Abstract
Systematic screening of single-gene knockout collection of Escherichia coli BW25113 (the Keio collection) was performed to select mutants that could enhance the deethylation of 7-ethoxycoumarin catalyzed by CYP154A1. After 96-well plate high-throughput screening followed by test tube assays, four mutants (ΔcpxA, ΔgcvR, ΔglnL, and an unknown-gene-deleted one (Δuk)) were able to increase the CYP154A1 activity by approximately 1.4–1.7 times compared with that of the control strain. When new mutants were constructed by disrupting individually the cpxA, gcvR, glnL, and uk genes in E. coli BW25113, three of them (ΔcpxA, ΔgcvR, and ΔglnL) showed high levels of CYP154A1 activity. However, the uk-disruptant failed to enhance the CYP154A1 activity, suggesting that the high CYP154A1 activity of the Δuk mutant in the Keio collection was due to a spontaneous mutation in the chromosome. In-frame deletion mutants of ΔcpxA, ΔgcvR, and ΔglnL also exhibited high enzyme activity, and complementation of these mutations could decrease CYP154A1 activity. These results indicated that the enhancement of the enzyme activity was not caused by polar effects on their neighbor genes. To our knowledge, this is the first report on a genome-wide screening of the genes for deletion to improve the activity of a recombinant whole-cell biocatalyst.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Biocatalysis has matured to a standard technology in the fine chemicals industry, which is reflected by the number of biotransformation processes running on a commercial scale (Straathof et al. 2002). Both isolated enzymes and whole cells have been used as industrial catalysts in commercial biotransformations. Compared with isolated enzymes, whole-cell biocatalysts can be cost-effectively prepared. Whole-cell biocatalysis is particularly advantageous for bioproductions in which cofactors for oxidative and reductive reactions are required, since metabolically active cells can recycle cofactors with existing machinery (Cirino and Sun 2008; Duetz et al. 2001; Goldberg et al. 2007; Ishige et al. 2005). In addition, recombinant DNA techniques have allowed the overproduction of a desired enzyme in various heterologous hosts and the use of cells as “bags stuffed with catalysts”.
The activity of a whole-cell catalyst may be affected by cellular components involved in substrate transport, product accumulation, energy metabolism, and cofactor regeneration. Fujii et al. (2009) have found that disruptions in the AcrB-Tol efflux pump system of Escherichia coli strains could increase the intracellular amounts of substrates and the concentrations of active enzymes. By using E. coli strains with tolC acrAB mutations expressing cytochrome P450 monooxygenase (P450) genes, the production levels of pravastatin, 25-hydroxy vitamin D3, and 25-hydroxy 4-cholesten 3-one were enhanced substantially. Ni and Chen (2005) have reported that a lipoprotein mutation altered the outer membrane structure of E. coli and enhanced the membrane permeability to hydrophobic substrates. When toluene dioxygenase was expressed in this mutant, the bioconversion of toluene to o-cresol was more effective than in the wild-type strain. Xie et al. (2007) have identified that BioH, a carboxylesterase involved in the biotin biosynthesis, could hydrolyze dimethylbutyryl-S-methyl mercaptopropionate during the production of simvastatin acid using E. coli as a whole-cell catalyst. A ΔbioH strain was effective in eliminating the degradation of dimethylbutyryl-S-methyl mercaptopropionate and significantly increased the product yield.
Recently, the collection of single-gene knockout (SGK) mutants of E. coli K-12 (the Keio collection) has been constructed (Baba et al. 2006) and used for identifying unknown gene functions (Melnick et al. 2004) and for analyzing the dynamics of metabolic pathways (Hua et al. 2003; Yang et al. 2003; Zhao et al. 2004). The Keio collection has also been successfully used to substantiate the value of systematic approaches for the understanding of the linkage between different cellular systems (Eydallin et al. 2007; Inoue et al. 2007; Ito et al. 2005; Niba et al. 2008; Perrenoud and Sauer 2005; Tenorio et al. 2003). However, to the best of our knowledge, there has been no report on systematic screening of SGK mutants to identify cellular components that affect the activity of a recombinant whole-cell catalyst.
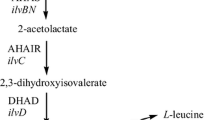
In this study, we designed a method to systematically screen SGK mutants that can enhance the activity of a recombinant whole-cell catalyst using the Keio collection. The deethylation of 7-ethoxycoumarin, a model reaction used in numerous P450 studies, was employed. The oxidative O-dealkylation of coumarins can be catalyzed by a number of mammalian and bacterial P450 enzymes. Here, CYP154A1, a P450 derived from Streptomyces coelicolor A3(2) was used. Most prokaryotic P450s require specific redox partners, ferredoxin and ferredoxin reductase, for electron transfer, and NAD(P)H as a cofactor. Therefore, P450s are usually used in metabolically active cells for industrial purpose and are likely the most suitable model enzymes for exploring cellular components that affect the activity of a recombinant whole-cell catalyst.
In this article, systematic screening of SGK collection of E. coli BW25113 transformed with a plasmid harboring the CYP154A1 gene with respect to their activity towards 7-ethoxycoumarin was performed. Three E. coli mutants, including ΔcpxA, ΔgcvR, and ΔglnL, exhibited 1.4–1.7 times higher biotransformation activity compared with that of the wild-type E. coli strain. We excluded the polar effect of the introduced kanamycin resistance gene cassette on the expression of downstream genes by construction of in-frame deletion mutants of ΔcpxA, ΔgcvR, and ΔglnL. We also confirmed that the observed enhanced activities truly depends on the respective deleted genes by construction of new deletion mutations and observing complementation by wild-type genes on a low copy vector. The systematic approach described here would enable the rapid identification of genes responsible for the desired phenotypes and contribute to the breeding of improved microbial catalysts.
Materials and methods
Materials, strains, and plasmids
E. coli BW25113, the Keio collection, and helper plasmids (pKD46 and pCP20) were described previously (Baba et al. 2006). 7-Ethoxycoumarin, 7-hydroxycoumarin, and 5-aminolevulinic acid were purchased from Wako Pure Chemical (Osaka, Japan). DNA polymerase, restriction enzymes, and other DNA-modifying enzymes were purchased from Takara Bio (Shiga, Japan). All other reagents are commercially available and of analytical grade. A P450 expression vector pET-cyp154a1-camAB, carrying the S. coelicolor A3(2) cyp154a1 and the Pseudomonas putida camA and camB, was described previously (Agematsu et al. 2006). A single-copy vector, pBAC-Lac (Asakawa et al. 1997), was a gift from Dr. S. Asakawa of Keio University.
Construction of CYP154A1 expression vector
The cyp154a1 and camAB were amplified from pET-cyp154a1-camAB with the primers CYP-P1 and CYP-P2 shown in Table 1. The PCR product was gel-purified, digested with EcoRI, and ligated to the corresponding site of the expression vector pKK223-3 (Brosius and Holy 1984). The nucleotide sequence was analyzed to confirm the direction of inserted genes and to verify the absence of PCR mutations. The resulting plasmid was designated as pKK-CYP-camAB.
Transformation of SGK mutants
The transformation of 3,985 individual SGK mutants with pKK-CYP-camAB was carried out in 96-well plates (U96 DeepWell™, Nunc International, Tokyo, Japan) in accordance with the method described by Hanahan (1983). SGK mutants, which had been kept as glycerol stocks at −80°C, were inoculated to 500 μl of Luria-Bertani (LB) medium containing kanamycin (20 µg/ml) in a 96-well plate. The plate was sealed with a Breathe-EASIER™ sealing film (Diversified Biotech, Boston, MA) for cultivation. The cells were precultured aerobically at 30°C for 24 h with orbital shaking (1,300 rpm) on a plate shaker (Mix-EVR, TAITEC, Tokyo, Japan). An aliquot (10 μl) of the fully grown culture was then inoculated to 500 µl of SOB medium (Hanahan 1983) in a 96-well plate and incubated at 30 °C with shaking to the mid-log phase of growth. The cells were collected by centrifugation at 3,000×g for 10 min at 4°C. After removing the supernatant, the cells were resuspended in 170 µl of ice-chilled transformation buffer (TB) (Hanahan 1983) and kept on ice for 15 min. The cells were then collected by centrifugation and resuspended in 40 µl of TB buffer containing 50 pg of pKK-CYP-camAB. The plate was kept on ice for 30 min and soaked in a waterbath at 42°C for 1 min. Then, 960 µl of fresh SOC medium (Hanahan 1983) was added to each well, and the plate was incubated at 37°C for 1 h. An aliquot of the mixture (10 μl) was transferred to 400 µl of LB medium supplemented with ampicillin (50 µg/ml) and kanamycin (20 µg/ml). The culture was incubated overnight at 30°C with shaking. The overnight-grown cultures were mixed with an equal volume of glycerol and stored at −80°C.
High-throughput screening
An autoinduction medium was prepared using the Overnight Express™ Auto Induction System (Novagen, Madison, WI) in accordance with the manufacturer's protocol. The glycerol stock of a transformant was inoculated with a sterilized toothpick to 400 µl of LB medium containing ampicillin (50 µg/ml) and kanamycin (20 µg/ml). Cultivation was carried out in a 96-well plate with orbital shaking (1,300 rpm) for 24 h at 30°C. An 8-µl aliquot of the preculture was then inoculated to 400 µl of the autoinduction medium containing 5-aminolevulinic acid (80 μg/ml), ampicillin (50 µg/ml), and kanamycin (20 µg/ml) in a 96-well plate. Further cultivation was carried out at 30°C with shaking for 24 h. The cells were harvested by centrifugation (3,000×g, 10 min, 4°C) and suspended in 400 μl of a reaction mixture consisting of 50 mM potassium phosphate buffer (pH 7.4), 2% (w/v) glycerol, 50 µg/ml ampicillin, 0.1 mM IPTG, and 100 μM 7-ethoxycoumarin. The reaction was carried out at 30°C with shaking (1,300 rpm) on the plate shaker. After 24 h of incubation, the reaction mixture was centrifuged to remove bacterial cells. The supernatant was subjected to high performance liquid chromatography (HPLC) analysis on a COSMOSIL 5C18–AR-II Waters column (4.6 mm × 50 mm, Nacalai Tesque, Japan). The column was eluted by a linear gradient of methanol (30–100 vol.%) in 0.1 vol.% orthophosphate at a flow rate of 1 ml/min. The eluates were detected using an SPD-20A UV/VIS detector (Shimadzu, Kyoto, Japan) at 325 nm. E. coli BW25113 (pKK-CYP-camAB) and E. coli BW25113 (pKK223-3) were used as positive and negative controls, respectively.
Test tube CYP154A1 assay
An overnight-grown culture of an E. coli transformant (40 μl) was inoculated into a test tube (180 × 18 mm) containing 2 ml of the autoinduction medium supplemented with 5-aminolevulinic acid (80 µg/ml) and adequate antibiotics (50 µg/ml ampicillin, 20 µg/ml kanamycin, and/or 20 µg/ml chloramphenicol). The cells were grown at 30°C with reciprocal shaking (200 rpm) for 12 h. After the incubation, 1 ml of the culture was sampled from the test tube. The cells were harvested by centrifugation, transferred to a test tube containing 2 ml of the reaction mixture for enzyme assay, and incubated at 30°C with reciprocal shaking (200 rpm). After 24 h of incubation, an aliquot of the reaction mixture was centrifuged to obtain the supernatant for HPLC analysis. The test tube culture was also adequately diluted with 50 mM potassium phosphate buffer, and the optical density at 600 nm (OD600) was measured using a UV150-02 spectrophotometer (Shimadzu). An OD600 of 1.0 corresponded to 0.61 mg dry cells per milliliter. Total enzyme activity was expressed as picomoles of 7-hydroxycoumarin produced per hour per milliliter of the culture. Specific activity was calculated from total activity divided by milligram dry cells of 1-ml culture. At least three independent cultures were subjected to CYP154A1 assay.
Gene disruption
The genes encoding CpxA, GcvR, GlnL, and an unknown (uk) gene were disrupted in accordance with the method described by Datsenko and Wanner (2000). The uk gene was located at a nucleotide position ranging from 1,079,456 to 1,079,647 on the chromosomal DNA of E. coli K-12 W3110 (Riley et al. 2006). Although this gene was formerly named yneE encoding a possible protein, it has been eliminated from the list of ORFs in the re-annotation published in 2006 (Riley et al. 2006). The chromosomal DNA of the respective disruptant in the Keio collection was used as the PCR template to amplify a kanamycin resistance (kmr) gene, which was flanked by Flippase (FLP) recognition target sites and approximately 200 bp of the adjacent chromosomal sequences. An adequate set of primers listed in Table 1 (P1 and P2 primers) were used as the PCR primers. The PCR product was introduced by electroporation into E. coli BW25113 carrying a Red helper plasmid pKD46, and kmr transformants were selected. The Red helper plasmid was removed by growing the transformant at 43°C. Replacement of a target gene with the kmr-gene was confirmed by colony PCR. Primers designed according to the kmr-gene cassette were used as common primers. Two PCRs were conducted using locus-specific primers approximately 500 bp upstream and downstream of the target gene (F and R primers in Table 1) with the respective common primers.
Excision of the kmr-gene cassette was performed using a helper plasmid pCP20, as described by Datsenko and Wanner (2000), to generate an in-frame deletion mutant. Excision of the cassette was confirmed by PCR using respective F and R primers (Table 1).
Double-knockout strains were constructed by disrupting the second gene of the in-frame deletion strain of the first gene disruptant using the abovementioned method.
Construction of complementary plasmids
The araC-P araBAD -MCS region of the plasmid pBAD33 (Guzman et al. 1995) was prepared by digestion with ClaI and HindIII. This DNA fragment was inserted into the corresponding sites of the pBAC-Lac plasmid to construct pBAC-CM. The genes encoding CpxA, GcvR, and GlnL were amplified from the genomic DNA of E. coli strain BW25113 by PCR using primers SD-P1 and SD-P2 in Table 1. The amplified DNA fragments were first cloned into the EcoRV site of the pBlueScriptII SK(+) plasmid and sequenced. After digesting the resulting plasmids with NheI and HindIII, the individual cloned genes were inserted into pBAC-CM to make expression vectors. The expression vectors were then transformed into in-frame deletion mutants. For complementary tests, 1 mM L-arabinose and 12 µg/ml chloramphenicol were added to the autoinduction medium.
Results
SGK mutants with enhanced CYP154A1 activity
When 96 randomly chosen mutants were inoculated to a 96-well plate, approximately 70–80% of the mutants could be simultaneously transformed with pKK-CYP-camAB. Untransformed mutants were subjected to another trial. By repeating this process, 3,978 SGK mutants could be transformed with pKK-CYP-camAB. Seven SGK mutants failed to be transformed, since they hardly grew in SOB medium at 30°C.
The 3,978 mutants were examined for their ability to produce 7-hydroxycoumarin in 96-well plates. Deethylation of 7-ethoxycoumarin has been widely employed as a model P450 reaction in various studies, since 7-hydroxycoumarin could be easily quantified by fluorescence measurement. However, since background fluorescence was not negligible in the 96-well plate assay, the reaction product was quantified by HPLC. The total activity of the wild-type strain harboring pKK-CYP-camAB (positive control) reached a maximum at approximately 24 h on the 96-well plate, while no significant activity was detected with that harboring pKK223-3 (data not shown). After 24 h of incubation, 81 SGK mutants were found to exhibit total enzyme activities of more than two-fold greater than that of the wild-type (Fig. 1). It was also observed that some mutants had a distinctly low CYP154A1 activity.
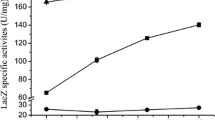
The 81 SGK mutants were subjected to test tube assays. Aside from enzyme activity, bacterial growth was also monitored in test tube assay, and mutants were selected based on the criterion that a certain strain could exhibit not only an enzyme activity which is appreciably higher than that of the wild-type, but also a growth rate similar to or higher than that of the wild-type. The positive control exhibited a total and specific enzyme activity of 2.12 ± 0.20 nmol/h/ml of the culture and 0.27 ± 0.03 nmol/h/mg dry cells, respectively. Many mutants showed CYP154A1 activity at levels similar to that of the wild-type (Fig. 2). However, four mutants, ΔcpxA, ΔgcvR, ΔglnL, and Δuk, exhibited a total activity of more than 1.4-fold greater than that of the wild-type. The ΔcpxA mutant showed growth similar to that of the positive control (Fig. 3). The growth rates of the ΔgcvR, ΔglnL, and Δuk mutants were somewhat slower than that of the positive control. This indicated that these mutants had specific CYP154A1 activity higher than that of the positive control in test tube assays. Figure 3 also shows that all of the four strains reached stationary phase after 12 h of incubation, and their highest enzyme activity appeared around 12 h. SGK mutants, including Δfbp, ΔnudE, ΔyejO, ΔyiaK, and ΔyrbL, showed a specific enzyme activity of approximately two-fold greater than that of the wild-type strain (Fig. 2). However, since these mutants showed poor growth in the autoinduction medium, they were not used for further studies.
Total and specific CYP154A1 activities detected with test tube cultures. CYP154A1 activity was assessed with the wild-type (open diamond) and SGK mutants (black circles) of E. coli BW25113. All experiments were conducted at least in triplicate. Mean values are shown in this figure. The standard deviations are within ±11.2% of their respective means
Time course of growth (empty squares) and total CYP154A1 activity (empty diamonds) of the wild-type strain and SGK mutants carrying pKK-CYP-camAB in test tube assays. The wild-type (a) and ΔcpxA (b), ΔgcvR (c), ΔglnL (d), and Δuk (e) mutants of E. coli BW25113 were cultivated in test tubes containing 2 ml of autoinduction medium. All experiments were conducted at least in triplicate. Mean values with the standard deviations (error bars) are depicted
Genes responsible for enhancing CYP154A1 activity
To further confirm if the deletion of the selected genes was responsible for the increased CYP154A1 activity, kmr-gene replacement mutants were constructed from E. coli BW25113 in the same manner as that used for the Keio collection. Kmr-gene replacement mutants in the cpxA, gcvR, and glnL genes showed CYP154A1 activity at levels similar to those detected with the corresponding SGK mutants in the Keio collection (Table 2). By contrast, the kmr-gene replacement mutant of uk failed to show high enzyme activity. This suggests that an unexpected spontaneous mutation was responsible for the enhancement of the CYP154A1 activity in the Δuk mutant in the Keio collection. The ΔcpxA, ΔgcvR, and ΔglnL mutants were used for further studies.
Complementation analysis of the in-frame deletion mutants was performed using single-copy expression plasmids. The araC-P araBAD -MCS region of the pBAD33 was introduced into pBAC-Lac to construct a single-copy expression vector, pBAC-CM. The cpxA, gcvR, or glnL gene was then placed under the control of P araBAD . When the resulting plasmid was transformed into the in-frame deletion mutants, ΔglnL and ΔcpxA mutants showed CYP154A1 activity at a level similar to that of the wild-type (Table 2). The introduction of pBAC-CM without the gcvR gene to ΔgcvR mutant significantly decreased CYP154A1 activity. However, when ΔgcvR mutant was transformed with pBAC-CM carrying the gcvR gene, the CYP154A1 activity was further decreased to 37% of the positive control. Although the reason remains unknown, the constitutive expression of the gcvR gene is likely to reduce the CYP154A1 enzyme activity.
Discussion
CYP154A1 activity of double-knockout strains
To pursue synergistic effects between each gene disruption, double-knockout strains were constructed. The combination of cpxA- and gcvR-deficient mutant could not be generated by the method used in this study. This combination might have a lethal effect on E. coli BW25113. The enzyme activities of ΔcpxA/ΔglnL and ΔgcvR/ΔglnL mutants were tested by test tube assay (Table 3). The ΔcpxA/ΔglnL mutant showed 1.52 and 1.47 times higher specific activity compared with ΔcpxA and ΔglnL SGK mutants, respectively. Similarly, the specific activity of ΔgcvR/ΔglnL mutant was 1.47 and 1.75 times higher than that of ΔgcvR and ΔglnL SGK mutants, respectively. However, due to their poor growth, the total activities of both double-knockout mutants were modest.
In the present study, among the 81 mutants selected through the first screening at 96-well plate scale, three mutants were finally confirmed to exhibit significantly higher total enzyme activities than that of the wild-type. Compared with the high-throughput assay in the 96-well plate, the test tube assay is a more reliable screening method. Furthermore, it is of practical importance to reproduce the high activities at larger scales. In fact, when the selected mutants and the wild-type harboring pKK-CYP-camAB were cultivated in a 500-ml Erlenmeyer flask containing 100 ml of the autoinduction medium, the maximum total CYP154A1 activities of ΔcpxA, ΔgcvR, and ΔglnL were 1.65, 1.56, and 1.28 times higher than that of the wild-type, respectively.
Although 33 mutants showed both relatively higher total and specific activity compared to that of the wild-type, the rest of them showed total activities lower than or similar to that of the wild-type (Fig. 2). This might be attributed to insufficient aeration in 96-well plates, unexpected changes caused by sample preparation and manipulation at a small scale (400 μl of the reaction mixture), changes in the expression profile of a specific enzymatic system under different culture conditions, etc.
Replacement of a target gene with a selectable marker gene often causes a polar effect on the expression of downstream genes, since the selectable marker gene has its own terminator which prevents downstream genes in the same operon from being transcribed (Tunca et al. 2007; Zhang et al. 2009). In SGK mutants of the Keio collection, the kmr-gene cassette was flanked by FLP recognition target sites. The excision of the kmr-gene cassette with FLP recombinase could create an in-frame deletion of the respective genes (Baba et al. 2006). To examine polar effect, in-frame deletion mutants of ΔcpxA, ΔgcvR, and ΔglnL were constructed by the method described previously (Datsenko and Wanner 2000). After transformation with pKK-CYP-camAB, the total CYP154A1 activity of the resulting mutants was examined using test tube assays. All the in-frame deletion mutants exhibited P450 activity at levels similar to those detected with the SGK mutants of the Keio collection (Table 2). This result implies that the enhanced enzyme activity was not caused by a polar effect.
In order to further clarify the underlying mechanisms for the improved CYP154A1 activity, the concentration of active CYP154A1 in the recombinant cells was determined by CO differential spectral analysis (Omura and Sato 1963). However, the concentration was too low for accurate quantification (≈35pmol/mg dry cells). Although we also tried to quantify the mRNA expression level of CYP154A1 by quantitative RT-PCR, no statistically significant difference was found between the mutants and the wild-type (data not shown).
CpxA is a membrane-bound sensory histidine kinase in a two-component regulatory system with the CpxR cognate response regulator in E. coli (Wulf et al. 2002). The physiological functions associated with the Cpx system (e.g., resistance to hostile conditions, mobility, adherence factors, and metabolism) are reported to be markedly diverse and complex (Dorel et al. 2006; Wulf et al. 1999; Zhou et al. 2003). It is reported that constitutive activation of E. coli Cpx pathway significantly depressed the active uptake of the lactose analog thiomethyl-β-D-galactopyranoside (Mileykovskaya and Dowhan 1997). Until now, it has only been identified that the Cpx system could be activated by either overexpression of the outer membrane lipoprotein NlpE (Gupta et al. 1995; Snyder et al. 1995) or mutational activation of CpxA (Danese et al. 1995; Snyder et al. 1995). Suppose that the Cpx system of E. coli BW25113 could be activated under the experimental conditions used in this study. This will lead to a difference in the uptake of lactose, which is an inducer for the promoter used in this study, between the wild-type strain and the cpxA-deficient mutant expressing CYP154A1.
Both gcvR and glnL are involved in the regulation of amino acid metabolism in E. coli. GcvR is described as a negative regulator of the glycine cleavage enzyme system, which is induced in the presence of glycine and repressed in the presence of purines (Ghrist and Stauffer 1995). Ghrist and Stauffer (1995) reported that the inactivation of gcvR led to a high-level constitutive expression of a glycine cleavage enzyme. This might cause changes in the metabolic balance, and thus the cellular energy and redox status might shift to that suitable for a CYP154A1-catalyzed reaction. In the present study, the CYP154A1 activity of the ΔgcvR mutant was significantly decreased by the introduction of pBAC-CM. In general, plasmids can impose a metabolic burden on the host cell. The introduction of pBAC-CM into the ΔgcvR cells might deteriorate the suitable cellular status for CYP154A1. GlnL is responsible for the regulation of glnA encoding glutamine synthetase and other genes involved in the assimilation of a wide range of nitrogen sources (Ginsburg and Stadtman 1973). Disruption of glnL directly or indirectly causes changes in the expression levels of these genes (Atkinson and Ninfa 1992; Chen et al. 1982). Enhancement of the CYP154A1 activity in the ΔglnL mutant might also be attributed to an alteration of the cellular energy and redox status resulting from metabolic abnormalities. Prospective studies will be required to reveal the detailed mechanisms for the enhanced CYP154A1 activities in the selected mutants.
In the field of white biotechnology, the breeding of production strains has been traditionally depended on repeated random mutation and selection (Ikeda and Nakagawa 2003; Sakuradani et al. 2009). By stepwise assembly of beneficial phenotypes with the use of mutagenic approach, many commercially potent producers have been developed. However, the classical method based on random mutation often results in the accumulation of detrimental mutations in the genome. The systematic screening of SGK mutants can avoid introducing unnecessary mutations and minimize the number of mutant strains to be investigated without losing the comprehensiveness. Furthermore, the reverse genetic approach enables rapid identification of genes responsible for the desired phenotypes. Although the present work has been done exclusively with one enzyme, other enzymes could be addressed using the same method as shown in this work.
References
Agematsu H, Matsumoto N, Fujii Y, Kabumoto H, Doi S, Machida K, Ishikawa J, Arisawa A (2006) Hydroxylation of testosterone by bacterial cytochromes P450 using the Escherichia coli expression system. Biosci Biotechnol Biochem 70:307–311
Asakawa S, Abe I, Kudoh Y, Kishi N, Wang Y, Kubota R, Kudoh J, Kawasaki K, Minoshima S, Shimizu N (1997) Human BAC library: construction and rapid screening. Gene 191:69–79
Atkinson MR, Ninfa AJ (1992) Characterization of Escherichia coli glnL mutations affecting nitrogen regulation. J Bacteriol 174:4538–4548
Baba T, Ara T, Hasegawa M, Takai Y, Okumura Y, Baba M, Datsenko KA, Tomita M, Wanner BL, Mori H (2006) Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol Syst Biol 2:2006.0008. doi:10.1038/msb4100050
Brosius J, Holy A (1984) Regulation of ribosomal RNA promoters with a synthetic lac operator. Proc Natl Acad Sci USA 81:6929–6933
Chen YM, Backman K, Magasanik B (1982) Characterization of a gene, glnL, the product of which is involved in the regulation of nitrogen utilization in Escherichia coli. J Bacteriol 150:214–220
Cirino PC, Sun LH (2008) Advancing biocatalysis through enzyme, cellular, and platform engineering. Biotechnol Prog 24:515–519
Danese PN, Snyder WB, Cosma CL, Davis LJ, Silhavy TJ (1995) The Cpx two-component signal transduction pathway of Escherichia coli regulates transcription of the gene specifying the stree-inducible periplasmic protease, DegP. Genes Dev 9:387–398
Datsenko KA, Wanner BL (2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA 97:6640–6645
Dorel C, Lejeune P, Rodrigue A (2006) The Cpx system of Escherichia coli, a strategic signaling pathway for confronting adverse conditions and for settling biofilm communities? Res Microbiol 157:306–314
Duetz WA, Beilen JB, Witholt B (2001) Using proteins in their natural environment: potential and limitations of microbial whole-cell hydroxylations in applied biocatalysis. Curr Opin Biotechnol 12:419–425
Eydallin G, Viale AM, Moran-Zorzano MT, Munoz FJ, Montero M, Baroja-Fernandez E, Pozueta-Romero J (2007) Genome-wide screening of genes affecting glycogen metabolism in Escherichia coli K-12. FEBS Lett 581:2947–2953
Fujii T, Fujii Y, Machida K, Ochiai A, Ito M (2009) Efficient biotransformations using Escherichia coli with tolC acrAB mutations expressing cytochrome P450 genes. Biosci Biotechnol Biochem 73:805–810
Ghrist AC, Stauffer GV (1995) Characterization of the Escherichia coli gcvR gene encoding a negative regulator of gcv expression. J Bacteriol 177:4980–4984
Ginsburg A, Stadtman ER (1973) Regulation of glutamine synthetase in Escherichia coli. In: Prusiner S, Stadtman ER (eds) The enzymes of glutamine metabolism. Academic, New York, pp 9–43
Goldberg K, Schroer K, Lütz S, Liese A (2007) Biocatalytic ketone reduction—a powerful tool for the production of chiral alcohols—part II: whole-cell reductions. Appl Microbiol Biotechnol 76:249–255
Gupta SD, Lee BT, Camakaris J, Wu HC (1995) Identification of cutC and cutF (nlpE) genes involved in copper tolerance in Escherichia coli. J Bacteriol 177:4207–4215
Guzman LM, Belin D, Carson MJ, Beckwith J (1995) Tight regulation, modulation, and high-level expression by vectors containing the arabinose P BAD promoter. J Bacteriol 177:4121–4130
Hanahan D (1983) Studies on transformation of Escherichia coli with plasmids. J Mol Biol 166:557–580
Hua Q, Yang C, Baba T, Mori H, Shimizu K (2003) Responses of the central metabolism in Escherichia coli to phosphoglucose isomerase and glucose-6-phosphate dehydrogenase knockouts. J Bacteriol 185:7053–7067
Ikeda M, Nakagawa S (2003) The Corynebacterium glutamicum genome: features and impacts on biotechnological progress. Appl Microbiol Biotechnol 62:99–109
Inoue T, Shingaki R, Hirose S, Waki K, Mori H, Fukui K (2007) Genome-wide screening of genes required for swarming motility in Escherichia coli K-12. J Bacteriol 189:950–957
Ishige T, Honda K, Shimizu S (2005) Whole organism biocatalysis. Curr Opin Chem Biol 9:174–180
Ito M, Baba T, Mori H, Mori H (2005) Functional analysis of 1440 Escherichia coli genes using the combination of knock-out library and phenotype microarrays. Metab Eng 7:318–327
Melnick J, Lis E, Park JH, Kinsland C, Mori H, Baba T, Perkins J, Schyns G, Vassieva O, Osterman A, Begley TP (2004) Identification of the two missing bacterial genes involved in thiamine salvage: thiamine pyrophosphokinase and thiamine kinase. J Bacteriol 186:3660–3662
Mileykovskaya E, Dowhan W (1997) The Cpx two-component signal transduction pathway is activated in Escherichia coli mutant strains lacking phosphatidylethanolamine. J Bacteriol 179:1029–1034
Ni Y, Chen RR (2005) Lipoprotein mutation accelerates substrate permeability-limited toluene dioxygenase-catalyzed reaction. Biotechnol Prog 21:799–805
Niba ET, Naka Y, Nagase M, Mori H, Kitakawa M (2008) A genome-wide approach to identify the genes involved in biofilm formation in E. coli. DNA Res 14:237–246
Omura T, Sato R (1963) The carbon monoxide-binding pigment of liver microsomes. J Biol Chem 239:2370–2378
Perrenoud A, Sauer U (2005) Impact of global transcriptional regulation by ArcA, ArcB, Cra, Crp, Cya, Fnr, and Mlc on glucose catabolism in Escherichia coli. J Bacteriol 187:3171–3179
Riley M, Abe T, Arnaud MB, Berlyn MKB, Blattner FR, Chaudhuri RR, Glasner JD, Horiuchi T, Keseler IM, Kosuge T, Mori H, Perna NT, Plunkett G 3rd, Rudd KE, Serres MH, Thomas GH, Thomson NR, Wishart D, Wanner BL (2006) Escherichia coli K-12: a cooperatively developed annotation snapshot—2005. Nucl Acids Res 34:1–9
Sakuradani E, Ando A, Ogawa J, Shimizu S (2009) Improved production of various polyunsaturated fatty acids through filamentous fungus Mortierella alpine breeding. Appl Microbiol Biotechnol 84:1–10
Snyder WB, Davis LJ, Danese PN, Cosma CL, Silhavy TJ (1995) Overproduction of NlpE, a new outer membrane lipoprotein, suppresses the toxicity of periplasmic LacZ by activation of the Cpx signal transduction pathway. J Bacteriol 177:4216–4223
Straathof AJ, Panke S, Schimid A (2002) The production of fine chemicals by biotransformation. Curr Opin Biotechnol 13:548–556
Tenorio E, Saeki T, Fujita K, Kitakawa M, Baba T, Mori H, Isono K, Horiuchi T, Wada C, Kanaya S, Kitagawa M, Ara T, Ohshima H, Miki T (2003) Systematic characterization of Escherichia coli genes/ORFs affecting biofilm formation. FEMS Microbiol Lett 225:107–114
Tunca S, Barreiro C, Sola-Landa A, Coque JJ, Martin JF (2007) Transcriptional regulation of the desferrioxamine gene cluster of Streptomyces coelicolor is mediated by binding of DmdR1 to an iron box in the promoter of the desA gene. FEBS J 274:1110–1122
Wulf PD, Kwon O, Lin ECC (1999) The CpxRA signal transductionsystem of Escherichia coli: growth-related autoactivation and control of unanticipated target operons. J Bacteriol 181:6772–6778
Wulf PD, McGuire AM, Liu X, Lin ECC (2002) Genome-wide profiling of promoter recognition by the two-component response regulator CpxR-P in Escherichia coli. J Biol Chem 277:26652–26661
Xie X, Wong WW, Tang Y (2007) Improving simvastatin bioconversion in Escherichia coli by deletion of bioH. Metab Eng 9:379–386
Yang C, Hua Q, Baba T, Mori H, Shimizu K (2003) Analysis of Escherichia coli anaplerotic metabolism and its regulation mechanisms from the metabolic responses to altered dilution rates and phosphoenol-pyruvate carboxykinase knockout. Biotechnol Bioeng 84:129–144
Zhang G, Tian Y, Hu K, Feng C, Tan H (2009) SCO3900, co-transcripted with three downstream genes, is involved in the differentiation of Streptomyces coelicolor. Curr Microbiol. doi:10.1007/s00284-009-9536-2
Zhao J, Baba T, Mori H, Shimizu K (2004) Effect of zwf gene knockout on the metabolism of Escherichia coli grown on glucose or acetate. Metab Eng 6:164–174
Zhou L, Lei XH, Bochner BR, Wanner BL (2003) Phenotype microarray analysis of Escherichia coli K-12 mutants with deletions of all two component systems. J Bacteriol 185:4956–4972
Acknowledgments
This work was partially supported by a Grant-in-Aid for Scientific Research from the Japan Society for the Promotion of Science and by the New Energy and Industrial Technology Development Organization (NEDO). We thank Dr. H. Mori, Nara Institute of Science and Technology, for offering the Keio collection, E. coli BW25113, pKD46, and pCP20; Dr. A. Arisawa, Mercian Co., for plasmid pET-CYP-camAB; and Dr. S. Asakawa for pBAC-Lac.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhou, Y., Minami, T., Honda, K. et al. Systematic screening of Escherichia coli single-gene knockout mutants for improving recombinant whole-cell biocatalysts. Appl Microbiol Biotechnol 87, 647–655 (2010). https://doi.org/10.1007/s00253-010-2505-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-010-2505-7