Abstract
In the cyanobacterium Spirulina platensis, the desaturation process is carried out by three desaturases: the Δ9, Δ12 and Δ6 desaturases, encoded by desC, desA and desD, respectively. The Δ6 desaturase is responsible for the catalysis of linoleic acid, yielding γ-linolenic acid (18:3Δ9,12,6), the end-product of the process. In this study, the desD gene was expressed in Escherichia coli using a pTrcHisA expression system. In order to identify the amino acid residues involved in the enzymatic activity, a sequence comparison was performed using various organisms. The alignment revealed three conserved histidine clusters, a number of conserved residues among all listed organisms and a few conserved residues among cyanobacterial species possibly involved in the desaturation activity. A series of site-directed mutations were generated in the desD gene to evaluate the role of these residues vis-à-vis the enzyme function. This approach revealed that: (1) H313 is involved in the regioselectivity of the enzyme, (2) the three histidine clusters together with H313, H315, D138 and E140 are required for enzymatic activity, most likely as providers of the catalytic Fe center and (3) W294 is also essential for the activity of Δ6 desaturase, possibly by forming part of the substrate-binding pocket.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Desaturases are known for their ability to catalyze the formation of unsaturated fatty acids. These enzymes act by introducing a double bond at a specific defined position in the hydrocarbon chain (Holloway 1983) and are grouped into three types. Cyanobacterial fatty acid desaturases are categorized as being part of the family of acyl-lipid desaturases or plant-type desaturases (Murata and Wada 1995), which are membrane-bound enzymes (Mustady et al. 1996). Acyl-lipid desaturases introduce double bonds into fatty acids that have been esterified to glycerolipids (Schmidt et al. 1993).
The Δ6 desaturase of Spirulina platensis converts linoleic acid (18:2) to γ-linolenic acid (GLA). The isolation of the desD gene encoding the Δ6 desaturase of S. platensis C1 was reported by Murata et al. (1996). A study conducted by our group on the regulation of desaturase gene expression in S. platensis revealed a 30% increase in the level of γ-linolenic acid after reducing the growth temperature from 35°C to 22°C (Deshnium et al. 2000). We also found that this enzyme is present in both the plasma and the thylakoid membranes and that the response to immediate temperature reduction differs in accordance with the membrane in question (Hongsthong et al. 2003).
The presence of conserved histidine clusters has been reported in other membrane-bound desaturases in mammals, fungi, insects, higher plants and cyanobacteria (Bloomfield and Bloch 1960; Avelange-Macherel et al. 1991; Diaz et al. 2002). Site-directed mutagenesis studies carried out with these conserved histidine residues (Shanklin et al. 1994) demonstrate that they play a crucial role, possibly by providing iron-active sites (Sundberg and Martin 1974). As a consequence, membrane-bound desaturases are classified into a superfamily of membrane iron proteins. It is also proposed that the hydrophobic stretch located between the two first histidine clusters and the other, positioned closer to the third histidine cluster, might be involved in substrate recognition (Diaz et al. 2002).
Integral membrane or membrane-bound desaturases, including Spirulina desaturases, are less well understood than soluble desaturases, due to the difficulties in obtaining large quantities of purified membrane-bound proteins (Shanklin and Cahoon 1998). A biochemical study of the Spirulina desaturases is one of the keys to understand how the cells survive under low-temperature conditions, when the cells synthesize more of the end-product of desaturation: GLA.
In many studies, such as those involving the analysis of an enzyme-active site with regioselectivity and substrate specificity, a site-directed mutagenesis approach has been applied to accomplish these goals (Cahoon et al. 1997a, b; Mcguire et al. 2001). In the present study, to explore the role of the residues involved in the catalytic activity of Δ6 desaturase, we conducted site-directed mutagenesis, substituting the histidine residues in the tripartite conserved histidine motif with arginine. The side-chain of histidine is an imidazole group (pK a=6), which can donate a proton to serve as a ligand to chelate metal ions. The arginine side-chain, in contrast, is a guanidinium group (pK a=12.5), which is a strong base. The evolutionarily conserved residues of Δ6 desaturase in all organisms listed in Fig. 1 (D138 and W294, which have the most acidic and the most complex aromatic side-chain, respectively) were changed to N and G (which have the most basic and the most simple side-chain, respectively). In addition, other evolutionarily conserved residues among the cyanobacteria (R123, G136, E140, H313, H315) were substituted with N, H, Q, R and N, found at the corresponding position of Mucor rouxii, Borago officinalis or Homo sapiens.
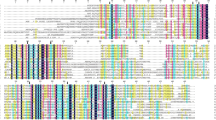
Comparison of the deduced amino acid sequence of Δ6 desaturase from M. rouxii, B. officinalis, Synechocystis, Spirulina platensis and H. sapiens. The signature motifs common to fatty acid desaturase are highlighted in dark gray, while the other residues subjected to mutagenesis are highlighted in light gray
The expression of mutant enzymes was detected by Western blot analysis; and the effects of mutation were analyzed by determination of the end-product. The analyses of the mutant forms of the enzyme permitted classification into three groups: (1) those with an undetectable level of Δ6 desaturase activity and the presence of an unusual fatty acid, (2) those with an undetectable level of Δ6 desaturase activity and (3) those with reduced Δ6 desaturase activity. The resulting data confirmed the hypothesis that the three conserved histidine motifs play a crucial role in enzymatic activity, most probably by building the iron active site. Moreover, the results demonstrated the requirement of a number of conserved amino acid residues, in addition to the three histidine motifs, for the enzymatic function. Interestingly, the single substitution mutation of H313 with arginine caused an alteration in the fatty acid desaturation position of the enzyme.
Materials and methods
Bacterial strains and plasmids
The bacterial strains used in this study were Escherichia coli DH5α chemically competent cells and E. coli XL-mut-S and XL-1-Blue competent cells. The plasmid pTrcHisA (Invitrogen, USA) was used for cloning the desD gene for the construction of an expression vector in E. coli. E. coli strains were grown routinely at 37°C in Luria–Bertani broth (LB) or LB agar (Sambrook et al. 1989), supplemented with appropriate antibiotics when necessary.
Chemicals and enzymes
All the chemicals used for the in vitro assay of enzyme activity were purchased from Sigma (USA), while the enzymes for DNA manipulations were purchased from New England Biolabs (USA), Promega (USA) and Invitrogen (USA) and used according to the manufacturers’ instructions. A Chameleon double-stranded site-directed mutagenesis kit from Stratagene (USA) was used for generating site-directed mutations in the desD gene.
DNA manipulation
Plasmid DNA from E. coli was isolated using the alkaline lysis method (Sambrook et al. 1989). DNA sequences were determined using standard methodologies for double-stranded plasmid DNA on an automated DNA sequencer (Applied Biosystem, USA) at the Bio Service Unit, National Science and Technology Development Agency, Bangkok, Thailand. The analysis of DNA sequences was carried out using the Genetyx package program.
Construction of plasmid for desD gene expression in E. coli
The open reading frame of the Δ6 desaturase of S. platensis C1 was amplified from the desD clone (Murata et al. 1996) by PCR using two oligonucleotide primers, the synthesis of which was based on the desD coding sequence. The two oligonucleotide primers used for PCR amplification were the desD forward primer (5′-TGGGGATCCTTAATGACATCAACAAC-3′) containing the BamHI restriction site (italics) upstream of the start codon, together with the desD reverse primer (5′- GGGAAGCTTAAGGTTAGTGATGATT-3′) containing the HindIII restriction site (italics) downstream of the stop codon. The amplification of DNA fragments was performed using taq DNA polymerase and a corresponding buffer (Invitrogen, USA) in a final volume of 100 μl in a DNA thermal cycler (Bio-Rad). The amplification process began with 3 min at 94°C, followed by 30 cycles of 94°C for 40 s, 50°C for 1 min and 72°C for 2 min, concluding with a final extension at 72°C for 10 min. The PCR-amplified products were digested with BamHI and HindIII, followed by gel purification. After elution, the respective DNA fragments were subcloned into the pTrcHisA expression vector, thus generating plasmid pTrcHisA-desD. This plasmid was transformed into E. coli DH5α host cells using the CaCl2 transformation method (Sambrook et al. 1989) for Δ6 desaturase expression studies.
Construction of site-directed mutants
Single- and double-point mutations encoding single amino acid substitutions were introduced directly into pTrcHisA-desD. Site-directed mutagenesis was carried out using the method employed by Deng and Nickoloff (1992), with the Chameleon double-stranded site-directed mutagenesis kit (Stratagene, USA) used according to the manufacturer’s instructions. Two synthetic oligonucleotide primers were used for the desired mutation. One primer (henceforth referred to as the mutagenic primer) was used to introduce the desired mutation, while the other (the selection primer) was used to mutagenize a unique restriction site in the plasmid for subsequent selection. The nucleotide sequences of the mutagenic primers used for mutagenesis are shown in Table 1. The mutations were H89R, H93R, H124R, H128R, H129R, H305R, H306R (conserved histidine residues of three histidine motifs), R123N, G136H, E140Q, W294G, H313R, H315N, D138N (conserved amino acids outside the histidine motifs). The letters in italics and underlined indicate altered nucleotides and codons, respectively. All the mutations were verified by sequencing. The resulting plasmids were used for E. coli transformation.
Culture conditions
E. coli DH5α was used as the host for expression of the desD gene. Cells containing pTrcHisA-desD were grown in LB in the presence of 100 μg ml−1 of ampicillin at 37°C, and shaken at 200 rpm. In order to assay Δ6 desaturase activity in vitro, E. coli DH5α was grown in M9 medium supplemented with 4 mg ml−1 glucose, 0.1 mM MgSO4, 1 mg ml−1 casamino acids, 10 μM FeCl3, 0.5 μg ml−1 vitamin B1, 100 μg ml−1 ampicillin (Wada et al. 1993) and sodium linoleate (a substrate of Δ6 desaturase) at concentrations of 0, 100, 200, 300, 400, 500, 600, and 800 μM. The culture was incubated at 37°C and shaken at 200 rpm. When the optical density of the culture at 600 nm reached 0.6, isopropyl-β-D-thiogalactopyranoside was added to yield a final concentration of 1 mM. The culture was further incubated at 30°C and shaken at 200 rpm for 3 h, until the optical density of the culture at 600 nm reached 0.7–0.75. The cells were harvested by centrifugation at 9,000 rpm for 10 min. The pellet was then washed twice with 30 ml of re-suspended buffer containing 50 mM MOPS-NaOH, pH 7.5, 10 mM MgCl2 and stored at −20°C until used.
In vitro assay
In a previous study, we showed that Δ6 desaturase cannot function in vivo (data not shown), possibly due to the absence of essential cofactors in the host cells. We thus modified the method employed by Wada et al. (1993) to conduct an in vitro assay in order to detect the enzymatic activity of Spirulina Δ6 desaturase expressed in E. coli.
The cell pellet was re-suspended in buffer containing 50 mM MOPS-NaOH, pH 7.5, 10 mM MgCl2, 300 mM sorbitol, 10 ng of DnaseI and 10,081 units of catalase (Wada et al. 1993). The cells were disrupted by passage through a chilled French pressure cell operated at 3.45 MPa, with this process repeated three times in order for each sample to achieve complete disruption. The cell debris was then removed by centrifugation at 10,000 rpm for 10 min at 4°C and the homogenate immediately subjected to the assay for Δ6 desaturase activity. An aliquot of the homogenate corresponding to 2–4 mg of total protein (quantified using Lowry’s 1951 method) was mixed with the following components in a total volume of 1.2 ml: 40 mM tricine-KOH, pH 8.0, 10 mM MgCl2, 100 μg of ferredoxin (F3013, Sigma), 5 mM β-NADPH (N1630, Sigma), 50,405 units of catalase (C3155, Sigma) and 200 milliunits of Fd–NADP+ oxidoreductase (F0628, Sigma). The reaction was incubated at room temperature (approximately 25°C) for 15 min—the appropriate time for incubation according to time-course studies (data not shown). The lipid was then extracted from the reaction mixture and analyzed using gas chromatography.
The substrate used in this assay was 18:2Δ9,12 in the form of a sodium salt. Since the desaturases of S. platensis are acyl-lipid desaturases, the free fatty acids taken up must be incorporated into glycerolipids, the form of substrate that these desaturases can utilize. In general, E. coli is able to incorporate exogenously added fatty acids into glycerolipids (Esfahani et al. 1969), so the substrate must be added to the culture medium.
Western blot analysis
Detection of the Δ6 desaturase was achieved by Western blot analysis (Gravel and Golaz 1996), using a monoclonal antibody against the 6× histidine tag, as described by the manufacturer (Amersham Biosciences, Sweden). The protein samples, which were separated by 12% SDS-PAGE, were transferred onto a nitrocellulose membrane using a semi-dry electroblotter (Bio-Rad) at a constant voltage of 20 V for 30 min at room temperature. A Western blot detection kit with an alkaline-phosphatase detection system was used (Zymed, USA) as recommended by the manufacturer.
Fatty acid extraction and analysis
After the lipid was extracted from the assay mixture, the extract was transmethylated using 5% HCl in methanol and stirred in the dark at 85°C for 150 min (Wada et al. 1993). The methylated fatty acids were then separated and analyzed using gas chromatography (Shimadzu GC 17-A). The capillary column used in the analysis was a fused silica glass column (30 m, OMEGAWAX 250; Supelco, USA) with a film thickness of 0.25 μm. The injector temperature was 205°C and the split ratio was 1:10.
The positions of the double bonds of the fatty acids were identified by 4,4-dimethyloxazoline (DMOX) derivatization prior to analysis by GC-MS (Fay and Richli 1991).
GC-MS analysis
The qualitative analysis of the fatty acid products, synthesized by heterologously expressed desaturases, was conducted using a TRACE GC/PolarisQ GC-MS (Thermo Finnigan, USA) in order to assure the presence of the products. GC was performed on a fused silica glass column (30 m, OMEGAWAX 250; Supelco, USA) with a film thickness of 0.25 μm. The column temperature, flow rate and split ratio were 205°C, 1.0 ml min−1 and 1:10, respectively. The GC was directly interfaced (with a transfer-line temperature of 300°C) to a PolarisQ mass selective detector (Thermo Finnigan) run in full-scan mode. The electron beam energy was 70 eV. Qualitative analysis was performed using a library search. The libraries used in this experiment were the National Institute of Standards and Technology library and a library constructed using standard fatty acids (Sigma, USA). The analysis of these standards was performed on this instrument under the conditions described.
The double-bond position of the fatty acids synthesized by the heterologously expressed desaturases was determined by GC-MS analysis of the DMOX derivative of the fatty acids in question. The GC-MS conditions were as follows: a fused silica glass column (30 m, OMEGAWAX 250; Supelco) with a film thickness of 0.25 μm, an oven temperature program at 80°C (3 min), increasing at 20°C min−1 to 180°C (15 min) and then at 2°C min−1 to 280°C (15 min), an ion-source temperature of 200°C and ionization at 70 eV.
Results
Detection of Δ6 desaturase expressed in E. coli
The Δ6 desaturase expressed in E. coli using the pTrcHisA system contained a 6× histidine tag at the N-terminus of the polypeptide. Detection was achieved by Western blot analysis, using a monoclonal antibody against the 6× histidine tag. Results showed that the Δ6 desaturase was present in all mutants (Fig. 2) with an approximate molecular mass of 47 kDa.
Detection of the heterologously expressed Δ6 desaturase from pTrcHis-desD and its mutants. Total cell extracts were prepared and total protein (15–20 μg protein per lane) was analyzed as described in the Materials and methods. M Pre-stained molecular weight markers. Lane 1 pTrcHisA, lane 2 pTrcHis-desD, lane 3 pTrcHis-desD H89R, lane 4 pTrcHis-desD H93R, lane 5 pTrcHis-desD H124R, lane 6 pTrcHis-desD H128R, lane 7 pTrcHis-desD H129R, lane 8 pTrcHis-desD H305R, lane 9 pTrcHis-desD H306R, lane 10 pTrcHis-desD H313R, lane 11 pTrcHis-desD H315N, lane 12 pTrcHis-desD R123N, lane 13 pTrcHis-desD G136H, lane 14 pTrcHis-desD E140Q, lane 15 pTrcHis-desD W294G, lane 16 pTrcHis-desD D138N
In vitro assay of Δ6 desaturase
In general, E. coli is able to incorporate exogenously added fatty acids into glycerolipids (Esfahani et al. 1969). When the host cells containing the designated plasmid with the insert are grown in the presence of sodium-linoleic acid, they can take up the exogenous substrate and transform it into a glycolipid, the only form that the Spirulina acyl-lipid Δ6 desaturase can utilize (Mustady et al. 1996). Also, it has been reported that cyanobacterial desaturation reactions require the reduced form of ferredoxin (Tocher et al. 1998). We therefore conducted our in vitro assay of the cell extract containing a substrate in the form of glycolipid and either Δ6 desaturase or modified Δ6 desaturase in the presence of exogenously provided cofactors. The resulting product was subsequently identified by GC (Fig. 3). In addition, it should be noted that GC-MS analysis of the two unknown peaks appearing after the peak of GLA in Fig. 3b was performed and the results showed that these peaks are not α-linolenic acid (ALA).
GC profiles of total fatty acids extracted from a E. coli containing pTrcHisA (control), b pTrcHis-desD and c pTrcHis-desD H313R in the presence of the substrate 18:2Δ9,12; and GC profile of d standard fatty acids, including ALA. The asterisk indicates the unusual fatty acid peak found in pTrcHis-desD H313R
Site-directed mutagenesis of Δ6 desaturase
Mutations of histidine residues in the three conserved histidine clusters that are purportedly involved in providing the catalytic iron center should theoretically diminish the ability of the enzyme to bind iron and thus eliminate desaturase activity (Murata and Wada 1995). To determine the role of these three histidine motifs in Δ6 desaturase, the mutated desD gene was expressed in E. coli, with the assay carried out in vitro. The resulting product was then analyzed.
The amino acid sequence alignment shown in Fig. 1 demonstrates that 14 amino acid residues were changed one-by-one using a Chameleon site-directed mutagenesis kit. Nine histidine residues were selected. Seven residues (H89, H93, H124, H128, H129, H305, H306) within the three conserved histidine motifs were mutated to arginine. In addition, two residues (H313, H315, which are conserved among cyanobacterial species and located adjacent to the third cluster) were mutated to arginine and asparagine, respectively. All the site-directed mutations of histidine residues within the three clusters caused an 80–100% reduction in the in vitro activity (Table 2); and mutation of the two conserved histidine residues (H313, H315) showed a complete elimination of desaturase activity (Table 2).
Interestingly, one of these mutants (H313R) revealed the synthesis of an unusual fatty acid with the same retention time as that of ALA (18:3Δ9,12,15; Fig. 3c,d). GC-MS analysis of the methyl-ester derivative of the fatty acid revealed that it was likely to be ALA, a conclusion drawn from the fact that the product obtained from the mutant enzyme possessed m/z 292, the parent ion of 18:3 methyl-ester. Moreover, analysis of the DMOX derivative of the unusual fatty acid synthesized by the H313R mutant enzyme showed m/z 196, 208, 236, 248, 276 and 288 (Fig. 4a), indicating double-bond positions at Δ9, Δ12 and Δ15, using the rule of 12 atomic mass unit intervals (Fay and Richli 1991). Analysis of the DMOX derivative fatty acid synthesized by the heterologously expressed Δ6 desaturase, however, showed m/z 194, 206, 234, 246, 220, 274, 166 and 167, indicating double-bond positions at Δ6, Δ9 and Δ12 (Fig. 4b).
Mass spectrum obtained from the GC-MS analysis of the DMOX derivative of a the unusual fatty acid extracted from pTrcHis-desD H313R, b the fatty acid GLA extracted from pTrcHis-desD in the presence of linoleic acid (18:2Δ9,12) and c the unusual fatty acid extracted from pTrcHis-desD H313R in the presence of linolenic acid (18:3Δ9,12,6)
In addition, since the H313R mutant enzyme has the ability to introduce a double bond at the Δ15 position, 10 μmol of GLA (18:3Δ9,12,6) was added to the culture medium of E. coli containing pTrcHis-desD H313R. The substrate was in the form of free fatty acid, due to the fact that GLA in the form of a sodium salt is not commercially available. The data showed that: (1) the cells were able to take up the substrate in the form of free fatty acid and (2) the mutant enzyme produced an extremely small peak of fatty acid product, which had a higher retention time than that of GLA. Then, a GC-MS analysis of the fatty acid was performed and the spectrum showed that this fatty acid was possibly stearidonic acid (SDA, 18:4Δ9,12,6,15) due to the presence of m/z 166, 167, 194, 206, 220, 234, 246, 260, 274 and 286 (Fig. 4c).
The single mutation of residue G136, E140, W294 or D138 totally eliminated the Δ6 desaturase activity, whereas mutation of R123 resulted in activity close to that of the wild type in the presence of 300 μM substrate (Table 2). However, the K m and V max of R123N (defined using a Lineweaver–Burk plot) were found to be 500 μM and 5 mg GLA mg protein−1 h−1, respectively. This mutation of a positively charged and non-polar residue (arginine) to an uncharged and polar residue (asparagine) led to a reduction in V max by approximately 50%, while the amount of K m remained unaltered (Table 3).
Discussion
This study was performed with the aim of identifying those amino acid residues within the Spirulina Δ6 desaturase required for its catalytic activity. The amino acid sequence alignment of various groups of Δ6 desaturase from various organisms demonstrates three histidine motifs conserved among all Δ6 desaturases, together with two histidine residues adjacent to the third cluster which are conserved among the acyl-lipid desaturases of cyanobacteria. In addition, there is a set of residues which we propose are located in the cytoplasmic phase and can be divided into three groups: (1) G135 and E140, conserved among cyanobacterial species, (2) D138 and W294, conserved among all listed organisms and (3) R123, conserved among cyanobacterial species and H. Sapiens (Fig. 1).
The Spirulina Δ6 desaturase was expressed as approximately 2% of the total protein in E. coli. Prior to the assay for enzyme activity being carried out in vitro, it was performed in vivo followed by GC analysis for the enzyme product, GLA. The enzyme product was not detected. However, the fatty acid GLA was detected after the in vitro assay in the presence of exogenously provided cofactors, including a photosynthetic form of ferredoxin from spinach. This indicated that the heterologously expressed enzyme was in a functional form. This finding also corresponded to studies involving desaturase activity influenced by ferredoxin.
In E. coli, an extremely low level of ferredoxin protein was expressed, as approximately 0.05% of the cell protein (Ta and Vickery 1992). Thus, an in vivo system for the characterization of desaturase activity in E. coli was developed by the co-expression of the ferredoxin gene in this organism (Cahoon et al. 1996). Otherwise, the activity would have to be assayed by the in vitro system. Moreover, Schultz et al. (2000) studied the influence of two isoforms of ferredoxin on the plant acyl–acyl carrier protein (ACP) desaturase: a photosynthetic form from Arabidopsis and spinach and a heterotrophic form from Impatiens balsamina. The results revealed that heterotrophic ferredoxins are 10- to 20-fold more effective than photosynthetic ferredoxins. Taking all of these data together, it is clear that ferredoxin plays a critical role in the desaturation activity of various desaturases.
The results obtained in the present study using the site-directed mutagenesis approach reveal that the three histidine-rich motifs are required for Δ6 desaturase activity, most likely by providing the catalytic iron center. Besides these motifs, there is a small set of histidine residues, H313 and H315, located adjacent to the proposed iron ligand. These residues were also found to be critical for catalytic activity.
Interestingly, an unusual fatty acid was synthesized in substantial quantities during the in vitro assay of the mutant H313R expressed in E. coli. This fatty acid has the same retention time as that of standard ALA (18:3Δ9,12,15). It has been reported that S. platensis contains a trace amount of ALA (Murata and Nishida 1987). Analysis by GC-MS showed that this fatty acid has a parent ion of 18:3 methyl-ester, m/z 292, as with GLA. Conducting a GC-MS analysis of the DMOX-derived fatty acid, we performed a test to locate the double bonds (Fig. 4). The spectrum showed that this unusual fatty acid is in fact ALA, a conclusion drawn from the presence of m/z 196, 208, 236, 248, 276 and 288 (Fay and Richli 1991), indicating double bonds at Δ9, Δ12 and Δ15 (Fig. 4a). Moreover, this mutant enzyme is also able to introduce a double bond at the Δ15 position in the other fatty acid substrate, GLA, and synthesizes SDA (18:4Δ9,12,6,15), as shown in Fig. 4c. These results strongly suggest that this residue might be involved either in the regioselectivity of the enzyme or in the alteration of enzyme function.
The evolutionary study of desaturases by Sperling et al. (2003) similarly reported that the creation of a new regioselectivity of a desaturase affects the amino acid sequence adjacent to the active site, which forms the substrate channel. Moreover, a study recently performed by Cahoon et al. (1998) showed that the single mutation of L118W causes a shift in the substrate specificity of acyl-ACP Δ9 desaturase. Whittle and Shanklin (2001) used a combinatorial saturation mutagenesis approach to identify two key residues that play a substantial role in the substrate specificity of Δ9 ACP desaturase. Broadwater et al. (2002) have meanwhile reported that the substitution of 4–7 residues in A. thaliana FAD2 (with desaturation activity) with residues from Ricinus communis LFAH (with both hydroxylase and desaturase activity) results in a substantial hydroxylase activity of the mutated FAD2. Interestingly, they also demonstrate that a single mutation of methionine at position 324 to isoleucine can cause a substantial shift in catalytic specificity.
In spite of mutations of the conserved histidine motifs, the Δ6 desaturase activity was also diminished by the single mutation of residue G136, E140, W294 or D138, indicating the critical role of these residues in the enzyme function. The amino acids E, W and D are known to be capable of binding to metal ligands and forming a metal catalytic center (Creighton 1993); and these residues are also located close to the second histidine cluster. They might thus be involved either in providing the catalytic iron center or in the maintenance of the structural integrity of the active site pocket. A similar finding was reported by Zámocky et al. (2001), who found that the site-directed mutagenesis of the distal active site, a conserved triad of arginine–tryptophan–histidine, revealed that these residues were essential for the catalysis of KatGs (plant peroxidase superfamily).
Moreover, W294, which is conserved in the Δ6 desaturase of all listed organisms, is positioned within the hydrophobic portion (residues 292–294), located close in sequence to the third histidine-rich region. This residue is located in the soluble portion of the enzyme. The discovery of a hydrophobic stretch within the soluble portion close to the di-iron active site can be expected, as it possibly plays a role in interacting with the acyl chains of membrane lipids (Diaz et al. 2002).
In addition, the substitution of R123, which is conserved among cyanobacterial species, to N, which is found conserved at the corresponding position in M. rouxii and B. officinalis, led to a reduction of approximately 50% in the level of V max, whereas the level of K m remained constant in comparison with the wild type. This V max/K m value points to the fact that the substitution of arginine at position 123 with asparagine causes a 50% reduction in enzyme efficiency, again in comparison with the wild type. Interestingly, Diaz et al. (2002) report that the hydropathy plots of several acyl-lipid desaturases show the presence of a hydrophobic segment located between the first two histidine clusters. They thus proposed that this hydrophobic stretch might be involved in substrate recognition. The R123 of Spirulina Δ6 desaturase is also located between the two first histidine clusters and the substitution of this residue causes a partial deficiency of the enzyme. This demonstrates that R123 is also important for the enzyme function, possibly by playing a part in the active site pocket.
This work provides crucial information on the amino acid residues required for the desaturation reaction of Spirulina Δ6 desaturase and identifies the role of three conserved histidine motifs, based on the results of our experimentation. Our data also allow us to delineate the topology of this membrane-bound desaturase more accurately. The three histidine motifs, as proposed by Murata and Wada (1995) and the amino acids H313, H315, R123, G136, E140, W294 and D138 are critical for desaturase activity and are thus likely to be located on the cytoplasmic phase of the membrane. We propose that these residues play a part in forming the active site, while the tripartite motif also possibly plays a role in providing the iron catalytic center.
The most important result to emerge from this study is the revelation for the first time of the role of H313 in the regioselectivity of the enzyme. At the same time, our work raises questions about the residues involved in regioselectivity and substrate specificity; and we propose that further experiments are conducted to elucidate the residues involved.
References
Avelange-Macherel M, Macherel D, Wada H, Murata N (1991) Site-directed mutagenesis of histidine residues in the delta 12 acyl-lipid desaturase of Synechocystis. FEBS 361:111–114
Bloomfield DK, Bloch K (1960) Formation of unsaturated fatty acids. J Biol Chem 235:337–345
Broadwater JA, Whittle E, Shanklin J (2002) Desaturation and hydroxylation; residue 148 and 324 of Arabidopsis FAD2, in addition to substrate chain length, exert a major influence in partitioning of catalytic specificity. Biol Chem 277:15613–15620
Cahoon EB, Mills LA, Shanklin J (1996) Modification of the fatty acid composition of Escherichia coli by coexpression of a plant acyl–acyl carrier protein desaturase and ferredoxin. J Bacteriol 178:936–939
Cahoon EB, Coughlan SJ, Shanklin J (1997) Characterization of a structurally and functionally diverged acyl–acyl carrier protein desaturase from milkweed seed. Plant Mol Biol 33:1105–1110
Cahoon EB, Lindqvist Y, Schneider G, Shanklin J (1997) Redesign of soluble fatty acid desaturases from plants for altered substrate specificity and double bond position. Proc Natl Acad Sci USA 94:4872–4877
Cahoon EB, Shah S, Shanklin J, Browse J (1998) A determinant of substrate specificity predicted from the acyl–acyl carrier protein desaturase of developing cat’s claw seed. Plant Physiol 117:593–598
Creighton TE (1993) Proteins: structures and molecular properties. Freeman, New York
Deng WP, Nickoloff JA (1992) Site directed mutagenesis of virtually any plasmid by eliminating a unique site. Anal Biochem 200:81–88
Deshnium P, Paithoonrangsarid K, Suphratrakul A, Meesapyodsuk D, Tanticharoen M, Cheevadhanarak S (2000) Temperature-independent and dependent expression of desaturase genes in filamentous cyanobacterium Spirulina platensis C1 (Arthrospira sp. PCC 9438). FEMS Microbiol Lett 184:207–213
Diaz AR, Mansilla MC, Vila AJ, Mendoza D (2002) Membrane topology of the acyl-lipid desaturase from Bacillus subtilis. Biol Chem 277:48099–48106
Esfahani M, Barnes EMJ, Wakil S (1969) Control of fatty acid composition in phospholipids of Escherichia coli: response to fatty acid supplements in a fatty acid auxotroph. Proc Natl Acad Sci USA 64:1057–1064
Fay L, Richli U (1991) Location of double bonds in polyunsaturated fatty acids by gas chromatography-mass spectrometry after 4,4-dimethyloxazoline derivatization. J Chromatogr 541:89–98
Gravel P, Golaz O (1996) Protein blotting by the semi-dry method. In: Walker JM (ed) The protein protocols handbook. Humana, New Jersey, pp 249–260
Holloway PW (1983) Fatty acid desaturation. In Boyer PD (ed) The enzymes, vol 16. Academic, Orlando, pp 63–83
Hongsthong A, Deshnium P, Paithoonrangsarid K, Cheevadhanarak S, Tanticharoen M (2003) Differential responses of the three acyl-lipid desaturases to immediate temperature reduction occurred in two lipid membranes of Spirulina platensis strain C1. J Biosci Bioeng 96:519–524
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with the Folin phenol reagent. J Biol Chem 193:269–275
Mcguire KA, Siggaard-Andersen M, Bangera MG, Olsen JG, Wettstein-Knowles P (2001) β-Ketoacyl-(acyl carrier protein) synthase I of Escherichia coli: aspects of the condensation mechanism revealed by analyses of mutations in the active site pocket. Biochemistry 40:9836–9845
Murata N, Nishida I (1987) Lipids of blue-green algae. In: Stumpf PK (ed) The biochemistry of plants, vol 9. Academic, Orlando, pp 315–347
Murata N, Wada H (1995) Acyl-lipid desaturases and their importance in the tolerance and acclimation to cold of cyanobacteria. Biochem J 308:1–8
Murata N, Deshnium P, Tasaka Y (1996) Biosynthesis of γ-linolenic acid in the cyanobacterium Spirulina platensis. In: Huang YS, Mills DE (eds) γ-Linolenic acid metabolism and its roles in nutrition and medicine. AOCS, Urbana-Champaign, pp 22–32
Mustady L, Los DA, Gombos Z, Murata N (1996) Immunocytochemical location of acyl-lipid desaturases in cyanobacterial cells: evidence that both thylakoid membranes and cytoplasmic membranes are the sites for lipid desaturation. Proc Natl Acad Sci USA 93:10524–10527
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y.
Schmidt H, Heinz E (1993) Direct desaturation of intact galactolipids by a desaturase solubilized from spinach (Spinacia oleracea) chloroplast envelopes. Biochem J 289:777–782
Schmidt H, Sperling P, Heinz E (1993) Biochemistry and molecular biology of membrane and storage lipids of plants. American Society for Plant Physiologists, Rockville
Schultz DJ, Chung Suh M, Ohlrogge JB (2000) Stearoyl–acyl carrier protein and unusual acyl–acyl carrier protein desaturase activities are differentially influences by ferredoxin. Plant Physiol 124:681–692
Shanklin J, Cahoon EB (1998) Desaturation and related modifications of fatty acids. Annu Rev Plant Physiol Plant Mol Biol 49:611–641
Shanklin J, Whittle E, Fox BG (1994) Eight histidine residues are catalytically essential in a membrane-associated iron enzyme, steroyl-CoA desaturase and are conserved in alkene hydroxylase and xylene monooxygenase. Biochemistry 33:12787–12794
Sperling P, Ternes P, Zank TK, Heinz E (2003) The evolution of desaturases. Prostaglandins Leukot Essent Fatty Acids 68:73–95
Sundberg RJ, Martin RB (1974) Interactions of histidine and other imidazole derivatives with transition metal ions in chemical and biological systems. Chem Rev 74:471–517
Ta TD, Vickery LE (1992) Cloning, sequencing and overexpression of a [2Fe–2S] ferredoxin gene from Escherichia coli. J Biol Chem 267:11120–11125
Tocher DR, Leaver MJ, Hodgson PA (1998) Recent advances in the biochemistry and molecular biology of fatty acyl desaturases. Prog Lipid Res 37:73–117
Wada H, Avelange-Macherel M, Murata N (1993) The desA gene of the cyanobacterium Synechocystis sp. strain PCC 6803 is the structural gene for Δ12 desaturase. J Bacteriol 175:6056–6058
Whittle E, Shanklin J (2001) Engineering delta9-16:0-acyl carrier protein (ACP) desaturase specificity based on combinatorial saturation mutagenesis and logical redesign of the castor delta9-18:0-ACP desaturase. Biol Chem 276:21500–21505
Zámocky M, Gunther R, Jakopitsch C, Obinger C (2001) The molecular peculiarities of catalase–peroxidase. FEBS 492:177–182
Acknowledgements
This research was funded by a grant from the National Center for Genetic Engineering and Biotechnology (BIOTEC), Bangkok, Thailand. The experiments comply with the current laws of Thailand.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hongsthong, A., Subudhi, S., Sirijuntarat, M. et al. Mutation study of conserved amino acid residues of Spirulina Δ6-acyl-lipid desaturase showing involvement of histidine 313 in the regioselectivity of the enzyme. Appl Microbiol Biotechnol 66, 74–84 (2004). https://doi.org/10.1007/s00253-004-1655-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-004-1655-x