Abstract
The Harbor Branch Marine Microbial Database (HBMMD) provides preliminary taxonomic identifications and features of microorganisms maintained in the Harbor Branch Oceanographic Institution Marine Microbial Culture Collection. The microbes are primarily derived from marine invertebrates such as sponges (phylum Porifera) and soft corals (phylum Cnidaria) found in deep water environments [>120 feet (>35 m) seawater]. The microbes isolated from within marine invertebrates represent some unique taxa and phylogenetic signatures. The database provides a user-friendly method to systemically search or sort a desired input. The site allows a powerful search for multiple parameters of any entry. Images of the microbes are contained within the database and can be accessed from the website. The HBMMD homepage is located at http://www.hboi.edu/dbmr/dbmr_hbmmd.html.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Microbes were the first living organisms to colonize the Earth and now actively thrive in almost every habitat, including lakes, oceans, sediments and deep-sea vents. Many microorganisms are found in association with marine invertebrate species and can comprise a significant percentage (up to 60%) of the biomass within marine sponges (Vacelet and Donaday 1977; Wilkinson 1987; Giovannoni and Rappé 2000). Microbes present within many marine sponges appear to represent unique taxa (Lopez et al. 1999; Hentschel et al. 2002; Hentschel et al. 2003; Fieseler et al. 2004; Sandell et al. 2004), which may be adapted to specific sponge microcosms. For over two decades, the Division of Biomedical Marine Research (DBMR) at the Harbor Branch Oceanographic Institution (HBOI) has been conducting autonomous underwater vehicle- and remote-operated vehicle-assisted deep-sea collections in various oceanic basins with a primary emphasis on the discovery of novel compounds from marine invertebrates with potential therapeutic uses (http://www.hboi.edu/dbmr/dbmr_home.html). Using specialized underwater vehicles such as Johnson-Sea-Link submersibles, which can dive to 3,000 feet seawater (fsw; approx. 915 m), DBMR has collected over 30,000 invertebrate samples. Over 17,000 marine eubacterial and fungal isolates have been cultured from the invertebrate samples and now comprise the HBOI Marine Microbial Culture Collection (HBMMCC).
The Harbor Branch Marine Microbial Database (HBMMD) exhibits a cross-section of the HBMMCC and highlights recent molecular taxonomic analyses carried out by DBMR (Lopez et al. 1999; Olson et al. 2000, 2002; Sandell 2003; Sandell et al. 2004). The molecular markers used to characterize microorganisms in the collection are the 16S and 18S small subunit (SSU) ribosomal (r)RNA genes (Pace 1997). SSU rRNA sequences, which are theoretically specific to a particular microbial species, can be used to identify microbes and to characterize the microbial diversity within a marine sponge either with or without cultivation (DeLong and Pace 2001). Furthermore, amplified rDNA restriction analysis of SSU rRNA can be applied as a rapid screen of diversity and describe many prokaryotic organisms to the genus and/or species level (Sandell et al. 2004). SSU rRNA analysis is currently the most widely used molecular method for the identification of microbial taxa, using only the SSU gene sequence, rather than an entire cell or organism, to provide preliminary taxonomic and phylogenic information (Hentschel et al. 2002; Bull et al. 2000). The need to expand the understanding of marine microbial diversity is very important, since the loss of even a single host species could translate into a greater loss of uncharacterized microbial diversity. It is currently estimated that only 1% of the microbes from aquatic environment have been cultured (Colwell et al. 1996). The HBMMD was originally designed for the following purposes:
-
1.
To organize microorganisms associated with deeper water (>120 fsw; >35 m ) marine invertebrates and cultured at Harbor Branch Oceanographic Institution
-
2.
To present specific details and characteristics for each microbe (e.g., rRNA-based taxonomy, geographic source, description, depth, preliminary taxonomy of invertebrate source, GenBank accession number, colony morphology, gram stain, cell morphology) as part of a National Science Foundation biotic survey and inventory (BSI)
-
3.
To provide the public and scientific communities with internet access to the inventoried section of HBMMCC
-
4.
To provide a user-friendly method to search and sort BSI results
Database and homepage
The HBMMD was created using Filemaker Pro 6 software (Filemaker, Santa Clara, Calif.). All microbiological and molecular taxonomic data were imported from a Microsoft Access 97 database (Microsoft Corp., Redmond, Wash.). The HBMMD homepage was created using Microsoft Frontpage (Microsoft Corp.) software.
Weblinks
The introductory HBMMD homepage is accessible via http://www.hboi.edu/dbmr/dbmr_hbmmd.html. The main HBMMD homepage (http://hbmmd.hboi.edu/index.htm) provides the following hypertext links:
-
1.
Database provides the main entry to the HBMMD
-
2.
Uncultured set is a link which includes comparable data from culture-independent studies
-
3.
Microbe Images accesses selected images in the database, alphabetically and numerically ordered
-
4.
Abbreviation Glossary provides a link to an explanation page containing all abbreviations used in the HBMMD
-
5.
A description of the major empirical protocols is provided, as described by Sandell et al. (2004), Sfanos et al. (in press) and Sandell (2003)
-
6.
Gel Examples show representative RFLP bands using (a) RsaI and (b) HaeIII
-
7.
Latest Phylogenetic trees are provided, based on representative isolate 16S rRNA sequences
-
8.
A link is provided to the DBMR homepage
The above weblinks are centralized through the HBMMD homepage and are updated periodically. Upon entering the main database from item 1 above, the first entry is shown in Form View. Users can toggle between Form and Table View. Individual microbial isolate forms can be scrolled one by one via the Next button or by entering the Record number of the isolate at the top of the Form View page.
Data fields and tabs
The primary data fields within HBMMD Form View consist of microbe identification number (ID), bacterium or fungus designations (B or F, respectively), physical description, preliminary taxonomy of source family and GenBank accession number. At present, approximately 357 HBMMCC isolates have been characterized by 16S SSU rDNA sequencing, which allows the taxonomic inference of 2,274 isolates (Sandell 2003). The HBOI ID consists of four characters (such as “K486”), which is a HBOI-specific designation and is not universal. “Physical Description” gives gross colony color and shape. “Family of Source” gives the current taxonomic identification of the respective source invertebrate, although identifications are continually updated (Hooper and Van Soest 2002). Some invertebrate taxonomic identifications only reach class level, although marine sponges comprise the majority of hosts and many have been identified to family level. A “?” in this field indicates ongoing taxonomic assessment. Taxonomic updates will be posted to the HBMMD website.
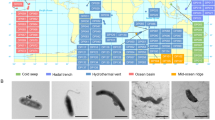
The design of the Form View chosen consists of four tabs, as shown in Fig. 1 with a picture inset showing colony morphology of the microbe at the right-hand side of the description tab. This layout design selection made the viewable area very succinct. The four tabs are described in the following sections.
Description tab
This tab includes fields for colony morphology, geographic source, gram stain, depth and JPEG images showing microbial colony morphology, as shown in Fig. 1. Presently, not all isolates in HBMMD have associated images, but these may be added in future updates. “Colony” is a description of the colony morphology as it appears on marine agar plates or slants. Keys to all abbreviations are listed from the HBMMD homepage link. “Geographic source” indicates the approximate geographic location of the host organism. The “Gram” field shows the results of a standard gram stain reaction. “Depth” indicates the collection depth of the source organism (in feet seawater). A photo in JPEG format shows the colony grown on marine agar.
SSU Profile tab
This tab denotes RFLP data (in base pairs; bp) generated by restriction digestion of 16S SSU rDNA fragments using RsaI, HaeIII, or HhaI. The bands encompass the first 700–800 bp of variable regions V1–V4 in the 5′ portion of the eubacterial 16S SSU rRNA gene (Escherichia coli positions 9–804) obtained by PCR amplification with the primers E. coli 9-forward (5′ GAT TTT GAT CCT GGC TCAG 3′) and Loop27rc (5′ GAC TAC CAG GGT ATC TAA TC 3′; Lopez et al. 1999; Sandell et al. 2004; Sandell 2003; Sfanos et al., in press).
ID tab
The temporary microbe ID (Temp Microbe ID), “broadly common”, “intermediate”, or “unique” designations (B, M, or U, respectively), major microbial group and percent sequence identity are located within this tab. B, M and U refer to an isolate’s relative representation in the HBOI collection and are based on whether an isolate was broadly common (>10 isolates), intermediate (5 isolates), or unique (<5 isolates) within the HBMMCC. “Temp Microbe ID” provides the closest genus and species designation of the microbe inferred from SSU data (SSU rRNA RFLP, or sequences) after a BLAST search (Sandell 2003). Some isolate entries display more than one value for percent sequence identity, indicating multiple sequenced reactions for the isolate. “Major Microbe Group” provides information on a specific isolate’s major eubacterial subdivision, again updated as required.
Other tab
This tab includes miscellaneous data on cell morphology, phylum of source invertebrate, base pairs sequenced and any other relevant comments on the isolate. “Cell Morph” is the morphology of the cell determined by phase contrast microscopy and includes the size of cells in microns. “Phylum of source” details the phylum of the source host invertebrate, such as Porifera (POR), Cnidaria (CNI) or Echinoderma (ECH). The “Comments” section details any other pertinent information.
Searches
To search for an entry in the database click “Search”:
-
1.
Enter the desired input (lowercase or uppercase) with or without any optional search operators in the desired field
-
2.
Click “Start Search,” “Go Back” or “Clear Form”
Possible search operators include the descriptors in the four tabs described above (e.g., colony morphology, B/F, Temp Microbial ID, etc.). Keywords or key numbers can be used in multiple fields to target a specific microbe. For example, when keyword “m” and key number <2,000 are entered in Temp Microbe ID and Depth fields, respectively, at least 24 possible entries appear which have a Temp Microbe ID starting with or containing “m”, while also being isolated under 2,000 fsw (583 m). To clear a searched group, click “Show All” which brings the Table View to the first entry of the database. The searched results are initially presented in the Table View. In this view, all fields of each entry are linkable to their respective Form View via a mouse click. The “Table View” button directs to a Table View of the results in increments of 25. The “Sort” link allows the user to sort the searched results or all the entries into two criteria, ascending or descending, by any field. Users can also sort entries according to the size of the RFLP bands, although only the samples that have been fully sequenced have the most accurate band sizes. Theoretically, a user could compare RFLP patterns of their own unknown isolate’s 16S rRNA fragments, generated with the PCR primers described above (Lopez et al. 1999; Sandell et al. 2004). A user-friendly Help page link based on the FileMaker web companion is provided at the top right (as shown in Fig. 1) to guide new users. Specific commands are laid out to guide the user through searches, sorting, etc. The “Home” link directs the user to the main HBMMD homepage.
Availability
For the public and scientific community, general information on HBMMD and the database itself are available at the HBMMD homepage: http://www.hboi.edu/dbmr/dbmr_hbmmd.html. Updates to the database, such as the addition of new isolates and SSU rRNA sequences from the culture-independent and fungal components of marine sponge microcosms are made periodically. More refined taxonomic analyses of existing and new microbial isolates and invertebrate hosts is expected and will add value to the online HBMMD in the future.
References
Bull AT, Ward AC, Goodfellow M (2000) Search and discovery strategies for biotechnology: the paradigm shift. Micro Mol Biol Rev 64:573–606
Colwell RR, Simidu U, Ohwada K (1996) Microbial diversity in time and space. Plenum, New York
DeLong EF, Pace NR (2001) Environmental diversity of bacteria and archaea. Syst Biol 50:470–478
Fieseler L, Horn M, Wagner M, Hentschel U (2004) Discovery of novel candidate phylum “Poribacteria” in marine sponges. Appl Environ Microbiol 70:3724–3732
Gich FB, Amer E, Figueras JB, Abella CA, Balaguer MD, Poch M (2000) Assessment of microbial community structure changes by amplified ribosomal DNA restriction analysis (ARDRA). Int Microbiol 3:103–106
Giovannoni S, Rappé M (2000) Evolution, diversity, and molecular ecology of marine prokaryotes. Wiley–Liss, New York
Hentschel U, Hopke J, Horn M, Friedrich AB, Wagner M, Hacker J, Moore BS (2002) Molecular evidence for a uniform microbial community in sponges from different oceans. Appl Environ Microbiol 68:4431–4440
Hentschel U, Fieseler L, Wehrl M, Gernert C, Steinert M, Hacker J, Horn M (2003) Microbial diversity of marine sponges. In: Müller WEG (ed) Molecular marine biology of sponges. Springer, Berlin Heidelberg New York, pp 60–88
Hooper JNA, Van Soest RWM (eds) (2002) Systema porifera: a guide to the classification of sponges. Kluwer, Dordrecht
Lopez JV, McCarthy PJ, Janda KE, Willoughby R, Pomponi SA (1999) Molecular techniques reveal wide phyletic diversity of heterotrophic microbes associated with the sponge genus Discodermia (Porifera: Demospongiae). Mem Queensl Mus 44:329–341
Olson JB, Lord CC, McCarthy PJ (2000) Improved recoverability of microbial colonies from marine sponge samples. Microb Ecol 40:139–147
Olson JB, Harmody DK, McCarthy PJ (2002) Alpha-proteobacteria cultivated from marine sponges display branching rod morphology. FEMS Microbiol Lett 211:169–173
Pace NR (1997) A molecular view of microbial diversity and the biosphere. Science 276:734–740
Sandell KA (2003) A molecular systematic survey of cultured microbial associates of deep water marine invertebrates. MSc thesis, Florida Institute of Technology, Melbourne
Sandell KA, Peterson CL, Harmody DK, McCarthy PJ, Pomponi SA, Lopez JV (2004) Molecular systematic survey of sponge-derived marine microbes. Boll Mus Ist Univ Genova 68:579–585
Sfanos KAS, Harmody DK, McCarthy PJ, Dang P, Pomponi SA, Lopez JV. A molecular systematic survey of cultured microbial associates of deep water marine invertebrates. System Appl Microbiol (in press)
Vacelet J, Donaday C (1977) Electron microscopic study of the association between some sponges and bacteria. J Exp Mar Biol Ecol 30:301–314
Wilkinson CR (1987) Significance of microbial symbionts in sponge evolution and ecology. Symbiosis 4:135–146
Acknowledgements
We are grateful to the Link Foundation of HBOI for supporting the work through an HBOI summer internship to A.S.G. We are grateful to Tom Smoyer for photographic images on the website and database and to Bailey Kessing for his assistance with Filemaker Pro. This material is based upon work supported by the National Science Foundation under grant DEB-0103668 to J.V.L. and P.J.M. Any opinions, findings and conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of the National Science Foundation. This manuscript is HBOI contribution 1566.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Gunasekera, A.S., Sfanos, K.S., Harmody, D.K. et al. HBMMD: an enhanced database of the microorganisms associated with deeper water marine invertebrates. Appl Microbiol Biotechnol 66, 373–376 (2005). https://doi.org/10.1007/s00253-004-1763-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-004-1763-7