Abstract
Mycobacterium tuberculosis is a bacterial pathogen that can persist for decades in an infected patient without causing a disease. In vivo, the tubercle bacillus present in the lungs store triacylglycerols in inclusion bodies. The same process can be observed in vitro when the bacteria infect adipose tissues. Indeed, before entering in the dormant state, bacteria accumulate lipids originating from the host cell membrane degradation and from de novo synthesis. During the reactivation phase, these lipids are hydrolysed and the infection process occurs. The degradation of both extra and intracellular lipids can be directly related to the presence of lipolytic enzymes in mycobacteria, which have been ignored during a long period particularly due to the difficulties to obtain a high expression level of these enzymes in M. tuberculosis. The completion of the M. tuberculosis genome offered new opportunity to this kind of study. The aim of this review is to focus on the recent results obtained in the field of mycobacterium lipolytic enzymes and although no experimental proof has been shown in vivo, it is tempting to speculate that these enzymes could be involved in the virulence and pathogenicity processes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Although tuberculosis is one of the oldest plagues and has probably attracted more research than any other disease, it is still the most deadly infectious disease in the world. It has been estimated that one third of the world’s population has been contaminated by Mycobacterium tuberculosis (M. tuberculosis), the etiologic agent responsible for this disease. Despite the existence of various treatments for this disease, it leads to the death of 2.5–3 million of people every year (WHO 2004).
The present upsurge of tuberculosis is directly linked to the spread of AIDS and the emergence of new multi-drug resistant (MDR) and extensively drug-resistant (XDR) strains (CDC 2006). The apparition of MDR and XDR strains is a serious barrier to controlling this disease because these strains can resist to the first and second line of drug treatment, respectively.
Unlike most bacterial pathogens, M. tuberculosis establishes persistent infection in the host and is resistant to the host’s immune system. The strategy used by the host to prevent the growth of M. tuberculosis during persistent infection consists in forming granulomas, which are aggregates of immune cells around foci and infected tissues (Fig. 1). Granulomas are thought to contain high levels of carbon dioxide and aliphatic organic acids (Wayne and Sohaskey 2001). The state of dormancy thus induced in the pathogen is probably triggered by the environmental stress exerted by the host’s immune response. The bacterium is able to survive in this dormant state for decades until the host’s immune system is weakened, due to AIDS disease for example. In these conditions, bacteria reactivate and cause the infectious disease. Unfortunately, the anti-mycobacterial drugs available so far mostly target the actively replicating mycobacteria and do not affect the latent bacteria (persisters) or MDR and XDR strains (Goldman et al. 2007; Gomez and McKinney 2004). Due to this dormant phase, tuberculosis control is still a major problem.
Model of a granuloma inspired from (Honer zu Bentrup and Russell 2001; Russell 2007). The center consists of a cheese-like, semi-solid structure with a low oxygen content, which is richly endowed with lipids and proteins originating from host cells and bacteria. The center is surrounded by activated or partially activated macrophages as well as non-activated macrophages and lymphocytes. Persistent bacteria can survive there for decades. In the event that liquefaction of the center occurs, water is absorbed from the surrounding tissues, oxygen enters, and an excellent culture medium for the extracellular growth of the surviving mycobacteria is formed
Achieving a better understanding of the biochemistry of enzymes involved in various metabolic pathways and their physiological role, and identifying the genes required for the pathogenic processes to occur is a high priority issue in tuberculosis research. Studies on these lines could lead to the development of new therapeutic agents to combat M. tuberculosis. The recent determination of the complete genome sequence of the most widely known h37Rv strain of M. tuberculosis (Cole et al. 1998) has opened new perspectives for developing novel drugs to treat tuberculosis. The current genome annotation consists of ~4,000 predicted proteins, classified into 11 distinct protein groups (Camus et al. 2002), 48% of which have no known function. Comparative sequence analysis of the M. tuberculosis genome has shown that M. tuberculosis encodes a remarkably wide range of proteins involved in lipid metabolism, which is unrivalled in the whole prokaryotic world. In Escherichia coli, for example, approximately 50 enzymes are involved in lipid metabolism, whereas the M. tuberculosis genome contains at least 250 genes encoding for these enzymes. Since lipids account for 60% of the dry weight of the bacillus, lipid metabolism can be assumed to be one of the major pathways in mycobacteria and needs to be studied in detail.
This review focuses on lipolytic enzymes involved in lipid degradation processes in mycobacteria. After describing the patterns of lipids distribution in this microorganism, we will present the first results obtained in this field and explain why these enzymes are liable to contribute importantly to the physiology of these bacteria.
Lipid distribution in mycobacteria
Considerable research has focused on the lipids present in mycobacteria, especially those involved in the cell wall. The importance of lipids in the M. tuberculosis life cycle has been established and the cell envelope has been found to have a most unusual lipid composition, including mycolic acids, phenolphthiocerol alcohol, phthiocerol dimycocerosate (PDIM) and glycolipids (LAM, PIM), which are known to be involved in the virulence of the bacteria and their pathogenicity (Daffe and Draper 1998; Kremer et al. 2005; Minnikin 1982; Minnikin et al. 2002).
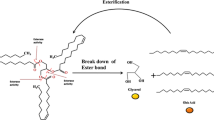
In addition, the cytosol has been found to be an important lipid storage location: an intracellular lipid storage process was recently described in vivo in the cytosol of M. tuberculosis, forming organites known as lipid inclusion bodies. These organites are essentially composed of triacylglycerols (TAG) and could provide the carbon source required for bacterial growth to occur during the chronic infection phase (Fig. 2a; Alvarez and Steinbüchel 2002; Dawes and Senior 1973; Garton et al. 2002; Koval’chuk et al. 1973; Wältermann and Steinbüchel 2005). The accumulation of TAG in lipid inclusion bodies has also been observed directly in vitro by infecting adipocytes with M. tuberculosis (Fig. 2b; Neyrolles et al. 2006). Although these particular tissues could be a potential niche for the bacteria, there is no evidence so far in vivo that this location might contain viable bacteria. TAG are convenient carbon and energy storage compounds because they show low rates of oxidation and a high calorific value. The storage of these lipids in vivo could be important for the bacteria and enables them to adapt to the environment by preparing them to enter in the dormant phase on similar lines to what occurs in hibernating animals, plant seeds and microbial spores. Under starvation conditions, these TAG contained in these lipid inclusion bodies are hydrolysed reflecting the presence of an intracellular lipase (Deb et al. 2006; Mishra et al. 2008) that could provide free fatty acid (FA) useful for many other metabolic functions yet unknown in vivo (Fig. 3). For example, based on in vitro studies, it seems that stored TAG may provide FA useful not only for the constitution of membrane lipids (Kremer et al. 2005) but also for the synthesis of wax ester under iron limitation (Bacon et al. 2007; Tsujita et al. 1999). However, due to the absence of in vivo study, there is no information whether this enzyme plays a role in the viability or in the infection processes (persistence or reactivation).
a Image of an acid-fast cell containing intracellular lipid inclusions in a sputum sample from a infected patient with clinical tuberculosis (Garton et al. 2002; reproduction with permission). The cell was dual-labelled with Auramine O and Nile red (Scale bar: 2 µm). b Electron microscopic view of mycobacteria growing on adipocytes loaded in small vesicles resembling lipid inclusions (Neyrolles et al. 2006; reproduction with permission)
M. tuberculosis genome contains 15 triacylglycerol synthase genes belonging to a non-classical diacylglycerol acyltransferase family. Some of these genes have been induced under in vitro conditions simulating dormancy and a concomitant accumulation of TAG inside bacterial cells was observed (Daniel et al. 2004). In vivo, FA may result from the degradation of phospholipids forming the membrane of the host cells, particularly during the formation of the granuloma (Fig. 1). In these histologic structures, these phospholipids may be hydrolysed by either phospholipases A (PLA) or phospholipases C (PLC), resulting in the release of lysophospholipids and diacylglycerols (DAG), respectively (Fig. 3; Honer zu Bentrup and Russell 2001). At this stage, the resulting DAG may therefore be hydrolysed by an excreted lipase and the FA released may then be absorbed by the microorganism. A synergy of this kind between lipases and phospholipases has been previously described in the case of another pathogen, Pseudomonas aeruginosa, which degrades lung surfactants (phospholipids) via the combined action of a PLC and a lipase (Konig et al. 1994). Excreted PLA and PLC have been previously described in M. tuberculosis (Raynaud et al. 2002; Stonehouse et al. 2002). However, only one intracellular and one extracellular lipase have been identified, purified and characterised so far in M. tuberculosis (Côtes et al. 2007; Deb et al. 2006; Mishra et al. 2008).
Lipolytic enzymes
The disappearance of lipid inclusion bodies during the reactivation phase and the degradation of the host cell membranes therefore suggests the existence of intra and extracellular lipases. Thanks to the sequence analysis of the M. tuberculosis genome, it was observed that M. tuberculosis possesses an unusual large number of gene-encoding enzymes involved in lipid degradation. This feature suggests that the use of host lipids for growth in vivo may be one of the major characteristics of the M. tuberculosis life cycle. From the genome annotation, it has been concluded that 24 genes may encode lipolytic enzymes: these proteins have been classified in a family of enzymes called the “Lip family”. However, this classification is only based on the presence of the consensus sequence GXSXG, which is characteristic of esterases and members of the α/β hydrolase fold family. This classification does not allow the distinction between lipolytic enzymes (lipase) and non-lipolytic enzymes (esterase). Using the genome sequence analysis cannot be the sole criterion for classification purposes and, up to now, only biochemical characterization has provided the means of discriminating between lipolytic and non-lipolytic enzymes. Studies of this kind have shown that the “Lip family” defined by blast analysis is composed of both lipolytic and non-lipolytic enzymes (Camus et al. 2002; Canaan et al. 2004; Cole et al. 1998; Daniel et al. 2004; Deb et al. 2006; Mishra et al. 2008; Zhang et al. 2005).
The idea that lipases and phospholipases can be involved in pathogenicity has been suggested by the results of several studies on non-mycobacterial pathogens such as Vibrio cholerae, Pseudomonas cepacia and P. aeruginosa (Jaeger et al. 1994). These enzymes purified from these organisms have been found to affect eukaryotic cells in several ways in vitro. They inhibit the phagocytic functions of alveolar macrophages and modulate the release of inflammatory mediators by various kinds of cells in the immune system. Another possible function of these lipases might consist in degrading host lipids, thus providing energy in the form of fatty acids to the pathogen.
To date, only two lipases present in M. tuberculosis have been purified and characterised, and no soluble phospholipase C has been produced and purified so far. Therefore, further investigations on the genome are now required to search for new genes encoding lipolytic enzymes.
Lipases
Intracellular lipase
Using E. coli as the expression system, Deb et al. cloned 24 genes from M. tuberculosis encoding putative lipase/esterase belonging to the “Lip family” with a view to determine whether they show any hydrolytic activity on long-chain TAG (Deb et al. 2006). Total cell lysates from recombinant E. coli were assayed in order to measure the lipolytic activity and no recombinant protein was purified. Only the Rv3097c gene product, which was denoted LipY, was expressed on a large scale, but the recombinant protein was found insoluble and only present in inclusion bodies. Insoluble recombinant LipY was solubilised in phosphate buffer containing 1% (w/v) sodium lauroyl sarcosine. Results showed that Rv3097c is active on radiolabelled long-chain TAG. Although LipY was the sole enzyme from the “Lip family” to exhibit lipolytic activity, the possibility that the absence of lipolytic activity in other members of this family might be simply due to the misfolding of recombinant proteins in the inclusion bodies cannot be ruled out. No experiments have been carried out so far to check whether these recombinant proteins are expressed in soluble form or not, since only the total fractions have been assayed to measure hydrolytic activity after the induction procedure. Further studies are now required on these proteins to establish whether they are involved in the lipid metabolism.
LipY belongs to the hormone-sensitive lipase (HSL) family (Deb et al. 2006; Mishra et al. 2008) and is also a member of the PE-PGRS protein family. The PGRS domain is a polymorphic GC-rich repetitive sequence found in M. tuberculosis for molecular fingerprinting of clinical isolates and particularly rich in Gly and Ala. The PE domain is an homologous domain found in almost 100 unique genes containing a signature Pro-Glu (PE) amino acid sequence in N-terminal position of the protein (Brennan and Delogu 2002; Delogu et al. 2006). LipY was found to be up-regulated to a higher level than the transcripts of other selected proteins in mycobacteria subjected to TAG-hydrolysing conditions after emerging from a quasi-dormant state in vitro. The ability to hydrolyse stored TAG was found to have dramatically decreased in a LipY-disrupted mutant (Fig. 4). This was the first study in which a long-chain TAG lipase from M. tuberculosis was characterised and functional evidence was obtained experimentally. Using an in vivo expression technology based on green fluorescent protein and kanamycin resistant, Srivastava et al. (2007) showed that LipY was up-regulated in the macrophages with a maximal expression after 24h. Recently, Mishra et al. (2008) investigated the role of the PE domain in LipY by overexpressing an active form of LipY and of LipY lacking the PE domain (LipY(ΔPE)) in Mycobacterium smegmatis and Mycobacterium bovis BCG. Surprisingly, this study showed that in absence of the PE domain, the TAG degradation was higher, indicating that the PE domain could be involved as a modulator of the LipY activity. Fractionation studies, western blot analysis and electron microscopy studies performed by these authors showed that both enzymes (LipY and LipY(ΔPE)) can be found in the cytosol and in the cell wall of M. smegmatis and M. bovis BCG suggesting that the PE domain is not involved in the translocation to the cell wall. Similar experiments have been performed with the LipY M. marinum homolog in which the PE domain was replaced by a PPE domain and similar results have been obtained. Finally, the serological analysis of tuberculosis-infected patients clearly proved that LipY as well as LipY(ΔPE) are similarly recognised by the antibodies present in the sera of these patients. Therefore, PE domain does not induce a specific immunogenic process.
TAG accumulated during 12 days of culture in hypoxic conditions of M. tuberculosis and hydrolysed by wild-type and disrupted mutant (Δ-lipY) cells at 0 h and after 6 h incubation in nutrient-rich (7H9) and PBS media. Lipids were separated on Silica gel TLC using n-hexane/diethyl ether (9:1, v/v) as a solvent and TLC chromatogram was charred after dichromate:sulphuric acid treatment. The intensity of TAG was determined by the AlphaImager 2200 Gel Doc system (Deb et al. 2006)
Results obtained from this work provide clear evidence that Lip Y can be considered as a good candidate for vaccine development. However the role of the PE or PPE domains is still unclear and has to be investigated in details.
Extracellular lipase
No extracellular lipases have been characterised so far in M. tuberculosis. We recently succeeded in cloning, expressing and characterizing the first exported lipase to be identified in M. tuberculosis, which was annotated Rv0183 (Côtes et al. 2007). This enzyme shows 34% to 36% amino acid sequence identity with human and rat monoglyceride lipases, respectively, and homologues were found in M. bovis (Mb0189, 100% sequence identity), Mycobacterium leprae (Ml2603, 79% sequence identity) and M. smegmatis (68% sequence identity; Fig. 5). The gene coding for the Rv0183 protein was expressed in E. coli and the high level of expression obtained in a soluble fraction made it possible to easily purify the recombinant Rv0183 (rRv0183). The pure enzyme was also used to perform biochemical studies to determine the substrate specificity, pH stability, pH and temperature dependence as well as inhibition studies, and to produce polyclonal antibodies (pAbs). The genome annotation suggested that Rv0183 was a lysophospholipase, but it has been demonstrated that rRv0183 does not hydrolyse lysophospholipids. Actually, rRv0183 shows a preference for monoacylglycerols (MAG), but it is also able to weakly hydrolyse DAG and TAG. Studies using pAbs directed against rRv0183 showed that Rv0183 was located in the culture medium and in the M. tuberculosis cell wall fraction (Fig. 6). It seems likely that Rv0183 can be exported from the cell wall of M. tuberculosis to the culture medium. This enzyme may contribute to the growth of the bacilli and be involved in the hydrolysis of host cell lipids during the infection process. Rv0183 may play a physiological role during the first step of the infection process, as well as after the reactivation step, when mycobacteria escape from the granulomas and are disseminated throughout the organism. Although many investigations on the physiological role of Rv0183 still remain to be carried out before we can fully understand how it contributes to the lipid metabolism of M. tuberculosis, it seems clear that Rv0183 does not play the same physiological role in vivo as LipY. Likewise, an Rv0183 homolog (Ml2603) showing 79% identity in its amino acid sequence has been identified in M. leprae, which is known to possess only the few necessary enzymes for its replication in the host cells. This finding suggests that this enzyme plays an important role in the lifecycle of both M. tuberculosis and M. leprae.
Amino-acid sequence alignment of the monoglyceride lipase Rv0183 with three homologous proteins from Mycobacterium: bovis (Mb0189, 100%), leprae (ML2603, 79%) and smegmatis (68%) and two other mammalian proteins: monoglyceride lipases from humans (34%) and rats (36%; Côtes et al. 2007). This figure was generated by the ESPrit programs (available from the ExPASy web site) using the Clustal W alignment (from the EBI web site) and the secondary structural elements obtained from the PredictProtein website (Rost et al. 2004). Conserved residues are shown with a black background. Residues involved in the catalytic triad are indicated by a dark triangle. Two lipase motifs are shown by dark circles. The secondary structural elements (β-strand and α-helix) are indicated at the top of each column
Rv0183 localization in M. tuberculosis cellular compartments (Côtes et al. 2007). Aliquots (15 µg of protein per well) of M. tuberculosis cellular compartments were subjected to immunoblotting analysis using purified polyclonal antibodies directed against rRv0183. Lane 1: rRv0183 purified protein (400 ng); lane 2: culture medium passed through a 0.22 µm filter; lane 3: culture medium; lane 4: cell wall; lane 5: membrane; lane 6: Triton X-114 extracted proteins; lane 7: cytosol. M. tuberculosis cellular samples were kindly provided by the TB Vaccine Testing and Research Materials Centre funded by the NIH and NIAD (Colorado State University)
Phospholipases
The phospholipase family is composed of phospholipases A, C and D, which hydrolyse phospholipids at different positions (Fig. 3). These enzymes have been isolated from a wide variety of gram-positive and gram-negative bacteria such as Bacillus cereus, Clostridium perfringens, Listeria monocytogenes, Pseudomonas aeruginosa and Legionella pneumophila (Schmiel and Miller 1999; Songer 1997). These phospholipases are important virulence factors in bacterial species and they are responsible for a whole range of pathological syndromes from infection and massive tissue destruction, as in the case of the gas gangrene caused by C. perfringens and the skin and lung infection due to P. aeruginosa (Songer 1997; Titball 1998). Although phospholipases have been widely studied in other pathogenic bacteria, little is known to date about the M. tuberculosis phospholipases.
The degradation of host cell (phospho) lipid membranes during macrophage infection and granuloma formation (Fig. 1) suggests the involvement of secreted or excreted phospholipases (Carvalho et al. 1998; Raynaud et al. 2002; Stonehouse et al. 2002).
Parker et al. recently described a phospholipase A activity in M. tuberculosis, which was associated with a putative mycobacterial cutinase (Parker et al. 2007). This study showed, for the first time, that phospholipase A activity can be associated to the enzyme family of cutinase. Cutinases are well characterised as being produced by phytopathogenic fungi during the colonization process; these enzymes may assist in penetration of the cuticle by hydrolysing cutin polymer into monomers (Kolattukudy et al. 1995; Sieber et al. 2000). Moreover, the substrates specific to cutinase also include polyesters, TAG and DAG and some nitrophenyl esters (Carvalho et al. 1999; Degani et al. 2002, 2004, 2006).
The fact that the M. tuberculosis genome possesses seven genes encoding for putative cutinases suggests that cutinases play an important role in mycobacteria. As this particular pathogen has little chance to encounter cutin under natural conditions, these enzymes may hydrolyse alternative substrates such as phospholipids.
Evidence has been presented for the occurrence of PLA activity in M. tuberculosis, and this activity was found to be associated with the cell membrane and cell wall fractions (Parker et al. 2007). These enzymes are probably excreted from the cell wall in M. tuberculosis and they may trigger the release of FA by hydrolysing host cell membrane phospholipids, thus providing a carbon source and contributing to cell growth processes. PLA activity may also contribute to the progression of damaging inflammatory responses by activating the arachidonic acid cascade as previously found to occur in C. perfringens (Fujii and Sakurai 1989; Gustavsson and Alling 1987; Stonehouse et al. 2002).
In addition to PLA activity, PLD and PLC activities have been found to occur in M. tuberculosis. The PLD activity detected in many species of mycobacteria, including pathogenic and non-pathogenic strains, suggests that this enzyme may not be directly involved in virulence but that it may play an important biological role in this genus (Gomez et al. 2001; Johansen et al. 1996). Studies in which phospholipase activity was characterised in pathogenic bacteria showed that PLC are the most active virulence factor (Songer 1997). The toxicity of PLC is linked to their cytolytic and haemolytic activity, which is presumed to be directly due to their ability to hydrolyse phospholipids located in the host cell membrane (Schmiel and Miller 1999; Titball 1993). The genome sequence of M. tuberculosis has been found to contain four closely related genes encoding for PLC (Camus et al. 2002; Cole et al. 1998), namely PLC-A, PLC-B, PLC-C and PLC-D. These genes show 69% amino acid sequence identity with each other. The PLC-A, PLC-B and PLC-C genes are full-length genes occurring in clusters on the chromosome, whereas PLC-D is truncated at the N-terminus and located in a different region. PLC-ABC show 30% and 40% overall amino acid identity with the PLC-H and PLC-N identified in P. aeruginosa, respectively. However, these PLC show low identity, 12% and 15% respectively, with the α-toxin of C. perfringens and PLC of B. cereus that are the best PLC characterised up to now. Upon constructing four single mutants of M. tuberculosis, in each of which one of the PLC genes was inactivated, a triple PLC-ABC mutant and a quadruple PLC-ABCD mutant, it was observed that all the mutants showed lower levels of PLC activity than the wild-type, which indicates that the four PLC genes encode a functional PLC in M. tuberculosis. Functional complementation of the PLC-ABC triple mutant with the individual PLC-A, PLC-B and PLC-C genes restored about 20% of the total PLC activity detected in the wild-type in each case, which suggests that these three enzymes contribute equally to the overall PLC activity of M. tuberculosis (Raynaud et al. 2002). The results of this study also showed that the growth of triple mutant and quadruple mutant in lungs of mice were attenuated in comparison with the wild-type strain (1–1.5 log reduction in lung CFU burden). However, the pulmonary infection by the quadruple mutant was attenuated similarly to that observed with the triple mutant in this model. This study suggests that PLC-ABC contribute to the infection process in contrast with the PLC-D. However, there is no experimental evidence that these phospholipases are involved in the persistence of the bacteria since the growth kinetics with these mutants have been performed during the active phase of the infection (≤6 weeks).
In view of the above findings, the fact that M. tuberculosis contains three or four PLC that contribute to the virulence of this bacterium in mice raises questions about the functional redundancy of these enzymes. As noted earlier, other organisms such as L. monocytogenes contain two PLC that hydrolyse different substrates (Mengaud et al. 1991). The M. tuberculosis PLC may therefore have different affinities for different phospholipid substrates. An alternative hypothesis is that the PLC genes may be regulated differently and that they may act at different stages in the host-infection process. It was recently reported that PLC-B and PLC-C were strongly induced in M. tuberculosis grown in vitro under iron-replete conditions stimulating macrophage infection (Bacon et al. 2007). PLC expressed in host cells might serve several functions relating to virulence: phospholipid hydrolysis by these enzymes might release DAG (Fig. 3). This compounds may be hydrolysed by lipase, resulting in the release of fatty acids, which may serve either as carbon source or cell wall lipid precursors. Biochemical studies have suggested that in chronically infected lung tissues, fatty acids might constitute a major source of carbon and energy for M. tuberculosis (Wheeler et al. 1990).
The exact biochemical mechanism whereby these enzymes hydrolyse host cell membrane phospholipids still remains to be elucidated. For this purpose, large amounts of active PLC are required, but unfortunately no soluble active protein has been obtained when E. coli was used as the expression system with the expression vector pET-23a (Leao et al. 1995) or other vectors containing either Thioredoxin (TRX), Glutathione-S-Transferase (GST), Maltose Binding Protein (MBP) or NuSA as fusion helpers (unpublished data). Obtaining soluble active recombinant PLC on a large scale is therefore still a challenge. Actually, the fact that active PLC can be obtained using M. smegmatis as the expression system suggests that this microorganism is a suitable bacterium for producing recombinant PLC (Raynaud et al. 2002). The amount of recombinant protein produced in this way is still very low and the expression plasmids need to be improved to increase the yield of recombinant PLC production.
Further biochemical characterisation particularly on the specificity of these PLC could help to understand how M. tuberculosis escapes from the phagolyzosome and the pathogenic processes for which it is responsible. In the long term, these studies may contribute to the rational design of better candidate vaccines or enzyme inhibitors with potential therapeutic effects on M. tuberculosis.
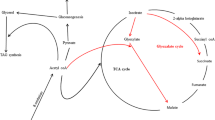
Conclusion
Although lipids play an important role in the life cycle of M. tuberculosis, the role of lipolytic enzymes has been neglected for many years, probably because of the low levels at which these enzymes are expressed. To date, no evidence in vivo has shown that this family of enzymes is clearly involved in the virulence and pathogenicity processes. However, it now seems quite likely that these enzymes may be involved in many different infection steps. Further research is now required on the role of lipolytic enzymes in M. tuberculosis. The genome sequence of M. tuberculosis provides a unique opportunity for studying all putative ORFs by producing recombinant protein. This approach is of particular interest when native proteins are expressed in low levels by the bacteria, as it is often the case. Finally, it can be concluded that these different lipolytic enzymes can interact with each other and several hypotheses can be put forward, as follows: (a) Rv0183 may be able to hydrolyse extracellular TAG, DAG and MAG with different rates, leading to the uptake of free fatty acids (b) Rv0183 may hydrolyse DAG released by PLC and then the resulting MAG, (c) Rv0183 may mainly hydrolyse the MAG resulting from the action of another lipase located in the cell wall or exported. In this hypothesis, LipY could be this lipolytic enzyme. The latter scheme is known to occur in human and mouse adipocytes, where the homologous monoglyceride lipase catalyses the hydrolysis of monoacylglycerols after the action of adipose triglyceride lipase (ATGL) and hormono-sensitive lipase (HSL) on the TAG and DAG, respectively (Haemmerle et al. 2006; Zimmermann et al. 2004).
The study of lipolytic enzymes could open a new approach to drug targeting, where the primary targets are no longer crucial enzymes involved in lipid biosynthesis but rather enzymes involved in lipid degradation, such as lipases and phospholipases C and A. It can be considered that new drugs targeting these enzymes could be used to alter the metabolism of the mycobacteria in order to allow a better efficiency of the drugs actually available to treat the disease. However, we have to keep in mind that these enzymes belong to a large family of protein present also in the host cells and it could be particularly difficult to discover specific inhibitors of mycobacterial lipolytic enzymes. Therefore, many investigations are still required on the physiological role of these lipolytic enzymes to understand how they contribute to the lipid metabolism of M. tuberculosis and how they can interact with each other.
References
Alvarez HM, Steinbüchel A (2002) Triacylglycerols in prokaryotic microorganisms. Appl Microbiol Biotechnol 60:367–376
Bacon J, Dover LG, Hatch KA, Zhang Y, Gomes JM, Kendall S, Wernisch L, Stoker NG, Butcher PD, Besra GS, Marsh PD (2007) Lipid composition and transcriptional response of Mycobacterium tuberculosis grown under iron-limitation in continuous culture: identification of a novel wax ester. Microbiology 153:1435–1444
Brennan MJ, Delogu G (2002) The PE multigene family: a ‘molecular mantra’ for mycobacteria. Trends Microbiol 10:246–249
Camus J, Pryor M, Medigue C, Cole S (2002) Re-annotation of the genome sequence of Mycobacterium tuberculosis H37Rv. Microbiology 148:2967–2973
Canaan S, Maurin D, Chahinian H, Pouilly B, Durousseau C, Frassinetti F, Scappuccini-Calvo L, Cambillau C, Bourne Y (2004) Expression and characterization of the protein Rv1399c from Mycobacterium tuberculosis. A novel carboxyl esterase structurally related to the HSL family. Eur J Biochem 271:3953–3961
Carvalho CM, Aires-Barros MR, Cabral JM (1998) Cutinase structure, function and biocatalytic applications. Electron J Biotechnol 1:160–173
Carvalho CM, Aires-Barros MR, Cabral JM (1999) Cutinase: from molecular level to bioprocess development. Biotechnol Bioeng 66:17–34
CDC (2006) Emergence of Mycobacterium tuberculosis with extensive resistance to second line drug world wide, 2000–2004. Morb Mortal Wkly Rep 55:301–305
Cole ST, Brosch R, Parkhill J, Garnier T, Churcher C, Harris D, Gordon SV, Eiglmeier K, Gas S, Barry CE 3rd, Tekaia F, Badcock K, Basham D, Brown D, Chillingworth T, Connor R, Davies R, Devlin K, Feltwell T, Gentles S, Hamlin N, Holroyd S, Hornsby T, Jagels K, Barrell BG et al (1998) Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature 393:537–544
Côtes K, Dhouib R, Douchet I, Chahinian H, de Caro A, Carriere F, Canaan S (2007) Characterization of an exported monoglyceride lipase from Mycobacterium tuberculosis possibly involved in the metabolism of host cell membrane lipids. Biochem J 408:417–427
Daffe M, Draper P (1998) The envelope layers of mycobacteria with reference to their pathogenicity. Adv Microb Physiol 39:131–203
Daniel J, Deb C, Dubey VS, Sirakova TD, Abomoelak B, Morbidoni HR, Kolattukudy PE (2004) Induction of a novel class of diacylglycerol acyltransferases and triacylglycerol accumulation in Mycobacterium tuberculosis as it goes into a dormancy-like state in culture. J Bacteriol 186:5017–5030
Dawes EA, Senior PJ (1973) The role and regulation of energy reserve polymers in micro-organisms. Adv Microb Physiol 10:135–266
Deb C, Daniel J, Sirakova TD, Abomoelak B, Dubey VS, Kolattukudy PE (2006) A novel lipase belonging to the hormone-sensitive lipase family induced under starvation to utilize stored triacylglycerol in Mycobacterium tuberculosis. J Biol Chem 281:3866–3875
Degani O, Gepstein S, Dosoretz CG (2002) Potential use of cutinase in enzymatic scouring of cotton fiber cuticle. Appl Biochem Biotechnol 102–103:277–289
Degani O, Gepstein S, Dosoretz CG (2004) A new method for measuring scouring efficiency of natural fibers based on the cellulose-binding domain-beta-glucuronidase fused protein. J Biotechnol 107:265–273
Degani O, Salman H, Gepstein S, Dosoretz CG (2006) Synthesis and characterization of a new cutinase substrate, 4-nitrophenyl (16-methyl sulfone ester) hexadecanoate. J Biotechnol 121:346–350
Delogu G, Sanguinetti M, Pusceddu C, Bua A, Brennan MJ, Zanetti S, Fadda G (2006) PE_PGRS proteins are differentially expressed by Mycobacterium tuberculosis in host tissues. Microbes Infect 8:2061–2067
Fujii Y, Sakurai J (1989) Contraction of the rat isolated aorta caused by Clostridium perfringens alpha toxin (phospholipase C): evidence for the involvement of arachidonic acid metabolism. Br J Pharmacol 97:119–124
Garton N, Christensen H, Minnikin D, Adegbola R, Barer M (2002) Intracellular lipophilic inclusions of mycobacteria in vitro and in sputum. Microbiology 148:2951–2958
Goldman RC, Plumley KV, Laughon BE (2007) The evolution of extensively drug resistant tuberculosis (XDR-TB): history, status and issues for global control. Infect Disord Drug Targets 7:73–91
Gomez JE, McKinney JD (2004) M. tuberculosis persistence, latency, and drug tolerance. Tuberculosis (Edinb) 84:29–44
Gomez A, Mve-Obiang A, Vray B, Rudnicka W, Shamputa IC, Portaels F, Meyers WM, Fonteyne PA, Realini L (2001) Detection of phospholipase C in nontuberculous mycobacteria and its possible role in hemolytic activity. J Clin Microbiol 39:1396–1401
Gustavsson L, Alling C (1987) Formation of phosphatidylethanol in rat brain by phospholipase D. Biochem Biophys Res Commun 142:958–963
Haemmerle G, Lass A, Zimmermann R, Gorkiewicz G, Meyer C, Rozman J, Heldmaier G, Maier R, Theussl C, Eder S, Kratky D, Wagner EF, Klingenspor M, Hoefler G, Zechner R (2006) Defective lipolysis and altered energy metabolism in mice lacking adipose triglyceride lipase. Science 312:734–737
Honer zu Bentrup K, Russell DG (2001) Mycobacterial persistence: adaptation to a changing environment. Trends Microbiol. 9:597–605
Jaeger K-E, Ransac S, Dijkstra BW, Colson C, Vanheuvel M, Misset O (1994) Bacterial lipases. FEMS Microbiol Rev 15:29–63
Johansen K, Gill R, Vasil M (1996) Biochemical and molecular analysis of phospholipase C and phospholipase D activity in mycobacteria. Infect Immun 64:3259–3266
Kolattukudy PE, Rogers LM, Li D, Hwang CS, Flaishman MA (1995) Surface signaling in pathogenesis. Proc Natl Acad Sci USA 92:4080–4087
Konig B, Jaeger K, Konig W (1994) Induction of inflammatory mediator release (12-hydroxyeicosatetraenoic acid) from human platelets by Pseudomonas aeruginosa. Int Arch Allergy Immunol 104:33–41
Koval’chuk LP, Donets AT, Razumovskii PN (1973) Lipid biosynthesis by actinomycetes cultured on different media. Mikrobiologiia 42:637–642
Kremer L, de Chastellier C, Dobson G, Gibson KJ, Bifani P, Balor S, Gorvel JP, Locht C, Minnikin DE, Besra GS (2005) Identification and structural characterization of an unusual mycobacterial monomeromycolyl-diacylglycerol. Mol Microbiol 57:1113–1126
Leao SC, Rocha CL, Murillo LA, Parra CA, Patarroyo ME (1995) A species-specific nucleotide sequence of Mycobacterium tuberculosis encodes a protein that exhibits hemolytic activity when expressed in Escherichia coli. Infect Immun 63:4301–4306
Mengaud J, Braun-Breton C, Cossart P (1991) Identification of phosphatidylinositol-specific phospholipase C activity in Listeria monocytogenes: a novel type of virulence factor? Mol Microbiol 5:367–372
Minnikin D (1982) Lipids: complex lipids, their chemistry, biosynthesis and roles. In: Ratledge C, Standford J (eds) The Biology of Mycobacteria. Academic, London, pp 95–184
Minnikin DE, Kremer L, Dover LG, Besra GS (2002) The methyl-branched fortifications of Mycobacterium tuberculosis. Chem Biol 9:545–553
Mishra KC, de Chastellier C, Narayana Y, Bifani P, Brown AK, Besra GS, Katoch VM, Joshi B, Balaji KN, Kremer L (2008) Functional role of the PE domain and immunogenicity of the Mycobacterium tuberculosis triacylglycerol hydrolase LipY. Infect Immun 76:127–140
Neyrolles O, Hernandez-Pando R, Pietri-Rouxel F, Fornes P, Tailleux L, Payan JA, Pivert E, Bordat Y, Aguilar D, Prevost MC, Petit C, Gicquel B (2006) Is adipose tissue a place for Mycobacterium tuberculosis persistence? PLoS ONE 1:e43
Parker SK, Curtin KM, Vasil ML (2007) Purification and characterization of mycobacterial phospholipase a: an activity associated with mycobacterial cutinase. J Bacteriol 189:4153–4160
Raynaud C, Guilhot C, Rauzier J, Bordat Y, Pelicic V, Manganelli R, Smith I, Gicquel B, Jackson M (2002) Phospholipases C are involved in the virulence of Mycobacterium tuberculosis. Mol Microbiol 45:203–217
Rost B, Yachdav G, Liu J (2004) The PredictProtein server. Nucleic Acids Res. 32:321–326
Russell DG (2007) Who puts the tubercle in tuberculosis? Nat Rev Microbiol 5:39–47
Schmiel DH, Miller VL (1999) Bacterial phospholipases and pathogenesis. Microbes Infect 1:1103–1112
Sieber P, Schorderet M, Ryser U, Buchala A, Kolattukudy P, Metraux JP, Nawrath C (2000) Transgenic Arabidopsis plants expressing a fungal cutinase show alterations in the structure and properties of the cuticle and postgenital organ fusions. Plant Cell 12:721–738
Songer JG (1997) Bacterial phospholipases and their role in virulence. Trends Microbiol 5:156–161
Srivastava V, Rouanet C, Srivastava R, Ramalingam B, Locht C, Srivastava BS (2007) Macrophage-specific Mycobacterium tuberculosis genes: identification by green fluorescent protein and kanamycin resistance selection. Microbiology 153:659–666
Stonehouse MJ, Cota-Gomez A, Parker SK, Martin WE, Hankin JA, Murphy RC, Chen W, Lim KB, Hackett M, Vasil AI, Vasil ML (2002) A novel class of microbial phosphocholine-specific phospholipases C. Mol Microbiol 46:661–676
Titball RW (1993) Bacterial phospholipase C. Microbial Reviews 57:347–366
Titball RW (1998) Bacterial phospholipases. Symp Ser Soc Appl Microbiol 27:127S–137S
Tsujita T, Sumiyoshi M, Okuda H (1999) Wax ester-synthesizing activity of lipases. Lipids 34:1159–1166
Wältermann M, Steinbüchel A (2005) Neutral lipid bodies in prokaryotes: recent insights into structure, formation, and relationship to eukaryotic lipid depots. J Bacteriol 187:3607–3619
Wayne LG, Sohaskey CD (2001) Nonreplicating persistence of Mycobacterium tuberculosis. Annu Rev Microbiol 55:139–163
Wheeler PR, Bulmer K, Ratledge C (1990) Enzymes for biosynthesis de novo and elongation of fatty acids in mycobacteria grown in host cells: is Mycobacterium leprae competent in fatty acid biosynthesis? J Gen Microbiol 136:211–217
WHO (2004) Tuberculosis. World Health Organization http://www.who.int/mediacentre/factsheets/fs104/en/print.html
Zhang M, Wang JD, Li ZF, Xie J, Yang YP, Zhong Y, Wang HH (2005) Expression and characterization of the carboxyl esterase Rv3487c from Mycobacterium tuberculosis. Protein Expr Purif 42:59–66
Zimmermann R, Strauss JG, Haemmerle G, Schoiswohl G, Birner-Gruenberger R, Riederer M, Lass A, Neuberger G, Eisenhaber F, Hermetter A, Zechner R (2004) Fat mobilization in adipose tissue is promoted by adipose triglyceride lipase. Science 306:1383–1386
Acknowledgments
We are indebted to Dr. Jessica Blanc for revising the English manuscript. This work was supported partly by the CNRS and partly by the GIP ANR contract 06-JCJC-0067-01.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Côtes, K., Bakala N’Goma, J.C., Dhouib, R. et al. Lipolytic enzymes in Mycobacterium tuberculosis . Appl Microbiol Biotechnol 78, 741–749 (2008). https://doi.org/10.1007/s00253-008-1397-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-008-1397-2