Abstract
We investigated the composition of soil-extracted solubilized organic and inorganic matter (SESOM) prepared from three different soils. Growth of various bacterial strains in these soil extracts was evaluated to find appropriate conditions for ecophysiological approaches. Analysis of SESOM by 1H-NMR and gas chromatography/mass spectrometry revealed a complex mixture of organic compounds. An oak forest SESOM supported the growth of several gram-positive and gram-negative soil-derived heterotrophic bacteria, whereas beech forest and grassland soil extracts did not. A metabolomic approach was performed by determining the extracellular metabolite profile of Bacillus licheniformis in SESOM. The results demonstrated that determination of the organic composition of SESOM during batch culturing is feasible. This makes SESOM amenable to studying the ecophysiology of a range of soil bacteria growing on soil-dissolved organic matter under more defined laboratory conditions. SESOM may also increase success in isolating previously uncultured or novel soil bacteria. Cell populations and the corresponding extracellular medium can be obtained readily and specific components extracted, paving the way for proteomic, transcriptomic, and metabolomic analyses. The synthetic carbon mixture based on SESOM, which mimics soil abilities, shows a positive impact on higher cell yields and longer cultivation time for biotechnological relevant bacteria.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Soils support a wide array of bacterial species, so a single gram of soil may contain between 1,000 and 1,000,000 taxa (Torsvik et al. 2002; Gans et al. 2005). The total density may reach 1.5 × 1010 bacteria per gram (Torsvik et al. 1990; Torsvik and Øvreås 2002). Key questions in soil microbiology are: “who is out there” and “what are they doing” (Urich et al. 2008). Recent molecular approaches allow us to gauge the extent of soil bacterial diversity, but the majority of this diversity remains uncharacterized beyond its 16SrRNA gene pool (Pace 1997; Rappe and Giovannoni 2003; Fierer et al. 2007). While taxa involved in select processes such as the nitrogen cycle and methane production and consumption have been studied extensively in situ, the ecophysiological characteristics of the majority of soil bacteria are largely unknown (Fierer et al. 2007). This is especially true of the array of largely uncultivated Acidobacteria (Tringe et al. 2005) and Crenarchaeota (Schleper et al. 2005) occurring in soils. Metagenomic approaches have shed light on the diversity and distribution of biocatalytic potential through the available gene pool in soils, but not on biocatalytic activity (Daniel 2004; Schloss and Handelsman 2006). DNA stable-isotope probing has been instrumental in defining members of specific physiological groups such as methanotrophs and acetate-metabolizing or glucose-metabolizing bacteria (Radajewski et al. 2003), yet little is known regarding the spectrum of functional processes that occur in soils (Fierer and Jackson 2006). Ecophysiological approaches to soil bacteria have been hampered by inadequate cultivation methods. This is due in part not only to the extremely slow growth rate and yield of many bacterial taxa (Bollmann et al. 2007), but also to the use of inappropriate culture media (Joseph et al. 2003; Ellis 2004). Soil extract as a culture medium to support otherwise unculturable bacteria or increase the culturable count has been revisited periodically over the years (James 1958; Bakken 1985; Ellis 2004; Davis et al. 2005). Detailed knowledge of the composition of a soil-derived culture medium would be beneficial to ecophysiological studies.
Soils are three-phase porous medium systems composed of solid, liquid, and gaseous phases (Stotzky and Burns 1982; Sharma 2005). Historically, the organic component of soil is divided into solid soil organic matter (SOM) and dissolved organic matter (DOM). The DOM is a cocktail of organic aromatic compounds derived from lignin, fulvic acids, some oligomeric and monomeric sugar derivatives, amino acids, and fatty acids between C14 and C54 believed to derive from both plant wall material and dead bacteria (Huang et al. 1998; Kalbitz et al. 2000). Fulvic acids comprise the bulk of DOM, but soils also contain a range of low-molecular-weight organic substances (LMWOS) such as monosaccharides and disaccharides, amino sugars, organic acids, and amino acids (Van Hees et al. 2005; Fischer et al. 2007). Heterotrophic soil bacteria are dependent for nutrition on LMWOS, obtained either directly from solution or following excretion and activity of degradative enzymes. A growing group of reports describes the spectrum and concentrations of LMWOS soils (Kaiser et al. 2001; Strobel 2001; Van Hees et al. 2005; Pizzeghello et al. 2006), but few studies have reported on their concentrations in soil solution (Fischer et al. 2007).
The study of bacteria and their physiology in soils suffers various constraints, including the sorption of cells to particles, presence of interfering substances in macromolecular extracts, and optical constraints (Benndorf et al. 2007). We have initiated a program to investigate the ecophysiology of heterotrophic soil bacteria. Heterotrophic bacteria obtain organic nutrients by uptake across the cell envelope, restricting nutrition largely to the nutrient pool occurring in solution. This prompted us to take a simplified approach to study the ecophysiology of soil bacteria by preparing aqueous extracts of soil: soil-extracted solubilized organic matter (SESOM) (Vilain et al. 2006). So, we were able to demonstrate that Bacillus cereus ATCC 14579 is able to grow and perform a full life cycle, from germination through exponential growth to sporulation in SESOM prepared from garden and deciduous forest soil (Vilain et al. 2006; Luo et al. 2007). SESOM lends itself to ecophysiological studies of soil bacteria, such as via a proteomic approach (Luo et al. 2007). An important aspect of such ecophysiological studies is information on the respective concentrations of the various organic and inorganic substances in SESOM. In this study, we offer a rigorous chemical analysis by gas chromatography/mass spectrometry (GC-MS) and 1H-NMR of SESOM from three different soils derived from an oak forest (oak SESOM), a beech forest (beech SESOM), and a wet grassland (grassland SESOM). An intensive inorganic characterization of the three extracts was also performed. The oak forest SESOM supported the growth of a range of gram-positive and gram-negative soil bacteria and is, therefore, amenable to future studies of bacteria growing on dissolved organic matter in soil. The beech and grassland SESOM were nongrowth-supporting. A consumption profile (extracellular metabolites) of Bacillus licheniformis in oak SESOM is presented to demonstrate the applicability of SESOM to one possible part of an ecophysiology approach of soil bacteria. This work opens the door for critical revision of media used for culturing soil-derived bacteria in biotechnology fermentations or new cultivation possibilities to find appropriate growth conditions and possibly enhance yields of the target product by knowledge of the soil-influenced physiology. A first experiment in this direction was the evaluation of a SESOM mimicking synthetic medium with a carbon source cocktail that reflects the composition of growth-supporting oak SESOM. It was successfully applied to support growth of B. licheniformis DSM 13 in a manner comparable to growth on standard culture media.
Materials and methods
Preparation of soil media
Soil for oak SESOM was taken during the summer from the top 5 cm of a deciduous forest dominated by bur oak, green ash, peach willow, and wild plum (Oak Lake Field Station, Brookings County, South Dakota, USA). Material for the second and third soil media was collected during spring from the top 5 cm of a beech forest and wet grassland (region around the Bay of Greifswald, Greifswald, Germany). The samples were dried at 50 °C, and stored at 4 °C. Soil media were prepared as an aqueous extract as described previously (Vilain et al. 2006). Briefly, 100 g of air-dried topsoil were suspended in 500 mL of sterile MOPS buffer (10 mM, pH 7.0) at 40 °C with shaking at 200 rpm for 1 h. The extract was filtered sequentially through filter paper (Whatman) and filters with a pore size of 5 and 0.45 µM in order to remove particulate matter. Directly after preparation, the extract was filtered to sterility using a 0.22-µm pore size and stored at 4 °C until use. For analytic purposes, MOPS buffer was replaced with water in consideration of possible overlapping signals in the 1H-NMR spectra. All extracts for chemical analysis were done at least in triplicate; for the growth experiments, each cultivation was conducted using a freshly extracted soil medium.
Bacterial strains and growth conditions
The soil-isolated strains Arthrobacter aurescens TC1 (ATCC BAA-1386), Pseudomonas ADP, B. licheniformis DSM 13, and B. cereus 20 were cultured at 30 °C, and Salmonella enterica ser.Typhimurium (Salmonella Typhimurium) was cultured at 37 °C overnight in Luria–Bertani (LB) broth while shaking at 200 rpm (for strain characteristics, see Table 1). Exponentially growing cells were harvested by centrifugation, washed in soil medium, and inoculated into corresponding soil medium to an initial absorbance (540 nm) of 0.005. Duplicate cultures were incubated while shaking at 200 rpm and the absorbance (540 nm) measured every 30 min. Cultivation of B. licheniformis in Belitzki minimal medium (adjusted to glucose limitation conditions) (Voigt et al. 2007) and synthetic medium was performed as described above (for concentrations of ingredients, see Tables 4 and 5). All used strains are deposited in public strain collections; Pseudomonas ADP, B. cereus 20, and Salmonella Typhimurium can be obtained after request from the authors.
Sample preparation
A fixed volume of 10 mL soil medium was frozen at −80 °C for 2 days. Samples were lyophilized with a Christ®beta 1–8 lyophilizer at −52 °C and 0.25 mbar. After determining the dry mass, the sample was redissolved in 0.5 mL deionized water. Particulate matter was removed by centrifugation at 5,000×g for 5 min.
For the analysis of soil compounds by GC-MS, 2 mg of the residues were redissolved and derivatized for 90 min at 37 °C in 40 µL of 20 mg/mL methoxyamine hydrochloride in pyridine, followed by a 30-min treatment with 80 µL N-methyl-N-(trimetylsilyl)trifluoroacetamide (MSTFA) at 37 °C as described previously (Liebeke et al. 2008). Eight microliters of a retention time standard mixture (0.5% (w/v) n-decane, n-dodecane, n-pentadecane, eicosane, n-octacosane, n-dotriacontane dissolved in hexan) was added to the samples.
Detection of SESOM compounds with GC-MS
Soil medium compounds were detected by GC-MS as described by Liebeke et al. (2008). The detected compounds were identified by processing the raw GC-MS data with ChemStation G1701CA software and comparing with the NIST mass spectral database 2.0 d (National Institute of Standards and Technology, Gaithersburg, USA) and from retention times and mass spectra of standard compounds (predictions without standard are asterisked in Table 2). The semiquantification of SESOM components was carried out by integrating the chromatographic peaks. Quantification was done using MetaQuant 1.2 (Bunk et al. 2006) with ribitol (adonitol) as internal standard.
Detection of amino acids in SESOM with GC-MS
Levels of free amino acids in the soil media samples were measured using the phenomenex® EZ:faast kit (EZ:faast GC/MS free [physiological] amino acid kit) and GC-MS. Prederivatization of amino acids was according to Makita et al. (1976). Injection into the GC-MS system was done using an Agilent®7683 Series injector (split 1:15 at 250 °C, 2.0 µL; carrier gas, helium 1.1 mL/min (60 kPa) at 110 °C; pressure rise, 6 kPa/min; oven program, increasing at 20 °C/min from 110 °C to 320 °C followed by a 2-min hold at 320 °C; detector, MSD at 250 °C). Full-scan mass spectra were acquired from m/z 45 to 450 at a rate of two scans per second with 1.50 min solvent delay. The amino acids were identified by processing the raw GC-MS data with ChemStation G1701CA software and comparing with NIST and EZ:faast reference library, as well as retention times of standard compounds. Quantification was done using MetaQuant 1.2 with norvaline as internal standard.
Detection of SESOM compounds with 1H-NMR
Lyophilized samples (from 10 mL SESOM) were redissolved in 500 µL water, and 300 µL were buffered to pH 7.0 by addition of 200 µL of a sodium hydrogen phosphate buffer (0.1 mM, pH 7.0) made up with 25% (v/v) D2O to provide a NMR lock signal; samples from synthetic media were prepared and analyzed as previously described (Hochgrafe et al. 2008) and were transferred to 5 mm NMR glass tubes (length 7 in.). Spectral referencing was relative to 1 mM sodium 3-trimethylsilyl-[2,2,3,3-D4]-1-propionic acid (TMSP) in phosphate buffer. All NMR spectra were obtained at 600.27 MHz at a nominal temperature of 298.5 K on a Bruker AVANCE-II 600 NMR spectrometer operated by TOPSPIN 2 software (both Bruker Biospin, Rheinstetten, Germany). We used the “noesypresat” pulse sequence with water presaturation during both the relaxation delay and the mixing time (Nicholson et al. 1995). One hundred twenty-eight free induction decay scans were collected into 64 k data points using a spectral width of 20 ppm for a one-dimensional spectrum. After Fourier transformation with 0.3-Hz line-broadening and a single zero-filling, spectra were automatically phased and baseline-corrected using the baseopt process, and the chemical shift scale was set by assigning the value of δ = 0.00 ppm to the signal from the added TMSP. Compound identification was done by matching the obtained spectra with a 1H-NMR spectra databank using AMIX®Viewer Version 3.8.2 Bruker Biospin and comparing with spectra of standard compounds. Quantification was done by integration of designated peaks and comparing with the added standard TMSP.
Detection of inorganic SESOM compounds with ICP, IC, and AAS
Soil media were analyzed by inductively coupled plasma atomic emission spectroscopy (ICP-AES) on an ICP-AES 3410 (Fision Instruments) for PO4, SO4, Zn, Si, Cu, Sr, Ba, Mn, Fe, Mg, and Ca. The anion content of the soil medium was analyzed by ion chromatography (IC) with a 761 Compact IC (Metrohm, Germany). Atomic absorption spectroscopy (AAS) was done according to DIN 38405-D 35.
Results
SESOM of Oak Lake forest soil contained a wide variety of carbon and nitrogen sources that were able to support the growth of a range of heterotrophic bacteria isolated from soil (Figs. 1 and 2a). We were able to detect a wide range of LMWOS by 1H-NMR (Fig. 3) and GC-MS (Fig. 4), including organic acids, amino acids, monosaccharides and disaccharides, sugar alcohols, and fatty acids (Table 2). This wide selection of carbon and nitrogen sources would support the growth of bacteria with different nutritional requirements. For the beech forest soil medium (beech SESOM) and the wet grassland soil medium (grassland SESOM), the same chemical classes were detected in both, but in lesser amounts than in the chemically more diverse and richer oak SESOM.
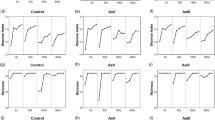
Growth of B. licheniformis DSM 13 cultured at 30 °C while shaking in a oak SESOM (filled squares), beech SESOM (filled triangles), and grassland SESOM (open circles); b Belitzki minimal medium (reduced glucose concentration) (filled triangles); c M9 medium with low carbon content, only glucose (filled triangles) and carbon source cocktail (open squares); and d M9 medium with high carbon content, glucose (filled triangles) and with carbon source cocktail (open squares)
1H-NMR spectrum of oak SESOM (compound abbreviations are listed in Table 2). Chemical shifts (in parts per million) referred to TMSP as internal standard
GC-MS total ion chromatogram of oak SESOM (compound abbreviations are listed in Table 2)
Bacterial growth in SESOMs
The five soil-derived bacterial strains (Table 1) all adapted readily to growth in oak SESOM following transfer from peptide-rich LB broth, as evidenced by the short lag periods (Figs. 1 and 2a). This indicated that oak SESOM contained nutrients that are readily taken up and metabolized, mimicking at least in part the available nutrient pool in the natural environment of these strains. The generation times in oak SESOM varied from 67 min (A. aurescens TC1), 65 min (B. licheniformis DSM 13), 63 min (P. ADP), 45 min (B. cereus 20), and 42 min (Salmonella Typhimurium). The biomass yields observed also varied between isolates, reflecting varying abilities to utilize the spectrum of available carbon sources in the oak SESOM. These and other soil bacterial isolates are also able to grow in SESOM prepared from soils such as Eastern South Dakota corn field and garden soil (data not shown). B. licheniformis was not able to grow in beech and grassland SESOM (Fig. 2a), but in oak SESOM, a growth until OD600 = 0.2 was observed (Fig. 2a).
The LMWOS composition of SESOM
GC-MS analysis of the amino acid composition of soil media revealed a high total concentration of 136 µM or 18 mg L−1 in oak SESOM. Fourteen of the 20 proteinogenic amino acids were detected (Table 2), and 13 of these could be quantified by GC-MS (Table 3). The predominant amino acid was glutamic acid, constituting 50% of the amino acid pool on a mass basis. This was followed by leucine, aspartic acid, valine, and phenylalanine. The only sulfur-containing amino acid detected was methionine at 2.15 µM. Eight of the amino acids detected are nonpolar, indicating that their concentrations in soil may be higher than detected. The soil media from beech forest and grassland showed markedly less diversity of components and lower quantities. Glutamic acid, leucine, and valine were also the top three amino acids, but with concentrations between 1 and 2 µM. No aromatic and sulfur-containing amino acids were detected in beech and grassland SESOM.
The predominant saccharides quantified were trehalose, a 1,1-disaccharide of α-glucose, present at 246 µM in oak SESOM, at 51 µM in beech, and at 49 µM in grassland SESOM, respectively. Sucrose, a 1,2-disaccharide of glucose and fructose was only detectable in traces in SESOM of beech forest and grassland, but very prominent at 121 µM in oak SESOM (Table 4). These were followed by α/β-glucose levels at 141 µM in oak, at 15.6 µM in beech, and at 3.2 µM in grassland SESOM.
The monosaccharides mannose, xylose, and myo-inositol were detectable as trace amounts but could not be quantified. The same holds for the sugar alcohols mannitol, sorbitol, and glycerol. Four quantifiable low-molecular-weight organic acids (LMWOA) known to support growth of bacterial heterotrophs, which included acetic, pyruvic, succinic, and fumaric acids, were analyzed (Table 4). Formic acid was present at 224 µM in oak SESOM and may be used by C1-compound-utilizing bacteria. A further number of LMWOA were detected in trace amounts. A number of aliphatic organic acids up to a chain length of 23 C atoms were also detected in trace amounts (Table 2). Due to their low water solubility, these may be present in much greater amounts in the forest soil. Breakdown products of amino acids like isovaleric acid and amines like putrescine and cadaverine were detected in SESOM with small abundances.
Other organic compounds
We detected many unknown signals in the 1H-NMR spectra within the chemical shift region of aromatic molecules (6.8–8.0 ppm) (Fig. 3), possibly components of humic acid, flavonoids, etc., but no matching reference spectra are available. Similarly, the gas chromatogram showed peaks without a NIST databank hit, most of them with m/z greater than 300, which indicates more complex molecules.
Inorganic composition of SESOM
SESOM contained a spectrum of inorganic compounds, many of which are obligately required for bacterial growth (Table 5). The concentrations of the main anions and cations within the tested soil media varied only marginally (onefold to fourfold). Phosphate was determined at 8.81 mg L−1in oak SESOM, while the corresponding forest soil contained a total of 2.4 g kg−1 soil (data not shown). Only 2.2% of the total phosphorus occurred as extractable water-soluble phosphate, indicating that the bulk of phosphorus was bound in organic and/or insoluble form. Other important inorganic extracted components were sulfate, magnesium, iron, manganese, calcium, and zinc.
Consumption of oak SESOM compounds by B. licheniformis DSM 13
B. licheniformis DSM 13 displayed diauxic growth in oak SESOM, while unable to grow on the low concentration of organic substrates available in beech and grassland SESOM (Fig. 2a). Determination of the temporal composition of SESOM during growth indicated sequential but overlapping consumption of carbohydrate substrates over time (Fig. 5). While glucose was consumed first, sucrose and trehalose consumption started before depletion of glucose. The fumarate concentration only decreased after the onset of the stationary phase. The amino acid concentrations decreased gradually during growth.
Consumption of selected oak SESOM metabolites by B. licheniformis DSM 13 along the growth curve (x). Values for metabolites are presented in percent, whereas the pure oak SESOM is 100% (n = 3, independent cultivations). Metabolites: open squares glucose, open circles trehalose, open triangles sucrose, open diamonds fumaric acid, filled squares leucine, filled circles proline, filled triangles alanine
Growth of B. licheniformis DSM 13 in different synthetic media consisting of glucose in comparison to artificial carbon source mixtures
B. licheniformis DSM 13 was cultivated in two common synthetic minimal culture media, Belitzki minimal medium and M9 medium (Paliy and Gunasekera 2007). Both were supplemented with low and high glucose concentrations as single carbon source or with a cocktail of carbon sources to mimic oak SESOM at low and high molarities. The composition of this cocktail is described in Table 4. Amino acids, present in very low micromolar concentrations in oak SESOM, were excluded for this synthetic medium. Growing on the carbon source cocktail, B. licheniformis reached at similar stoichiometric carbon input higher optical densities (∼+30%) and a longer stable stationary phase (see Fig. 2c, d). Analyzing the extracellular metabolites concentrations during growth, a clear consumption priority can be stated. First of all, glucose is metabolized by B. licheniformis, than sucrose and followed by the other carbon sources. Interestingly, formate is cometabolized by B. licheniformis DSM 13 in the exponential growth phase. In the stationary phase, acetic acid is the most important carbon source after most of the added nutrients are nearly consumed (cf. supporting information).
Discussion
The aqueous extract of Oak Lake forest soil, oak SESOM, is able to support exponential growth of a range of gram-positive and gram-negative heterotrophic soil bacteria, including various Bacillus sp. (Vilain et al. 2006). In addition, the chemical cues in SESOM can influence the bacterial phenotype. SESOM plays a key role in the switch to multicellular growth of B. cereus (Vilain et al. 2006). Similarly, the collection of chemical cues in cystic fibrosis (CF) lung sputum affect the physiology, competitiveness, and biofilm formation of P. aeruginosa in ways not observed during growth on laboratory culture media (Palmer et al. 2005, 2007). In a more specific approach, Watanabe et al. (2008) designed an artificial medium that imitates photoautotroph–heteroautotroph interaction in algal cultures. Genome sequences and metagenomic data reveal that several autochthonous soil bacteria are metabolically highly diverse (Daniel 2004; Mongodin et al. 2006). The diverse nutritional cues in soil will most likely influence the phenotype of many soil bacteria by modulating gene expression patterns. Our understanding of the spectrum of biocatalytic reactions that occur in soils and the ecophysiology of soil bacteria can be advanced through using either SESOM or defined media that are designed to mimic the chemical composition of soluble material in soils.
The rigorous chemical analysis of all three SESOMs reported in this study reflects a wide range of carbon and nitrogen sources that are known to support the growth of a wide array of heterotrophic bacteria. The majority of bacteria grow by taking up water-soluble small molecules across their cell envelope. By using a warm aqueous shaking approach, we obtained a much higher concentration than when using an elution approach designed to sample only the solution in macropores and mesopores that would be in equilibrium with sorbed solutes (Fischer et al. 2007). SESOM, especially the oak forest-derived one, contained a much wider range of compounds than the majority of laboratory culture media, such as BMM (Tables 2 and 3) and R2A (Reasoner and Geldreich 1985; Stulke et al. 1993). A versatile biochemical response may, therefore, be anticipated in soil bacteria when growing under soil conditions. Similarly, CF lung sputum contains a range of amino acids, salts, and other organic substrates in millimolar concentrations and ratio much more diverse than most defined media used (Palmer et al. 2007). A. aurescens grew to a higher final yield than the other three strains, displaying a high degree of metabolic diversity apparent from the genome sequence (Mongodin et al. 2006). The growth of A. aurescens, Pseudomonas ADP, and the Bacillus isolates in oak SESOM is not surprising as these species are autochthonous to soil. The Salmonella Typhimurium strain used is one of many isolates obtained from river sediment in a temperate region subject to high population densities and poor sanitation (Burke et al. 2008) and is, therefore, more likely allochthonous to soil. Yet, it was able to grow in oak SESOM under noncompetitive conditions. The fact that beech and grassland SESOM did not support the growth of B. licheniformis implicates a lack of growth-supporting nutrients. This was not due to a lack of available phosphate (Hoi et al. 2006) as all SESOM contained sufficient phosphate and other cations and anions required for growth. Supplementation of beech and grassland SESOM with phosphate did not restore growth (data not shown). The lack of growth in these SESOMs is ascribed to the extremely low concentration of organic substrates. Specifically, beech and grassland SESOM contained one tenth the carbon of oak SESOM (Table 4). Belitzky minimal medium used to grow B. licheniformis under glucose-limiting conditions contains 65-fold the carbon of grassland or beech SESOM (Voigt et al. 2007).
The results demonstrated that oak SESOM is a suitable system for studying the ecophysiology of autochthonous and allochthonous soil bacteria under more defined laboratory conditions. Not all soil-derived media support the growth of soil bacteria like B. licheniformis DSM 13, so a prior evaluation is required.
Amino acids
An elution approach to amino acid analysis in actively farmed soil yielded a 1,000-fold lower concentration, albeit containing a wide spectrum of both polar and nonpolar amino acids (Fischer et al. 2007). This may be due in part to a higher amino acid concentration in the forest soil, but it also may be due to the more rigorous extraction approach used for preparing SESOM. Shaking may, therefore, be a more suitable approach than water elution for extracting the water-soluble fraction of LMWOS.
The amino acid analysis reflected a high concentration and diverse array of 14 amino acids, including polar and apolar amino acids for oak SESOM. This soil medium contained an organic sulfur source in methionine, but no basic amino acids were detected. The free amino acids in SESOM are likely due to proteolytic degradation of native proteins derived from detritus. Common amino acid biomarkers like the nonprotein amino acid ornithine for bacteria or hydroxyproline for plants (Allard 2006) were not detected in SESOM, perhaps because these were associated in some way with humic acids or peptides. The main amino acid in SESOM was glutamic acid, also described for two soils (Amelung and Zhang 2001), so it may serve as easily accessible energy and nitrogen source for soil-dwelling bacteria. A recent proteomic analysis of B. cereus growing in SESOM indicated the uptake and usage of amino acids (Luo et al. 2007).
Carbohydrates
Monosaccharides, disaccharides, oligosaccharides, and polysaccharides are the major forms of photosynthetically assimilated carbon in the biosphere and are widespread among life forms. The primary saccharides in oak SESOM were the monosaccharide glucose, the disaccharides trehalose (mycose), and sucrose (saccharose), all together at 0.15% w/v (Table 4). B. cereus growing in oak SESOM demonstrated glycolytic activity (Luo et al. 2007), indicating uptake and growth on sugars occurring in this SESOM. It furthermore showed signs of interconversion of sugars into the core glycolytic pathway. Sugar alcohols such as sorbitol and further hexoses like myo-inositol and mannose were detected in SESOMs in trace amounts (Table 2). Microbial consumption of these diverse carbohydrates is likely as soil bacteria such as B. subtilis encode a diversity of phosphotransferase systems for carbohydrate uptake (Reizer et al. 1999). Yet, the data may represent a snapshot of the natural occurrence of saccharides, as carbohydrates have a high flux in soil due to seasonal input and degradation (Kaiser et al. 2001; Van Hees et al. 2005). The relatively high concentration of glucose in oak SESOM was surprising as this is widely viewed as the preferred carbon and energy source to a host of heterotrophic bacteria (Tobisch et al. 1999; Shivers et al. 2006). As glucose uptake systems are widespread across the bacterial domain, any residual glucose in soil should be attributed to either sorption or location at a site distal to the closest active cell. Polysaccharides such as cellulose and xylan possibly derived from plants were not found as their detection requires different analytical methods to those used in this study.
Low-molecular-weight organic acids
LMWOAs are common in soils, especially in the zone of the soil–root interface. Concentrations of aliphatic organic acids in soil solutions generally range from less than 1 µM to 10 mM (Strobel 2001; Sandnes et al. 2005). B. cereus growing in oak SESOM showed a response to the fatty acid pool by expressing a range of proteins involved in fatty acid metabolism (Luo et al. 2007). Organic acids represent a readily utilizable carbon source for microorganisms in the soil and contribute, therefore, for facilitation of microbial processes of great significance for soil nutrient cycling. Soil solutions are often characterized by higher concentrations of monocarboxylic acids than of dicarboxylic and tricarboxylic acids (Strobel 2001), and oak SESOM conforms with this trend, whereas beech and grassland SESOM do not. We identified monocarboxylic acids (e.g., formic, pyruvic, acetic, butyric, lactic, and tetronic acids) at higher concentration than the dicarboxylic and tricarboxylic acids fumaric, malic, succinic, and citric acids. Most of these LMWOA are intermediates of carbon catabolism in bacteria. They can also derive from old and damaged root cells (Pizzeghello et al. 2006). Plants and microorganisms efflux these LMWOAs in times of nutrient overflow or seasonal changes (Pizzeghello et al. 2006), leading to a residual pool in soil that is available for consumption by other bacteria. SESOM had a neutral pH (data not shown), so no influence of the LMWOAs on pH was observed, possibly caused by the dissolved salts and organic substances like amino acids which can serve as a natural buffer.
Inorganic composition of SESOM
The main cations in SESOM are Ca and Mg, also described by Hafner et al. (2005) for a leachate of a forest soil. Anions like phosphate, sulfate, and nitrate are present at intermediate abundance, but they play a distinct role in environmental microbiology; these nitrogen and sulfur sources are essential for bacterial growth. Elements like Mn, Zn, and others play a crucial role as cofactors for enzymes and for microbial soil life. On the other hand, toxic elements like arsenic and bromine are present and able to disturb growth of bacteria, but it was shown for a Bacillus sp. that resistance mechanisms can occur (Shivaji et al. 2005). No apparent growth inhibition was observed for the bacterial strains tested. Compared to the soil media, BMM contains a 100-fold higher concentration of inorganic solutes. The requirement for these high salt levels has to be reconsidered in light of the growth yields achieved using oak SESOM.
Growth of the various bacterial strains in SESOM comprised of a range of LMWOS and inorganic components, each of low concentration, indicated that no one compound was solely responsible for supporting growth. Growth of soil-derived bacteria of biotechnological importance should, therefore, be revisited from an ecophysiological perspective by culturing in media composed of an array of LMWOS at low concentrations to more closely mimic soil conditions. Looking on the growth of biotechnologically important B. lichenifromis utilizing synthetic carbon mixture, which mimics soil abilities, a clear positive impact on higher cell yields and longer cultivation time compared to media which contains only glucose as single carbon source was observed.
We show in this study that SESOM prepared from certain soils supports the growth of a range of soil-derived bacteria. The extract contains a diverse array of organic and inorganic matter. SESOM can, therefore, be used for detailed ecophysiological studies of a range of soil bacteria in culture, and possibly also previously uncultured species, under defined laboratory conditions. SESOM was amenable to determination of the extracellular metabolite pool, indicating the usefullness of this approach for more complex ecophysiological studies of soil bacteria growing on the soluble substrates present in soil, such as integrated proteomic, transcriptomic, or metabolomic analyses. SESOM could also be investigated for increased success in isolating previously uncultured or novel soil bacteria.
References
Allard B (2006) A comparative study on the chemical composition of humic acids from forest soil, agricultural soil and lignite deposit—bound lipid, carbohydrate and amino acid distributions. Geoderma 130(1–2):77–96
Amelung W, Zhang X (2001) Determination of amino acid enantiomers in soils. Soil Biol Biochem 33(4–5):553–562
Bakken LR (1985) Separation and purification of bacteria from soil. Appl Environ Microbiol 49(6):1482–1487
Benndorf D, Balcke GU, Harms H, Von Bergen M (2007) Functional metaproteome analysis of protein extracts from contaminated soil and groundwater. ISME J 1(3):224–234
Bollmann A, Lewis K, Epstein SS (2007) Incubation of environmental samples in a diffusion chamber increases the diversity of recovered isolates. Appl Environ Microbiol 73(20):6386–6390
Bunk B, Kucklick M, Jonas R, Munch R, Schobert M, Jahn D, Hiller K (2006) MetaQuant: a tool for the automatic quantification of GC/MS-based metabolome data. Bioinformatics 22(23):2962–2965
Burke L, Brozel V, Venter S (2008) Construction and evaluation of a gfp-tagged Salmonella Typhimurium strain for environmental applications. Water SA 34(1):19–24
Daniel R (2004) The soil metagenome—a rich resource for the discovery of novel natural products. Curr Opin Biotechnol 15(3):199–204
Davis KE, Joseph SJ, Janssen PH (2005) Effects of growth medium, inoculum size, and incubation time on culturability and isolation of soil bacteria. Appl Environ Microbiol 71(2):826–834
de Souza ML, Wackett LP, Boundy-Mills KL, Mandelbaum RT, Sadowsky MJ (1995) Cloning, characterization, and expression of a gene region from Pseudomonas sp. strain ADP involved in the dechlorination of atrazine. Appl Environ Microbiol 61(9):3373–3378
Ellis RJ (2004) Artificial soil microcosms: a tool for studying microbial autecology under controlled conditions. J Microbiol Methods 56(2):287–290
Fierer N, Jackson RB (2006) The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci USA 103(3):626–631
Fierer N, Bradford MA, Jackson RB (2007) Toward an ecological classification of soil bacteria. Ecology 88(6):1354–1364
Fischer H, Meyer A, Fischer K, Kuzyakov Y (2007) Carbohydrate and amino acid composition of dissolved organic matter leached from soil. Soil Biol Biochem 39(11):2926–2935
Gans J, Wolinsky M, Dunbar J (2005) Computational improvements reveal great bacterial diversity and high metal toxicity in soil. Science 309(5739):1387–1390
Hafner SD, Groffman PM, Mitchell MJ (2005) Leaching of dissolved organic carbon, dissolved organic nitrogen, and other solutes from coarse woody debris and litter in a mixed forest in New York State. Biogeochemistry 74(2):257–282
Hochgrafe F, Wolf C, Fuchs S, Liebeke M, Lalk M, Engelmann S, Hecker M (2008) Nitric oxide stress induces different responses but mediates comparable protein thiol protection in Bacillus subtilis and Staphylococcus aureus. J Bacteriol 190(14):4997–5008
Hoi LT, Voigt B, Jurgen B, Ehrenreich A, Gottschalk G, Evers S et al (2006) The phosphate-starvation response of Bacillus licheniformis. Proteomics 6(12):3582–3601
Huang Y, Eglinton G, Van der Hage ERE, Boon JJ, Bol R, Ineson P (1998) Dissolved organic matter and its parent organic matter in grass upland soil horizons studied by analytical pyrolysis techniques. Eur J Soil Sci 49(1):1–15
James N (1958) Soil extract in soil microbiology. Can J Microbiol 4(4):363–370
Joseph SJ, Hugenholtz P, Sangwan P, Osborne CA, Janssen PH (2003) Laboratory cultivation of widespread and previously uncultured soil bacteria. Appl Environ Microbiol 69(12):7210–7215
Kaiser K, Guggenberger G, Haumaier L, Zech W (2001) Seasonal variations in the chemical composition of dissolved organic matter in organic forest floor layer leachates of old-growth Scots pine (Pinus sylvestris L.) and European beech (Fagus sylvatica L.) stands in northeastern Bavaria, Germany. Biogeochemistry 55(2):103–143
Kalbitz K, Solinger S, Park JH, Michalzik B, Matzner E (2000) Controls on the dynamics dissolved organic matter in soils: a review. Soil Sci 165(4):277–304
Kovats E (1958) Gas-Chromatographische Charakterisierung Organischer Verbindungen .1. Retentionsindices Aliphatischer Halogenide, Alkohole, Aldehyde Und Ketone. Helv Chim Acta 41(7):1915–1932
Liebeke M, Pother DC, van Duy N, Albrecht D, Becher D, Hochgrafe F et al (2008) Depletion of thiol-containing proteins in response to quinones in Bacillus subtilis. Mol Microbiol 69(6):1513–1529
Luo Y, Vilain S, Voigt B, Albrecht D, Hecker M, Brozel VS (2007) Proteomic analysis of Bacillus cereus growing in liquid soil organic matter. FEMS Microbiol Lett 271(1):40–47
Makita M, Yamamoto S, Kono M (1976) Gas–liquid chromatographic analysis of protein amino acids as N-isobutyloxycarbonylamino acid methyl esters. J Chromatogr 120(1):129–140
Mongodin EF, Shapir N, Daugherty SC, DeBoy RT, Emerson JB, Shvartzbeyn A et al (2006) Secrets of soil survival revealed by the genome sequence of Arthrobacter aurescens TC1. PLoS Genet 2(12):e214
Nicholson JK, Foxall PJ, Spraul M, Farrant RD, Lindon JC (1995) 750 MHz 1H and 1H-13C NMR spectroscopy of human blood plasma. Anal Chem 67(5):793–811
Pace NR (1997) A molecular view of microbial diversity and the biosphere. Science 276(5313):734–740
Paliy O, Gunasekera TS (2007) Growth of E. coli BL21 in minimal media with different gluconeogenic carbon sources and salt contents. Appl Microbiol Biotechnol 73(5):1169–1172
Palmer KL, Mashburn LM, Singh PK, Whiteley M (2005) Cystic fibrosis sputum supports growth and cues key aspects of Pseudomonas aeruginosa physiology. J Bacteriol 187(15):5267–5277
Palmer KL, Aye LM, Whiteley M (2007) Nutritional cues control Pseudomonas aeruginosa multicellular behavior in cystic fibrosis sputum. J Bacteriol 189(22):8079–8087
Pizzeghello D, Zanella A, Carletti P, Nardi S (2006) Chemical and biological characterization of dissolved organic matter from silver fir and beech forest soils. Chemosphere 65(2):190–200
Radajewski S, McDonald IR, Murrell JC (2003) Stable-isotope probing of nucleic acids: a window to the function of uncultured microorganisms. Curr Opin Biotechnol 14(3):296–302
Rappe MS, Giovannoni SJ (2003) The uncultured microbial majority. Annu Rev Microbiol 57:369–394
Reasoner DJ, Geldreich EE (1985) A new medium for the enumeration and subculture of bacteria from potable water. Appl Environ Microbiol 49(1):1–7
Reizer J, Bachem S, Reizer A, Arnaud M, Saier MH Jr, Stulke J (1999) Novel phosphotransferase system genes revealed by genome analysis - the complete complement of PTS proteins encoded within the genome of Bacillus subtilis. Microbiology 145:3419–3429
Sandnes A, Eldhuset TD, Wollebaek G (2005) Organic acids in root exudates and soil solution of Norway spruce and silver birch. Soil Biol Biochem 37(2):259–269
Schleper C, Jurgens G, Jonuscheit M (2005) Genomic studies of uncultivated archaea. Nat Rev Microbiol 3(6):479–488
Schloss PD, Handelsman J (2006) Toward a census of bacteria in soil. PLoS Comput Biol 2(7):e92
Sharma PD (2005) Terrestial environments. In Environmental Microbiology. Alpha Science International, Harrow, Middlesex, UK, pp 27–51
Shivaji S, Suresh K, Chaturvedi P, Dube S, Sengupta S (2005) Bacillus arsenicus sp nov., an arsenic-resistant bacterium isolated from a sidente concretion in West Bengal, India. Int J Syst Evol Microbiol 55:1123–1127
Shivers RP, Dineen SS, Sonenshein AL (2006) Positive regulation of Bacillus subtilis ackA by CodY and CcpA: establishing a potential hierarchy in carbon flow. Mol Microbiol 62(3):811–822
Stotzky G, Burns RG (1982) The soil environment: clay–humus–microbe interactions. In: Burns RG, Slater JH (eds) Experimental microbial ecology. Blackwell Scientific Publishing, Oxford, p 100110
Strobel BW (2001) Influence of vegetation on low-molecular-weight carboxylic acids in soil solution—a review. Geoderma 99(3–4):169–198
Stulke J, Hanschke R, Hecker M (1993) Temporal activation of beta-glucanase synthesis in Bacillus subtilis is mediated by the Gtp pool. J Gen Microbiol 139:2041–2045
Tobisch S, Zuhlke D, Bernhardt J, Stulke J, Hecker M (1999) Role of CcpA in regulation of the central pathways of carbon catabolism in Bacillus subtilis. J Bacteriol 181(22):996–7004
Torsvik V, Øvreås L (2002) Microbial diversity and function in soil: from genes to ecosystems. Curr Opin Microbiol 5(3):240–245
Torsvik V, Goksoyr J, Daae FL (1990) High diversity in DNA of soil bacteria. Appl Environ Microb 56(3):782–787
Torsvik V, Ovreas L, Thingstad TF (2002) Prokaryotic diversity—magnitude, dynamics, and controlling factors. Science 296(5570):1064–1066
Tringe SG, von Mering C, Kobayashi A, Salamov AA, Chen K, Chang HW et al (2005) Comparative metagenomics of microbial communities. Science 308(5721):554–557
Urich T, Lanzen A, Qi J, Huson DH, Schleper C, Schuster SC (2008) Simultaneous assessment of soil microbial community structure and function through analysis of the meta-transcriptome. PLoS ONE 3(6):e2527
van Hees PAW, Jones DL, Finlay R, Godbold DL, Lundstomd US (2005) The carbon we do not see—the impact of low molecular weight compounds on carbon dynamics and respiration in forest soils: a review. Soil Biol Biochem 37(1):1–13
Veith B, Herzberg C, Steckel S, Feesche J, Maurer KH, Ehrenreich P et al (2004) The complete genome sequence of Bacillus licheniformis DSM13, an organism with great industrial potential. J Mol Microbiol Biotechnol 7(4):204–211
Vilain S, Luo Y, Hildreth MB, Brozel VS (2006) Analysis of the life cycle of the soil saprophyte Bacillus cereus in liquid soil extract and in soil. Appl Environ Microb 72(7):4970–4977
Voigt B, Hoi LT, Jurgen B, Albrecht D, Ehrenreich A, Veith B et al (2007) The glucose and nitrogen starvation response of Bacillus licheniformis. Proteomics 7(3):413–423
Watanabe K, Imase M, Aoyagi H, Ohmura N, Saiki H, Tanaka H (2008) Development of a novel artificial medium based on utilization of algal photosynthetic metabolites by symbiotic heterotrophs. J Appl Microbiol 105(3):741–751
Acknowledgements
We are grateful to S. Seefeld for performing the ICP and IC. We thank K. Surmann for the assistance and Dr. M. Sadowsky for donating the bacterial strains. This research was funded by a grant from the Apotheker-Paul-Marschall-Stiftung to M. Liebeke and by the South Dakota Agricultural Experiment Station and the State of South Dakota. VSB was the recipient of a fellowship from the Stiftung Alfried Krupp Kolleg Greifswald.
Author information
Authors and Affiliations
Corresponding author
Additional information
Journal series publication 3622 from the South Dakota Agricultural Experiment Station.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
Growth of Bacillus licheniformis DSM 13 in synthetic media and appending metabolite concentrations [in millimolars] of culture supernatants (determined by 1H-NMR measurements) for M9 medium with a carbon source cocktail in low concentrations (a) and high concentrations (b). Also displayed is growth in M9 medium with glucose in low concentrations (c) and high concentrations (d) as carbon source. All experiments were done in triplicate and a representative experiment is shown (DOC 221 kb)
Rights and permissions
About this article
Cite this article
Liebeke, M., Brözel, V.S., Hecker, M. et al. Chemical characterization of soil extract as growth media for the ecophysiological study of bacteria. Appl Microbiol Biotechnol 83, 161–173 (2009). https://doi.org/10.1007/s00253-009-1965-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-009-1965-0