Abstract
The gene coding for an aerobic azoreductase was cloned from Pigmentiphaga kullae K24, which is able to grow with the carboxylated azo compound 1-(4′-carboxyphenylazo)-4-naphthol (carboxy-Orange I) as sole source of carbon and energy. The gene encoded a protein with a molecular weight of 20,557 Da, with a conserved putative NAD(P)H-binding site in the amino-terminal region. The deduced amino acid sequence showed no further significant sequence homologies to previously studied aerobic azoreductases. The azoreductase was heterologously expressed in Escherichia coli and shown to convert the sulfonated azo dye Orange I and furthermore Magneson II [4-(4-nitrophenylazo)-1-naphthol].
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Azo dyes are widely used in the textile, leather, food, and cosmetics industries (Anliker 1979; Zollinger 1991). There are only very few bacteria known that are able to grow aerobically on azo compounds. These bacteria initially cleave the azo group (–N=N–) reductively and then utilize the amines formed in the course of the reaction as carbon, energy, and nitrogen source (Blümel et al. 1998; Bumpus 1995; Stolz 2001). The best studied examples of this metabolic trait are Xenophilus azovorans KF46 (previously Pseudomonas sp. KF46) and Pigmentiphaga kullae K24 (previously Pseudomonas sp. K24), which can grow aerobically with the model azo dyes 1-(4′-carboxyphenylazo)-2-naphthol (carboxy-Orange II) or 1-(4′-carboxyphenylazo)-4-naphthol (carboxy-Orange I) (Blümel et al. 2001a, 2001b; Kulla 1981; Kulla et al. 1984). X. azovorans KF46, P. kullae K24 and also some more recently described bacterial strains (e.g. Bacillus strain OY1-2, Paenibacillus azoreducens and Sphingomonas sp. 1CX) are also able to decolorize a wide range of more complex sulfonated azo dyes under aerobic conditions. In general, these dyes cannot be used by the bacteria as carbon and energy sources, probably because they are unable to productively metabolize the sulfonated or highly substituted amines formed as reduction products (Coughlin et al. 1999; Meehan et al. 2001; Suzuki et al. 2001).
The aerobic reductive metabolism of azo dyes requires specific enzymes—aerobic azoreductases—that catalyze the NAD(P)H-dependent reduction of azo compounds to the corresponding amines (Zimmermann et al. 1982, 1984). Recently, the genes encoding aerobic azo reductases have been cloned from Bacillus strain OY1-2 and X. azovorans KF46F (Blümel et al. 2002; Suzuki et al. 2001). Furthermore, a protein with the ability to decolorize the simple azo dye Methyl Red [4′-(dimethylamino)-azobenzene-2-carboxylic acid] has been cloned from Escherichia coli (Nakanishi et al. 2001). These three enzymes did not show any significant sequence homologies with each other and therefore presumably are the result of convergent evolution. In order to obtain more information about the evolution of aerobic azoreductases we decided to study the aerobic azo reductase from P. kullae K24, which was previously shown to differ significantly in its substrate specificity from the aerobic azo reductases from X. azovorans KF46F and Bacillus strain OY1-2 (Suzuki et al. 2001; Zimmermann et al. 1984), on the molecular level.
Materials and methods
Bacterial strains, media, and plasmids
Strain K24 was previously isolated by Kulla et al. (1984) after enrichment with the model azo dye carboxy-Orange I as sole source of carbon and energy. The strain was originally described as Pseudomonas sp. K24 and recently reclassified as Pigmentiphaga kullae, representing a new genus within the family Alcaligenaceae (Blümel et al. 2001b). The strain has been deposited at the Deutsche Sammlung für Mikroorganismen und Zellkulturen (DSMZ) as DSM 13608. P. kullae K24 was routinely cultivated in LB-medium plus Orange I (50 mg/l).
For recombinant DNA work, E. coli DH5α and E. coli BL21(DE3)pLysS were used. The E. coli strains were cultured at 37°C in Luria-Bertani (LB) medium supplemented with ampicillin (100 μg/ml) if appropriate.
The plasmids pBluescript II KS(+) (Alting-Mees et al. 1992) and pET11a (Studier and Moffatt 1986) were used for cloning experiments and for expression, respectively, of the recombinant azo reductase in E. coli.
Standard assay for the determination of enzyme activities
Azoreductase activity was determined spectrophotometrically with cell extracts or purified enzyme fractions at room temperature at λ= 482 nm. The test cuvettes contained, in 1.0 ml 87 µmol potassium phosphate buffer (pH 7.1): 1 µmol NADH, 8 nmol Orange I (ε482=22.3 mM−1 cm−1) and different amounts of protein (1–600 µg). One unit of enzyme activity was defined as the amount of enzyme catalyzing the decolorization of 1 µmol substrate per minute. The protein content of cell extracts and purified enzyme fractions was determined by the method of Bradford (1976).
Enzyme purification
Protein purification was performed at room temperature using a fast-performance liquid chromatography system, which consisted of a LCC 500 controller, pump P-500, UV-1 monitor, conductivity monitor, REC-482 recorder, and FRAC autosampler from Amersham Pharmacia (Uppsala, Sweden). For purification of azoreductase, cells of P. kullae K24 were grown in LB medium with Orange I (0.15 mM) at 30°C to the end of the exponential growth phase. The cells were harvested by centrifugation and a crude extract prepared using a French press. This cell extract (189 mg protein) was loaded onto a Blue Sepharose Cibachrome Blue 3GA column (column volume 50 ml; Sigma, Deisenhofen, Germany). The column was initially washed with 0.1 M K-phosphate buffer (pH 8.5) and the proteins subsequently eluted with an increasing linear gradient of NADH (in 0.1 M K-phosphate buffer). The azoreductase was eluted as a single peak at a concentration of about 17–20 mM NADH. The active fractions (4 ml each) were pooled and NADH removed by repeated ultrafiltration and dilution with 0.1 M K-phosphate buffer (pH 8.5). The concentrated protein solution was finally applied to a Red Sepharose CL-6B column (column volume 32 ml; Amersham Pharmacia). The column was initially washed with 250 ml 0.1 M K-phosphate buffer (pH 8.5) and afterwards with 60 ml 10 mM NADH in K-phosphate buffer (0.1 M, pH 8.5). Finally, bound proteins were eluted with 60 ml 18 mM NADH in the same phosphate buffer. Fractions containing azoreductase were concentrated by ultrafiltration (Centricon 30, Amicon, Danvers, Mass.) and contained only a single protein as judged by SDS-PAGE.
Protein analysis
Preparation of cell-free extracts, polyacrylamide gel electrophoresis, determination of molecular weight, protein cleavage, isolation of peptides, and sequencing of peptides and N-termini were performed as described previously (Blümel et al. 2002).
DNA manipulation techniques
Genomic DNA was isolated as described by Ausubel et al. (1987). Plasmid DNA from E. coli DH5α was isolated with a Flexi-Prep kit (Amersham Pharmacia) or a Qiaprep Spin Miniprep kit (Qiagen, Hilden, Germany). Digestion of DNA with restriction endonucleases (Gibco BRL, New England Biolabs), electrophoresis, purification of DNA fragments, and ligation with T4 DNA ligase (Gibco BRL) were performed according to standard procedures (Sambrook et al. 1989). Transformation of E. coli with plasmid DNA was carried out as described by Inoue et al. (1990). DNA hybridization, DNA sequencing and nucleotide sequence analysis were performed as described previously (Blümel et al. 2002).
Polymerase chain reaction
PCR experiments and the cloning of PCR products were performed as described previously (Marchuk et al. 1991; Blümel et al. 2002). For the amplification reaction, the following oligonucleotide primers (deduced from the amino-terminus of azoreductase) were used: primer AR-Nterm-2 (corresponding to amino acids 3–10 of the amino terminus) 5′-ATC GCG ATC ATC GGC GCC ACT GGC-3′ and primer AR-Nterm-30–25 (corresponding to amino acids 25–30 of the amino terminus): 3′-GT(AG) GT(AG) CAN TGN CGN TA-5′.
Expression of azoreductase in E. coli
The azoreductase gene was amplified by PCR using the following oligonucleotide primers: 5′-CCG CCA TAT GAA TAT CGC CAT CAT CGG-3′ and 5′-AAT TGG ATC CTG GGA GAG GCA TCA CGT TA-3′. This resulted in the introduction of an NdeI site upstream and a BamHI site downstream of the azoreductase gene azoB. The amplified products were cleaved with NdeI and BamHI and ligated into pET11a (Studier and Moffatt 1986). The resulting plasmid was used to transform E. coli DH5α.
Chemicals
Acid Orange I was obtained from Fluka (Buchs, Switzerland) and [4-(4-nitrophenylazo)-1-naphthol] (Magneson II) from Aldrich (Steinheim, Germany). The sources of all other chemicals and the oligonucleotides have been described previously (Blümel et al. 2002).
Nucleotide sequence accession number
The sequence data reported in this article will appear in the GenBank nucleotide sequence database under the number AY165002.
Results
Purification of aerobic azoreductase
Azoreductase was purified from P. kullae K24 cells cultivated in LB medium with Orange I (0.15 mM). The azoreductase was purified from cell extracts by two subsequent affinity chromatographies using Blue Sepharose Cibachrom Blue 3GA and Red-Sepharose CL-6B (see Materials and methods), which resulted in an approximately 40-fold purification, giving a specific activity of 2.8 U/mg protein (Table 1). The purified enzyme appeared as a single band on SDS-PAGE with a molecular weight of approximately 20,000 Da. Previously, Zimmermann et al. (1984) determined by SDS gel electrophoresis a molecular weight of 20,200±1,200 for the azoreductase from strain K24.
Determination of the NH2-terminal amino acid sequence
The NH2-terminal amino acid sequence was determined by Edman degradation as MNIAIIGATGNVGSRLVNEALARGHHVTAIARHASRLAARP. The sequence of one internal peptide fragment was determined, after trypsin digestion of the enzyme and HPLC purification of the peptides, as ALARGHHVTAIARHASRLAARGLDTLDIDLA. These two sequences obviously overlapped, thus allowing 51 amino acids from the amino terminus of the azoreductase to be deduced.
Cloning of the azoreductase gene
The amino acids at positions 3–10 and 25–30 of the amino terminal amino acid sequence served for the design of oligonucleotide primers. The utilization of these primers and genomic DNA of strain K24 as template resulted in the amplification of a DNA-fragment with the expected size of 83 bp. This PCR-product was cloned into a pBluescript II KS(+) T-vector, sequenced, labeled using DIG-labelled dUTP residues and used as a probe in Southern hybridization to identify the complete azoreductase gene. The hybridization experiments resulted in the identification of an EcoRI fragment of approximately 6 kb and a 3 kb PstI fragment from the total DNA of strain K24, and finally in the cloning of a 12 kb EcoRI fragment into plasmid pBluescript II SK(+) giving pBlue-OI-E1-2.
Determination of the nucleotide sequence of the azoreductase gene and surrounding DNA fragments
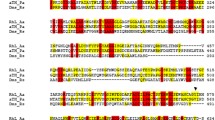
We searched for the azoreductase gene (azoA) by sequencing a continuous 11,821 bp DNA insert in plasmid pBlue-OI-E2 and identified the gene on this fragment by its amino-terminal region (Fig. 1). The gene encoded a protein consisting of 200 amino acids, which corresponded to a molecular weight of 20,557 kDa. Thus the protein was significantly smaller than the aerobic azoreductase from X. azovorans (30,278 Da) that was previously cloned in our laboratory. An NADPH binding site was identified near the amino-terminus of the deduced protein sequence (Fig. 1). A BLAST search using the deduced amino acid sequence suggested that AzoA possessed the highest degree of sequence identity (49%) to a hypothetical protein (Q8XSF1) from Ralstonia eutropha. No significant sequence similarities were detected with the aerobic azoreductases from Bacillus sp. OY1-2 and X. azovorans KF46F or the Methyl Red decolorizing activity from E. coli (Blümel et al. 2002; Nakanishi et al. 2001; Suzuki et al. 2001).
The azoreductase gene was present between positions 1,056–1,658 of the sequenced 11,821 bp DNA fragment. The sequenced DNA-fragment contained some more putative ORFs downstream of the azoreductase gene, with no obvious functional relationship to the azoreductase (Table 2).
Expression of the azoreductase in E. coli
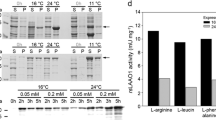
The azoreductase gene was amplified by PCR from the genomic DNA of strain K24 and functionally expressed in E. coli using a phage T7-promotor system as described previously (Blümel et al. 2002). Cell extracts from IPTG-induced cultures of E. coli BL21(DE3)pLysS carrying the relevant expression plasmid demonstrated an azoreductase activity (0.42 U/mg protein) that was about 6-fold higher than the activity observed in P. kullae K24.
Substrate specificity of the azoreductase
It was previously shown that the azoreductase from strain K24 requires the presence of a hydroxy-group on the naphthalene ring system in ortho-position towards the azogroup (Zimmermann et al. 1984). This structural element is very rare in azo dyes other than Orange I, and could only be detected in the commercially available monoazo dye Magneson II. This dye was converted with about 13% of the activity found with Orange I.
Discussion
P. kullae K24 was originally isolated and characterized by Kulla, Leisinger, Zimmermann and coworkers in a project that also resulted in the isolation of X. azovorans KF46. Both strains were isolated by long-term enrichments in continuous culture systems but with the difference that P. kullae K24 was enriched with carboxy-Orange I, while strain X. azovorans was isolated with carboxy Orange II. The results of some immunological experiments had already suggested that Orange I and Orange II azoreductases would be rather different, because no cross-reactivity was observed using polyclonal antibodies raised against the purified enzymes. This initial suggestion has now been proven by the determination of the nucleotide sequences of both genes (azoA and azoB), which clearly demonstrates that the enzymes share significant sequence similarity only in the amino-terminus, which in both enzymes contains an NAD(P)H-binding site.
Recently, two more bacterial genes encoding enzymes with the ability to aerobically reduce and decolorize azo dyes have been cloned and sequenced from other bacterial sources. Thus, an aerobic azoreductase, which decolorizes Acid Red 88 [2′-hydroxy-(1,1′)-azonaphthalene-4-sulfonic acid; C.I. 15620], Acid Orange 7 [C.I. 15510] and a series of proprietary reactive dyes, was described from a Bacillus strain (OY1-2) (Suzuki et al. 2001). More recently, a putative aerobic azoreductase (acpD) was cloned from E. coli (Nakanishi et al. 2001). This enzymatic activity was originally detected as a Methyl Red decolorizing activity in cell extracts of E. coli. In contrast to the other three characterized aerobic azoreductases, this enzyme activity contained a flavine cofactor (FMN). It was suggested that this enzymatic activity is evolutionarily related to NAD(P)H:quinone acceptor oxidoreductases (Nakanishi et al. 2001) and therefore will presumably convert only Methyl Red and structurally related azo dyes.
Thus, currently the sequences of four genes that encode proteins with the ability to decolorize azo dyes under aerobic conditions are known. Surprisingly, no significant sequence homologies can be found among these proteins, which suggests that they have evolved the ability to reduce azo compounds independently. Furthermore, this suggests that several possibilities exist in nature to develop catalysts with azoreductase activity and that the emergence of aerobic azoreductases has not been a single event in evolution. This appears to contrast the observation that it has been extremely difficult to isolate bacteria with the ability to grow aerobically with sulfonated azo dyes and may indicate that the aerobic degradation of sulfonated azo compounds is not restricted by the metabolism of the azo group.
References
Alting-Mees MA, Sorge JA, Short JM (1992) pBluescript II: multifunctional cloning and mapping vectors. Methods Enzymol 216:483–495
Anliker R (1979) Ecotoxicology of dyestuffs—a joint effort by industry. Ecotoxicol Environ Saf 3:59–74
Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl K (1987) Current protocols in molecular biology, vol 1, chapter 2.4. Wiley, New York
Blümel S, Contzen M, Lutz M, Stolz A, Knackmuss H-J (1998) Isolation of a bacterial strain with the ability to utilize the sulfonated azo compound 4-carboxy-4′-sulfoazobenzene as sole source of carbon and energy. Appl Environ Microbiol 64:2315–2317
Blümel S, Busse H-J, Stolz A, Kämpfer P (2001a) Xenophilus azovorans gen. nov., sp. nov., a soil bacterium able to degrade azo dyes of the Orange II type. Int J Syst Evol Bacteriol 51:1831–1837
Blümel S, Mark B, Busse H-J, Kämpfer P, Stolz A (2001b) Pigmentiphaga kullae gen. nov., sp. nov., a new member of the family Alcaligenaceae with the ability to decolorize aerobically azo dyes. Int J Syst Evol Bacteriol 51:1867–1871
Blümel S, Knackmuss H-J, Stolz A (2002) Molecular cloning and characterization of the gene coding for the aerobic azoreductase from Xenophilus azovorans KF46F. Appl Environ Microbiol 68:3948–3955
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Bumpus JA (1995) Microbial degradation of azo dyes. In: Singh VP (ed) Microbial degradation of health risk compounds. Elsevier, Amsterdam, pp 157–167
Coughlin MF, Kinkle BK, Bishop PL (1999) Degradation of azo dyes containing aminonaphthol by Sphingomonas sp. strain 1CX. Ind Microbiol Biotechnol 23:341–346
Inoue H, Nojima H, Okayama H (1990) High efficiency transformation of Escherichia coli with plasmids. Gene 96:23–28
Kulla HG (1981) Aerobic bacterial degradation of azo dyes. In: Leisinger T, Cook AM, Nüesch J, Hütter R (eds) Microbial degradation of xenobiotics and recalcitrant compounds. Academic, London, pp 387–399
Kulla HG, Krieg R, Zimmermann T, Leisinger T (1984) Experimental evolution of azo dye-degrading bacteria. In: Klug MJ, Reddy CA (eds) Current perspectives in microbial ecology. American Society of Microbiology, Washington, D.C., pp 663–667
Marchuk D, Drumm M, Saulino A, Collins FS (1991) Construction of T-vectors, a rapid and general system for direct cloning of unmodified PCR products. Nucleic Acids Res 19:1154
Meehan C, Bjourson AJ, McMullan G (2001) Paenibacillus azoreducens sp. nov., a synthetic azo dye decolorizing bacterium from industrial wastewater. Int J Syst Evol Microbiol 51:1681–1685
Nakanishi M, Yatome C, Ishida N, Kitade Y (2001) Putative ACP phosphodiesterase gene (acpD) encodes an azoreductase. J Biol Chem 276:46394–46399
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
Stolz A (2001) Basic and applied aspects in the microbial degradation of azo dyes. Appl Microbiol Biotechnol 56:69–80
Studier FW, Moffatt BA (1986) Use of bacteriophage T7 RNA polymerase to direct selective high-level expression of cloned genes. J Mol Biol 189:113–130
Suzuki Y, Yoda T, Ruhul A, Sugiura W (2001) Molecular cloning and characterization of the gene coding for azoreductase from Bacillus sp. OY1-2 isolated from soil. J Biol Chem 276:9059–9065
Zimmermann T, Kulla HG, Leisinger T (1982) Properties of purified Orange II azoreductase, the enzyme initiating azo dye degradation by Pseudomonas KF46. Eur J Biochem 129:197–203
Zimmermann T, Gasser F, Kulla HG, Leisinger T (1984) Comparison of two azoreductases acquired during adaptation to growth on azo dyes. Arch Microbiol 138:37–43
Zollinger H (1991) Color chemistry, 2nd edn. VCH, Weinheim
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Blümel, S., Stolz, A. Cloning and characterization of the gene coding for the aerobic azoreductase from Pigmentiphaga kullae K24. Appl Microbiol Biotechnol 62, 186–190 (2003). https://doi.org/10.1007/s00253-003-1316-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-003-1316-5