Abstract
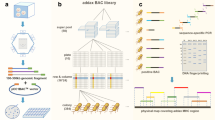
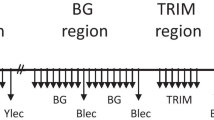
Bacterial artificial chromosome (BAC) clones were assigned within the pig major histocompatibility complex (Mhc) by polymerase chain reaction-screening and Southern blot hybridization using sequence-tagged site (STS) markers and BAC end-rescued sequences. In all, 35 BAC clones were discovered containing 12 anchor genes of the SLA class I region and two genes of the SLA class III region. Twenty of these 35 clones comprised two distinct class I gene clusters, each spanning about 100 kilobases. One cluster enclosed three class I related genes (SLA-6 to -8) and two genes (MIC-1 and MIC-2) more distantly related to class I. The other cluster enclosed typical class I genes, of which three (SLA-1, -2, and -3) were transcribed by fibroblasts homozygous for the H01 haplotype which we used to construct a pig BAC library. Ordered clones are certainly helpful in isolating agronomically, biologically, and medically important genes. They would also be useful for inducing genetic modifications in pig cell lines.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Author information
Authors and Affiliations
Consortia
Additional information
Received: 1 December 1998 / Revised: 5 May 1999
Rights and permissions
About this article
Cite this article
Velten, F., Renard, C., Rogel-Gaillard, C. et al. Spatial arrangement of pig MHC class I sequences. Immunogenetics 49, 919–930 (1999). https://doi.org/10.1007/s002510050575
Issue Date:
DOI: https://doi.org/10.1007/s002510050575