Abstract
The efe gene encoding an ethylene-forming enzyme from Pseudomonas syringae pv. glycinea has been expressed for the first time under the control of Trichoderma reesei cbh1 promoter in Trichoderma viride. Reverse transcription polymerase chain reaction analysis showed that transformant Y2 produced mRNA of the efe gene. Southern blot analysis showed that there was one copy of efe gene which was integrated into the chromosomal DNA of T. viride. Ethylene production by transformant Y2 was efficiently induced by cellulose, while very low level of ethylene was produced when sodium carboxymethyl cellulose or lactose was used as carbon source. Peptone exerted a much greater stimulatory effect on ethylene production. A high level of ethylene was produced when transformant Y2 was cultured in solid fermentation medium containing wheat straw, indicating that plant wastes could be directly converted to ethylene by the recombinant filamentous fungus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ethylene is the most important starting material in petroleum chemistry and it is also a gaseous plant hormone involved in the regulation of numerous physiological processes such as fruit ripening and senescence. It is well-known that ethylene is not only produced by plants, but also by microorganisms (Chalutz and Lieberman 1977; Nagahama et al. 1992; Sato et al. 1997; Chague et al. 2002). However, for large-scale industrial application, ethylene is made from hydrocarbon feedstocks such as natural gas liquids or crude oil. As the global fossil fuel reserves decline, it is desirable to develop a process for the microbial production of ethylene from various renewable resources.
Microorganisms produce ethylene via two different biochemical pathways. Both pathways are distinct from that of higher plants, which use only one pathway from methionine via 1-aminocyclopropane-1-carboxylic acid (Jiao et al. 1986; Hall and Smith 1995; Weingart et al. 1999). Most microorganisms produce only trace amounts of ethylene from methionine via 2-keto-4-methyl-thiobutyric acid by an NADH:Fe(III)EDTA oxidoreductase (Weingart et al. 1999). Four pathovars of Pseudomonas syringae and a few species of fungi such as Penicillium digitatum produce efficiently ethylene from 2-oxoglutarate by an ethylene-forming enzyme (EFE) (Nagahama et al. 1991; Völksh and Weingart 1997). The efe gene from P. syringae pv. phaseolicola PK2 has been cloned and expressed in Escherichia coli at high level (Ishihara et al. 1995). In order to make the biological conversion of carbon dioxide into ethylene, the efe gene from P. syringae pv. phaseolicola PK2 has been introduced into cyanobacterium Synechococcus sp. PCC 7942 and the recombinant cyanobacterial strains produce ethylene at a low level (Sakai et al. 1997; Takahama et al. 2003).
The filamentous Trichoderma species, such as Trichoderma reesei and Trichoderma viride, are known to be good cellulase producers and they produce a complete set of cellulases consisted of three general classes of enzymes: cellubiohydrolases (CBHI and CBHII), endoglucanases (EGI, EGII, and EGIII), and β-glucosidase, which act synergistically to hydrolyze crystalline cellulose to glucose (Henrique-Silva et al. 1996; Ilmen et al. 1997; Sun et al. 1997; Van Wyk and Mohulatsi 2003). Homologous and heterologous genes under the strong main cellobiohydrolase (cbh1) promoter of T. reesei have been successfully overexpressed in T. reesei and the related species Trichoderma harzianum (Joutsjoki et al. 1993; Miettinen-Oinonen et al. 1997; Margolles-Clark et al. 1996a, b). Extensive studies showed that a high level of cellulase gene expression of T. reesei was induced by insoluble polymer such as cellulose and plant material (Ilmen et al. 1997). Thus it will be a very important potential value to introduce efe gene into Trichoderma species and to enable the recombinant Trichoderma species to produce ethylene from agricultural wastes.
In this study, efe gene encoding EFE was cloned and sequenced from P. syringae pv. glycinea ICMP2189 and was then successfully expressed under the control of T. reesei cbh1 promoter in T. viride. We further characterized the effect of different carbon sources or nitrogen source peptone on efe gene expression.
Materials and methods
Microorganisms and plasmids
P. syringae pv. glycinea ICMP2189 was obtained from Plant Protection Institute of Chinese Academy of Agricultural Science. T. viride TL124, which has a strong cellulolytic activity, was isolated by our lab. Transformant Y2 carrying efe gene from P. syringae pv. glycinea ICMP2189 was developed in this study. E. coli DH5α was used as a host for cloning and sequencing. Plasmid pMD18-T was used as a cloning vector and pTRIL, which carries T. reesei cbh1 promoter and terminator, was used as an expression vector in T. viride (Wang et al. 2004).
Cloning of efe gene and construction of recombinant plasmid pTRIL-efe
A 1,071-bp fragment including the efe coding region, was amplified with primers P1 (5’-ccgtcgacatgaccaacctacagactte-3’ [a SalI site italicized and the efe start codon indicated by bold letters]) and P2 (5’-tatggatccaactcatgagcctgtcgcg-3’ [a BamHI site italicized and the efe stop codon indicated by bold letters]) from P. syringae pv. glycinea ICMP2189. The polymerase chain reaction (PCR) product was ligated to vector pMD18-T and then sequenced. At the same time, the amplified fragment was digested with SalI and BamHI and ligated to vector pTRIL digested with the same enzymes, yielding recombinant plasmid pTRIL-efe (Fig. 1).
Protoplast preparation and transformation of T. viride TL124
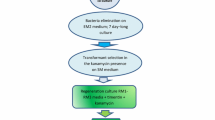
Protoplasts of T. viride TL124 were prepared as described by Penttila et al. (1987). The recombinant plasmid pTRIL-efe was cotransformed with plasmid pAN7–1 carrying a hygromycin resistance cassette into the protoplasts of T. viride TL124 as described by Wang et al. (2004). The hygromycin-resistant transformants were selected on minimal medium containing 250 μg ml−1 hygromycin. The positive transformant Y2 was obtained and identified by PCR amplification with the above-mentioned primers P1 and P2 for efe gene and by Southern blot analysis.
Southern blot analysis
The entire efe gene fragment was recovered from the recombinant plasmid pTRIL-efe and then was labeled using DIG-DNA labeling and detection kit purchased from Roche (Germany). The chromosomal DNA of transformant Y2 was isolated by using a modified CTAB procedure as described by Tel-Zur et al. (1999). Southern blot analysis was carried out using the labeled efe gene as a probe according to the instructions of the DIG-DNA labeling and detection kit.
RT-PCR amplification
Total RNAs were isolated from the fresh mycelia of transformant Y2 using the Trizol reagent (Invitrogen) according to the manufacturer’s protocol, and then reverse-transcribed to cDNA by using oligo-dT primers in the reverse transcription (RT)-PCR with the instructions of AMV reverse transcriptase (Calo et al. 2006). The resulting cDNA was amplified with primers P1 and P2 for efe gene under the above-mentioned PCR conditions.
Ethylene production of transformant Y2 on cellulose-induction medium
T. viride strains were maintained on potato dextrose agar. For the inducible production of ethylene under the control of T. reesei cbh1 promoter, cellulose-inducing (CI) medium was used, which was Trichoderma minimal medium (Ilmen et al. 1997), in which glucose was substituted with 0.5% cellulose (Penttila et al. 1987).
To determine ethylene production of transformant Y2 on CI medium, experiments were carried out as follows: 0.5 ml of 107 ml−1 spore suspension, collected from cultures grown on potato dextrose agar, was inoculated into 50 ml of the minimal medium in a 250-ml flask and grown at 30 °C with shaking at 200 rpm. At 48 h after incubation, mycelia were collected, washed, and transferred into 2 ml cellulose-induction medium in 12 × 120 mm test tube covered with a rubber stopper and grown with shaking at 200 rpm at 30 °C. After 36 h of incubation, gas samples were then removed from test-tube using a syringe and the dry weight of mycelia was determined following drying for 48 h at 70 °C. Ethylene levels were determined by injecting 0.1 ml gas samples into a HP6890 gas chromatograph as described by Chalutz and Lieberman (1977). Ethylene production rates were calculated as nanoliter ethylene produced in 1 h per gram of dry weight of mycelium (nl h−1 g−1 dry wt) (Chalutz and Lieberman 1977; Chague et al. 2002).
Effects of carbon sources and peptone on ethylene production
To determine the effect of different carbon sources and peptone on ethylene production, Trichoderma minimal medium was used, in which 2% glucose was substituted with different carbon sources such as 0.5% cellulose, 2% cellulose, 0.5% sodium carboxymethyl cellulose, or 3% lactose. Nitrogen source such as 0.2% peptone was added to the medium containing different carbon sources when indicated.
Ethylene production cultured in solid state fermentation
To evaluate the effect of plant material as substrate on ethylene production, the solid fermentation medium containing wheat straw is here developed. The wheat straw used here was washed, chopped, dried (70 °C to constant weight), and ground to 40-mesh size. The solid fermentation medium in 25-ml serum bottle contains pretreated wheat straw, 0.72 g; bran, 0.08 g; (NH4)2SO4, 0.02 g; and water, 2.0 ml, being autoclaved at 121 °C for 30 min. Then 0.08 ml of 107 ml−1 spore suspension was inoculated into 25-ml serum-bottle containing the above-described fermentation medium and grown at 30 °C under stationary conditions. At 48 h after incubation, cotton plugs covered in serum bottle was replaced with rubber stoppers and the cultures were further grown for 48 h and then the total amount of ethylene that accumulated in the flask was determined. The total amount of ethylene per flask was calculated by multiplying the free volume in the flask (25-ml) by the amount of ethylene in 1 ml gas sample as determined by gas chromatographic analysis (Chague et al. 2002).
During the determination of ethylene production, all treatments were in three replicates and all the experiments were repeated three times.
Results
Cloning and sequencing of efe gene
A 1,071-bp fragment including the complete efe coding region was PCR-amplified from P. syringae pv. glycinea ICMP2189 using primers designed according to the known efe genes of P. syringae. The GenBank accession number for the efe gene isolated in this study is EF17587. Sequence alignments showed that the sequence of the open reading frame from P. syringae pv. glycinea strain ICMP2189 was completely identical to those derived from P. syringae pvs. glycinea 7a/90, cannabina GSPB2553 and sesami 962 and differed in only two single-base pair substitutions from the efe nucleotide sequence of P. syringae pv. phaseolicola PK2. Our data were in agreement with Weingart et al’s report (1999, 2001) that a high conservation of the efe gene among P. syringae pvs. phaseolicola, glycinea, cannabina and sesami, all of which were efficient ethylene-producers (Sato et al. 1997; Weingart and Völksch 1997).
Transformation of T. viride with expression plasmid carrying efe gene
For overexpression of the efe gene in T. viride, the powerful promoter of the T. ressei cbh1 gene was used. The plasmid pTRIL-efe containing the entire coding region of the P. syringae pv. glycinea efe gene linked to the chb1 promoter and terminator was constructed. The constructed plasmids pTRIL-efe was cotransformed into T. viride TL124 with plasmid pAN7–1, which contains a hygromycin resistance cassette. Transformants were selected on plates containing hygromycin. About 200 hygromycin-resistant transformants obtained here were further identified by PCR analysis using P1 and P2 primers for efe gene. The PCR product of plasmid DNA (pTRIL-efe) was used as a positive control. PCR analysis showed that five potential transformants contained efe gene, while no corresponding amplified fragment could be detected in the wild-type T. viride.
Southern blot analysis
Southern blot analysis using the labeled efe gene as a probe was further performed with genomic DNA isolated from the wild-type T. viride and from the potential transformants identified by PCR analysis. The results showed that efe gene in transformant Y2 was integrated into the genome of the recipient, whereas no hybridization signals were observed in the other four transformants or wild-type T. viride in this study. Figure 2 showed that HindIII- and SalI-digested genomic DNA isolated from transformant Y2 gave a different single positive signal band at the positions of 5 kb and 1.2 kb, respectively. Since the length of the coding region of the P. syringae pv. glycinea efe gene is 1.053 kb, thus we deduce from the current results that there is one copy of efe gene which was integrated into the chromosomal DNA of transformant Y2.
Southern blot analysis of genomic DNA isolated from transformant Y2. Lane 1 DNA molecular weight marker (λDNA/HindIII+EcoRI); Lanes 2 and 3 genomic DNA of transformant Y2 digested respectively with HindIII and SalI; Lane 4 genomic DNA of non-transformated T. viride digested with HindIII. The whole coding sequence of efe gene (1,053 bp) was used as a hybridizing probe
RT-PCR analysis
RT-PCR analysis was performed with total RNA isolated from transformant Y2 that were confirmed by Southern blot analysis. The total RNA of the wild-type T. viride was analyzed as a parallel control. The data showed that accumulation of mRNA of the efe gene could only be detected in transformant Y2 (Fig. 3). No detectable level of transcript, however, was observed in the recipient T. viride.
RT-PCR analysis of EFE mRNA in transformant Y2. Lane 1 1 kb DNA ladder; Lane 2 RT-PCR amplification of first-strand cDNA from transformant Y2; Lane 3 PCR product from plasmid pTRIL-efe as a positive control; Lane 4 PCR amplification of total RNA treated with DNaseI as a negative control. Lack of a signal in this lane indicates that PCR product in lane 2 are due to the amplification of cDNA, rather than low levels of genomic DNA; Lane 5 RT-PCR amplification of the wild-type T. viride as a negative control
Ethylene production of transformant Y2
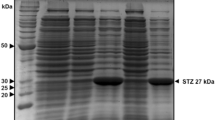
Previous reports have demonstrated that cellulose induces the expression of T. reesei cbh1 promoter in T. reesei or T. harzianum (Margolles-Clark et al. 1996a, b). To determine if ethylene was produced by transformant Y2, CI medium which contains 0.5% cellulose as the sole carbon source was used. Before ethylene assay, it has been determined that there were no differences in growth rate or morphology between transformant Y2 and the wild-type T. viride, indicating that integration of efe gene into the genome of T. viride does not seem to be harmful to the host. Gas chromatograph assay showed that transformant Y2 produced ethylene efficiently (the ethylene production rate was 1,059.7 nl h−1 g−1 dry wt), while no production of ethylene was observed in the wild-type T. viride (Table 1). The ethylene level produced by transformant Y2 was similar to that produced by P. digitatum, which is the most efficient ethylene-producer among fungi (Spalding and Lieberman 1964). The results indicate that the efe gene from P. syringae pv. glycinea was successfully expressed in T. viride. This is the first time to express the efe gene of P. syringae under the control of T. reesei cbh1 promoter in Trichoderma species.
Effects of carbon sources and peptone on ethylene production
As different carbon sources and peptone affect the expression of cbh1 promoter, we here determined the ethylene production rates in media containing different carbon sources or peptone as described in “Materials and methods”. Our results, presented in Fig. 4, revealed that 2% cellulose induced ethylene production more efficiently than 0.5% cellulose did. Very low level of ethylene was produced when sodium carboxymethyl cellulose or lactose was used as carbon source. The results are a little different from the previous report that lactose induced a moderate expression of cbh1 in T. reesei (Ilmen et al. 1997). Nitrogen source peptone exerts a much greater stimulatory effect on ethylene production when added to minimal medium supplemented with different carbon sources, especially in which containing 2% cellulose, in agreement with the report that peptone provoked production of cellulases in Chaetomium globosu (Umikalsom et al. 1997).
Ethylene production cultured in solid state fermentation
To determine if transformant Y2 could utilize plant wastes to produce ethylene, here we developed a solid fermentation medium which contains wheat straw as the main carbon source. Transformant Y2 was found to produce ethylene at the amount of 2,280 nl when cultured in this solid fermentation medium for 48 h at 30 °C. The result suggests that plant wastes could be directly converted to ethylene by the recombinant filamentous fungus.
Discussion
In this study, we have established a high expression system of P. syringae pv. glycinea efe gene using T. viride as a host. We further analyzed the effects of different carbon and nitrogen sources on ethylene production by transformant Y2.
So far, overexpression of P. syringae pv. phaseolicola efe gene has been performed by using E. coli and Synechoccus elongates as hosts (Ishihara et al. 1995; Sakai et al. 1997; Takahama et al. 2003). However, expression of P. syringae efe gene has not been carried out in filamentous fungi. In this study, the bacterial efe gene was linked to the T. reesei cbh1 promoter and then transformed to T. viride. The positive transformant Y2 carrying P. syringae pv. phaseolicola efe gene was obtained from about 200 hygromycin-resistant transformants and identified by PCR amplification and by Southern blot analysis. Initially, five potential transformants containing efe gene were identified by PCR amplification. However, Southern blot analysis using the labeled efe gene as a probe showed that only transformant Y2 gave a positive signal while the other four potential transformants did not. The data suggested that efe gene was integrated into the genome in transformant Y2 while efe gene was not in the other four hygromycin-resistant transformants. Thus we chose transformant Y2 for further experiments. Since Southern blot analysis showed that HindIII- and SalI-digested genomic DNA gave a different single positive signal band at the positions of 5 kb and 1.2 kb, we deduce that there is one copy of efe gene integrated into the chromosomal DNA of transformant Y2. RT-PCR and ethylene assay demonstrated that the efe gene encoding an EFE has been successfully expressed in transformant Y2. The current results that cellulose and peptone induced a high level of ethylene production indicated that expression of the bacterial efe gene in transformant Y2 was strictly under the control of T. reesei cbh1 promoter. The data are consistent with the report that cellulose or plant materials induces expression of the T. reesei cbh1 promoter in T. reesei or in T. harzianum (Margolles-Clark et al. 1996a, b; Ilmen et al. 1997). Our results also showed that transformant Y2 produced efficiently ethylene on the solid fermentation medium which contains wheat straw as a main carbon source. Since T. viride has a very strong cellulolytic activity, using T. viride as a host to produce ethylene seems to be attractive in industrial use.
References
Calo L, García I, Gotor C, Romero LC (2006) Leaf hairs influence phytopathogenic fungus infection and confer an increased resistance when expressing a Trichoderma α-1,3-glucanase. J Exp Bot 57:3911–3920
Chague V, Elad Y, Barakat R, Tudzynski P, Sharon A (2002) Ethylene biosynthesis in Botrytis cinerea. FEMS Microbiol Ecol 40:143–149
Chalutz E, Lieberman M (1977) Methionine-induced ethylene production by Penicillium digitatum. Plant Physiol 60:402–406
Hall MA, Smith AR (1995) Ethylene and the responses of plants to stress. Bulg J Plant Physiol 21:71–79
Henrique-Silva F, El-Gogary S, Carle-Urioste JC, Matheucci EJ, Crivellaro O, El-Dorry H (1996) Two regulatory regions controlling basal and cellulose-Induced expression of the gene encoding cellobiohydrolase I of Trichoderma reesei are adjacent to Its TATA Box. Biochem Biophys Res Commun 228:229–237
Ilmen M, Saloheimo A, Onnela ML, Penttila ME (1997) Regulation of cellulase gene expression in the filamentous fungus Trichoderma reesei. Appl Environ Microbiol 63:1298–1306
Ishihara K, Matsuoka M, Inoue Y, Tanase S, Ogawa T, Fukuda H (1995) Overexpression and in vitro reconstitution of the ethylene-forming enzyme from Pseudomonas syringae. J Ferment Bioeng 79:205–211
Jiao XZ, Philosoph-Hadas S, Su LY, Yang SF (1986) The Conversion of 1-(malonylamino)cyclopropane- l-carboxylic acid to 1-aminocyclopropane-1-carboxylic acid in plant tissues. Plant Physiol 81:637–641
Joutsjoki V, Torkkeli T, Nevalainen H (1993) Transformation of Trichoderma reesei with the Hormoconis resinae glucoamylase P (gamP) gene: production of a heterologous glucoamylase by Trichoderma reesei. Curr Genet 24:223–229
Margolles-Clark E, Hayes CK, Harman GE, Penttila M (1996a) Improved production of Trichoderma harzianum endochitinase by expression in Trichoderma reesei. Appl Environ Microbiol 62:2145–2151
Margolles-Clark E, Harman GE, Penttila M (1996b) Enhanced expression of endochitinase in Trichoderma harzianum with the cbh1 Promoter of Trichoderma reesei. Appl Environ Microbiol 62:2152–2155
Miettinen-Oinonen A, Torkkeli T, Paloheimo M, Nevalainen H (1997) Overexpression of the Aspergillus niger pH 2.5 acid phosphatase gene in a heterologous host Trichoderma reesei. J Biotechnol 58:13–20
Nagahama K, Ogawa T, Fujii T, Tazaki M, Tanase S, Morino Y, Fukuda H (1991) Purification and properties of ethylene-forming enzyme from Pseudomonas syringae pv. phaseolicola PK2. J Gen Microbiol 137:2281–2286
Nagahama K, Ogawa T, Fujii T, Fukuda H (1992) Classification of ethylene producing bacteria in terms of biosynthetic pathways to ethylene. J Ferment Bioeng 73:1–5
Penttila M, Nevalainen H, Ratto M, Salminen E, Knowles J (1987) A versatile transformation system for the cellulolytic filamentous fungus Trichoderma reesei. Gene 61:155–164
Sakai M, Ogawa T, Matsuoka M, Fukuda H (1997) Photosynthetic conversion of carbon dioxide to ethylene by the recombinant cyanobacterium, Synechococcus sp. PCC 7942, which harbours a gene for the ethylene-forming enzyme of Pseudomonas syringae. J Ferment Bioeng 84:434–443
Sato M, Watanabe K, Yazawa M, Takikawa Y, Nishiyama K (1997) Detection of new ethylene-producing bacteria, Pseudomonas syringae pvs. cannabina and sesami, by PCR amplification of genes for the ethylene-forming enzyme. Phytopathology 87:1192–1196
Spalding DH, Lieberman M (1964) Factors affecting the production of ethylene by Penicillium digitatum. Plant Physiol 40:645–648
Sun T, Liu BH, Liu DM, Li ZH (1997) Effect of elevated temperature on Trichoderma viride SL-1 in solid state fermentations. Biotechnol Lett 19:171–174
Takahama K, Matsuoka M, Nagahama K, Ogawa T (2003) Construction and analysis of a recombinant cyanobacterium expressing a chromosomally inserted gene for an ethylene-forming enzyme at the psbA1 locus. J Ferment Bioeng 95:302–305
Tel-Zur N, Abbo S, Myslabodski D, Mizrahi Y (1999) Modified CTAB procedure for DNA isolation from epiphytic cacti of the genera Hylocereus and Selenicereus (cactaceae). Plant Mol Biol Report 17:249–254
Umikalsom MS, Ariff AB, Zulkifli HS, Tong CC, Hassan MA, Karim MIA (1997) Production of cellulase by a wild strain of Chaetomium globosum using delignified oil palm empty-fruit-bunch fibre as substrate. Appl Microbiol Biotechnol 47:590–595
Van Wyk JPH, Mohulatsi M (2003) Biodegradation of wastepaper by cellulase from Trichoderma viride. Bioresource Technol 86:21–23
Völksh B, Weingart H (1997) Comparison of ethylene-producing Pseudomonas syringae strains isolated from kudzu (Pueraria lobata) with Pseudomonas syringae pv. phaseolicola and Pseudomonas syringae pv. glycinea. Eur J Plant Pathol 103:795–802
Wang TH, Liu L, Wu ZH, Liu SL, Qu YB (2004) Novel cellulase profile of Trichoderma reesei strains constructed by cbh1 gene replacement with eg3 gene expression cassette. Acta Biochim Biophys 36:667–672
Weingart H, Völksch B (1997) Ethylene production by Pseudomonas syringae pathovars in vitro and in planta. Appl Environ Microbiol 63:156–161
Weingart H, Völksch B, Ullrich MS (1999) Comparison of ethylene production by Pseudomonas syringae and Ralstonia solanacearum. Phytopathology 88:360–365
Weingart H, Ullrich MS, Geider K, Völksch B (2001) The role of ethylene production in virulence of Pseudomonas syringae pvs. glycinea and phaseolicola. Phytopathology 91:511–518
Acknowledgements
We wish to thank Dr. Qun He for discussions. This work was supported by China high Technology (863) Project (Grant No. 2006AA10A213).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tao, L., Dong, HJ., Chen, X. et al. Expression of ethylene-forming enzyme (EFE) of Pseudomonas syringae pv. glycinea in Trichoderma viride . Appl Microbiol Biotechnol 80, 573–578 (2008). https://doi.org/10.1007/s00253-008-1562-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-008-1562-7