Abstract
The 5-aminolevulinate (ALA) synthase gene (hemA) containing several codons rarely used by Escherichia coli was cloned from the genome of Rhodobacter sphaeroides and optimized in two strains of Escherichia coli: BL21(DE3) and Rosetta(DE3), which is a rare codon optimizer strain. The effects of initial isopropyl-β-d-thiogalactopyranoside (IPTG) concentration, induction time, and temperature on enzyme activity were studied and compared for two strains. The results indicated that the ALA synthase expressed by Rosetta(DE3)/pET-28a(+)-hemA was higher than that by BL21(DE3)/pET-28a(+)-hemA. The initial precursors, glycine and succinate, and initial glucose, which is an inhibitor for both ALA synthase and dehydratase, were observed to be the key factors affecting ALA production. ALA synthase activity was generally higher with Rosetta(DE3) than with BL21(DE3), so was ALA biosynthesis. Based on the optimal culture system using Rosetta(DE3), the yield of ALA achieved 3.8 g/l (29 mM) under the appropriate conditions in fermenter.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
ALA (5-aminolevulinic acid or 5-aminolevulinate) is a kind of photodynamic chemical. A drug mainly consisted of ALA has been approved by the Food and Drug Administration (FDA, USA) in 1999 as a photodynamic medicine for the treatment of skin cancer. ALA is also of potential application for other cancers (De Dominicis et al. 2001), oral errucous hyperplasia (Chen et al. 2004), and so on. In agriculture, ALA can be used as a selective and biodegradable herbicide or insecticide (Sasikala et al. 1994; Sasaki et al. 2002).
Currently, ALA is mainly produced by chemical synthesis and is difficult to satisfy increasing various commercial applications due to the numerous reaction steps required (Sasaki et al. 2002), relatively low yields, and its noxious byproducts.
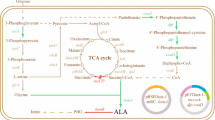
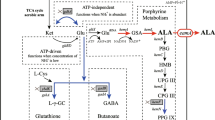
ALA is generally known as an essential intermediate in the biosynthesis of tetrapyrrole, which is found in porphyrins, heme, and vitamin B12 (Vladimir et al. 1997). ALA can be synthesized biologically in cells by two kinds of distinguished metabolic pathways (Vladimir et al. 1997): C4 and C5 pathways. In the C4 pathway, present in mammals, birds, yeast, some protozoa, and purple non-sulfur photosynthetic bacteria such as Rhodobactersphaeroides, the pyridoxal phosphate-dependent enzyme, ALA synthase, catalyzes the condensation of succinyl-CoA and glycine to yield ALA. In the C5 pathway, which occurs in higher plants, algae, and in many bacteria including Escherichia coli and archaeabacteria, ALA synthesis is related to three enzymatic catalyzed reactions. The formation of ALA in cells is considered a rate-limiting step for the biosynthesis of tetrapyrroles, and it is tightly regulated by feedback inhibition (Lascelles 1978). It is reported that the addition of levulinic acid (Sasaki et al. 1987) and d-glucose (Lee et al. 2003) as competitive inhibitors for ALA dehydratase (ALAD) can greatly improve the accumulation of ALA, and d-glucose may also be an inhibitor (Xie et al. 2003) for ALA synthase (ALAS).
Own to the presence of these pathways, ALA can be produced by fermentation with natural selected bacteria and algae (Sasikala and Ramana 1995), mutants of photosynthetic bacteria (Kamiyama et al. 2000), and recombinant organisms. Among them, gene recombination technique offers a new choice to increase the expression of ALA synthase and to enhance the productivity of ALA. As the C4 pathway involves only one gene (hemA) for ALA synthase, it is easier to construct a recombinant cell for ALA production.
E. coli is the most popular host cell for genetic engineering application. The production of ALA using recombinant E. coli has been studied by several groups (Lee et al. 2005; Liu et al. 2005, 2006; Chung et al. 2005). The hemA gene has been cloned mainly from photosynthetic bacterium R. sphaeroides (Suwanto 1995; Edward et al. 1999; Xie et al. 2003). It was found that the expression of ALA synthase and the productivity of ALA by genetic engineering E. coli were still very low. Using factorial design optimization to improve ALA synthase activity, the yield of ALA reached only about 39 mM using recombinant E. coli (Xie et al. 2003).
Each kind of microbes has their own favorable codons. Comparing codons in the ALA gene cloned from R. sphaeroides with those used by E. coli, it is clear that several codons are underrepresented. In particular, Arg codons AGA, AGG, CGG, CGA, Ile codon AUA, Leu codon CUA, Gly codon GGA, and Pro codon CCC are rarely used by E. coli. Excessive rare codon usage in the target gene has been implicated as a cause for low level expression (Sorensen et al. 1989; Zhang et al. 1991) as well as truncation products. The effect seems to be most severe when multiple rare codons occur near the amino terminus (Chen and Inouye 1990). Several laboratories have shown that the yield of protein whose genes contain rare codons can be improved when the cognate transfer RNA (tRNA) is increased within the host (Brinkmann et al. 1989; Rosenberg et al. 1993). A complete compilation of codon usage of the sequences placed in the GenBank database can be found at http://www.kazusa.or.jp/codon/ (Nakamura et al. 2000), and E. coli displays frequencies of about 0.6% of rare codon CCC. As there are five rare codons (CCC) in the hemA gene (408 codons) from R. sphaeroides (about 1.2% frequencies), it may be beneficial to apply a rare codon optimizer strain, such as E. coli Rosetta(DE3), which is a derivative of BL21(DE3) and contains a plasmid of pRARE(CmR) encoding rare codon tRNAs including CCC.
The aim of this work is to increase the ALA productivity via the adaptation of a rare codon optimizer strain and the optimization of cultivation conditions of the new recombinant E. coli.
Materials and methods
Strains
The host strains applied in this work are E. coli BL21(DE3) (Novagen, Germany) [(F-ompT hsdSB(\( {\text{r}}^{ - }_{{\text{B}}} {\text{m}}^{ - }_{{\text{B}}} \)) gal dcm (DE3)] and E. coli Rosetta(DE3) (Novagen, Germany) [F-ompT hsdSB(\( {\text{r}}^{ - }_{{\text{B}}} {\text{m}}^{ - }_{{\text{B}}} \)) gal dcm lacY1(DE3) pRARE(CmR)].
Construction of pET-28a (+)-hemA
The genome of R. sphaeroides 2.4.1 (wild type) served as the DNA template. Primers were designed based on the published R. sphaeroideshemA gene sequence (GenBank, GeneID, 3720398), and an EcoRI (GAATTC) restriction site at the beginning of the amplified fragment and a HindIII (AAGCTT) restriction site at the end of the amplified fragment were involved. The forward and reverse primers were 5′-GGATCCGAATTCATGG ACTACAATCTGGCACTC-3′ and 5′-AAGCTTTCAGGCAACGACCTCGGCGCGA-3′ (ATG start, EcoRI sites and HindIII sites are underlined), respectively. The hemA gene was amplified using polymerase chain reaction (PCR), where Pfu DNA polymerase was used instead of Taq DNA polymerase. The resulting 1.3-kb PCR product was gel-isolated (Qiagene GmbH, Germany) and then restricted with EcoRI and HindIII and ligated into a pET-28a(+) (Novagen, Germany) expression vector, which had been restricted with the same two enzymes.
Growth conditions
All fermentation experiments were conducted in 250-ml shake flasks containing 50 ml medium. The Luria–Bertani (LB) medium was used containing (per liter): 5 g yeast extract, 10 g tryptone, and 10 g NaCl. Also in the medium, when necessary, 30 mg/l kanamycin for E. coli containing plasmid pET-28a(+) and 34 mg/l chloramphenicol for E. coli Rosetta(DE3) containing plasmid pRARE were added, respectively, to inhibit the growth of plasmid-free cells.
Seed cultures were incubated for 8 h at 200 rpm at 37°C, and then 1 ml seed culture was transferred into shake flasks containing 50 ml LB medium. The effects of carbon sources as well as IPTG addition on the ALA yield were evaluated.
The optimal conditions were verified and amplified in a 5 l fermenter (KoBio.Tech, Korea). Three liters of sterilized medium was added, and 100-ml seed culture was inoculated to provide an initial optical density at 600 nm (OD600) of approximately 0.1. The fermenter was operated at 400 rpm. The air flow rate was regulated at 3 l/min, and pH value was controlled at 6.5 with 10% H2SO4 (v/v).
Analysis
Cell density in the medium was represented by measuring OD600 (Ultrospec 3300 pro, Amersham Biosciences, Sweden). Succinate was analyzed by high-performance liquid chromatography (HPLC; Yang et al. 1998) using an Agilent SB-Aq column with 0.05 M phosphate buffer (pH 2.6) as eluent and detection at 210 nm using a UV detector (Agilent 2410, G1314A VWD). Reversed phase liquid chromatography (RP-HPLC) for analyzing glycine concentration was based on a dansyl chloride pre-column derivatization (Fan et al. 2006). ALA was analyzed using modified Ehrlich’s reagent (Burnham 1970). Specifically, 2 ml of sample or standard was mixed with 1 ml 1-M sodium acetate (pH 4.6) in a cuvette, and 0.5 ml acetylacetone (2,4-pentanedione) was added to each cuvette. Then, the mixtures were heated at 100°C for 15 min. After cooling for 15 min, 2 ml of the reaction mixture and 2 ml freshly prepared modified Ehrlich’s reagent (Burnham 1970) were mixed together. After 30 min, the absorbance at 554 nm was measured.
Enzyme assay
Ten milliliters of fermentation broth was centrifuged (10,000×g for 20 min at 4°C). The cells were resuspended in 3 ml 100-mM Tris-buffer (pH 7) and then disrupted by an ultrasonic instrument (Ningbo Scientz Biotechnology, China) for 6 min. After recentrifugation (10,000×g for 10 min at 4°C), ALA synthase activity in the supernatant was measured (Burnham 1970). One unit of ALA synthase activity was defined as the amount of enzyme needed to produce 1 nmol ALA in 1 min. Protein was measured using a Pierce BCA protein assay kit (Pierce, USA).
Results
Effects of induction time, culture temperature, and IPTG concentration on ALA synthase expressed by two strains
The effect of IPTG induction time on the ALA synthase activity for both E. coli BL21(DE3)/pET-28a(+)-hemA and E. coli Rosetta(DE3)/pET-28a(+)-hemA was evaluated, and the results were shown in Fig. 1a. In all experiments, 1 mM IPTG was added to induce the expression of ALA synthase, and the cultivation temperature was lowered from 37 to 28°C after induction. ALA synthase activity of the cells was measured after cultivation for 8 h. The results indicated that the highest ALA synthase activity appeared when IPTG was added at the early exponential phase when the cells were cultivated for 2 h and OD600 was approximate 1. There was no significant difference in ALA synthase activity when the induction was taken at the beginning of the fermentation or early exponential phase (after 2 h cultivation) for both strains, but the cells grew much better when IPTG was added at the early exponential phase than when IPTG was added at the beginning of fermentation (Fig. 1b).
From Fig. 1, it is obvious that the ALA synthase activity expressed in E. coli Rosetta(DE3)/pET-28a(+)-hemA is much higher than that in E. coli BL21(DE3)/pET-28a(+)-hemA. The results showed that the application of rare codon optimizer as host cell was favorable for the expression of genes with rare codons.
The effect of IPTG concentration on the ALA synthase activity was studied, and the results are summarized in Fig. 2. IPTG was added after batch culture were carried out for 2 h and cultivated at 28°C. It was obvious that without IPTG induction, ALA synthase activities for both E. coli BL21(DE3)/pET-28a(+)-hemA and E. coli Rosetta(DE3)/pET-28a(+)-hemA were less than 2 U/mg protein (Fig. 2). At low IPTG concentration, ALA synthase activities of E. coli Rosetta(DE3)/pET-28a(+)-hemA were higher than that of E. coli BL21(DE3)/pET-28a(+)-hemA, whereas at high IPTG concentration, the trend was reversed. The possible reason is that more insoluble protein was formed in E. coli Rosetta(DE3)/pET-28a(+)-hemA than in E. coli BL21(DE3)/pET-28a(+)-hemA at high IPTG concentration due to the higher and more stable expression rate of ALA synthase in E. coli Rosetta(DE3)/pET-28a(+)-hemA. The highest ALA synthase activities were 34.3 and 30.1 U/mg protein for E. coli BL21(DE3)/pET-28a(+)-hemA and E. coli Rosetta(DE3)/pET-28a(+)-hemA, respectively.
The effects of culture temperature after IPTG induction on the ALA synthase activity are shown in Fig. 3. The results indicated that lower cultivation temperature after IPTG induction, such as 28°C, was favorable for higher expression of ALA synthase.
Effects of glucose and precursors on the ALA yield
According to the preliminary experiments, in the absence of precursors (glycine and succinic acid) in the medium, ALA production with two strains was too low (less than 0.15 g/l) to make a significant difference in ALA yield when changing the cultivation conditions.
When the precursors, 1 g/l glycine and 3 g/l succinic acid, respectively, were added in the medium, the effects of initial glucose concentration on the ALA yield was examined, and the results are shown in Fig. 4. It seemed that the ALA production with rare codon optimizer E. coli Rosetta(DE3)/pET28a(+)-hemA was higher than that with E. coli BL21(DE3) /pET28a(+)-hemA, and an optimal initial concentration of glucose existed for ALA production for both strains. When initial glucose concentration was 2 g/l, the highest ALA concentration in the final fermentation broth reached 1 and 0.83 g/l for Rosetta(DE3)/pET28a(+)-hemA and E. coli BL21(DE3)/pET28a(+)-hemA, respectively.
The effects of precursors, glycine and succinic acid, and their ratio on the ALA production are shown in Fig. 5. In all the cases, the initial glucose concentration in the medium was kept at 2 g/l.
The effect of initial precursors concentration on ALA production in E. coli BL21(DE3)/pET28a(+)-hemA and Rosetta(DE3)/pET28a(+)-hemA (control) without glycine and succinic acid. A represents 1 g/l glycine and 3 g/l succinic acid. B represents 1 g/l glycine and 6 g/l succinic acid. C represents 1 g/l glycine and 10 g/l succinic acid. D represents 2 g/l glycine and 3 g/l succinic acid. E represents 2 g/l glycine and 6 g/l succinic acid. F represents 2 g/l glycine and 10 g/l succinic acid. G represents 1 g/l glycine and no succinic acid. H represents 3 g/l succinic acid and no glycine
The results indicated that, if only one precursor was added (G and H in Fig. 5), the ALA yield was very low, only about 0.3–0.4 g/l. The addition of both precursors was necessary for high ALA yield. It was obvious that the concentration of glycine in the medium was more important than that of succinic acid on ALA yield. When glycine concentration was increased from 1 g/l (A, B, and C in Fig. 5) to 2 g/l (D, E, and F in Fig. 5), the ALA yield increased 20 to 30% correspondingly. However, when succinic acid concentration was increased from 3 to 10 g/l, the ALA yield was almost unchanged.
Also from Fig. 5, it is clear that ALA yields with strain E. coli Rosetta(DE3)/pET28a(+)-hemA were always much higher than that with strain E. coli BL21(DE3)/pET28a(+)-hemA.
ALA production with E. coli Rosetta(DE3)/pET28a(+)-hemA in a fermenter
From the above results of shake flask experiments, rare codon optimizer E. coli Rosetta(DE3)/pET28a(+)-hemA was a better ALA producer, and the optimal conditions for the ALA production were 10 g/l succinic acid, 2 g/l glycine, and 2 g/l glucose added initially. Two hours after incubation, IPTG was added and culture temperature was adjusted to 28°C. It was desirable to perform the fermentation in a fermenter. In this work, a 5-l fermenter was applied, and batch fermentation was carried out using E. coli Rosetta(DE3)/pET28a(+)-hemA. The pH value of fermentation broth was kept at 6.5 by adding 10% H2SO4.
An additional dose of 1 g/l glycine was added at 9 h and 15 h, respectively, to resupply the glycine that had been consumed, and 4 g/l glucose was added at 9, 12, and 15 h, respectively, as an inhibitor for ALA dehydratase. The fermentation was terminated at 22 h. The concentration of ALA reached 3.8 g/l at 18 h (Fig. 6) when glycine was exhausted.
Fermentation of E. coli Rosetta(DE3)/pET28a(+)-hemA. An additional 1 g/l glycine was added at 9 and 15 h, respectively, and 4.00 g/l glucose was added at 9, 12, and 15 h, respectively, as an inhibitor of ALA dehydratase. Filled traingle, glycine; inverted filled triangle, succinic acid; filled square, ALA; open square, OD; open circle, ALA synthase activity. The pH was controlled at 6.5 with 10% H2SO4. The data express the averages of triplicates
Discussion
In this work, a rare codon optimizer, E. coli Rosetta(DE3), was applied to express the hemA gene from R. sphaeroides. The results showed that the ALA synthase activity as well as ALA productivity with E. coli Rosetta(DE3)/pET28a(+)-hemA was higher than that with ordinary E. coli BL21(DE3)/pET28a(+)-hemA, which explained that the codon favorability of the host strain was an important factor for enzyme activity expressed by a foreign gene, and the higher ALA synthase activity was able to promote the ALA synthesis.
The existence of rare codons in foreign gene can cause insufficient tRNA pools, which will lead to translational stalling, premature translation termination, translation frameshifting, and amino acid mis-incorporation. Consequently, one of the effective ways to get high expression levels of proteins was to improve the abundance of tRNAs for rare codons in the host cells. Several groups have shown that the yield of protein whose genes contain rare codons can be improved when the cognate tRNA is increased within the host (Brinkmann et al. 1989; Rosenberg et al. 1993). Our results also showed that the highest ALA synthase activity in E. coli Rosetta(DE3) was 10% higher than in E. coli BL21(DE3).
The adaption of rare codon optimizer strain, E. coli Rosetta (DE3), improved the expression of ALA synthase; however, it was not enough for higher ALA production because the host cells were unable to synthesize enough precursors (both glycine and succinic acid) for high ALA productivity. Thus, it is necessary to supply these precursors in the medium. Our results indicated that the addition of both precursors was necessary for high ALA yield, and it was obvious that the concentration of glycine in the medium was more important than that of succinic acid on ALA yield.
Glucose was regarded as an inhibitor for both ALA synthase and ALA dehydratase (Lee et al. 2003; Xie et al. 2003). The results in this study indicated that low initial glucose concentration (2 g/l) was favorable for ALA production due to elevated glucose inhibition for fed-batch fermentation. The increased production of ALA presumably occurred through the inhibition of ALA dehydratase (Lee et al. 2003). Then, the addition of glucose in the late stage of fermentation acted as the inhibitor of ALA dehydratase.
In the preliminary study of batch fermentation in a 5-l fermenter, the final ALA yield reached 3.8 g/l. Further study on the optimization of fed-batch culture system is in progress.
References
Brinkmann U, Mattes RE, Buckel P (1989) High-level expression of recombinant genes in Escherichia coli is dependent on the availability of the dnaY gene product. Gene 85:109–114
Burnham BF (1970) δ-Aminolevulinic acid synthase (from Rhodopseudomonas sphaeroides). Methods Enzymol 17A:195–204
Chen GF, Inouye M (1990) Suppression of the negative effects of arginine codons on gene expression; preferential usage of minor codons within the first 25 codons of the E. coli genes. Nucleic Acids Res 18:1465–1473
Chen HM, Chen CT, Yang H, Kuo MYP, Kuo YS (2004) Successful treatment of oral verrucous hyperplasia with topical 5-aminolevulinic acid-mediated photodynamic therapy. Oral Oncol 40:630–637
Chung SY, Seo KH, Rhee JI (2005) Influence of culture conditions on the production of extra-cellular 5-aminolevulinic acid (ALA) by recombinant E. coli. Process Biochem 40:385–394
De Dominicis C, Liberti M, Perugia G, De Nunzio C, Sciobica F, Zuccala A, Sarkozy A, Iori F (2001) Role of 5-aminolevulinic acid in the diagnosis and treatment of superficial bladder cancer: improvement in diagnosis sensitivity. Urology 57:1059–1062
Edward LB, Luiza K, Jill ZR, Peter MSJ, Martin JW (1999) Characterization of the Rhodobacter sphaeroides 5-aminolaevulinic acid synthase isoenzymes, hemA and hemT, isolated from recombinant Escherichia coli. Eur J Biochem 265:290–299
Fan L, Lin JP, Liu XX, Qin G, Cen PL (2006) Determination of 5-aminolevulinic acid in E. coli fermentation broth by reversed phase high performance liquid chromatography with precolumn derivatization. Chin J Anal Chem 7:937–940
Kamiyama H, Hotta Y, Tanaka T, Nishikawa S, Sasaki K (2000) Production of 5-aminolevulinic acid by a mutant strain of a photosynthetic bacteria. Seibutu-Kougaku 78:48–55
Lascelles J (1978) Regulation of pyrrole synthesis. In: Clayton RK, Sistrom WR (eds) Photosynthetic Bacteria. Plenum, New York, p 795
Lee DH, Jun WJ, Kim KM, Shin DH, Cho HY, Hong BS (2003) Inhibition of 5-aminolevulinic acid dehydratase in recombinant Escherichia coli using d-glucose. Enzyme Microb Technol 32:27–34
Lee DH, Jun WJ, Shin DH, Cho HY, Hong BS (2005) Effect of culture conditions on production of 5-aminolevulinic acid by recombinant Escherichia coli. Biosci Biotechnol Biochem 69:470–476
Liu XX, Lin JP, Qin G, Cen PL (2005) Expression a new hemA gene from Agrobacterium radiobacter in Escherichia coli for 5-aminolevulinate production. Chin J Chem Eng 13:522–528
Liu XX, Lin JP, Cen PL (2006) Effect of inducers on the production of 5-aminolevulinic acid by recombinant Escherichia coli. Prep Biochem Biotechnol 36:223–233
Nakamura Y, Gojobori T, Ikemura T (2000) Codon usage tabulated from international DNA sequence databases: Status for the year 2000. Nucleic Acids Res 28:292
Rosenberg AH, Goldman E, Dunn JJ, Studier FW, Zubay G (1993) Effects of consecutive AGG codons on translation in Escherichia coli, demonstrated with a versatile codon test system. J Bacteriol 175:716–722
Sasaki K, Ikeda S, Nishizawa Y, Hayashi M (1987) Production of 5-aminolevulinic acid by photosynthetic bacteria. J Ferment Technol 65:511–515
Sasaki K, Watanabe M, Tanaka T, Tanaka T (2002) Biosynthesis, biotechnological production and applications of 5-aminolevulinic acid. Appl Microbiol Biotechnol 58:23–29
Sasikala CH, Ramana V, Raghuveer RP (1994) 5-Aminolevulinic acid: a potential herbicide/insecticide from micro-organism. Biotechnol Prog 10:451–459
Sasikala CH, Ramana CH (1995) Biotechnological potentials of anoxygenic phototrophic bacteria II: biopolyesters, biopesticide, biofuel, and biofertilizer. Adv Appl Microbiol 41:227–278
Sorensen MA, Kurland CG, Pedersen S (1989) Codon usage determines translation rate in Escherichia coli. J Mol Biol 207:365–377
Suwanto A (1995) Genetically engineered Rhodobacter sphaeroides for the overproduction of aminolevulinic acid. Appl Microbiol Biotechnol 51:794–799
Vladimir YB, Demain AL, Zaitseva NI (1997) The crucial contribution of starved resting cells to the elucidation of the pathway of vitamin B12 biosynthesis. Crit Rev Biotechnol 17:21–37
Xie L, Hall D, Eiteman MA, Altman E (2003) Optimization of recombinant aminolevulinate synthase production in Escherichia coli using factorial design. Appl Microbiol Biotechnol 63:267–273
Yang RY, Li M, Chen CQ (1998) RP-HPLC rapid analysis of organic acids during high cell density culture of recombinant Escherichia coli YK537pSB-TK. Chin J Indust Microbiol 28:29–31
Zhang SP, Zubay G, Goldman E (1991) Low-usage codons in Escherichia coli, yeast, fruit fly and primates. Gene 105:61–72
Acknowledgments
This work was supported by a grant from the National Natural Science Foundation of China (20306026) and by a grant from the Ministry of Science and Technology of China (National Basic Research Program of China, 2007CB707805).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Fu, W., Lin, J. & Cen, P. 5-Aminolevulinate production with recombinant Escherichia coli using a rare codon optimizer host strain. Appl Microbiol Biotechnol 75, 777–782 (2007). https://doi.org/10.1007/s00253-007-0887-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-007-0887-y