Abstract
This study presents a new coupon sampling device that can be inserted directly into the pipes within water distribution systems (WDS), maintaining representative near wall pipe flow conditions and enabling simultaneous microscopy and DNA-based analysis of biofilms formed in situ. To evaluate this sampling device, fluorescent in situ hybridization (FISH) and denaturing gradient gel electrophoresis (DGGE) analyses were used to investigate changes in biofilms on replicate coupons within a non-sterile pilot-scale WDS. FISH analysis demonstrated increases in bacterial biofilm coverage of the coupon surface over time, while the DGGE analysis showed the development of increasingly complex biofilm communities, with time-specific clustering of these communities. This coupon design offers improvements over existing biofilm sampling devices in that it enables simultaneous quantitative and qualitative compositional characterization of biofilm assemblages formed within a WDS, while importantly maintaining fully representative near wall pipe flow conditions. Hence, it provides a practical approach that can be used to capture the interactions between biofilm formation and changing abiotic conditions, boundary shear stress, and turbulent driven exchange within WDS.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The provision of microbiologically safe supplies of potable water, following treatment, represents one of the cornerstones for maintenance of good public health (Szewzyk et al. 2000). Disinfectant residuals, typically chlorine based, are routinely used to reduce the numbers of microorganisms in water distribution systems (WDS). Nevertheless, increases in microbial numbers during distribution of potable water have long been recognized (Baylis et al. 1930) with microbially mediated processes contributing to the deterioration of water quality. Although disinfection significantly reduces the numbers of planktonic bacteria in WDS, multi-species biofilms form on the internal surfaces of WDS, serving as the primary source of microorganisms in the WDS (Batté et al. 2004; LeChevallier et al. 1987), and acting as refugia for bacteria, including pathogens, against disinfectant residuals (Berry et al. 2006; Gagnon et al. 2005). While numerous abiotic factors will influence biofilm formation in WDS, including temperature, disinfectant type and residuals (Gagnon et al. 2005; Lund and Ormerod 1995), organic matter (Norton and LeChevallier 2000), nutrient concentrations (Chu et al. 2005; Volk and LeChevallier 1999), substratum (Camper et al. 1996), and hydraulics (Lehtola et al. 2006), a holistic understanding of how these factors act in concert to influence and control compositional changes during biofilm formation and detachment within WDS remains a key challenge.
One of the major barriers to studying biofilms within WDS is the lack of suitable experimental systems that both represent conditions within real pipe networks and enable the effects of abiotic factors to be explored in a controlled environment. Ideally, such systems should permit replicate sampling to allow structural (e.g., via microscopy) and compositional (e.g., DNA-based fingerprint and/or sequencing-based) characterization of biofilms over space and time. Several bench-top laboratory biofilm reactor systems, such as the rotating disc reactor (Murga et al. 2001), the CDC biofilm reactor (Goeres et al. 2005), the biofilm annular reactor (Batté et al. 2003a, b), and the Propella™ reactor (Appenzeller et al. 2001), have been used to develop and investigate physico-chemical effects upon biofilms. However, these systems poorly reflect the conditions of real pipe networks and cannot be used to study in situ biofilm formation (see Table 1). Other systems such as the Robbins device (Manz et al. 1993), the modified Robbins device (Kharazmi et al. 1999), the Prévost coupon (Prévost et al. 1998), the Bioprobe monitor (LeChevallier et al. 1998), the pipe sliding coupon holder (Chang et al. 2003), or the biofilm sampler (Juhna et al. 2007) are more amenable to deployment directly within WDS or experimental pipe systems (Table 1). However, all these devices critically alter the flow patterns local to the sampling points, leading to non-representative boundary layer flow conditions (i.e., shear stress and turbulence) which will affect formation and detachment of biofilms and nutrient exchange rates therein.
The aim of this paper therefore is to present the design and application of a new, Pennine Water Group (PWG) coupon that can be deployed within a pilot-scale test facility representative of a full-scale WDS, enabling simultaneous quantitative (microscopy based) and qualitative (DNA-based) compositional characterization of in situ biofilm assemblages, without disruption to the boundary layer flow conditions.
Material and methods
PWG coupon design
PWG coupons (Fig. 1) were cut directly from 90-mm diameter high pressure polyethylene (HPPE) SDR17 pipe, the current UK standard for new and replacement WDS pipes. The coupons are comprised of two parts; an “outer coupon” (Fig. 1A) used for harvesting biomass for nucleic acid extraction and community analyses, and an “insert” (Fig. 1B) which can be removed from within the body of the outer coupon, and used directly for microscopic analysis including quantifying biofilm depth and coverage. Fig. 1C highlights how the coupon and insert fit together. The “outer coupon” retains the curvature of the pipe from which it was removed and fits precisely into an aperture made in a removable and flanged identical pipe section (Fig. 1D). The coupon is fixed with a gasket to a section of 110 mm SDR 11 medium density polyethylene pipe with equal internal diameter of 90 mm (see Fig. 1C). Only the 4.5-mm-wide top surface of the “insert” is machined flat to enable microscopy. This design results in a maximum deviation from curvature of 0.064 mm, which is of the same order of magnitude of the surface roughness coefficient used in hydraulic modeling to represent smooth plastic pipes, and hence minimal planned distortion of the boundary flow layer. The pipe sizes and material used here are suited to the full-scale pilot test facility at the University of Sheffield. Other sizes and materials can be readily machined or formed as the design does not rely on threads, geometries, etc. that are only applicable to one material.
PWG coupons for in situ analysis of biofilms in drinking water distribution systems. Coupons have the same internal diameter and curvature as the distribution pipe and fit flush with the internal pipe surface. A Schematic representation of coupon showing “outer coupon” (surface area 224 mm2) with trapezoidal “insert” (surface area 90 mm2). B Schematic of insert. C Coupon and coupon mounting showing partial insertion of the insert into the outer coupon; coupon mounting includes a sealing gasket. D Coupon mounting within pipe section and E pipe sections with multiple coupon sample points/holes in full-scale laboratory pipe loop. Dimensions as indicated
Pilot-scale WDS and coupon sampling
Trials reported here were conducted on a pilot-scale WDS consisting of a 90-m-long coil of pipe into which coupons were inserted. The facility was supplied via a 1-m3 enclosed reservoir tank, filled from the local WDS, with water re-circulated through the system. During operation there was a gradual turnover of water in the facility from the local WDS. This ensured hydraulic retention times of 48 h within the pipe loop (in addition to the system time to reach the laboratory), maintenance of disinfection residual and nutrient feed consistent within full-scale WDS systems. In contrast to bench scale experiments the larger pipe surface area of the test loop facility also enables sufficient exchange processes and interactions between the bulk fluid and the pipe wall to occur, replicating conditions in a typical WDS (Husband et al. 2008). The system was run at a constant flow rate of 0.4 l s−1 (velocity = 0.08 m s−1, boundary shear stress = 0.03 N m−2, with flow rates up to 8 l s−1 tested) and a pressure of 4 bar (tested up to 7 bar), typical of operating conditions in similar size pipes within UK WDS. Average water temperature during the trials was 25 °C. Coupon removal was facilitated by first closing an upstream valve, removing the pressurizing effects of the system pump, and then closing a downstream valve, to isolate the pipe section containing the coupons. By individual coupon removal, leakage and possible hydraulic disturbance of developing biofilm on the coupons was minimized. Following removal from the facility, coupons were placed in sterile Falcon tubes containing 25 ml of the reservoir water prior to microscopy/molecular analyses, which commenced within 30 min of sampling. Fig. 1E shows application of multiple coupons within a purpose built temperature controlled recirculating, full-scale test loop facility that can be controlled to represent three different time-variable hydraulic regimes. The three individual loops of the system each consist of nine-and-a-half coils (21.4 m long each) of 90-mm diameter HPPE pipe.
Catalyzed reported deposition and fluorescence in situ hybridization analysis and microscopy
Epifluorescent microscopy using 4′,6′-diamidino-2-phenylindole (DAPI) staining was used to visualize biofilm formation on coupon inserts with additional targeting of bacteria via catalyzed reported deposition and fluorescence in situ hybridization (CARD-FISH) analysis (Pernthaler et al. 2002). Briefly, biofilms present on coupon inserts were pre-fixed with 2% (w/v) formaldehyde solution for 12 h at 4 °C, washed with sterile distilled water, and stored at −20 °C until further processing. Cells were permeabilized with lysozyme followed by achromopeptidase as described previously (Pernthaler et al. 2002; Sekar et al. 2003). Hybridization of biofilm samples was performed using pooled 5′HRP-labeled (ThermoHybaid, Interactiva Division, Ulm, Germany) eubacterial (EUB I–III) oligonucleotide probes (Daims et al. 1999), with subsequent hybridization, washing, and tyramide signal amplification conducted as described previously for marine bacterioplankton (Pernthaler et al. 2002). Samples were visualized following counter staining with DAPI using an Olympus BX51 eipfluorescence microscope (Olympus UK Ltd., Watford) with a 100X oil immersion objective lens. Images of FITC and DAPI fluorescence were captured using CellB imaging software (Olympus UK Ltd., Watford, UK) at an XY resolution of 1,360 × 1,024 pixels. Biofilm coverage (% area) was determined via digital image analysis using ImageJ software (National Institutes of Health, USA). One-way analysis of variance (ANOVA) was used to investigate differences in biofilm coverage over time with post-hoc comparison of means using Tukey tests. Statistical analyses were performed using JMPTM, The Statistical Discovery Software, Version 5.0.1.2 (SAS Institute Inc., USA).
DNA extraction
Biofilms present on the “outer coupon” surface were aseptically harvested via three rounds of scraping using a sterile nylon brush following additions of ∼15 ml of sterile distilled water (Sharma et al. 1990). Epifluorescent microscopy showed this method removed >91% of cells from the coupon surface (data not shown). Pooled biofilm suspensions (∼45 ml) from the “outer coupon” and reservoir water samples (50 ml) were each vacuum filtered onto 0.22 µm pore size (25 mm diameter) polycarbonate membrane filters (Millipore UK Ltd., Watford, UK) and stored at −20 °C prior to DNA extraction. Filters were cut into pieces using a sterile scalpel and added directly to bead solution tubes from the Ultraclean Soil DNA Isolation kit (MoBio Laboratories Inc., Carlsbad, CA, USA) with DNA isolation following the manufacturer’s protocol. DNA was eluted in 30 µl of sterile nuclease free water.
Polymerase chain reaction amplification and denaturing gradient gel electrophoresis analysis
16S rRNA genes were amplified by the polymerase chain reaction (PCR) using primers 338F (with GC-clamp) (Lane 1991) and 530R (Muyzer et al. 1993). Reactions were performed in a total volume of 50 µl comprising 200 µM of each of the dNTPs, 0.3 µM of each primer, 1× PCR Buffer, 1× Q-Solution, and 2.5 U of Taq polymerase (Qiagen Ltd., Crawley, UK). Undiluted environmental DNA (1 µl) was used as template. Reactions were initially denatured at 95 °C for 2 min, followed by 35 cycles of 95 °C for 1 min, 60 °C for 1 min, and 72 °C for 2 min, and a final extension at 72 °C for 12 min. Denaturing gradient gel electrophoresis (DGGE) analysis (Muyzer et al. 1993) was performed using the Bio-Rad DCode System (Bio-Rad, Hercules, CA, USA). PCR products (20 or 30 µl depending on product yield) were loaded onto 8% (w/v) polyacrylamide gels containing a denaturant gradient of 40% to 70% (80% denaturant consisted of 5.6 M urea and 32% v/v deionized formamide). Electrophoresis was performed at 60 °C at 100 V for 16 h and gels then stained with SYBR-Gold (Invitrogen Ltd, Molecular Probes, Paisley, UK) for 45 min in the dark, prior to image capture using an EpiChemi II Darkroom imager (UVP Inc., Upland, CA, USA). DGGE gel image analysis was performed using Quantity One (Bio-Rad, Hercules, CA, USA). Subsequently, cluster analysis was conducted using the group average linking routine and based on the Bray–Curtis resemblance matrix derived from DGGE profiles and performed using Primer-E software (Plymouth Marine Laboratory, Plymouth, UK).
Results
The application of the PWG coupons to study quantitative and compositional changes during biofilm formation, via microscopy/molecular analyses, was evaluated using the pilot-scale WDS facility. The facility was operational for 11 days, with removal of coupons at days 0 (after 5 min), 3, 7, and 11 (n = 3 at each time point). Triplicate water samples (50 ml) were also taken from the reservoir tank at day 0. The pilot system was not sterile prior to conducting the tests; however, individual coupons were clean when inserted on day 0.
DAPI/CARD-FISH analyses of biofilm formation on PWG coupons
DAPI and CARD-FISH analyses of the inserts from the removed coupons showed bacterial cell attachment after only 5 min of exposure of the clean coupons in the non-sterile pilot WDS on day 0 (Fig. 2A). By day 3, a monolayer of microbial cells submerged in the biofilm matrix was observed (Fig. 2B). Images of day 7 (data not shown) and day 11 (Fig. 2C) revealed formation of increasingly complex structurally mature biofilms. CARD-FISH analysis with bacterial probes demonstrated that the biofilms were comprised primarily of bacterial cells (Fig. 2D–F). No significant differences (P > 0.05) were found in biofilm coverage between the triplicate PWG coupons sampled on individual days, suggesting that biofilm development proceeded at a similar rate on different coupons within the system. Significant increases in relative biofilm coverage (% area) were observed over time (R 2 = 0.99, F 3,8 = 384.37, P < 0.0001) reaching ∼40% (40.66 ± 5.62%) on day 11 (Fig. 2G).
Increase in biofilm formation on PWG coupons in a pilot-scale water distribution system over time. A–C DAPI-stained images and D–F CARD-FISH images of biofilms at day 0, 3, and 11, respectively. G Increase in biofilm coverage (% area) on the coupon inserts over time (days 0, 3, 7, and 11). Error bars represent standard deviations in area calculated from ten XY images on triplicate coupons sampled at each time point
DGGE-based analysis of biofilm community structure on PWG coupons
DGGE analysis of 16S rRNA gene sequences amplified from DNA isolated from the outer coupons showed formation of increasingly diverse biofilm assemblages over the 11-day experiment (Fig. 3). For coupons sampled after 5 min exposure on day 0, a single band was seen in all of the profiles derived from the three replicate coupons at this time point. Complex DGGE profiles were observed from coupons on day 3 with further increases in numbers of DGGE bands observed on day 7 and day 11, although some bands on day 3 were absent in profiles from coupons sampled on day 11 (Fig. 3A).
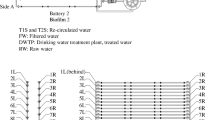
DGGE profiles and cluster analysis of bacterial 16S rRNA gene amplified from DNA extracted from biofilms on outer coupons within a pilot-scale distribution system over time and from the reservoir water. A DGGE profiles generated from triplicate coupons (C1–C3) at each time point and triplicate samples (R1–R3) within the tank reservoir water. Lanes as indicated. B Cluster analysis was calculated based on the Bray–Curtis similarity of DGGE profiles
Cluster analysis (Fig. 3B) showed time-specific clustering of the bacterial communities present on coupons and high reproducibility (>86.1% similarity) between replicate coupons at each time point. DGGE analysis showed that the majority of the bands observed in the profile of the bacterioplankton present within the recirculating WDS at day 0 were also seen within the coupon biofilm community profiles, albeit with differing intensities (Fig. 3).
Discussion
The experimental systems that are currently available for studying biofilm formation in WDS are either small scale laboratory reactor-based systems (e.g., Appenzeller et al. 2001; Murga et al. 2001; Batté et al. 2003a, b; Goeres et al. 2005), from which results are not readily transferable for the analysis of biofilm processes within real WDS, or consist of sampling devices that when placed in situ (e.g., Kalmbach et al. 1997; LeChevallier et al. 1998; Prévost et al. 1998; Kharazmi et al. 1999; Juhna et al. 2007) disrupt the local flow patterns within the studied WDS (Table 1). Consequently, there remains a need for improved biofilm sampling systems that maintain representative flow patterns within WDS experimental systems (and ideally that could be used in distribution networks) to enable improved investigation of the abiotic controls on biofilm formation and detachment processes. This research therefore reports the design and subsequent evaluation of the suitability of new PWG coupons that maintain representative boundary flow conditions and enable simultaneous quantitative and compositional characterization of microbial biofilms that form in situ within WDS. The trial reported herein utilized a pilot pipe loop system, constructed from actual WDS pipes and recreating system conditions including hydraulic and bulk water properties.
Following deployment of the PWG coupons within the non-sterile pilot rig system, CARD-FISH studies of biofilm formation on these coupons showed increasing bacterial biofilm coverage over the 11-day experiment (Fig. 2). In contrast, earlier FISH-based studies of biofilms on polyethylene slides using a Robbins device within a WDS (Kalmbach et al. 1997) showed a peak in bacterial biofilm coverage at 7 days, followed by a subsequent decline. Longer term studies of WDS biofilm assemblages in polyethylene pipe sections, via acridine orange-staining, showed cell numbers within biofilms continued to increase over a 100-day period (Lehtola et al. 2004). It should be stressed that operating conditions across these three studies were not comparable. Nevertheless, our data demonstrate that the PWG coupons allow quantitative changes in bacterial biofilm formation to be investigated. Moreover, via combinatorial application of additional FISH-probes, these coupons could be used in future studies to allow quantitative studies of archaea and eukarya and/or hierarchical targeting of particular phyla or species within WDS biofilms.
A key design feature of the PWG coupon is that it enables simultaneous quantitative and (qualitative) compositional characterization of biofilm assemblages formed within WDS. The “insert” within the coupon enables direct microscopic observation after fluorescence staining and FISH. Importantly, the use of a flat “insert” that can be placed directly on the microscope stage overcomes difficulties in imaging that would arise from studying a curved surface, but in addition the small dimensions of this flat insert do not result in changes to boundary flow conditions, enabling representative imaging of biofilm development and architecture under realistic in situ pipe conditions. The “outer coupon” was used for microbial community analysis. DGGE analysis of these biofilm communities demonstrated very rapid attachment of one particular bacterial phylotype to the coupon (Fig. 3A). Such rapid attachment is in agreement with culture-based studies on a variety of surfaces in which irreversible bacterial attachment can proceed within as little as a minute (reviewed in Palmer et al. 2007). Comparison of the DGGE profiles of the bacterioplankton present within the recirculating WDS water to those on the coupons showed that the majority of the taxa present in the coupon biofilms were also present within the planktonic community at day 0. This demonstrates the applicability for the PWG coupons to couple analysis of planktonic WDS assemblages (e.g., Eichler et al. 2006) and study in situ community dynamics within WDS biofilms.
Replicate coupons in the pilot-scale WDS showed time-specific clustering of biofilm community composition throughout the 11-day experiment (Fig. 3B), suggesting that these coupons can be used to reproducibly study temporal community changes in biofilms in situ. Time-specific clustering of biofilm bacterial communities has also previously been demonstrated via T-RFLP analysis of biofilms present on stainless steel coupons within a model WDS (Martiny et al. 2003), albeit over far longer periods revealing separation between communities sampled from “young” (1–94 days) and “old” (500–1,093 days) biofilms. However, communities sampled within the first 20 days in that system did not show temporal clustering. Again, these differences are likely to reflect the variation between the experimental conditions of these two studies.
In summary, this research has demonstrated the suitability of the PWG coupon to enable molecular-based characterization of both structural and compositional changes in biofilms within a pilot-scale WDS. The PWG coupon design allows direct insertion into the internal wall of WDS pipes, thereby minimizing the distortion of boundary layer conditions that may influence biofilm formation. Such a system facilitates future experimental investigations without disrupting boundary layer flow conditions and enables the impact of changing abiotic conditions, boundary shear stress, and turbulent driven exchange with the main water body upon the formation, growth, and detachment of biofilms to be fully captured under controlled conditions. Moreover, the coupon design offers the potential for deployment directly within distribution network infrastructure wherever the pipeline is accessible, enabling studies of biofilm dynamics and/or targeting and enumeration of specific microbial taxa therein.
References
Appenzeller BMR, Batté M, Mathieu L, Block J-C, Lahoussine V, Cavard J, Gate D (2001) Effect of adding phosphate in drinking water on bacterial growth in slightly and highly corroded pipes. Water Res 35:1100–1105
Batté M, Koudjonou B, Laurent P, Mathieu L, Coallier J, Prévost M (2003a) Biofilm responses to ageing and to a high phosphate load in a bench-scale drinking water system. Water Res 37:1351–1361
Batté M, Mathieu L, Laurent P, Prévost M (2003b) Influence of phosphate and disinfection on the composition of biofilms produced from drinking water, as measured by fluorescence in situ hybridization. Can J Microbiol 49:741–753
Batté M, Appenzeller BMR, Grandjean D, Fass S, Gauthier V, Jorand F, Mathieu L, Boualam M, Saby S, Block JC (2004) Biofilms in drinking water distribution systems. Rev Environ Sci Biotechnol 2:147–168
Baylis JR, Chase ES, Cox CR, Ellms JW, Emerson CA, Knouse HV, Streeter HW (1930) Bacterial aftergrowths in water distribution systems. Am J Public Health Nations Health 20:485–489
Berry D, Xi C, Raskin L (2006) Microbial ecology of drinking water distribution systems. Curr Opin Biotechnol 17:297–302
Camper AK, Jones JT, Hayes WL (1996) Effect of growth conditions and substratum composition on the persistence of coliforms in mixed-population biofilms. Appl Environ Microbiol 62:4014–4018
Chang YC, Puil ML, Biggerstaff J, Randall AA, Schulte A, Taylor JS (2003) Direct estimation of biofilm density on different pipe material coupons using a specific DNA-probe. Mol Cell Probes 17:237–243
Chu CW, Lu CY, Lee CM (2005) Effects of inorganic nutrients on the regrowth of heterotrophic bacteria in drinking water distribution systems. J Environ Manage 106:1328–1335
Daims H, Bruhl A, Amann R, Schleifer K-H, Wagner M (1999) The domain-specific probe EUB338 is insufficient for the detection of all bacteria: development and evaluation of a more comprehensive probe set. Syst Appl Microbiol 22:434–444
Eichler S, Ghristen R, Holtje C, Westphal P, Botel J, Brettar I, Mehling A, Höfle M (2006) Comparison and dynamics of bacterial communities of a drinking water supply system as assessed by RNA- and DNA-based 16S rRNA gene fingerprinting. Appl Environ Microbiol 72:1858–1872
Gagnon GA, Rand JL, O’Leary KC, Rygel AC, Chauret C, Andrews RC (2005) Disinfectant efficacy of chlorite and chlorine dioxide in drinking water biofilms. Water Res 39:1809–1817
Goeres DM, Loetterle LR, Hamilton MA, Murga R, Kirby DW, Donlon RM (2005) Statistical assessment of a laboratory method for growing biofilms. Microbiology 151:757–762
Husband PS, Boxall JB, Saul AJ (2008) Laboratory studies investigating the processes leading to discolouration in water distribution networks. Water Res 42:4309–4318
Juhna T, Birzniece D, Larsson S, Zulenkovs D, Sharipo A, Azevedo NF, Menard-Szczebara F, Castagnet S, Feliers C, Keevil CW (2007) Detection of Escherichia coli in biofilms from pipe samples and coupons in drinking water distribution networks. Appl Environ Microbiol 73:7456–7464
Kalmbach S, Manz W, Szewzyk U (1997) Dynamics of biofilm formation in drinking water: phylogenetic affiliation and metabolic potential of single cells assessed by formazan reduction and in situ hybridization. FEMS Microbiol Ecol 22:265–279
Kharazmi A, Giwercman B, Hoiby N (1999) The Robbins device in biofilm research. Meth Enzymol 310:207–215
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. John Wiley & Sons Ltd, West Sussex, UK, pp 115–175
LeChevallier MW, Babcock TM, Lee RG (1987) Examination and characterization of distribution system biofilms. Appl Environ Microbiol 53:2714–2724
LeChevallier MW, Norton CD, Camper A, Morin P, Ellis B, Jones W, Rompré A, Prévost M, Coallier J, Servais P, Holt D, Delanoue A, Colbourne J (1998) Microbial impact of biological filtration. AWWA Research Foundation, USA, p 180
Lehtola MJ, Miettinen IT, Keinãnen MM, Kekki T, Laine O, Hirvonen A, Vartiainen T, Martikainen P (2004) Microbiology, chemistry and biofilm development in a pilot drinking water distribution system with copper and plastic pipes. Water Res 38:3769–3779
Lehtola MJ, Laxander M, Miettinen IT, Hirvonen A, Vartiainen T, Martikainen PJ (2006) The effects of changing water flow velocity on the formation of biofilms and water quality in pilot distribution system consisting of copper or polyethylene pipes. Water Res 40:2151–2060
Lund V, Ormerod K (1995) The influence of disinfection processes on biofilm formation in water distribution systems. Water Res 29:1013–1021
Manz W, Szewzyk U, Ericsson P, Amann R, Schleifer KH, Stenstron TA (1993) In-situ identification of bacteria in drinking water and adjoining biofilms by hybridization with 16S ribosomal RNA directed and 23S ribosomal RNA directed fluorescent oligonucleotide probes. Appl Environ Microbiol 59:2293–2298
Martiny AC, Jørgensen TM, Albrechtsen H-J, Arvin E, Molin S (2003) Long-term succession in structure and diversity of a biofilm formed in a model drinking water distribution system. Appl Environ Microbiol 69:6899–6907
McCoy WF, Bryers JD, Robbins J, Costerton JW (1981) Observations of fouling biofilm formation. Can J Microbiol 27:910–917
Millar MR, Linton CJ, Sheriff A (2001) Use of continuous culture system linked to a modified Robbins device or flow cell to study attachment of bacteria to surfaces. Microbial Growth Biofilms 337:43–62
Möhle RB, Langemann T, Haesner M, Augustin W, Scholl S, Neu TR, Hempel DC, Horn H (2007) Structure and shear strength of microbial biofilms as determined with confocal laser scanning microscopy and fluid dynamics gauging using a novel rotating disc biofilm reactor. Biotechnol Bioeng 98:747–755
Murga R, Forster TS, Brown E, Pruckler JM, Fields BS, Donlan RM (2001) Role of biofilms in the survival of Legionella pneumophila in a model potable-water system. Microbiology 147:3121–3126
Muyzer G, de Waal EC, Uitterlinden AG (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59:695–700
Norton CD, LeChevallier MW (2000) A pilot study of bacteriological population changes through potable water treatment and distribution. Appl Environ Microbiol 66:268–276
Palmer J, Flint S, Brooks J (2007) Bacterial cell attachment, the beginning of a biofilm. J Ind Microbiol Biotech 34:577–588
Parent A, Fass S, Dincher ML, Readoner D, Gatel D, Block JC (1996) Control of coliform growth in drinking water distribution systems. J Chart Inst Water Environ Manage 10(6):442–445
Pernthaler A, Pernthaler J, Amann R (2002) Fluorescence in situ hybridization and catalyzed reporter deposition for the identification of marine bacteria. Appl Environ Microbiol 68:3094–3101
Prévost M, Rompré A, Coallier J, Servais P, Laurent P, Clément B, Lafrance P (1998) Suspended bacterial biomass and activity in full-scale drinking water distribution systems: impact of water treatment. Water Res 32:1393–1406
Sekar R, Pernthaler A, Pernthaler J, Warnecke F, Posch T, Amann R (2003) An improved protocol for quantification of freshwater Actinobacteria by fluorescence in situ hybridization. Appl Environ Microbiol 69:2928–2935
Sharma MO, Bhosle NB, Wagh AB (1990) Methods for removal and estimation of microfouling biomass. Ind J Mar Sci 19:174–176
Szewzyk U, Szewzyk R, Manz W, Schleifer K-H (2000) Microbiological safety of drinking water. Ann Rev Microbiol 54:81–127
Volk CJ, LeChevallier MW (1999) Impacts of the reduction of nutrient levels on bacterial water quality in distribution systems. Appl Environ Microbiol 65:4957–4966
Acknowledgments
This research was funded by a European Union FP6 Marie-Curie Transfer of Knowledge grant “Microbiology of Urban Water Systems” (grant number 42444) awarded to CAB, AMO, and JBB. CAB and JBB would also like to acknowledge the Engineering and Physical Sciences Research Council (EPSRC) for the provision of an Advanced Research Fellowship (EP/E053556/01) and Challenging Engineering award (EP/G029946/1), respectively.
Author information
Authors and Affiliations
Corresponding author
Additional information
P.D. and R.S. made equal contributions to this research.
Rights and permissions
About this article
Cite this article
Deines, P., Sekar, R., Husband, P.S. et al. A new coupon design for simultaneous analysis of in situ microbial biofilm formation and community structure in drinking water distribution systems. Appl Microbiol Biotechnol 87, 749–756 (2010). https://doi.org/10.1007/s00253-010-2510-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-010-2510-x