Abstract
Azotobacter vinelandii contains an iron-regulatory small RNA ArrF whose expression is dependent upon the levels of iron and ferric uptake regulator. The deletion of this ArrF-encoding gene resulted in a 300-fold increase in the production of poly-β-hydroxybutyrate (PHB), a polymer of industrial importance. This ∆arrF mutant exhibited wild-type growth and growth-associated PHB production. Limited iron and aeration elevated the PHB production in the mutant as well as wild type. Real-time RT-PCR revealed that phbB, phbA, and phbC were upregulated ∼61-, 18-, and eightfold, respectively, in the mutant. The phbR transcript of the activator PhbR for this operon was also ∼11 times more abundant. The analysis of phbR transcript predicted a region of complementarity near its Shine–Dalgarno sequence that could potentially basepair with the conserved region of ArrF. These results suggest that ArrF represses the expression of PhbR in an antisense manner and derepression of this activator in the mutant elevates the expression of phbB, phbA, and phbC, resulting in the PHB overproduction.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Poly-β-hydroxybutyrate (PHB) is of much commercial interest as a plastic material because it is an archetype of a natural biodegradable thermoplastic with similar chemical and physical properties as that of petrochemical-based plastics such as polyethylene and polypropylene (Anderson and Dawes 1990; Galindo et al. 2007).
Azotobacter vinelandii synthesizes PHB as a storage molecule of excess carbon in response to unbalanced growth conditions. Accumulation and degradation of PHB endow bacteria with enhanced survival, competition abilities, and stress tolerance, increasing fitness in changing environments (Kadouri et al. 2005). Biosynthesis of PHB starts with the condensation of two molecules of acetyl-CoA to give acetoacetyl-CoA by β-ketothiolase. Acetoacetyl-CoA is reduced by NADPH-dependent acetoacetyl-CoA reductase to β-hydroxybutyryl-CoA. This activated monomer is then polymerized by PHB synthase to form PHB. The genes involved in the PHB synthesis are organized in an operon phbBAC. The phbB, phbA, and phbC genes code for acetoacetyl-CoA reductase, β-ketothiolase, and PHB synthase, respectively. The upstream region of this operon contains phbR coding for an AraC-type transcriptional activator PhbR, phbP coding for a putative granule-associated protein, and phbF, a putative regulator of phbP (Galindo et al. 2007).
The expression of this phbBAC operon is driven by two overlapping promoters, pB1 and pB2 (Peralta-Gil et al. 2002). While PhbR promotes the transcription of this operon from pB1, RpoS, alternative sigma factor of RNA polymerase that is involved the expression of stationary genes (Hengge-Aronis 2002), stimulates the expression from pB2 during the stationary phase (Peralta-Gil et al. 2002). CydR (an Fnr-like regulatory protein) and GacS/GacA (a two-component regulatory system) also affect the PHB synthesis in negative and positive manners, respectively (Castaneda et al. 2000, 2001; Peralta-Gil et al. 2002; Wu et al. 2001). In addition to these regulators that control the expression of the phb operon enzymes, other factors such as growth conditions or mutations have a substantial impact on the PHB yield by affecting the activities of the PHB biosynthetic operon enzymes and/or the accessibility of their substrates (Anderson and Dawes 1990; Cereda et al. 2007; Galindo et al. 2007; Jackson and Dawes 1976; Page et al. 2001; Senior and Dawes 1973). For instance, the first enzyme of the PHB biosynthetic pathway, β-ketothiolase, is allosterically inhibited by CoASH, and acetyl-CoA releases this inhibition. Diminished carbon flux through the tricarboxylic acid (TCA) cycle by a mutation on the cycle enzymes would not only favor a higher acetyl-CoA availability but also diminish CoASH pool. This allosterically activates β-ketothiolase and eventually leads to higher PHB production. In addition, PHB can be synthesized through de novo fatty acid biosynthesis and β-oxidation pathways (Aldor and Keasling 2003).
Recently discovered families of iron-responsive small RNA have important regulatory roles by turning genes on and off in response to iron levels. Examples are RyhB from Enterobacteriaceae (Davis et al. 2005; Masse and Gottesman 2002; Mey et al. 2005) and PrrF from Pseudomonas aeruginosa (Vasil 2007; Wilderman et al. 2004). These small RNA-encoding genes have a classic “iron box” in their promoter region and are under the negative control of Fur, a global regulator that controls iron acquisition, metabolism, and storage. In iron-replete conditions, iron-bound Fur binds to the iron box of the small RNA genes, preventing their expression. Conversely, when iron is scarce, iron-free Fur does not bind to the iron box, resulting in the transcription of small RNA genes. The newly synthesized small RNA pairs with the mRNA of target genes and the sRNA–mRNA duplex is subsequently degraded by RNase E and/or RNase III (Afonyushkin et al. 2005; Masse et al. 2003). More than half of their target genes were shown to encode the proteins involved in central carbon metabolisms such as TCA cycle and aerobic respiration (Davis et al. 2005; Masse et al 2005; Mey et al. 2005; Vasil 2007).
A. vinelandii contains an iron-regulatory small RNA, ArrF (Jung and Kwon 2008). This ArrF-encding gene (arrF) has a conserved region that forms a potential stem-loop structure and an iron box in its promoter region that matches at 15 bases of the 19-base consensus sequence, GATAATGATAATCATTATC (Escolar et al. 1999). As expected, this gene was negatively regulated by iron and Fur (Jung and Kwon 2008). Although its target genes are yet to be identified, PrrF-like ArrF might regulate the genes involved in central carbon metabolisms. Since the metabolic activities of these pathways were known to have a substantial influence on the PHB synthesis (Page et al. 2001; Senior and Dawes 1973), we hypothesized that ArrF might control the PHB production capability of A. vinelandii.
In the present study, this hypothesis was tested by using an A. vinelandii mutant that had a deletion of the entire arrF gene.
Materials and methods
Bacterial strains
A. vinelandii strains used in this study are wild-type trans (Isas et al. 1995; Suh et al. 2002) (laboratory stock), and its isogenic ΔarrF mutant strain that had a deletion of the entire arrF gene. The construction and confirmation of this ΔarrF mutant were previously described by Jung and Kwon (2008).
A. vinelandii growth media and conditions
Overnight-grown culture was inoculated to the 400-ml fresh Burk’s medium at the ratio of 1:800 (v/v) in 1-l baffled flasks. The Burk’s medium was supplemented with 36 mM ammonium acetate and 100 μM FeCl3·6H2O. The culture was grown at 30ºC on Model G25 Controlled Environment Incubator Shaker (New Brunswick Scientific, Edison, NJ, USA) with shaking at 200 rpm. The growth was monitored by measuring optical density at 600 nm (OD600 nm) using a Cary 3C UV-visible spectrophotometer. Iron-limiting conditions were created by the omission of 100 μM FeCl3·6H2O from the medium and visually confirmed by appearance of a greenish color in the culture supernatant, due to the derepression of siderophore synthesis by iron-free Fur (Hantke 2001). To study the effect of aeration on the PHB yield, at constant agitation of 200 rpm, aeration was decreased by increasing the culture volume per flask volume from 20 ml to 100 ml culture medium in 125-ml Erlenmeyer flasks.
Determination of cell dry mass
Fifteen milliliters of bacterial culture was taken at defined time-points and placed into a previously dried and weighed 30-ml glass centrifuge tube. After centrifugation at 10,000 × g for 10 min at 4°C, the cell pellet was washed twice with deionized water. After the supernatant was decanted, the glass centrifuge tube was oven dried at 105°C to constant weight and later cooled in a desiccator and then weighed. The cell pellets were later used for determination of PHB.
Determination of PHB
PHB content was determined by converting PHB to crotonic acid by treatment with a hot concentrated sulfuric acid, as described by Huang and Reusch (1996). Concentrated sulfuric acid (0.5 ml) was added to a dry cell pellet and the mixture was stirred and heated on a dry heating block at 120°C for 40 min. After incubation, the mixture was cooled on ice, 1 ml of saturated sodium sulfate was added, and the solution was extracted four times with 3 ml of dichloromethane. One hundred microliters of 1 N NaOH was added to the dichloromethane extract and the dichloromethane was evaporated with a stream of nitrogen. The residue was mixed with distilled water, filtered with a 0.45 μm nylon syringe filter, and chromatographed on a HPLC Aminex HPX-87H ion exchange column (Bio-Rad, Hercules, CA, USA) in an Agilent HPLC 1100 series (Agilent Technologies, Santa Clara, CA, USA) equipped with a diode array detector. Mobile phase was 0.01 N H2SO4 at a flow rate of 0.5 ml/min. The crotonic acid content in the residue was estimated by evaluation of the peak area using pure crotonic acid (Sigma-Aldrich, St. Louis, MO, USA) as standard. Since the yield of crotonic acid from 1 µg of granule PHB is 0.5 µg, the crotonic acid measured in A. vinelandii cells represents 50% of actual PHB. PHB content was defined as the ratio of PHB to cell dry weight and expressed as a percentage. All measurements were done in triplicate for two independent determinations.
Transmission electron microscopy
Samples were fixed in 2.5% glutaraldehyde in 0.1 M sodium cacodylate buffer, pH 7.2 by changing solution six times every 10 min, and then post-fixed in 2% osmium tetroxide in 0.1 M sodium cacodylate buffer on ice for 1 h followed by the same solution at room temperature for 30 min. After washing 6 × 10 min with distilled water, the samples were en bloc stained with 2% aqueous uranyl acetate for overnight and wash 6 × 10 min with distilled water. The fixed cells were dehydrated by passing the cells through a series of increasing concentrations of ethanol until pure solvent replaced the water (35% 2 × 15 min, 50% 2 × 15 min, 70% overnight, 95% 2 × 15 min, and 100% 4 × 15 min). After washing 2 × 15 min with acetone/ethanol (1:1) followed by 2 × 15 min with acetone, the cells were passed through acetone–epoxy mixtures and the tissue is soaked in the liquid epoxy mixture for an extended period (several hours to overnight) to permit through infiltration of the cells with resin. The samples were then embedded and polymerized in an oven at 68–70°C overnight. The blocks containing the embedded tissue were then removed from oven and left for 24 h before trimming and thin sectioning on an ultramicrotome. Transmission electron microscopy (TEM) photograph were collected on JEOL JEM-100CX II TEM (JEOL, Peabody, MA, USA) operated at 80 kV in the Electron Microscopy Center on Campus.
Isolation of RNA, synthesis of cDNA, and real-time reverse transcription-PCR
For real-time reverse transcription (RT)-polymerase chain reaction-(PCR) experiments, Azotobacter vinelandii wild type and ΔarrF mutant strains were grown to their early stationary phase and the cells were spun down at 10,000×g for 2 min at 4ºC and quickly stored at −80ºC until use. Isolation of total RNA, synthesis of cDNA, and real-time reverse transcription-PCR were conducted as described by Park et al. (2007) and Jung and Kwon (2008). Total RNA was isolated from all samples using RNeasy Mini kit (Qiagen, Valencia, CA, USA) and further purified with RNase-free DNase kit (Qiagen). The synthesis of cDNA was carried out using ThermoScript RT-PCR system (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s protocol. Real-time RT-PCR was performed on a LightcyclerR 2.0 (Roche Applied Science, Indianapolis, IN, USA) using primer sets listed in Table 1. These primers were designed using algorithm Primer3 (Rozen and Skaletsky 2000). All measurements were done in triplicate on two independent runs. The 16 S rRNA gene was used as control. Relative expression ratio of a gene of interest in ∆arrF mutant versus wild type was calculated as described by Pfaffl (2001).
Results
Electron micrography of ∆arrF mutant strain revealed accumulation of intracellular granules in that mutant strain
As soon as arrF gene was knocked out from the Azotobacter vinelandii wild type, we immediately noticed that the resulting mutant had an opaque appearance on the Burk’s solid medium as well as in liquid culture. Transmission electron micrography revealed that the mutant at the stationary phase produced much larger cells containing many large intracellular granules (Fig. 1b), indicating the PHB overproduction.
Growth phase-dependent PHB production by A. vinelandii wild type and ∆arrF mutant strains
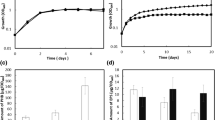
HPLC was used to quantitate PHB polymer in ∆arrF mutant and wild-type strains after converting the PHB to crotonic acid by treatment with a hot concentrated sulfuric acid. Both strains showed comparable growth rates in Burk’s medium supplemented with 36 mM ammonium acetate and 100 μM FeCl3·6H2O (Fig. 2a) and similar growth phase-dependent PHB production with a minimal yield at the exponential phase and a maximal yield at the stationary phase (Fig. 2b and inset). However, the mutant maintained the similar level of PHB polymer even at the late stationary phase, whereas the wild type showed decreased polymer level after early stationary phase, suggesting that wild-type cells used the stored polymer as carbon and energy source after sucrose depletion. The mutant might also use this polymer at the late stationary phase but the amount of PHB to be consumed might be neglectably small as compared to total PHB. Therefore, the PHB consumption by ∆arrF mutant at the late stationary phase was not reflected in Fig. 2b as a decrease in total PHB level. Despite of similar growth and growth phase-dependent PHB production, the mutant at the stationary phase overproduced PHB polymer 300 times higher than the corresponding wild type, reaching up to ∼500 µg of PHB/mg dry cell mass. Within our knowledge, this is the most dramatic change in the PHB production of A. vinelandii by a mutation in a gene apparently not directly involved in the PHB biosynthesis.
Growth (a) and PHB production (b) of wild-type trans (solid circle) and ∆arrF mutant (solid square) strains. Four-hundred-milliliter cultures of both strains were grown in 1 l baffled flasks. Growth conditions were 30ºC with shaking at 200 rpm in Burk’s minimal medium supplemented with 36 mM ammonium acetate and 100 μM FeCl3·6H2O. The data represent mean values from three independent determinations. Inset The figure showing growth-dependent PHB production of wild-type strain was enlarged
Effects of iron and aeration on the PHB production in the ΔarrF mutant
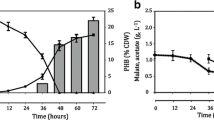
Iron limitation in the medium is known to increase the PHB yield in A. vinelandii (Reusch and Sadoff 1983). Consistent with this, Fig. 3b also showed the overproduction of PHB polymer by wild type in iron limitation. In Escherichia coli and P. aeruginosa, the function of iron-regulatory small RNAs is closely related with their iron metabolism (Masse et al. 2005; Vasil 2007). To see if this iron-dependent PHB production in A. vinelandii involves the action of ArrF, the effect of iron on the PHB production in the ∆arrF mutant was also investigated. The growth of the ΔarrF mutant was almost the same in the presence or absence of iron (Fig. 3c). Like the wild-type strain, the ∆arrF mutant strain also accumulated PHB under iron limitation (Fig. 3d), suggesting that the iron effect is not dependent on the presence of ArrF.
Aeration is known to be another factor that greatly influences the production of PHB: oxygen limitation initiates the PHB accumulation (Jackson and Dawes 1976). We investigated whether ΔarrF mutant shows similar aeration-dependent PHB production at two different growth phases. The degree of aeration was adjusted by changing the culture volume from 20 ml up to 100 ml in the flasks of 125 ml capacity at a constant agitation rate (200 rpm). At exponential phase, the PHB production of the ΔarrF mutant increased up to 60 ml culture volume and then slightly declined after that culture volume (Fig. 4), suggesting that limited aeration initiates the PHB synthesis in exponentially growing ΔarrF mutant. However, this aeration-dependent PHB production disappeared in the stationary mutant cells (OD600 nm = 2.05). In the same conditions, wild-type strain also showed similar pattern of PHB production (Fig. 4): PHB yield was high at low aeration and low at high aeration; but wild-type strain no longer showed the aeration-dependent PHB production at stationary phase. Forty milliliters ΔarrF mutant culture showed a PHB yield of ∼800 µg/mg dry cell weight to the end of culture (OD600 nm = ∼2.7) at an agitation of 200 rpm (data not shown). This yield is quite comparable to that of Ralstonia eutropha, the best PHB producer selected by Imperial Chemical Industries (ICI).
Effect of aeration on the PHB production of wild-type trans and ∆arrF mutant strains harvested at two different growth phase. White bars exponentially growing cells; grey bars stationary cells. At constant agitation of 200 rpm, aeration was decreased by increasing culture volume in 120-ml flask. Other growth conditions were 30ºC in Burk’s minimal medium supplemented with 36 mM ammonium acetate and 100 μM FeCl3·6H2O. The data represent mean values from three independent determinations
Increased expression of PhbR, PhbB, PhbA, and PhbC in the ΔarrF mutant
Real-time RT-PCR revealed that the PHB operon genes phbB, phbA, and phbC were upregulated in the mutant by ∼61-, 18-, and eightfold, respectively (Table 2). The transcript of the activator gene phbR was also ∼11 times more abundant in the mutant. This result suggests that an increase in the amount of the activator PhbR in the mutant elevates the expression of PHB operon genes, phbB, phbA, and phbC, which is responsible for the PHB accumulation in that strain.
Discussion
The results from this study showed that the disruption of the gene encoding small RNA ArrF clearly resulted in the overproduction of PHB polymer by more than 300 times. However, the ΔarrF mutant exhibited the same growth phase-, aeration-, and iron-dependent PHB productions as wild type, suggesting that these features are not dependent on the presence of ArrF.
Accumulation of PHB at the stationary phase of A. vinelandii cells was due to the elevated RpoS levels in that phase (Fig. 2). RpoS was known to enhance the transcription of phbR gene as well as the PHB biosynthetic operon (Peralta-Gil et al. 2002).
Figure 4 showed that PHB was accumulated when the aeration of culture was decreased. Under oxygen limitation, the respiratory chain activity of A. vinelandii cells slows down, and the ratio of NADH/NAD+ increases (Jackson and Dawes 1976). The abundance of NADH is a trigger for PHB formation (Anderson and Dawes 1990) and citrate synthase and isocitrate dehydrogenase activities are allosterically inhibited by high NADH/NAD+ ratio. Therefore, acetyl-CoA is no longer fed into the TCA cycle and instead, is converted to acetoacetyl-CoA by β-ketothiolase activated by acetyl-CoA, favoring the formation of PHB (Senior and Dawes 1973). However, this aeration-dependent PHB production was undetected in the stationary cells (Fig. 4). Although the reason why these stationary cells lack the aeration-dependent PHB production remains unknown, it is possible that the effect of aeration on the TCA cycle activity may be neglectable in the stationary cells, for the TCA cycle activity already significantly diminished as the cells enter the stationary phase.
Iron is an essential element for the survival and growth of A. vinelandii as a cofactor by many enzymes including TCA cycle enzymes and as a catalyst in many electron transport processes, such as respiration, photosynthesis, and nitrogen fixation. Thus, the lack of iron in the growth medium would expect to decrease the TCA cycle and respiratory chain activities of the A. vinelandii cells. Also, the lack of iron decreases the activity of antioxidant enzymes such as Fe-containing superoxide dismutase (Tindale et al. 2000), resulting in the accumulation of reactive oxygen species (ROS). Some of TCA cycle enzymes, for example, aconitase, are highly sensitive to these ROS, for they have a very labile [Fe–S] cluster (Gardner and Fridovich 1991). Thus, it appears that the oxidative stress and decreased TCA cycle activity due to ROS may favor the PHB formation in the A. vinelandii cells under iron-depleted growth conditions.
Real-time RT-PCR showed the upregulation of all three PHB operon genes. The gene encoding PhbR protein that functions as transcriptional activator for the PHB biosynthetic operon was also upregulated in the ΔarrF mutant. Therefore, it appeared that increased levels of PhbR in the ΔarrF mutant would elevate the expression of the operon enzymes, PhbB, PhbA, and PhbC, which leads to accumulation of PHB granules. Then, how does ArrF regulate the expression of activator PhbR of the phbBAC operon? In other organisms, the iron-regulatory small RNA downregulates its target gene in an antisense RNA mechanism: basepairing of the small RNA with a target mRNA fosters the degradation of the sRNA/mRNA complex by RNase E (Masse and Gottesman 2002; Wilderman et al. 2004). The analysis of phbR transcript predicted a region of sequence complementarity in the ribosome binding site that potentially can basepair with the highly conserved region of ArrF (Fig. 5), suggesting that ArrF might similarly downregulate the expression of this activator. Thus, the disruption of arrF gene would upregulate the activator gene expression.
Real-time RT-PCR experiments were conducted with the A. vinelandii cells grown in Burk’s medium supplemented with 100 μM FeCl3·6H2O, the conditions that normally repress the ArrF expression by iron-bound Fur (Jung and Kwon 2008). However, even though ArrF expression was repressed by iron repletion, ArrF was still present at a considerable amount in wild-type strain, as judged by real-time RT-PCR. Based on the crossing point values (the number of cycles required to produce a signal above background fluorescence) and PCR amplification efficiencies of arrF and phbR, the level of arrF transcript was calculated to be ca. 12 times higher than phbR transcript in wild type. This amount of arrF transcript might be sufficient to basepair with phbR transcripts, eventually repressing the production of PHB.
In some microorganisms (Davis et al. 2005; Masse et al. 2005; Mey et al. 2005; Vasil 2007), many of target genes of ArrF-like small RNA encoded the proteins that were involved in the central carbon metabolisms including TCA and respiratory chain that affects the PHB accumulation. It is also possible that ΔarrF deletion might change the proteomics of A. vinelandii, which might be responsible for the accumulation of PHB in the mutant. Further study is needed to clarify this issue.
In conclusion, iron-regulatory small RNA ArrF in A. vinelandii shows a strong regulatory effect on the synthesis of PHB, a polymer of industrial importance. The arrF deletion resulted in the overexpression of the phbBAC operon and its activator gene phbR and thereby the overproduction of PHB polymer, implying that ArrF functions as a negative regulator for the PhbR expression. The presence of a region of ArrF sequence complementary to near the Shine–Dalgarno sequence of phbR transcript suggests that the antisense mechanism might be involved in the regulation.
References
Afonyushkin T, Vecerek B, Moll I, Blasi U, Kaberdin VR (2005) Both RNase E and RNase III control the stability of sodB mRNA upon translational inhibition by the small regulatory RNA RyhB. Nucleic Acids Res 33:1678–1698
Aldor IS, Keasling JD (2003) Process design for microbial plastic factories: metabolic engineering of polyhydroxyalkanoates. Curr Opin Biotechnol 14:475–483
Anderson AJ, Dawes EA (1990) Occurrence, metabolism, metabolic role, and industrial uses of bacterial polyhydroxyalkanoates. Microbiol Rev 54:450–472
Castaneda M, Guzman J, Moreno S, Espin G (2000) The GacS sensor kinase regulates alginate and poly-β-hydroxybutyrate production in Azotobacter vinelandii. J Bacteriol 182:2624–2628
Castaneda M, Sanchez J, Moreno S, Nunez C, Espin G (2001) The global regulators GacA and σS form part of a cascade that controls alginate production in Azotobacter vinelandii. J Bacteriol 183:6787–6793
Cereda A, Carpen A, Picariello G, Iriti M, Faoro F, Ferranti P, Pagani S (2007) Effects of the deficiency of the rhodanese-like protein RhdA in Azotobacter vinelandii. FEBS Lett 581:1625–1630
Davis BM, Quinones M, Pratt J, Ding Y, Waldor MK (2005) Characterization of the small untranslated RNA RyhB and its regulon in Vibrio cholerae. J Bacteriol 187:4005–4014
Escolar L, Perez-Martin J, De Lorenzo V (1999) Opening the iron box: transcriptional metalloregulation by the Fur protein. J Bacteriol 181:6223–6229
Gardner PR, Fridovich I (1991) Superoxide sensitivity of the Escherichia coli aconitase. J Biol Chem 266:19328–19333
Galindo E, Pena C, Nunez C, Segura D, Epsin G (2007) Molecular and bioengineering strategies to improve alginate and polyhydroxyalkanoate production by Azotobacter vinelandii. Microbial Cell Factories 6:7
Hantke K (2001) Iron and metal regulation in bacteria. Curr Opin Microbiol. 4:172–177
Hengge-Aronis R (2002) Signal transduction and regulatory mechanism involved in control of sigma (S) (RpoS) subunit of RNA polymerase. Microbiol Mol Biol Rev 66:373–395
Huang R, Reusch RN (1996) Poly (3-hydroxybutyrate) is associated with specific proteins in the cytoplasm and membranes of Escherichia coli. J Biol Chem 271:22196–22202
Isas J, Yannone SM, Burgess BK (1995) Azotobacter vinelandii NADPH: ferredoxin reductase cloning, sequencing, and overexpression. J Biol Chem 270:21258–21263
Jackson FA, Dawes EA (1976) Regulation of the tricarboxylic acid cycle and poly-β-hydroxybutyrate metabolism in Azotobacter beijeinckii grown under nitrogen or oxygen limitation. J Gen Microbiol 97:303–312
Jung YS, Kwon YM (2008) Small RNA ArrF regulates the expression of sodB and fesII genes in Azotobacter vinelandii. Curr Microbiol 57:593–597
Kadouri D, Jurkevitch E, Okon Y, Castro-Sowinski S (2005) Ecological and agricultural significance of bacterial polyhydroxyalkanoates. Crit Rev Microbiol 31:55–67
Masse E, Gottesman S (2002) A small RNA regulates the expression of genes involved in iron metabolism in Escherichia coli. Proc Natl Acad Sci USA 99:4620–4625
Masse E, Escorcia FE, Gottesman S (2003) Coupled degradation of a small regulatory RNA and its mRNA targets in Escherichia coli. Genes Dev 17:2374–2383
Masse E, Vanderpool CK, Gottesman S (2005) Effect of RyhB small RNA on global iron use in Escherichia coli. J Bacteriol 187:6962–6971
Mey AR, Craig SA, Payne SM (2005) Characterization of Vibrio cholerae RyhB: the RyhB regulon and role of RyhB in biofilm formation. Infect Immun 73:5706–5719
Page WJ, Tindale A, Chandra M, Kwon E (2001) Alginate formation in Azotobacter vinelandii UWD during stationary phase and the turnover of poly-β-hydroxybutyrate. Microbiology 147:483–490
Park BS, Kwon YM, Pyla R, Boyle JA, Jung YS (2007) E1 component of pyruvate dehydrogenase complex does not regulate the expression of NADPH-ferredoxin reductase in Azotobacter vinelandii. FEMS Microbiol Lett 273:244–252
Peralta-Gil M, Segura D, Guzman J, Servin-Gonzalez L, Espin G (2002) Expression of the Azotobacter vinelandii poly-β-hydroxybutyrate biosynthetic phbBAC operon is driven by two overlapping promoters and is dependent on the transcriptional activator PhbR. J Bacteriol 184:5672–5677
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 29:2002–2007
Reusch RN, Sadoff HL (1983) D-(-)-Poly-β-hydroxybutyrate in membranes of genetically competent bacteria. J Bacterol 156:778–788
Rozen S, Skaletsky H (2000) Primer3 on the WWW for general users and for biologist programmers. Methods Mol Biol 132:365–386
Senior PJ, Dawes EA (1973) The regulation of poly-β-hydroxybutyrate metabolism in Azotobacter beijerinckii. Biochem J 134:225–238
Suh MH, Pulakat L, Gavini N (2002) Functional expression of the FeMo-cofactor-specific biosynthetic gene nifEN as a NifE-N fusion protein synthesizing unit in Azotobacter vinelandii. Biochem Biophys Res Commun 299:233–240
Tindale AE, Mehrotra M, Ottem D, Page WJ (2000) Dual regulation of catecholate siderophore biosynthesis in Azotobacter vinelandii by iron and oxidative stress. Microbiology 146:1617–1626
Vasil ML (2007) How we learnt about iron acquisition in Pseudomonas aeruginosa: a series of very fortunate events. Biometals 20:587–601
Wilderman PJ, Sowa NA, FitzGerald DJ, FitzGerald PC, Gottesman S, Ochsner UA, Vasil ML (2004) Identification of tandem duplicate regulatory small RNAs in Pseudomonas aeruginosa involved in iron homeostasis. Proc Natl Acad Sci USA 101:9792–9297
Wu G, Moir AJG, Sawers G, Hill S, Poole RK (2001) Biosynthesis of poly-β-hydroxybutyrate (PHB) is controlled by CydR (Fnr) is the obligate aerobe Azotobacter vinelandii. FEMS Microbiol Lett 194:215–220
Acknowledgements
The authors would like to thank Professors Scott Willard, Din-Pow Ma, and Ken Willeford for reviewing the paper. The authors thank Professor Jeff Wilkinson for allowing us to use LightcyclerR 2.0. This work was supported in part by a grant from Robert M Hearing Foundation, by the Mississippi Agricultural and Forestry Experiment Station (MAFES) Project Number MIS-401030, and by a grant of MAFES SRI. This paper was approved for publication as Journal Article No. J-11447 of the Mississippi Agricultural and Forestry Experiment Station, Mississippi State University.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pyla, R., Kim, TJ., Silva, J.L. et al. Overproduction of poly-β-hydroxybutyrate in the Azotobacter vinelandii mutant that does not express small RNA ArrF. Appl Microbiol Biotechnol 84, 717–724 (2009). https://doi.org/10.1007/s00253-009-2002-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-009-2002-z