes. Mol Microbiol 52(1):3–11
Real Farmacopea Española (1997) Análisis Estadístico, Segunda edn. Ministerio de Sanidad y Consumo, Madrid
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbour Laboratory, Cold Spring Harbour
Samuelsson E, Moks T, Nilsson B, Uhlen M (1994) Enhanced in vitro refolding of insulin-like growth factor I using a solubilizing fusion partner. Biochemistry 33(14):4207–4211
Shägger H, von Jagow G (1987) Tricine–sodium dodecyl sulfate–polyacrylamide gel electrophoresis for the separation of proteins in the range from 1 to 100 kDa. Anal Biochem 166:368–379
Shen SH (1984) Multiple joined genes prevent product degradation in Escherichia coli. Proc Natl Acad Sci U S A 81(15):4627–4631
Sung WL, Yao FL, Zahab DM, Narang SA (1986) Short synthetic oligodeoxyribonucleotide leader sequences enhance accumulation of human proinsulin synthesized in Escherichia coli. Proc Natl Acad Sci U S A 83(3):561–565
Talmadge K, Brosius J, Gilbert W (1981) An ‘internal’ signal sequence directs secretion and processing or proinsulin in bacteria. Nature 294(5837):176–178
Talmadge K, Gilbert W (1982) Cellular location affects protein stability in Escherichia coli. Proc Natl Acad Sci U S A 79(6):1830–1833
Tang JG, Hu MH (1993) Production of human proinsulin in Escherichia coli in a non-fusion form. Biotechnol Lett 15:661–666
Thim L, Hansen MT, Norris K, Hoegh I, Boel E, Forstrom J, Ammerer G, Fiil NP (1986) Secretion and processing of insulin precursors in yeast. Proc Natl Acad Sci U S A 83(18):6766–6770
Tikhonov RV, Pechenov SE, Belacheu IA, Yakimov SA, Klyushnichenko VE, Boldireva EF, Korobko VG, Tunes H, Thiemann JE, Vilela L, Wulfson AN (2001) Recombinant human insulin. VIII. Isolation of fusion protein--S-sulfonate, biotechnological precursor of human insulin, from the biomass of transformed Escherichia coli cells. Protein Expr Purif 21(1):176–182
Valdez SN, Iacono RF, Villalba A, Cardoso_Landaburu A, Ermacora MR, Poskus E (2003) A radioligand-binding assay for detecting antibodies specific for proinsulin and insulin using 35S-proinsulin. J Immunol Methods 279(1–2):173–181
Wilkinson DL, Ma NT, Haught C, Harrison RG (1995) Purification by immobilized metal affinity chromatography of human atrial natriuretic peptide expressed in a novel thioredoxin fusion protein. Biotechnol Prog 11(3):265–269
Winter J, Neubauer P, Glockshuber R, Rudolph R (2000) Increased production of human proinsulin in the periplasmic space of Escherichia coli by fusion to DsbA. J Biotechnol 84(2):175–185
Acknowledgements
We thank C. Mazza and A.G. Krochik at the J. P. Garrahan National Pediatrics Hospital and the Hemotherapy Division at the José de San Martín Clinical Hospital (Buenos Aires, Argentina) for collecting and providing the sera of diabetic patients and control individuals. We are grateful to Graeme Bell, University of Chicago, USA, for the gift of pBR328–preproinsulin vector. We also thank Eli Lilly for the generous supply of standard proinsulin. This work was supported in part by grants from FONCYT Programme of the Agency for Science and Technology Promotion (ANPCyT), National Research Council (CONICET) and the University of Buenos Aires, Buenos Aires, Argentina.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Trabucchi, A., Guerra, L.L., Faccinetti, N.I. et al. Expression and characterization of human proinsulin fused to thioredoxin in Escherichia coli . Appl Microbiol Biotechnol 94, 1565–1576 (2012). https://doi.org/10.1007/s00253-011-3721-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-011-3721-5