Abstract
The role of natural killer (NK) cells is tightly modulated by interactions of killer cell immunoglobulin-like receptors (KIR) with their ligands of the MHC class I family. Several characteristics of the KIR gene products are conserved in primate evolution, like the receptor structures and the variegated expression pattern. At the genomic level, however, the clusters encoding the KIR family display species-specific diversity, reflected by differential gene expansions and haplotype architecture. The human KIR cluster is extensively studied in large cohorts from various populations, which revealed two KIR haplotype groups, A and B, that represent more inhibitory and more activating functional profiles, respectively. So far, genomic KIR analyses in large outbred populations of non-human primate species are lacking. In this study, we roughly quadrupled the number of rhesus macaques studied for their KIR transcriptome (n = 298). Using segregation analysis, we defined 112 unique KIR region configurations, half of which display a more inhibitory profile, whereas the other half has a more activating potential. The frequencies and functional potential of these profiles might mirror the human KIR haplotype groups. However, whereas the human group A and B KIR haplotypes are confined to largely fixed organizations, the haplotypes in macaques feature highly variable gene content. Moreover, KIR homozygosity was hardly encountered in this panel of macaques. This study exhibits highly diverse haplotype architectures in humans and macaques, which nevertheless might have an equivalent effect on the modulation of NK cell activity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The killer cell immunoglobulin-like receptor (KIR) family comprises inhibitory and activating members that modulate the education and activity of NK cells through interactions with their ligands, mainly MHC class I allotypes (Parham et al. 2012; Ljunggren and Kärre 1990; Lanier 2005). The human KIR and MHC gene clusters are encoded on chromosomes 19 and 6, respectively, and are considered to represent the most complex regions in the primate genome. Different molecular mechanisms, including point mutations and chromosomal recombination events, propel a rapid evolution process that diversifies the KIR gene region at a species, population, and individual level (Bruijnesteijn et al. 2020b). This diversity involves allelic polymorphism recorded for the different KIR genes as well as gene content variation defined on the KIR haplotypes. Moreover, novel fusion KIR entities that might possess distinct functional characteristics are generated by intragenic recombination events, which extend the total set of genes (Shilling et al. 1998; Martin et al. 2003). Unequal crossing-over processes seem to impact the continuous expansion and contraction events observed for this component of the immune system and thus may alter the genetic architecture (Shilling et al. 1998; Wilson et al. 2000). A variegated expression pattern of these diverse KIR genes generates a NK cell repertoire that is highly unique to individuals. Different alternative splicing patterns extend the KIR receptor diversity even more (Bruijnesteijn et al. 2018a).
Although human KIR haplotypes vary in their gene content, a relatively simple classification distinguishes two groups that comprise A and B haplotypes (Uhrberg et al. 1997). The basis of these haplotypes consists of four framework genes that mark a centromeric (KIR3DL3 to KIR3DP1) and a telomeric (KIR2DL4 to KIR3DL2) region. The group A haplotypes contain a fixed set of seven KIR genes, of which a single gene encodes an activating receptor, KIR2DS4. In contrast, the group B haplotypes are more diverse in the number of genes they contain and may encode one or more activating members. These diverse group B haplotypes, however, display some consistency that is reflected by the strong linkage disequilibria involving several KIR genes, such as KIR2DL5-KIR2DS3/S5 (Vierra-Green et al. 2012; Gourraud et al. 2010). Both haplotype groups are distributed among all human populations, but the relative frequency of KIR genes and haplotypes displays variation among different ethnical and geographical populations (John et al. 2012; Gonzalez-Galarza et al. 2020). This suggests a balancing selection that results from a functional relevance of the different inhibitory (group A) and activating (group B) haplotype profiles.
In other primate species, such as chimpanzees, orangutans, and macaques, a selection for distinct groups of KIR haplotypes is not yet documented or evident. The most common KIR haplotypes in chimpanzees and orangutans largely resemble the organization of a supposed ancestral haplotype, including three framework genes and one to five KIR2D genes (Guethlein et al. 2017; Abi-Rached et al. 2010). This indicates that human KIR haplotypes culminated in the formation of functionally different haplotype groups after speciation and were shaped by differential selective factors. In macaques, a species commonly used as a model in preclinical studies to develop vaccines and medicines, the KIR haplotypes display extensive levels of expansion and contraction (Bruijnesteijn et al. 2018b; Blokhuis et al. 2010). Abundant chromosomal recombination events rapidly rearrange the haplotype architecture and expand the macaque KIR repertoire by generating novel fusion genes. The plasticity of the contemporary KIR gene cluster in macaques exceeds that observed in humans (Bruijnesteijn et al. 2018b). This difference in diversity is, for instance, demonstrated by the high frequency of the non-variable group A haplotypes in human populations, and the relatively large KIR gene repertoire encountered in rhesus macaques. The reason for a differential expansion of the KIR gene cluster in primate species and its subsequent selection forces is elusive. It might involve a balanced co-evolution with MHC class I ligands and thereby indirectly with the continuous arms race with pathogens, but distinct pathways to educate NK cells might also skew the expansion of KIR (Bruijnesteijn et al. 2020b). The diversification of the primate KIR system seems to be an ongoing process and may achieve an equilibrium at a given moment in evolution that serves different functions; this might involve not only antiviral defense but also successful reproductive biology. This equilibrium might have become more balanced in humans — evidenced by a relatively fixed set of KIR genes and two sets of categorized haplotype groups — than in rhesus macaques, which display extensive diversification in their KIR cluster.
The differential expansion and contraction of KIR haplotypes in primate species is in contrast to the similarities in the characteristics of the encoded receptors, such as their structure (KIR2D and KIR3D), variegated expression patterns, and MHC class I ligand specificity. A better understanding of the rearranging KIR repertoires and haplotype configurations might help in the elucidation and understanding of the different selective forces that drove rapid KIR gene evolution. However, the limited number of characterized KIR haplotypes in outbred non-human primate populations might hamper a comparative analysis. For instance, defining different haplotype groups, like groups A and B in humans, or identifying co-occurring KIR genes, such as the human KIR2DL5-KIR2DS3/S5 tandem, is unreliable in small cohorts. In this communication, we report the KIR region configurations of 298 rhesus macaques, which roughly quadrupled the total number of studied individuals. This robust dataset allowed us to comprehensively analyze the KIR repertoires and region configurations in rhesus macaques and to compare these data to the human KIR haplotype architecture.
Materials and methods
Animals and cells
A total of 298 rhesus macaques from a self-sustaining colony housed at the Biomedical Primate Research Centre (BPRC, Rijswijk, the Netherlands), comprising 77 families of at least three members, were selected based on their MHC-A, -B, and -DRB content. The selected animals possessed 113 different MHC haplotypes, ensuring an outbred study cohort. PBMCs were isolated from heparin whole blood samples, which were obtained during regular annual health checks.
KIR transcriptomes
The rhesus macaque KIR transcriptome samples were prepared as previously described (Bruijnesteijn et al. 2018b). In short, total RNA was extracted from PBMCs, and cDNA was synthesized with the RevertAid First Strand cDNA Synthesis Kit (Invitrogen, Carlsbad, CA) using oligo(dT)18 primers. Two sets of barcoded primers amplified all full-length KIR3D transcripts, and an additional set was specific for KIR2DL04. The amplicons generated by the different primer sets from at least 40 animals were purified and subsequently pooled. After an additional clean-up step, the pools were used to prepare SMRTbell libraries. Sequencing primer V4 was used in combination with binding kit 2.1 and sequencing kit 2.0. Prior to library loading on the Pacific Biosciences Sequel II system, unbound polymerase enzyme was removed by AMPure PB beads. Sequence data collection was performed with a 20–24-h movie time to obtain sufficient yields of high-quality circular consensus reads. Raw sequencing data was base called to optimize read accuracy using a CCS algorithm (version 6.0.0) in SMRT Link software (version 10). Fastq files were generated using the following setting: minimum number of 3 passes, average QV of 36. Data was demultiplexed based on barcodes using demultiplex software (version 1.0.0). In total, seven runs were performed on the Sequel II system to obtain the KIR transcriptomes of all 298 animals.
The primer sets were validated to amplify all lineage I, II, and V KIR genes by comparing the transcriptome of three unrelated individuals to their genomic KIR haplotypes that were previously characterized by Oxford Nanopore sequencing (Bruijnesteijn et al. 2021). This comparison of transcriptome and genome confirmed the expression of all KIR genes present on a haplotype in peripheral blood samples and illustrated that all genes were detected by the used primer sets, except for pseudogene KIR2DP. Nevertheless, as this validation is constrained to only three cases, we cannot exclude the possibility that in some cases transcripts are missed by this study set-up.
KIR allele calling and phasing of region configurations
Demultiplexed fastq files were analyzed using Geneious Prime software (version 2021.2). First, reads were mapped to a database that contained all published rhesus macaque KIR sequences from the IPD-NHKIR database (release 1.3.0.0) to identify 100% matched transcripts (0% mismatch, minimum mapping quality = 30, minimum overlap identity = 80%, minimum overlap = 400, maximum ambiguity = 1). Unused reads were de novo assembled using Geneious-integrated assemble software (minimum overlap = 400, minimum overlap identity = 100%, maximum ambiguity = 1), and consensus sequences of the generated contigs were used in phylogenetic analysis together with a macaque KIR reference database to define novel KIR alleles. Sequences that branched independently from all known macaque KIR genes were reviewed and received official gene designations according to the primate KIR nomenclature guidelines (Bruijnesteijn et al. 2020c). Pivot tables were generated, which included all KIR genes/alleles that were identified per individual and the number of reads that were called for a specific allele. The family-based study design allowed segregation analysis of the defined KIR alleles from one generation to the next, thereby phasing KIR region configurations at the transcription level. All obtained consensus sequences were aligned with published sequences from the NHKIR-IPD database, and phylogenetic trees were generated. Novel KIR genes, alleles, and region configurations were confirmed in at least two individuals or by two independent PCR amplifications. Three of the Mamu-KIR1D sequences were recently renamed by the IPD-NHKIR Database, namely KIR1D*003:04 to KIR1D*003:02, KIR1D*003:02 to KIR1D*010, and KIR1D*003:03 to KIR1D*011.
KIR gene linkage analysis: co-occurrence and co-exclusion associations
The frequency of the different rhesus macaque KIR genes was determined based on all previously published and novel KIR region configurations by defining the presence of at least one gene copy or its absence. For each possible pair of KIR genes, the expected co-occurrence was determined by multiplying the individual gene frequencies. The observed co-occurrence was corrected for the expected frequencies, which provides a pairwise matrix, in which positive values indicate co-occurrence and negative values indicate co-exclusion. A statistical significance of the deviations between the observed and the expected co-occurrence or co-exclusion was determined by a two-tailed Fisher’s exact test with a significance level of 0.05.
Results
Extensive KIR polymorphism in rhesus macaques
The complete KIR transcriptome of 298 Indian rhesus macaques was characterized using single-molecule real-time (SMRT) sequencing on a Pacific Biosciences (PacBio) Sequell II platform. To confirm the presence of novel KIR alleles and to deduce haplotypes by segregation analysis, the study was designed in a family set-up, with at least one relative for each individual included. The rhesus macaque cohort represents an outbred population that originates from 114 founder animals, which display largely unique MHC repertoires (MHC-A, -B, and -DRB).
From previous transcriptome studies, 604 rhesus macaque KIR alleles were documented, which could be designated to 55 distinct KIR genes (IPD-NHKIR, release 1.3.0.0). We discovered another 101 novel KIR alleles, six of which represented novel gene entities (Table 1 and Suppl. Table I). In addition, 27 alleles were extended to full-length transcripts (Suppl. Table II). Most novel KIR alleles were documented for KIR3DL07, which is a highly frequent KIR gene, whereas a relatively large number of novel alleles clustered to KIR3DL20, KIR3DL01, KIR3DL05, and KIR3DL08. Abundant alternatively spliced transcripts were also recorded, which largely followed the previously reported splicing patterns (Bruijnesteijn et al. 2018a), but were not included for further analysis in this communication. The high number of novel alleles in the present cohort reflects the extensive polymorphism of the KIR gene system in rhesus macaques.
A diverse set of KIR region configurations in an outbred rhesus macaque cohort
Rhesus macaque KIR haplotypes were phased by segregation analysis and could be defined at the allele level resolution (Suppl. Fig. 1) and as region configurations (Suppl. Table III). In this study, we define a KIR region configuration as a differential set of genes that segregate together, thereby disregarding its allelic polymorphism. Only when multiple copies of a specific gene were found to segregate on a single haplotype, allele level information was used to define the gene copy number for the region configurations. The studied cohort comprised 112 unique KIR region configurations, of which the majority — namely, 73 configurations — had not previously been documented (Suppl. Table III). Thirteen novel KIR configurations could not be confirmed in at least two individuals and were excluded from further analysis. Eight KIR region configurations displayed different combinations of alleles, and these are defined as distinct KIR haplotypes (Fig. S1, indicated with additional letters as suffix), whereas all other region configurations represent a specific combination of KIR alleles.
On the basis of data from the current communication and from previously published transcriptome and segregation reports, a total of 118 unique KIR region configurations are at present defined in rhesus macaques (Suppl. Table III) that originate from Indian (n = 88), Burmese (n = 9), and Chinese (n = 21) populations. This set of KIR region configurations is used for further in-depth analysis and for comparison to its human equivalent. No further distinction is made for the different geographical populations, as the Burmese and Chinese cohorts represent a relatively low number of studied individuals.
Homozygosity of KIR haplotypes is virtually absent in the rhesus macaque cohort
In humans, most KIR region configurations are structured around several common centromeric and telomeric segments that display a specific gene content (e.g., cA01, tA01, cB01) (Pyo et al. 2010). Different combinations of these segments result in diverse sets of KIR region configurations (e.g., cA01-tA01, cA01-tB01) (Pyo et al. 2010, 2013). In addition, contraction and expansion of centromeric and telomeric segments may generate diversity by deleting or introducing genes, but the frequency of these deviating KIR configurations is generally relatively low, with some exceptions where haplotypes seem to have been enriched due to selection in particular populations (Norman et al. 2009; Misra et al. 2018; Cisneros et al. 2020). This is in contrast to the high frequency of the more standard KIR configurations, with the group A configuration (cA01-tA01) reaching a frequency of 50% or higher in most populations (Solloch et al. 2020; Hollenbach et al. 2012). As a result, the fraction of human individuals that are homozygous for their KIR gene content is significant and reaches over 30% for the KIR AA genotypes, which indicates a relatively moderate level of diversity concerning gene content per individual.
In contrast, such largely fixed KIR haplotype segments as documented in humans are mostly absent in rhesus macaques. Only the centromeric region displays two fixed configurations: namely, KIR3DL20 with or without the presence of KIR1D. These two centromeric segments are more or less balanced, with KIR1D present on approximately 40% of all KIR haplotype configurations. As such, the diverse array of unique KIR configurations in rhesus macaques is mostly generated by extensive contraction and expansion of the telomeric segment, which can contain 2 to 15 different KIR genes from a total repertoire of 61 known gene entities (IPD-NHKIR, release 1.3.0.0 and this study). The frequency of the different rhesus macaque KIR configurations ranges from 0.13 to 6.05% (Suppl. Table III). Only three region configurations displayed a frequency of 5% or above, whereas 45 different configurations were unique to one or two individuals (frequency < 0.25%). All configurations that were identified in a single individual were reported in previous studies that involved Burmese and Chinese rhesus macaque populations (Bruijnesteijn et al. 2020a). The relatively low configuration frequencies are also reflected in the minimal number of rhesus macaques that are homozygous for their KIR gene cluster; only six individuals from the current cohort (2.0%) display the same KIR gene content on both haplotypes. However, none of these homozygous haplotype configurations are identical at the allele level (Fig. S1). This extensive heterozygosity of the KIR haplotype configurations in rhesus macaques suggests a selective pressure to maintain a diverse array of KIR gene content in the population.
Fusion KIR entities indicate abundant chromosomal recombination events
Even though the most frequent human KIR haplotypes follow relatively standard configurations, abundant deviations are documented (Pyo et al. 2013; Vendelbosch et al. 2015). These diverse configurations are generated by chromosomal recombination events that shuffle complete centromeric and telomeric segments (e.g., cA01-tB01), but might also introduce and delete one or more KIR genes. Hotspots for recombination, such as transposable elements, that map within introns may mediate the generation of fusion KIR genes, which contain segments from two independent gene entities (Bruijnesteijn et al. 2020b). This molecular mechanism diversifies the genetic content of human KIR configurations, although the frequency of non-standard configurations seems to be limited in most populations that have been subject to analysis.
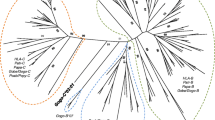
The reshuffling of centromeric and telomeric KIR region segments in rhesus macaques is hard to define due to the limited centromeric gene content. There are, however, KIR haplotypes that display identical telomeric segments at the allele level, combined with distinct centromeric regions. Haplotypes H5-A and H5-B, for instance, share seven alleles in their telomeric region but differ in their centromeric KIR allele content (Fig. S1), which hints at a chromosomal recombination event that interchanged complete centromeric and telomeric segments. A more prominent indication of chromosomal recombination events in rhesus macaques is provided by the relatively high number of fusion KIR genes that have been identified (Bruijnesteijn et al. 2020c). In total, 32 KIR gene entities are defined as fusion genes, which are distributed over 54 region configurations that together display a frequency of 35% (Suppl. Table III and Fig. S1). The only framework gene in rhesus macaques, KIR3DL20, is often substrate for recombination, either with KIR1D, which diminishes the extent of the centromeric region even more, or with KIR2DL04, which links the centromeric and telomeric region directly by the generation of a fusion gene (Fig. 1). Other frequent fusion KIR genes are KIR3DLW34 and KIR3DLW45, which consist of segments from an unknown donor and KIR3DLW03 and KIR3DL05 and KIR3DL10, respectively. The relatively high frequency of fusion KIR genes indicates that selection favors the maintenance of KIR gene cluster contraction and expansion, ultimately resulting in substantial diversification in rhesus macaques.
Generation of fusion genes by chromosomal recombination events involving KIR3DL20. Chromosomal recombination events might shuffle segments from different genes to generate a novel gene entity. Two distinct events are recorded for the framework gene KIR3DL20. One event involves the recombination with KIR1D, thereby contracting the centromeric haplotype region (blue lines and arrow), whereas the other event involves segment reshuffling with KIR2DL04, and fuses the centromeric and telomeric regions directly (red lines and arrow)
Rhesus macaque KIR region configurations might divide into functional haplotype groups
The presence of group A and B KIR haplotypes in all human populations and their differential frequencies suggests functional implications that are maintained by a balancing selection (Parham 2005). The functional properties of the different KIR haplotype groups might affect antiviral immunity and successful pregnancy, but the precise correlations are complex, and involve the presence of specific HLA epitopes (C1 and C2) as well (Hiby et al. 2004; Khakoo et al. 2004).
The large set of KIR region configurations in rhesus macaques could not be divided into distinct groups based on specific gene content. Similar to group A and B KIR haplotypes in humans, however, rhesus macaque KIR configurations might be categorized into more inhibitory profiles or profiles that display different levels of activating potential (Fig. 2 and Suppl. Table III). Region configurations with a more inhibitory profile are defined by the absence or the presence of one KIR gene that encodes for an activating receptor (hereinafter referred to, for sake of convenience, as an activating/inhibitory gene), whereas a more activating potential is distinguished by the presence of two or more activating KIR entities. The average number of inhibitory KIR genes present on a haplotype configuration varies for the different functional profiles (Fig. 2). Region configurations that lack an activating KIR gene display an average of 2.7 inhibitory KIR genes in their telomeric segment. In the presence of a single activating KIR gene, region configurations contain on average 3.4 inhibitory KIR genes. Most inhibitory entities are documented for KIR configurations with a more activating potential, with the presence of 4.6 inhibitory KIR genes on average.
Schematic overview of the KIR region architecture in rhesus macaques and humans. The organization of the most common KIR configurations in rhesus macaques and humans is schematically illustrated. Regions of diversity are in the branched lines. The centromeric and telomeric regions are divided by the double-lined break. Inhibitory and activating KIR genes are indicated by blue and red boxes, respectively. Lineage V and I KIR genes are indicated by yellow and green boxes, respectively, whereas grey boxes represent pseudogenes. In rhesus macaques, KIR3DL20 represents the only framework gene, whereas the presence of all other KIR genes is variable. The telomeric region in rhesus macaque is expanded, and the average number of inhibitory KIR genes is indicated in respect to the number of activating copies, representing the different functional haplotypes defined. As their telomeric KIR gene content displays great diversity, specific genes are not assigned to this region (empty blue and red boxes). The human haplotypes display four framework genes and several regions of diversity. The upper and lower content lines represent the more activating group B and more inhibitory group A KIR haplotypes, respectively. The human KIR haplotype representation is adapted from Abi-Rached and colleagues (Abi-Rached et al. 2010)
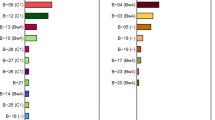
In total, 70 configurations display an inhibitory profile, of which 20 lack an activating KIR gene (Fig. 3). The more inhibitory group A KIR haplotypes in humans consistently encode KIR2DS4. In contrast, the inhibitory profiles in rhesus macaques display variability in their activating gene repertoire, of which KIR3DS05 is most prominently found (22%) (Fig. 4A). No specific KIR3DS05 alleles were associated with inhibitory profiles. The overall frequency of inhibitory profiles in the cohort studied was approximately 50% (Fig. 3), which is comparable to the frequency of group A KIR haplotypes in most human populations.
Distribution of the functional KIR region configuration profiles in rhesus macaques. The number of identified region configurations with a more inhibitory and more activating profile is designated by the blue bars, together with their subgroups that are distinguished based on the number of activating genes present on a configuration (none or 1 activating gene for inhibitory profiles and 2–5 activating genes for activating profiles). The relative frequencies of these region configuration categories are shown as a percentage by the red bars (right axis). More inhibitory than activating profiles are recorded, but their frequency is nearly equal in the population studied. Most inhibitory profiles contain a single activating entity, whereas the majority of the activating profiles are defined by two activating genes
Distribution of the different KIR genes on inhibitory and activating functional profiles. The frequency of KIR genes that could be identified on both inhibitory (blue bars) and activating (red bars) configuration profiles (A, upper panel) or on only one of the functional profiles (B, lower panel). The framework gene KIR3DL20 is always present
Activating profiles were evident for 48 KIR region configurations (Fig. 3 and Suppl. Table III), which contained two to five activating KIR genes. Half of these activating configurations contained two activating KIR genes, whereas three, four, and five activating copies were identified at 14, 4, and 6 region configurations, respectively (Fig. 3). The distribution of these different activating profiles in the present cohort displayed differential frequencies, with profiles containing two activating KIR genes being defined in 27% of the individuals, whereas region configurations with three, four, and five activating KIR genes were defined in 10.7%, 6.3%, and 6.1% of the rhesus macaques, respectively. On the region configurations with an activating profile, the most prominent activating gene is KIR3DS02, which is present with at least one copy on 68.7% of the configurations (Fig. 4A). Other activating genes that are frequently present on activating region configurations are KIR3DS05, which is prevalent on the inhibitory profiles as well, KIR3DS06, KIR3DS04, and KIR3DS01, with an occurrence of 37.5%, 25.0%, 18.8%, and 18.8%, respectively (Fig. 4). In total, 20 different activating KIR genes could be defined on activating region configurations, none of which displayed predominance for specific alleles. Half of those KIR genes were exclusively present on activating profiles and thereby absent from the inhibitory KIR region configurations, including one of the highly frequent activating genes, KIR3DS04 (Fig. 4B).
Human group A and B KIR haplotypes could also be distinguished by the presence of specific inhibitory genes, like KIR2DL5 on group B haplotypes. Similar distinctive inhibitory markers are absent from inhibitory and activating profiles on rhesus macaque KIR region configurations. Some inhibitory KIR genes are, however, more frequently present on a specific functional profile. For instance, the presence of KIR3DL05 is documented for 58% of the activating profiles compared to 20% of the inhibitory counterparts (Fig. 4A). KIR3DL07, KIR3DL10, and KIR3DL11 are also more prominent on activating profiles, whereas KIR3DLW03 is slightly more dominant on inhibitory region configurations. No specific alleles of those unevenly distributed KIR genes were defined for either activating or inhibitory profiles.
Whether the functional implications of inhibitory and activating KIR profiles in rhesus macaques relate to selective pressure that maintain group A and B KIR haplotypes in humans is unclear. There is no unambiguous categorization of the different KIR region configurations as in humans, but even the less stringent functional KIR gene profiles in rhesus macaques might impact their health and disease.
Sets of rhesus macaque KIR genes display non-random relationships
A non-random association, or linkage disequilibrium, is demonstrated for specific sets of KIR genes and alleles in different human populations. For instance, KIR2DS3 is in absolute linkage with KIR2DL5 in Caucasian populations (Pyo et al. 2013; Ordóñez et al. 2008). The presence of these neighboring KIR genes in both centromeric and telomeric haplotype segments indicates that they might duplicate as a tandem. Linkage disequilibrium calculations require sufficient data from different populations on specific gene loci, which is limited for the transcriptome study in our rhesus macaque cohort. Instead, we determined the co-occurrence and co-exclusion of KIR genes, which displays the pairwise relationship of genes, independent of their loci and population.
Most rhesus macaque KIR genes display a random relationship, where the observed and expected co-occurrence is similar (Fig. 5). Two sets of KIR genes display a statistically significant strong association (p < 0.05), where the observed co-occurrence is higher than the expected tandem frequency, which is the case for KIR3DL10-KIR3DS05 and KIR3DLW03-KIR3DSW39. However, these KIR gene tandems do not display an absolute co-occurrence, as 16% of the region configurations that contain KIR3DL10 lack the presence of a KIR3DS05 gene, whereas 28% of the KIR3DS05-positive configurations do not contain a KIR3DL10 gene. No specific KIR3DL10 or KIR3DS05 alleles were involved in this co-occurrence tandem. The mentioned other KIR tandem displays similar levels of co-occurrence, with 53% and 27% of the KIR3DLW03- and KIR3DSW39-positive haplotype configurations lacking its tandem partner, respectively. All configurations that contain KIR3DSW39*001 lack the presence of KIR3DLW03, whereas no specific alleles were defined for the KIR3DLW03-KIR3DSW39 tandem or KIR3DLW03 alone. The co-occurrence of other KIR gene tandems was not statistically significant, but trends for more non-random KIR gene couples were noted and included: for instance, KIR3DL05-KIR3DS02, KIR3DL07-KIR3DS02, KIR3DL05-KIR3DL07, and KIR3DL01-KIR3DL08. A triplet of genes including KIR3DL05, KIR3DL07, and KIR3DS02 might be co-occurring based on these trends, but the observed frequency of this triplet was only slightly above expected (0.085 versus 0.069). In addition to co-occurrence, sets of KIR genes display a non-random co-exclusion, in which the observed frequency of a gene tandem was lower than expected. Statically significant co-exclusions were evidenced for KIR3DL02-KIR3DL10 and KIR3DL02-KIR3DS05, whereas trends for co-exclusion were indicated for KIR3DL01-KIR3DL02, KIR3DL01-KIR3DL11, and KIR3DL02-KIR3DLW03. The presence of KIR3DL02 displays an absolute co-exclusion with KIR3DL10 and KIR3DS05.
KIR gene co-occurrence and co-exclusion matrix. The matrix displays the co-occurrence and co-exclusion for pairwise KIR genes. The expected frequency is subtracted from the observed frequency for all KIR gene tandems. White backgrounds indicate similar expected and observed frequencies of pairwise KIR genes, indicating a random association. Positive corrected frequencies demonstrate co-occurrence (green background), whereas negative corrected frequencies indicate co-exclusion (red background). KIR3DL20 is present on all haplotypes as a framework gene and therefore will always display identical observed and expected frequencies in relation to any other KIR gene. Only KIR genes that were documented on at least two KIR haplotype configurations were included in the matrix. Significant (p < 0.05) associations are in bold
Additional associations might be confirmed by elucidating more KIR region configurations in rhesus macaques. The relationships between genes might indicate selection for specific sets of KIR genes to balance synergistic or antagonistic functions or indicate a haplotype architecture that is structured around physical constraints.
Discussion
Human and rhesus macaque KIR systems display considerable similarities, especially in their receptor structures, MHC class I ligand recognition, and variegated expression patterns. As such, the functional role of KIR receptors in NK cell biology seems to be conserved in both species. However, since the separation from a common ancestor, approximately 25–33 million years ago, the genetic make-up of the region encoding the KIR receptors has deviated considerably in humans and rhesus macaques. With the emergence of MHC-C in an ancestor of humans and orangutans, after the split from the macaque lineage, a differential expansion of KIR genes occurred, with KIR2D (lineage III) dominating the hominoid KIR repertoire, whereas KIR3D (lineage II) features extensive expansion in macaques. In humans, this lineage III expansion affects both centromeric and telomeric regions, whereas the KIR3D layout in rhesus macaques is confined to the telomeric segment. Over decades, the organization of human KIR haplotypes has been extensively studied in different populations and manifested the existence of two groups, A and B, that are distinguished based on their gene content. These haplotypes are arranged around a standard framework structure complemented with an assortment of genes, of which some display strong linkage, such as KIR2DL5-KIR2DS3/S5. These characteristics of functional haplotype groups and strongly linked genes could only be defined in large datasets of outbred populations. Such comprehensive datasets are lacking for non-human primates, which limits the power to thoroughly compare the KIR cluster evolution during primate speciation. During this study, the number of rhesus macaques that are characterized for their KIR transcriptome was roughly quadrupled, and it now covers 118 thoroughly defined KIR region configurations (Suppl. Fig. 1 and Suppl. Table III). Although the number of studied human individuals from a wide range of populations exceeds the current rhesus macaque study, this large dataset allowed a comprehensive comparison of KIR haplotype characteristics.
The most prominent difference in the KIR haplotype organization is the relatively fixed organization that is reflected by the non-variable gene content of the two haplotype groups in humans as compared to the highly diverse and apparently less categorizable nature of the rhesus macaque KIR region configurations (Suppl. Figs. 1 and 2). The functional character of the different region configurations in rhesus macaques, based on the number of inhibitory and activating KIR genes, however, might be categorized in such a manner that it resembles the more inhibitory group A and the more activating group B KIR haplotype classification in humans. These functional profiles are evenly distributed in the rhesus macaque cohort studied, as is observed for the A and B KIR haplotypes in most human populations. Several common KIR2DS4 alleles on human group A haplotypes have a mutation that prevents expression of a functional receptor, which gives these haplotypes a solely inhibitory potential (Graef et al. 2009). These haplotypes are mirrored by at least 20 KIR region configurations in rhesus macaques, which do not display any activating gene entity. The conservation of these inhibitory profiles in two different primate species indicates that an activating KIR signaling potential is not essential for life. Nevertheless, the balancing selection that maintains activating KIR gene profiles across human and rhesus macaque populations suggests particular functions that might be beneficial. The precise biological consequences of having distinct functional KIR profiles is unclear and is probably dependent on the presence of relevant epitopes on MHC class I ligands. The KIR and MHC systems are embedded on different chromosomes and segregate independently. The different combinations of KIR and MHC, with specific tendencies towards activating or inhibitory signaling, may associate with a wide spectrum of biological or clinical scenarios, including successful pregnancy, cell and organ transplantation, viral infections, autoimmunity, and different forms of malignancies (Alecsandru et al. 2014; Johnsen et al. 2018; McQueen et al. 2007; Littera et al. 2017; Lin et al. 2016; Hernandez et al. 2018; Kulkarni et al. 2008; Aghaei et al. 2019; Matzaraki et al. 2017). There is, however, a lack of consensus, as these association studies display controversies. There may be multiple reasons for these contradicting association outcomes, including the extensive variation in MHC and KIR alleles, the uneven distribution of diversity across individuals and populations, the variegated expression patterns, inaccurate typing methodologies, and the limited sample sizes of some studies. In addition, most association studies define simplified structural motifs, such as centromeric and telomeric segments or gene presence/absence profiles, which underestimate the complexity of the MHC and KIR interplay. The lack of consensus in association studies limits our knowledge regarding the selective pressure to balance activating and inhibitory KIR haplotype profiles. The presence of a similar balance in rhesus macaques, however, might indicate that the selection to maintain distinct functional haplotype configurations is conserved and might be important in NK cell biology.
The KIR cluster diversity is mainly generated by chromosomal recombination events that reshuffle the KIR gene content across haplotypes. In humans, a major hotspot for recombination is located between the centromeric and telomeric haplotype segments. This transposon-rich stretch facilitates similar recombination events in rhesus macaques, as is identified in this study based on the allele content of centromeric and telomeric regions (Fig. S1). Other recombination hotspots are located within introns and generate not only reshuffled KIR haplotypes but also novel fusion genes. For example, one of the first extended human KIR haplotypes that was documented contains two copies of KIR2DL4 and KIR3DL1/S1 and a fusion gene that is composed of KIR2DL5A and KIR3DP1 (Martin et al. 2003). As such, the presence of a fusion KIR gene is often a marker for haplotype recombination events that map within introns. The total number of fusion KIR genes and the frequency of haplotypes that contain a fusion entity seem to be higher in rhesus macaques than in humans. One may translate this into the notion that the KIR system in rhesus macaques experiences stronger diversifying selection than humans. This conclusion is, however, difficult to substantiate and deserves to be handled with caution. In rhesus macaques, the novel KIR entities received new gene designations (e.g., KIR3DLW31, which represents a fusion of KIR3DL02 and KIR3DL01) and are easily recognized as entities that might encode a receptor with distinct characteristics, such as differential signaling potential, cellular localization, and ligand binding specificity (Bruijnesteijn et al. 2020c). In contrast, the human nomenclature guidelines simplify the recombination complexity, as human fusion KIR genes are named as alleles of the gene that was the donor of the largest fraction of the fusion entity (Marsh et al. 2003). For instance, KIR2DS2*005 is a fusion gene that was generated by a recombination event in intron 6 of KIR2DS2 and KIR2DS3 genes, thereby reshuffling the binding and signaling domains (Ordóñez et al. 2011). As such, KIR2DS2*005 might harbor functional properties that are different from other KIR2DS2 alleles. The current nomenclature standards might blur the visibility of the total number of fusion KIR genes in the human population as well as their differential functional capacities. An adaption of the nomenclature guidelines for fusion genes in humans, as is also proposed by Cisneros and colleagues, might improve the classification of KIR receptors (Cisneros et al. 2020). Previous KIR recombination studies and a phylogenetic analysis of all human KIR sequences present in the IPD-KIR Database (release 2.10.0) would estimate at least 10 different fusion KIR alleles (Pyo et al. 2013; Vendelbosch et al. 2015; Ordóñez et al. 2008; Cisneros et al. 2020; Jiang et al. 2012; Hou et al. 2012). These documented recombination events involve different KIR genes as donor and acceptor entities, including the pseudogenes KIR2DP1 and KIR3DP1. Even more, a recombination event that introduces a KIR2DL5 promoter in front of KIR3DP1 is documented, which results in transcription of this pseudogene (Gómez-Lozano et al. 2005). The event demonstrates that promoter regions might also be reshuffled from one gene to the other, thereby affecting the transcription status. Most frequent human KIR haplotypes, however, lack fusion genes, which is in contrast to some of the relatively high frequency of rhesus macaque KIR haplotypes that do contain a fusion entity (Fig. S1, H4 and H80). This suggests that selection for novel fusion KIR genes in rhesus macaques is probably an ongoing process, resulting in a continuous diversification of their gene repertoire, whereas the main repertoire in humans remains largely fixed in the standard set of 17 KIR genes. The predominance of such a fixed set of KIR genes substantiates the largely balanced equilibrium of common human KIR haplotypes, which is in contrast to the more fluent rhesus macaque KIR gene repertoire.
The KIR haplotype diversity in macaques is further illustrated by the few individuals that are homozygous for their KIR region configuration in the cohort studied. Even more, at the allele level, none of the individuals displayed a set of identical KIR haplotypes. This is in contrast with the relatively high frequency of humans that are homozygous for group A KIR haplotypes, and in particular populations, a selection for KIR homozygosity is even proposed (de Brito Vargas et al. 2021). This selection might be driven by a limited repertoire of MHC class I ligands, indicating that minimal KIR and MHC diversity is sufficient to maintain NK cell education and modulation in a population. On the opposite, differential selective pressure might shape the extensive KIR haplotype diversity in macaques, reflected by the lack of KIR homozygosity, to maintain a functional relationship with their highly expanded MHC class I repertoire.
Human KIR haplotypes may contain sets of KIR genes that display a strong linkage disequilibrium, of which some are even in an absolute pairwise relationship. The relatively large and outbred cohort presented in this study allowed for the first time a similar definition of co-occurring and co-excluding KIR genes in rhesus macaques. Although most of these non-random pairwise relationships only displayed trends for linkage, several gene combinations presented statistically significant associations, of which the pairwise linkage of KIR3DL02 with KIR3DL10 and KIR3DS05 involved absolute co-exclusions (Fig. 5). So far, only a few rhesus macaque KIR haplotypes are completely defined and annotated at the genomic level. The wide diversity of their haplotype configurations suggests extensive variation in haplotype architecture. Information on co-occurring and co-excluding KIR genes might help to resolve more genomic haplotypes, thereby elucidating the complex architecture. For example, most strong human linkage disequilibria involve neighboring KIR genes, such as KIR2DL5-KIR2DS3/5, which provide insights into these haplotype structures. Similarly, three of the eight completely defined rhesus macaque KIR haplotypes contain both KIR3DL10 and KIR3DS05, which are always located adjacent to each other (Bruijnesteijn et al. 2021). Larger cohorts might confirm more co-occurrence and co-exclusion KIR gene combinations, which would help to explain the complex rhesus macaque KIR cluster. However, since the majority of KIR genes seem to display a random pairwise relationship, a genomic characterization of haplotypes will still be required.
The relatively abundant linkage combinations defined for human KIR genes already constrain their potential haplotype diversity. The variation is limited even further by the strong associations of particular sets of alleles that are linked to haplotype segments. For example, the centromeric segment of the cB01-tA01 haplotype mainly contains the KIR2DL5*002-KIR2DS3*001 tandem, with only rare deviations in allele content (Solloch et al. 2020). In the rhesus macaque cohort studied, similar co-occurrences were not determined at the allele level. Only the co-exclusion of KIR3DSW39*001 with KIR3DLW03 displayed a tendency towards an allele-specific association, but such trends have to be confirmed by increasing the sample size. These linkage disequilibria put the extensive polymorphism documented for both humans and rhesus macaques into a different perspective. Whereas KIR alleles in rhesus macaques seem to combine more or less randomly on haplotypes, the linkage studies in humans might indicate more confined groups of alleles that segregate together. Even though additional KIR characterization studies at the allele level resolution might be required to confirm more combinations of specific alleles, the current indication for KIR gene and allele linkage further substantiates largely balanced KIR haplotypes in humans compared to rhesus macaques.
During evolution, the diversity of the human KIR gene cluster skewed towards a balanced equilibrium of inhibitory and activating haplotype profiles, which are structured around a more or less standard set of KIR genes. This balanced equilibrium might be largely achieved by an interplay of different selective factors, such as the emergence of HLA-C as a specialized KIR ligand, different pathogenic encounters, and increased demands for successful pregnancy (Parham and Moffett 2013). In comparison, the KIR gene diversity in rhesus macaques seems to display a less balanced equilibrium, which indicates differential selective forces that continuously shape their KIR gene repertoire and haplotypes. Several characteristics of the KIR cluster complexity are, however, shared in both species, such as distinct functional haplotype profiles and the association of particular gene combinations, but display different levels of plasticity. As such, the human KIR haplotypes are largely confined to common motifs with moderate deviations, whereas more freedom for diversifying selection seems to apply for the KIR cluster in rhesus macaques.
References
Abi-Rached L, Moesta AK, Rajalingam R, Guethlein LA, Parham P (2010) Human-specific evolution and adaptation led to major qualitative differences in the variable receptors of human and chimpanzee natural killer cells. PLoS Genet 6(11):e1001192-e. https://doi.org/10.1371/journal.pgen.1001192
Aghaei H, Mostafaei S, Aslani S, Jamshidi A, Mahmoudi M (2019) Association study between KIR polymorphisms and rheumatoid arthritis disease: an updated meta-analysis. BMC Med Genet 20(1):24. https://doi.org/10.1186/s12881-019-0754-6
Alecsandru D, Garrido N, Vicario JL, Barrio A, Aparicio P, Requena A et al (2014) Maternal KIR haplotype influences live birth rate after double embryo transfer in IVF cycles in patients with recurrent miscarriages and implantation failure. Hum Reprod 29(12):2637–2643. https://doi.org/10.1093/humrep/deu251
Blokhuis JH, van der Wiel MK, Doxiadis GG, Bontrop RE (2010) The mosaic of KIR haplotypes in rhesus macaques. Immunogenetics 62(5):295–306. https://doi.org/10.1007/s00251-010-0434-3
Bruijnesteijn J, de Groot N, van der Wiel MKH, Otting N, de Vos-Rouweler AJM, de Groot NG et al (2020a) Unparalleled rapid evolution of KIR genes in rhesus and cynomolgus macaque populations. J Immunol 204(7):1770–1786. https://doi.org/10.4049/jimmunol.1901140
Bruijnesteijn J, de Groot NG, Bontrop RE (2020b) The genetic mechanisms driving diversification of the KIR gene cluster in primates. Front Immunol 11:582804. https://doi.org/10.3389/fimmu.2020.582804
Bruijnesteijn J, de Groot NG, Otting N, Maccari G, Guethlein LA, Robinson J et al (2020c) Nomenclature report for killer-cell immunoglobulin-like receptors (KIR) in macaque species: new genes/alleles, renaming recombinant entities and IPD-NHKIR updates. Immunogenetics 72(1–2):37–47. https://doi.org/10.1007/s00251-019-01135-8
Bruijnesteijn J, van der Wiel M, de Groot NG, Bontrop RE (2021) Rapid Characterization of Complex Killer Cell Immunoglobulin-Like Receptor (KIR) Regions Using Cas9 Enrichment and Nanopore Sequencing. Front Immunol. https://doi.org/10.3389/fimmu.2021.722181
Bruijnesteijn J, van der Wiel MKH, de Groot N, Otting N, de Vos-Rouweler AJM, Lardy NM et al (2018a) Extensive alternative splicing of KIR transcripts. Front Immunol. https://doi.org/10.3389/fimmu.2018.02846
Bruijnesteijn J, van der Wiel MKH, Swelsen WTN, Otting N, de Vos-Rouweler AJM, Elferink D et al (2018b) Human and rhesus macaque KIR haplotypes defined by their transcriptomes. J Immunol 200(5):1692. https://doi.org/10.4049/jimmunol.1701480
Cisneros E, Moraru M, Gómez-Lozano N, Muntasell A, López-Botet M, Vilches C (2020) Haplotype-based analysis of KIR-gene profiles in a South European population-distribution of standard and variant haplotypes, and identification of novel recombinant structures. Front Immunol 11:440. https://doi.org/10.3389/fimmu.2020.00440
de Brito Vargas L, Beltrame MH, Ho B, Marin WM, Dandekar R, Montero-Martín G et al (2021) Remarkably low KIR and HLA diversity in Amerindians reveals signatures of strong purifying selection shaping the centromeric KIR region. Mol Biol Evol 39(1). https://doi.org/10.1093/molbev/msab298
Gómez-Lozano N, Estefanía E, Williams F, Halfpenny I, Middleton D, Solís R et al (2005) The silent KIR3DP1 gene (CD158c) is transcribed and might encode a secreted receptor in a minority of humans, in whom the KIR3DP1, KIR2DL4 and KIR3DL1/KIR3DS1 genes are duplicated. Eur J Immunol 35(1):16–24. https://doi.org/10.1002/eji.200425493
Gonzalez-Galarza FF, McCabe A, Santos Eduardo J Md, Jones J, Takeshita L, Ortega-Rivera Nestor D et al (2019) Allele frequency net database (AFND) 2020 update: gold-standard data classification, open access genotype data and new query tools. Nucleic Acids Res 48(D1):D783-D8. https://doi.org/10.1093/nar/gkz1029
Gourraud PA, Meenagh A, Cambon-Thomsen A, Middleton D (2010) Linkage disequilibrium organization of the human KIR superlocus: implications for KIR data analyses. Immunogenetics 62(11–12):729–740. https://doi.org/10.1007/s00251-010-0478-4
Graef T, Moesta AK, Norman PJ, Abi-Rached L, Vago L, Older Aguilar AM et al (2009) KIR2DS4 is a product of gene conversion with KIR3DL2 that introduced specificity for HLA-A*11 while diminishing avidity for HLA-C. J Exp Med 206(11):2557–2572. https://doi.org/10.1084/jem.20091010
Guethlein LA, Norman PJ, Heijmans CMC, de Groot NG, Hilton HG, Babrzadeh F et al (2017) Two orangutan species have evolved different KIR alleles and haplotypes. J Immunol 198(8):3157. https://doi.org/10.4049/jimmunol.1602163
Hernandez EG, Partida-Rodriguez O, Camorlinga-Ponce M, Nieves-Ramirez M, Ramos-Vega I, Torres J et al (2018) Genotype B of killer cell immunoglobulin-like receptor is related with gastric cancer lesions. Sci Rep 8(1):6104. https://doi.org/10.1038/s41598-018-24464-2
Hiby SE, Walker JJ, O’Shaughnessy KM, Redman CW, Carrington M, Trowsdale J et al (2004) Combinations of maternal KIR and fetal HLA-C genes influence the risk of preeclampsia and reproductive success. J Exp Med 200(8):957–965. https://doi.org/10.1084/jem.20041214
Hollenbach JA, Nocedal I, Ladner MB, Single RM, Trachtenberg EA (2012) Killer cell immunoglobulin-like receptor (KIR) gene content variation in the HGDP-CEPH populations. Immunogenetics 64(10):719–737. https://doi.org/10.1007/s00251-012-0629-x
Hou L, Chen M, Ng J, Hurley CK (2012) Conserved KIR allele-level haplotypes are altered by microvariation in individuals with European ancestry. Genes Immun 13(1):47–58. https://doi.org/10.1038/gene.2011.52
Jiang W, Johnson C, Jayaraman J, Simecek N, Noble J, Moffatt MF et al (2012) Copy number variation leads to considerable diversity for B but not A haplotypes of the human KIR genes encoding NK cell receptors. Genome Res 22(10):1845–1854. https://doi.org/10.1101/gr.137976.112
John E, Christiansen FT, Mueller I, Schofield L, Senitzer D, Siba P et al (2012) Distinct distribution of killer-cell immunoglobulin-like receptor genes in the Mugil and Ilaita areas of Papua New Guinea. Tissue Antigens 79(4):263–271. https://doi.org/10.1111/j.1399-0039.2012.01848.x
Johnsen GM, Størvold GL, Drabbels JJM, Haasnoot GW, Eikmans M, Spruyt-Gerritse MJ et al (2018) The combination of maternal KIR-B and fetal HLA-C2 is associated with decidua basalis acute atherosis in pregnancies with preeclampsia. J Reprod Immunol 129:23–9. https://doi.org/10.1016/j.jri.2018.07.005
Khakoo SI, Thio CL, Martin MP, Brooks CR, Gao X, Astemborski J et al (2004) HLA and NK cell inhibitory receptor genes in resolving hepatitis C virus infection. Science 305(5685):872–874. https://doi.org/10.1126/science.1097670
Kulkarni S, Martin MP, Carrington M (2008) The Yin and Yang of HLA and KIR in human disease. Semin Immunol 20(6):343–352. https://doi.org/10.1016/j.smim.2008.06.003
Lanier LL (2005) NK cell recognition. Annu Rev Immunol 23:225–74. https://doi.org/10.1146/annurev.immunol.23.021704.115526
Lin Z, Kuroki K, Kuse N, Sun X, Akahoshi T, Qi Y et al (2016) HIV-1 Control by NK cells via reduced interaction between KIR2DL2 and HLA-C(∗)12:02/C(∗)14:03. Cell Rep 17(9):2210–2220. https://doi.org/10.1016/j.celrep.2016.10.075
Littera R, Piredda G, Argiolas D, Lai S, Congeddu E, Ragatzu P et al (2017) KIR and their HLA Class I ligands: two more pieces towards completing the puzzle of chronic rejection and graft loss in kidney transplantation. PLoS One 12(7):e0180831. https://doi.org/10.1371/journal.pone.0180831
Ljunggren HG, Kärre K (1990) In search of the ‘missing self’: MHC molecules and NK cell recognition. Immunol Today 11(7):237–244. https://doi.org/10.1016/0167-5699(90)90097-s
Marsh SG, Parham P, Dupont B, Geraghty DE, Trowsdale J, Middleton D et al (2003) Killer-cell immunoglobulin-like receptor (KIR) nomenclature report, 2002. Hum Immunol 64(6):648–654. https://doi.org/10.1016/s0198-8859(03)00067-3
Martin MP, Bashirova A, Traherne J, Trowsdale J, Carrington M (2003) Cutting edge: expansion of the KIR locus by unequal crossing Over. J Immunol 171(5):2192. https://doi.org/10.4049/jimmunol.171.5.2192
Matzaraki V, Kumar V, Wijmenga C, Zhernakova A (2017) The MHC locus and genetic susceptibility to autoimmune and infectious diseases. Genome Biol 18(1):76. https://doi.org/10.1186/s13059-017-1207-1
McQueen KL, Dorighi KM, Guethlein LA, Wong R, Sanjanwala B, Parham P (2007) Donor-recipient combinations of group A and B KIR haplotypes and HLA class I ligand affect the outcome of HLA-matched, sibling donor hematopoietic cell transplantation. Hum Immunol 68(5):309–323. https://doi.org/10.1016/j.humimm.2007.01.019
Misra MK, Augusto DG, Martin GM, Nemat-Gorgani N, Sauter J, Hofmann JA et al (2018) Report from the killer-cell immunoglobulin-like receptors (KIR) component of the 17th International HLA and Immunogenetics Workshop. Hum Immunol 79(12):825–833. https://doi.org/10.1016/j.humimm.2018.10.003
Norman PJ, Abi-Rached L, Gendzekhadze K, Hammond JA, Moesta AK, Sharma D et al (2009) Meiotic recombination generates rich diversity in NK cell receptor genes, alleles, and haplotypes. Genome Res 19(5):757–769. https://doi.org/10.1101/gr.085738.108
Ordóñez D, Gómez-Lozano N, Rosales L, Vilches C (2011) Molecular characterisation of KIR2DS2*005, a fusion gene associated with a shortened KIR haplotype. Genes Immun 12(7):544–551. https://doi.org/10.1038/gene.2011.35
Ordóñez D, Meenagh A, Gómez-Lozano N, Castaño J, Middleton D, Vilches C (2008) Duplication, mutation and recombination of the human orphan gene KIR2DS3 contribute to the diversity of KIR haplotypes. Genes Immun 9(5):431–437. https://doi.org/10.1038/gene.2008.34
Parham P (2005) MHC class I molecules and kirs in human history, health and survival. Nat Rev Immunol 5(3):201–214. https://doi.org/10.1038/nri1570
Parham P, Moffett A (2013) Variable NK cell receptors and their MHC class I ligands in immunity, reproduction and human evolution. Nat Rev Immunol 13(2):133–144. https://doi.org/10.1038/nri3370
Parham P, Norman PJ, Abi-Rached L, Guethlein LA (2012) Human-specific evolution of killer cell immunoglobulin-like receptor recognition of major histocompatibility complex class I molecules. Philos Trans R Soc Lond B Biol Sci 367(1590):800–811. https://doi.org/10.1098/rstb.2011.0266
Pyo CW, Guethlein LA, Vu Q, Wang R, Abi-Rached L, Norman PJ et al (2010) Different patterns of evolution in the centromeric and telomeric regions of group A and B haplotypes of the human killer cell Ig-like receptor locus. PLoS One 5(12):e15115. https://doi.org/10.1371/journal.pone.0015115
Pyo C-W, Wang R, Vu Q, Cereb N, Yang SY, Duh F-M et al (2013) Recombinant structures expand and contract inter and intragenic diversification at the KIR locus. BMC Genomics 14(1):89. https://doi.org/10.1186/1471-2164-14-89
Shilling HG, Lienert-Weidenbach K, Valiante NM, Uhrberg M, Parham P (1998) Evidence for recombination as a mechanism for KIR diversification. Immunogenetics 48(6):413–416. https://doi.org/10.1007/s002510050453
Solloch UV, Schefzyk D, Schäfer G, Massalski C, Kohler M, Pruschke J et al (2020) Estimation of German KIR allele group haplotype frequencies. Front Immunol 11(429). https://doi.org/10.3389/fimmu.2020.00429
Uhrberg M, Valiante NM, Shum BP, Shilling HG, Lienert-Weidenbach K, Corliss B et al (1997) Human diversity in killer cell inhibitory receptor genes. Immunity 7(6):753–763. https://doi.org/10.1016/s1074-7613(00)80394-5
Vendelbosch S, de Boer M, van Leeuwen K, Pourfarzad F, Geissler J, van den Berg TK et al (2015) Novel insights in the genomic organization and hotspots of recombination in the human KIR locus through analysis of intergenic regions. Genes Immun 16(2):103–111. https://doi.org/10.1038/gene.2014.68
Vierra-Green C, Roe D, Hou L, Hurley CK, Rajalingam R, Reed E et al (2012) Allele-level haplotype frequencies and pairwise linkage disequilibrium for 14 KIR loci in 506 European-American individuals. PLoS One 7(11):e47491-e. https://doi.org/10.1371/journal.pone.0047491
Wilson MJ, Torkar M, Haude A, Milne S, Jones T, Sheer D et al (2000) Plasticity in the organization and sequences of human KIR/ILT gene families. Proc Natl Acad Sci U S A 97(9):4778–4783. https://doi.org/10.1073/pnas.080588597
Acknowledgements
We thank Prof. Dr. A. Sanchez-Mazas and Dr. J. M. Nunes from the Anthropology Unit of the Department of Genetics and Evolution (University of Geneva, Switzerland) for their advice on pairwise gene linkage analysis. We also thank D. Devine for editing the manuscript and F. van Hassel for preparing the figures.
Funding
This work was supported in part by the NIH/NIAID contract number HHSN272201600007C.
Author information
Authors and Affiliations
Contributions
NdG and AdV-R performed all practical work. NdG, AdV-R, and JB analyzed the data. JB wrote the manuscript. NGdG and RB supervised the project and edited the manuscript.
Corresponding author
Ethics declarations
Competing interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bruijnesteijn, J., de Groot, N., de Vos-Rouweler, A.J.M. et al. Comparative genetics of KIR haplotype diversity in humans and rhesus macaques: the balancing act. Immunogenetics 74, 313–326 (2022). https://doi.org/10.1007/s00251-022-01259-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-022-01259-4