Abstract
Sinorhizobium sp. C4 was isolated from a polycyclic aromatic hydrocarbon (PAH)-contaminated site in Hilo, HI, USA. This isolate can utilize phenanthrene as a sole carbon source. Sixteen metabolites of phenanthrene were isolated and identified, and the metabolic map was proposed. Degradation of phenanthrene was initiated by dioxygenation on 1,2- and 3,4-C, where the 3,4-dioxygenation was dominant. Subsequent accumulation of 5,6- and 7,8-benzocoumarins confirmed dioxygenation on multiple positions and extradiol cleavage of corresponding diols. The products were further transformed to 1-hydroxy-2-naphthoic acid and 2-hydroxy-1-naphthoic acid then to naphthalene-1,2-diol. In addition to the typical degradation pathways, intradiol cleavage of phenanthrene-3,4-diol was proposed based on the observation of naphthalene-1,2-dicarboxylic acid. Degradation of naphthalene-1,2-diol proceeded through intradiol cleavage to produce trans-2-carboxycinnamic acid. Phthalic acid, 4,5-dihydroxyphthalic acid, and protocatechuic acid were identified as probable metabolites of trans-2-carboxycinnamic acid, but no trace salicylic acid or its metabolites were found. This is the first detailed study of PAH metabolism by a Sinorhizobium species. The results give a new insight into microbial degradation of PAHs.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Polycyclic aromatic hydrocarbons (PAHs) are ubiquitous environmental contaminants which originated from petroleum industry, fossil fuel combustion, and waste incineration. PAHs pose a serious risk to the natural environment and humans because they have various types of toxicities (Hakura et al. 1998). Various microbial remediation and phytoremediation methods have been studied (Parrish et al. 2004; Pradhan et al. 1998). A number of bacterial species are able to utilize two- to five-ring PAHs as sole sources of carbon and energy (Johnsen et al. 1996; Stringfellow and Aitken 1995; Walter et al. 1991).
Bacteria in the class of Rhizobiaceae are found in contaminated environments where various toxic chemicals are present (Ahmad et al. 1997; Bodour et al. 2003). Among numerous rhizobial microorganisms, several bacterial species in the genus of Agrobacterium, Rhizobium, and Sinorhizobium are able to utilize PAHs, polychlorinated biphenyls (PCBs), or aromatic heterocycles (Aitken et al. 1998; Damaj and Ahmad 1996; Frassinetti et al. 1998). Recent biochemical studies showed that several putative genes, which are usually found in other PAH-degrading bacteria, are also present in Sinorhizobium meliloti (Ahmad et al. 1997; Galibert et al. 2001). Because of their possible symbiosis with plant, rhizobial bacteria have been studied in relation to phytoremediation (Johnson et al. 2004; Suominen et al. 2000).
In comparison with other bacteria (e.g., namely, Pseudomonas and Mycobacterium species), however, researches on metabolic pathways of PAHs or metabolic enzymes are very limited for Sinorhizobium species or their closely related rhizobial bacteria. A recent screening with Sinorhizobium sp. C4, isolated from PAH-contaminated soil, showed that this strain utilized phenanthrene as a sole carbon source (Seo et al., unpublished data). The objective of this study was to isolate and identify metabolites of phenanthrene by this strain. Novel metabolism pathways of phenanthrene by strain C4 are found in this study, in addition to those previously reported. Several new metabolic pathways are confirmed with authentic standards. To our knowledge, this is the first study of PAH metabolism by symbiotic bacteria.
Materials and methods
Chemicals
Phenanthrene (>98 % purity) was purchased from Sigma-Aldrich (Milwaukee, WI). Several metabolites were purchased from Sigma-Aldrich, Fisher Scientific (Morris Plains, NJ), or TCI America (Portland, OR). These are 1-hydroxy-2-naphthaldehyde, 2-hydroxy-1-naphthaldehyde, 1-hydroxy-2-naphthoic acid, 2-hydroxy-1-naphthoic acid, naphthalene-1,2-dicarboxylic acid anhydride, naphthalene-1,2-diol, 2-carboxycinnamic acid, phthalic acid, 2-formylbenzoic acid, salicylic acid, protocatechuic acid, gentisic acid, and catechol. Ethyl acetate and other solvents were the highest grade commercially available. Synthetic methods of metabolites, which are not commercially available, were described elsewhere (Keum et al. 2005). The carboxyl and phenolic metabolite standards were derivatized to corresponding methyl esters or ethers with diazomethane that was prepared from N-methyl-N-nitroso-p-toluenesulfonamide in a diazomethane generator (Aldrich).
Isolation and identification of the PAH-degrading strain
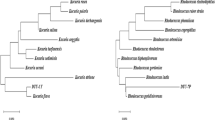
Soil contaminated by PAHs was collected from Hilo, HI, USA (Seo et al., unpublished data). Concentrations of PAHs, including phenanthrene, pyrene, anthracene, and benzo[a]pyrene ranged from 0.6 to 29.9 ppm each. Several metabolites of the abovementioned PAHs were also detected. Those were acenaphthenone, phthalic acid, gentisic acid, protocatechuic acid, o-hydroxynaphthoic acids, diphenic acid, naphthalene-1,2-dicarboxylic acid, 9-fluorenone-1-carboxylic acid, and phenanthrene-4,5-dicarboxylic acid. Strain C4 was isolated from a soil enrichment culture, established in a mineral medium (MM) with solid phenanthrene (50 mg/50 ml of MM) using the soil as inoculums. Strain C4 was identified as Sinorhizobium sp. (AY 943388) through comparison of 16S rRNA gene sequence with those in the National Center for Biotechnology Information basic local alignment search tool (NCBI BLAST) database. The closest neighbor was Sinorhizobium sp. HF6 (AB 195269) with 99% identity.
Utilization of phenanthrene
The bacterial cells were pregrown in MM supplied with phenanthrene to optical density of 0.3 at 580 nm. A 200-μl aliquot of each phenanthrene stock solution (1 mg/ml) was spiked into a sterilized culture tube. The MM (4 ml) and pregrown cell (1 ml) were added after evaporation of solvent. After incubation for 0, 1, 2, 5, and 7 days with shaking at 150 rpm, 28°C, pH values of the culture media were adjusted to 2.3 with 6 N hydrochloric acid and extracted with 4 ml of ethyl acetate. The concentration of phenanthrene was quantified with a gas chromatograph–flame ionization detector (GC-FID). Cultures inoculated with heat-sterilized cells were used as controls. All metabolism experiments were done in triplicate.
Growth of bacterium and extraction of metabolites
Strain C4 was grown in mineral media (Bastiaens et al. 2000) with phenanthrene (500 mg/1.5 l) as a sole source of carbon and energy at 28°C and 150 rpm (C24 Rotary shaker, New Brunswick Scientific, NJ). After the incubation for 4, 7, and 14 days, culture media were filtered through glass wool and centrifuged (6,000×g, 10 min), and the pH of supernatant was adjusted to 2.3 with 6 N hydrochloric acid and extracted three times with ethyl acetate (500 ml). The combined organic layer was extracted three times with aqueous sodium hydroxide (500 ml, 10 mM). The remaining organic phase was dried over anhydrous sodium sulfate and concentrated to 5 ml of ethyl acetate (neutral fraction). Aqueous layer was acidified to pH 2.3 and extracted with ethyl acetate (500 ml, three times, acidic fraction).
Gas chromatography–mass spectrometry (GC-MS) analysis of neutral fraction was done with/without derivatization. For the detection of diol or cis-dihydrodiols, ethyl acetate was removed, and the residue was dissolved in acetone (10 ml) with n-butylboronic acid (50 mg). After refluxing for 30 min, the mixture was concentrated to 1 ml and analyzed by GC-MS. Metabolites in acidic fraction were derivatized with diazomethane.
Analytical methods
Gas chromatography–mass spectrometry analysis was performed with a Varian QP-5000 GC with a Saturn-2000 mass spectrometer (Varian Inc., Palo Alto, CA) equipped with a ZB-1 column (60 m, 0.25 μm film thickness, Phenomenex Inc., Torrance, CA). Helium was the carrier gas at a rate of 2 ml/min. The column temperature was held at 120°C (2 min), programmed to 280°C at a rate of 2°C/min, and held at 280°C for 10 min. Injector and analyzer temperatures were 270 and 280°C, respectively. The mass spectrometer was operated in electron impact (EI) mode (70 eV).
Results
Sinorhizobium sp. C4 can utilize phenanthrene as a sole source of carbon and energy. A short lag period was observed at the initial incubation period (<2 days), but the concentration of phenanthrene rapidly decreased below the detection limit after 7 days of incubation (Fig. 1). Sixteen metabolites, ranging from cis-phenanthrenedihydrodiol to protocatechuic acid, in the culture supernatant were identified by comparison of their GC retention times (RTs) and mass spectra with those of authentic standards (Table 1).
Four metabolites (P1–P3 and P11) were identified in the neutral fraction (Figs. 2 and 4). Among several common dihydrodiols from phenanthrene via bacterial metabolism, two cis-dihydrodiols (P1 and P2) were found at 50.78 and 53.66 min (Fig. 2). The GC RT and mass spectrum of P1 were identical to those of the standard cis-1,2-phenanthrenedihydrodiol n-butylboronate. The mass spectrum of P2 was identical with that of cis-3,4-phenanthrenedihydrodiol described by Krivobok et al. (2003). It is noted that the mass spectra of P1 and P2 are very similar with that of the standard cis-9,10-dihydrodiol n-butylboronate but have different GC RTs (RT of cis-9,10-dihydrodiol n-butylboronate, 51.11 min). Metabolite P3 (RT 58.08 min) gave a mass spectrum consistent with that of standard 9,10-phenanthrenediol n-butylboronate (RT 58.45 min). The concentrations of P2, P7, and P9 were approximately four to five times higher than those of P1, P6, and P8, respectively, which suggests that P3 is 3,4-phenanthrenediol n-butylboronate and undergoes meta-cleavage to yield P7 rather than P6. P11 was identified as 1,2-naphthalenediol n-butylboronate. Its concentration reached a maximum at 3 days followed by a rapid decrease.
Twelve metabolites (P4–10, P12–16) were identified from acidic fractions (Figs. 3 and 4). The GC chromatograms of the acidic fractions showed that the upper metabolites (from P4 and P5 to P8, P9, and P10) of phenanthrene were abundant after 4 and 7 days of incubation; however, the lower metabolites became abundant after 14 days of incubation. Two benzocoumarins (P5 and P4; RTs 39.83 and 41.01 min, respectively) were detected and identified as 7,8-and 5,6-benzocoumarin, respectively. Both metabolites, however, completely disappeared at 14 days. Metabolites P6 and P7 were methyl ethers of 1-hydroxy-2-naphthaldehyde and 2-hydroxy-1-naphthaldehyde, respectively. Both o-hydroxynaphthaldehydes are known intermediates in phenanthrene metabolism (Pinyakong et al. 2000). The concentration of methyl 1-hydroxy-2-naphthoate (P8) reached a maximum after 7 days of incubation and then slowly decreased. P9 was methyl 2-hydroxy-1-naphthoate. Naphthalene-1,2-dicarboxylic acid was detected as its dimethyl ester at 39.53 min (P10). The concentration of P10 was negligible at the initial incubation period (3 days) but rapidly increased during the entire experiment. The mass spectrum and RT of P12 were consistent with trans-2-carboxycinnamic acid dimethyl ester. P12 was not detectable after 4 days of incubation, but its level increased up to 7% of total identifiable metabolites after 7 days of incubation. Metabolites P13–P16 were identified as methyl (or dimethyl) esters of 2-formylbenzoic acid, phthalic acid, 4,5-dihydroxyphthalic acid, and protocatechuic acid, respectively. The concentrations of these metabolites rapidly increased till 14 days of incubation, the amount of which accounted for approximately 40% of the total identifiable metabolites.
Metabolic pathway of phenanthrene by Sinorhizobium sp. P1-1. Carbon atoms in phenanthrene were numbered in Arabic numeral. Bold and thin arrows indicate major and minor pathways, respectively; dotted-line arrows indicate tentative pathways. Metabolites in brackets were proposed but were not detected
Discussion
Metabolic pathways of phenanthrene by strain C4 were constructed based on the metabolites isolated and identified with the authentic standards (Fig. 4). Bacterial degradation of PAHs is usually initiated by mono/dioxygenation (Kim et al. 2005). Naphthalene 1,2-dioxygenase or equivalent enzymes catalyze the incorporation of oxygen molecules on several different carbons of PAHs (Kribovok et al. 2003; Rehmann et al. 1998) to produce cis-dihydrodiols. Although genes or proteins with similar activities were not reported for Sinorhizobium species, a full genome analysis of S. meliloti showed the presence of many putative dioxygenases (Galibert et al. 2001), which may be relevant to metabolism of aromatic compounds. In addition, Damaj and Ahmad (1996) reported that several Rhizobium species have genes relevant to biphenyl dioxygenase and enzymes of subsequent reactions.
In this study, the detection of two dihydrodiols (P1 and P2) and one diol (P3) suggests an involvement of PAH dioxygenase. The presence of the benzocoumarins also supports the enzymatic dioxygenation of phenanthrene because 5,6- and 7,8-benzocoumarin (P4 and P5) are well-known metabolites of phenanthrene, which are produced from 1,2- and 3,4-dihydrodiols, respectively (Pinyakong et al. 2000). In the metabolic study of dibenzothiophene by S. meliloti strain Orange 1, Frassinetti et al. (1998) could detect benzothienopyren-2-one and some associated intermediates, of which the structures are close to PAH metabolites. These results support that at least some Sinorhizobium species have PAH dioxygenase and subsequent metabolic enzymes.
Concentrations of 2-hydroxy-1-naphthaldehyde (P6) and its carboxylic acid analog (P8) were much lower than that of 1-hydroxy-2-naphthoic acid (P9). The concentrations of their precursors, including 1,2-phenanthrenedihydrodiol (P1) and 5,6-benzocoumarin (P4), were also very low. These results suggest that 1,2-dioxygenation and its subsequent reaction play a minor role in phenanthrene degradation by strain C4. These products, however, were not dead-end metabolites because their concentrations decreased rapidly after 7 days.
We proposed that naphthalene-1,2-dicarboxylic acid (P10) be produced from an intradiol cleavage of 3,4- or 1,2-phenanthrenediol and consecutive metabolisms, which is the first example of an intradiol cleavage of PAH diols by Sinorhizobium species as Gram-negative bacteria. PAH diols are usually decomposed by extradiol cleavage to produce o-hydroxyaryl α-oxobut-3-enoic acid (Kim et al. 2005; Pinyakong et al. 2000). Several extradiol dioxygenases, which may catalyze the corresponding reactions, are reported (Saito et al. 2000; Stingley et al. 2004). Although various simple catechols can be metabolized through either intradiol or extradiol cleavage by the action of catechol-1,2-dioxygenase or catechol-2,3-dioxygenase (Maltseva et al. 1994), the intradiol cleavage of high-molecular-weight PAH diols are usually limited to K-region diols (Rehmann et al. 1998; Vila et al. 2001). Some Gram-positive bacteria produce intradiol cleavage products from non-K region diol of anthracene and phenanthrene (Dean-Ross et al. 2001; Kim et al. 2005). However, intradiol cleavage has not been reported in Gram-negative bacteria. 1-Carboxyvinyl-2-naphthoic acid and 2-carboxyvinyl-1-naphthoic acid (intermediates I and II, respectively, Fig. 4) are proposed as precursors of naphthalene-1,2-dicarboxylic acid (P10). Although these metabolites were not detected in the cultures in this study, Dean-Ross et al. (2001) reported the conversion of 2-carboxyvinyl-3-naphthoic acid to naphthalene-2,3-dicarboxylic acid. van Herwijnen et al. (2003) described a mass spectrum of an unknown metabolite of phenanthrene, of which the mass spectrum was well coincided with that of standard 2-carboxyvinyl-1-naphthoic acid synthesized in this study.
Two metabolic pathways are commonly reported for the degradation of 1-hydroxy-2-naphthoic acid (P9) catalyzed by different enzymes (Fig. 5). These are (a) dioxygenation and subsequent ortho-ring cleavage to produce 2-carboxybenzalpyruvate (pathway a in Fig. 5, Adachi et al. 1999) and (b) decarboxylation and hydroxylation to produce naphthalene-1,2-diol (P11) (pathway b, Balashova et al. 2001) that is further degraded through extradiol cleavage (pathway b2; Eaton and Chapman 1992). In this study, naphthalene-1,2-diol (P11) was found without any trace of 2-carboxybenzalpyruvate and 2-hydoxyl benzalpyruvate. This result suggests that strain C4 degrades 1-hydroxy-2-naphthoic acid solely by pathways b and b1 (Fig. 5). 2-Carboxycinnamic acid (P12) was reported in the biodegradation of naphthalene by Bacillus thermoleovorans (Annweiler et al. 2000) and phenanthrene by Mycobacterium sp. strain AP1 (Vila et al. 2001), but its metabolic origin and degradation pathway were not proposed. Detection of 2-carboxycinnamic acid (P12) in this study has clarified the transformation from naphthalene-1,2-diol (P11) to P12.
Rhizobial bacteria can utilize various aromatic acids (Vela et al. 2002). Salicylate and phthalate are among the preferred growth substrates. A recent genetic study showed that some Sinorhizobium species have many genes encoding simple aromatic acid assimilation systems (Galibert et al. 2001). 2-Formylbenzoic acid (P13) was rapidly transformed to phthalate (P14) by strain C4. Accumulation of 4,5-dihydroxyphthalate (P15) and protocatechuic acid (P16) indicates degradation of phthalate (P14) through a common phthalate degradation pathway (Keyser et al. 1976). However, no metabolites associated with salicylate metabolism (e.g., hydroxybenzalpyruvate, salicyladehyde, salicylic acid, and gentisic acid) were detected in the culture media. These results showed that this strain metabolized naphthalene-1,2-diol (P11) to protocatechuic acid through a single unbranched pathway (Fig. 4).
In summary, metabolism of phenanthrene by Sinorhizobium sp. C4 was initiated by incorporation of an oxygen molecule on 1,2- or 3,4-C positions. Phenanthrenediols were transformed to o-hydroxynaphthoates or naphthalene-1,2-dicarboxylic acid by extradiol or intradiol cleavages. Subsequent metabolisms of these acids produced naphthalene-1,2-diol, which was further degraded to protocatechuic acid. The detailed metabolite profiles indicate diversified PAH metabolic enzymes in this strain.
References
Adachi K, Iwabuchi T, Sano H, Harayama S (1999) Structure of the ring cleavage product of 1-hydroxy-2-naphthoate, an intermediate of the phenanthrene-degradative pathway of Nocardioides sp. strain KP7. J Bacteriol 181:757–763
Ahmad D, Mehmannavaz R, Damaj M (1997) Isolation and characterization of symbiotic N2-fixing Rhizobium meliloti from soils contaminated with aromatic and chloroaromatic hydrocarbons: PAHs and PCBs. Int Biodeterior Biodegrad 39:33–43
Aitken MD, Stringfellow WT, Nagel RD, Kazunga C, Chen SH (1998) Characteristics of phenanthrene-degrading bacteria isolated from soils contaminated with polycyclic aromatic hydrocarbons. Can J Microbiol 44:743–752
Annweiler E, Richnow HH, Antranikian G, Hebenbrock S, Grams C, Franke S, Franke W, Michaelis W (2000) Naphthalene degradation and incorporation of naphthalene-derived carbon into biomass by the thermophile Bacillus thermoleovorans. Appl Environ Microbiol 66:518–523
Balashova NV, Stolz A, Knackmuss HJ, Kosheleva IA, Naumov AV, Boronin AM (2001) Purification and characterization of a salicylate hydroxylase involved in 1-hydroxy-2-naphthoic acid hydroxylation from the naphthalene and phenanthrene-degrading bacterial strain Pseudomonas putida BS202-P1. Biodegradation 12:179–188
Bastiaens L, Springael D, Wattiau P, Harms H, deWachter R, Verachtert H, Diels L (2000) Isolation of adherent polycyclic aromatic hydrocarbon (PAH)-degrading bacteria using PAH-sorbing carriers. Appl Environ Microbiol 66:1834–1843
Bodour AA, Wang JM, Brusseau ML, Maier RM (2003) Temporal change in culturable phenanthrene degraders in response to long-term exposure to phenanthrene in a soil column system. Environ Microbiol 5:888–895
Damaj M, Ahmad D (1996) Biodegradation of polychlorinated biphenyls by rhizobia: a novel finding. Biochem Biophys Res Commun 218:908–915
Dean-Ross D, Moody JD, Freeman JP, Doerge DR, Cerniglia CE (2001) Metabolism of anthracene by a Rhodococcus species. FEMS Microbiol Lett 204:205–211
Eaton RW, Chapman PJ (1992) Bacterial metabolism of naphthalene: construction and use of recombinant bacteria to study ring cleavage of 1,2-dihydroxynaphthalene and subsequent reaction. J Bacteriol 174:7542–7554
Frassinetti S, Setti L, Corti A, Farrinelli P, Montevecchi P, Vallini G (1998) Biodegradation of dibenzothiophene by a nodulating isolate of Rhizobium meliloti. Can J Microbiol 44:289–297
Galibert F, Finan TM, Long SR, Puhler A, Abola P, Ampe F, Barloy-Hubler F, Barnett MJ, Becker A, Boistard P, Bothe G, Boutry M, Bowser L, Buhrmester J, Cadieu E, Capela D, Chain P, Cowie A, Davis RW, Dreano S, Federspiel NA, Fisher RF, Gloux S, Godrie T, Goffeau A, Golding B, Gouzy J, Gurjal M, Hernandez-Lucas I, Hong A, Huizar L, Hyman RW, Jones T, Kahn D, Kahn ML, Kalman S, Keating DH, Kiss E, Komp C, Lelaure V, Masuy D, Palm C, Peck MC, Pohl TM, Portetelle D, Purnelle B, Ramsperger U, Surzycki R, Thebault P, Vandenbol M, Vorholter FJ, Weidner S, Wells DH, Wong K, Yeh KC, Batut J (2001) The composite genome of the legume symbiont Sinorhizobium meliloti. Science 293:668–672
Hakura A, Tsutsui Y, Sonoda J, Kai J, Imade T, Shimada M, Sugihara Y, Mikami T (1998) Comparison between in vivo mutagenicity and carcinogenicity in multiple organs by benzo[a]pyrene in the lacZ transgenic mouse (Muta Mouse). Mutat Res 398:123–130
Johnsen K, Andersen S, Jacobsen CS (1996) Phenotypic and genotypic characterization of phenanthrene-degrading fluorescent Pseudomonas biovars. Appl Environ Microbiol 62:3818–3825
Johnson DL, Maguire KL, Anderson DR, McGrath SP (2004) Enhanced dissipation of chrysene in planted soil: the impact of a rhizobial inoculum. Soil Biol Biochem 36:33–38
Keyser P, Pujar BG, Eaton RW, Ribbons DW (1976) Biodegradation of the phthalates and their esters by bacteria. Environ Health Perspect 18:159–166
Keum YS, Seo JS, Li QX (2005) Synthesis of bacterial metabolites of polycyclic aromatic hydrocarbons: benzochromenones, o-carboxyvinylnaphthoates, and o-substituted aryl-a-oxobutenoates. Synth Commun 35:2685–2693
Kim YH, Freeman JP, Moody JD, Engesser KH, Cerniglia CE (2005) Effects of pH on the degradation of phenanthrene and pyrene by Mycobacterium vanbaalenii PYR-1. Appl Microbiol Biotechnol 67:275–285
Krivobok S, Kuony S, Meyer C, Louwagie M, Willison JC, Jouanneau Y (2003) Identification of pyrene-induced proteins in Mycobacterium sp. strain 6PY1: evidence for two ring-hydroxylating dioxygenases. J Bacteriol 185:3828–3841
Maltseva OV, Solyanikova IP, Golovleva LA (1994) Chlorocatechol 1,2-dioxygenase from Rhodococcus erythrpolis 1CP. kinetic and immunochemical comparison with analogous enzymes from Gram-negative strains. Eur J Biochem 226:1053–1061
Parrish ZD, Banks MK, Schwab AP (2004) Effectiveness of phytoremediation as a secondary treatment for polycyclic aromatic hydrocarbons (PAHs) in composted soil. Int J Phytoremediation 6:119–137
Pinyakong O, Habe H, Supaka N, Pinpanichkarn P, Juntongjin K, Yoshida T, Furihata K, Nojiri H, Yamane H, Omori T (2000) Identification of novel metabolites in the degradation of phenanthrene by Sphingomonas sp. strain P2. FEMS Microbiol Lett 191:115–121
Pradhan SP, Conrad JR, Paterek JR, Srivastava VJ (1998) Potential of phytoremediation for treatment of PAHs in soil at MGP sites. J Soil Contam 7:467–480
Rehmann K, Noll HP, Steinberg CEW, Kettrup AA (1998) Pyrene degradation by Mycobacterium sp. strain KR2. Chemosphere 36:2977–2992
Saito A, Iwabuchi T, Harayama S (2000) A novel phenanthrene dioxygenase from Nocardioides sp strain KP7. Expression in Escherichia coli. J Bacteriol 182:2134–2141
Stingley RL, Khan AA, Cerniglia CE (2004) Molecular characterization of a phenanthrene degradation pathway in Mycobacterium vanbaalenii PYR-1. Biochem Biophys Res Commun 322:133–146
Stringfellow WT, Aitken MD (1995) Competitive metabolism of naphthalene, methylnaphthalenes, and fluorene by phenanthrene-degrading pseudomonads. Appl Environ Microbiol 61:357–362
Suominen L, Jussila MM, Mäkeläinen K, Romantschuk M, Lindstrom K (2000) Evaluation of the Galega-Rhizobium galegae system for the bioremediation of oil-contaminated soil. Environ Pollut 107:239–244
van Herwijnen R, Wattiau P, Bastiaens L, Daal L, Jonker L, Springael D, Govers HAJ, Parsons JR (2003) Elucidation of the metabolic pathway of fluorene and cometabolic pathways of phenanthrene, fluoranthene, anthracene, and dibenzothiophene by Sphingomonas sp. LB126. Res Microbiol 154:199–206
Vela S, Häeggblom MM, Young LY (2002) Biodegradation of aromatic and aliphatic compounds by rhizobial species. Soil Sci 167:802–810
Vila J, López Z, Sabaté J, Minguillón C, Solanas AM, Grifoll M (2001) Identification of a novel metabolite in the degradation of pyrene by Mycobacterium sp. strain AP1: actions of the isolate on two- and three-ring polycyclic aromatic hydrocarbons. Appl Environ Microbiol 67:5497–5505
Walter U, Beyer M, Klein J, Rehm HJ (1991) Degradation of pyrene by Rhodococcus sp. UW1. Appl Microbiol Biotechnol 34:671–676
Acknowledgements
This work was supported in part by US-EPA award no. 989512-01-1 and USDA-TSTAR grants 0034135-9576, 2001-34135-11295, and 2002-34135-12724. We thank Michael Cripps of Hawaii Department of Health for his assistance on soil sample collection.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Keum, YS., Seo, JS., Hu, Y. et al. Degradation pathways of phenanthrene by Sinorhizobium sp. C4. Appl Microbiol Biotechnol 71, 935–941 (2006). https://doi.org/10.1007/s00253-005-0219-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-005-0219-z