Abstract
Rheumatoid arthritis (RA) and type 1 diabetes (T1D) are two autoimmune disorders that have been reported to co-occur in the same subjects or in different subjects from the same family. This suggests the sharing of disease susceptibility loci between RA and T1D. This study was aimed to find out such susceptibility loci that are common in both T1D and RA in Pakistani population. A total of 366 Pakistanis comprising related and unrelated RA cases and controls were recruited. Blood samples were collected from all patients followed by DNA isolation. Thirty-one single-nucleotide polymorphisms (SNPs) previously reported to be associated with T1D were genotyped in RA cases and controls using TaqMan SNP genotyping assays. Data was analyzed using FamCC software. We have identified seven SNP associations that survived multiple testing corrections using false discovery rate: SKAP2/rs7804356 (p = 2.47E-04), GLIS3/rs7020673 (p = 2.86E-04), GSDMB/rs2290400 (p = 23.48E-04), BACH2/rs11755527 (p = 9.16E-04), C6orf173/ rs9388489 (p = 3.11E-03), PRKCQ/DKFZp667F0711/ rs947474 (p = 4.53E-03), and DLK1/ rs941576 (p = 9.51E-03). Our results support the presence of overlapping loci between RA and T1D in Pakistani patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As a result of certain genetic and epigenetic variations, immune system undergoes dysregulation, which leads to the loss of self-tolerance and in turn to autoimmune diseases (Invernizzi and Gershwin 2009). Autoimmune disorders are usually multifactorial and thus various genetic and environmental factors contribute to the disease onset and progression (Boscolo et al. 2008; Lleo et al. 2008). Clusters of different autoimmune disorders have been reported to occur in same subjects and/or families (Torfs et al. 1986). Shared susceptibility loci have been reported for a number of autoimmune disorders (Becker et al. 1998). These observations suggest that clinically distinct autoimmune disorders might share common genetic background.
Rheumatoid arthritis (RA) and type 1 diabetes (T1D) are two autoimmune disorders that have been reported to co-occur in the same subjects or in different subjects from the same family (Liao et al. 2009). It has long been established that major genetic predisposition to both of these diseases is contributed by variants of the class II MHC gene, HLA DRB1 (Agrawal and Desai 2003; Munakata et al. 2005; Somers et al. 2006; Zonana et al. 2002). Recent genome-wide association and subsequent replication studies in these two diseases have revealed a large number of non-HLA genetic risk loci that provide an opportunity to explore the possibility of identifying overlapping susceptibility loci/genes between the two autoimmune diseases (Eyre et al. 2010).
Rheumatoid arthritis is a chronic, systemic autoimmune inflammatory disorder that primarily manifests as arthritis that clinically represents as joint pain, stiffness, and swelling. While about 60 % of the genetic contribution to RA pathogenesis seems to be mediated by human leukocytes antigen (HLA) variants, the non-HLA genes also contribute to disease manifestation (de Vries 2011). RA affects 1 % of the general population worldwide (Joshi 2003) and 0.5 % of the general population in Pakistan (Akhter et al. 2011).
Type 1 diabetes is caused by autoimmune destruction of the insulin-producing β cells in the pancreatic islets. While it affects approximately 0.4 % of the European populations, the incidence of T1D has been reported as 1.02 per 100,000 per year in Pakistan (Shera et al. 2008). The major susceptibility genes, the MHC class II genes, HLA-DQB1, and HLA-DRB1 on chromosome 6p21, act in combination with many other non-HLA loci/genes across the genome to influence the susceptibility to T1D (Nejentsev et al. 2007). Genetic predisposition as well as environmental factors contribute to its etiology. To date, more than 40 risk loci have been identified for T1D (Barrett et al. 2009).
The major genetic risk for both RA and T1D is the HLA class II loci (Smyth et al. 2008; Symmons 2002). HLA-DR3 and DR4 are strongly associated with T1D. In addition, DQ2 (DQB1*0201-DQA1*0501) and DQ8 (DQB1*0302-DQA1*0301), which are in strong linkage disequilibrium with DR3 and DR4, respectively, are also found to be associated with T1D in several Caucasian populations (Field 2002). Likewise, HLA-DRB1 antigen of MHC class II as well as MHC Class III region has been associated with susceptibility and disease severity of RA (Gregersen et al. 1987; du Montcel et al. 2005). In addition to HLA genes at 6p21, other genetic loci that have been reported to be shared between RA and T1D include the tyrosine kinase 2 gene (TYK2) located at 19p13 and the protein tyrosine phosphatase, non-receptor type 22 (PTPN22) located at 1p13 (Burn et al. 2011; Parkes et al. 2013). In this study, we investigated 31 susceptibility loci for T1D in a cohort of Pakistani RA patients and controls to further explore the genetic link between these two autoimmune disorders.
Materials and methods
Subjects
A total of 366 subjects, including 239 RA patients and 127 controls, were recruited from different rheumatology departments in Islamabad and Rawalpindi, Pakistan. Blood samples and relevant information were collected from recruited subjects. Because of the study protocol, both related and unrelated subjected were included (Table 1). All 239 RA patients were diagnosed following the American College of Rheumatology (ACR) 1987 classification criteria (Arnett et al. 1988). The mean age-at-onset of disease was 39.1 ± 13.0 years in RA cases (63 % females). The control group (n = 127; 54 % females) had no history of autoimmune diseases and their mean age was 41.2 ± 12.0 years (Table 1). The study was approved by Institutional Review Board (IRB) of Atta-ur-Rahman School of Applied biosciences (ASAB) National University of Sciences and Technology (NUST) Pakistan and University of Pittsburgh (USA). All participants provided written informed consent.
DNA extraction and quantification
DNA was extracted from whole blood using either a phenol chloroform based method or the Fermantas Whole Blood Genomic DNA Purification kit. DNA was quantified using Quant-iT™ PicoGreen® ds-DNA assay kit (Life Technologies, NY, USA).
SNP selection
Forty genetic loci have been reported to show statistically significant association with T1D (Barrett et al. 2009). Of these 40 loci, those that have already been reported to be associated with RA (e.g., CTLA4, PTPN22) were excluded from this study as well as those single-nucleotide polymorphisms (SNP) with less significant p values. Following these exclusions, we genotyped a total of 31 T1D-associated SNPs in our RA study sample (Table 2).
Genotyping
Genotyping was performed using TaqMan SNP genotyping assays (Life Technologies) following manufacturer’s protocol. PCR amplification was performed in 384 well plates on dual-block Gene-Amp ® PCR system 9700 (Life Technologies) and end-point readings were performed on ABI Prism 7900HT sequence detection system instrument (Life Technologies).
Statistical analysis
For the calculation of allele and genotype frequencies in the unrelated sample, the allele counting method was used. Chi-squared (Χ 2) goodness-of-fit test was used to check deviation from Hardy-Weinberg equilibrium. PedCheck (O’Connell and Weeks 1998) program was used to check Mendelian inconsistencies in the pedigree data of family-based samples (http:/Watson.hgen.pitt.edu).
Association of SNPs with RA was examined using Family Case Control (FamCC) software Ver 1.0. FamCC is a software designed to check the association by combining the family dataset and case/control dataset together, but can also analyze each dataset independently. In order to control for multiple tastings, we applied false discovery rate (FDR) value of <0.05 as statistically significant.
Results
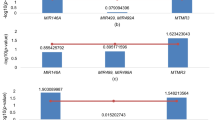
A total of 366 unrelated and family-based RA and control subjects were genotyped to analyze the association of 31 T1D-associated SNPs with RA in order to check common genetic link between these two diseases. The genotyping call rate was >98 % for all SNPs. Genotyping error was estimated by using 10 % of the samples as replicates and the discrepancy rate was found to be 0 % for all SNPs. In the unrelated case/control sample, all SNPs were found to be in Hardy-Weinberg equilibrium. Similarly, the family-based samples showed Mendelian consistency. For the combined analysis of the family-based and unrelated samples, we used FamCC software. Of 31 SNPs analyzed, 11 showed nominal association with RA in our Pakistani sample and 7 of them remained significant after controlling for multiple testing using FDR at <0.05 (Table 3). The most significant association was observed with SKAP2/rs7804356 (p = 2.47E-04) followed by GLIS3/rs7020673 (p = 2.86E-04), GSDMB/ rs2290400 (p = 3.48E-04), BACH2/rs11755527 (p = 9.16E-04), C6orf173/rs9388489 (p = 3.11E-03), PRKCQ/DKFZp667F0711/ rs947474 (p = 4.53E-03), and DLK1/ rs941576 (p = 9.51E-03).
Discussion
By investigating the T1D-associated loci/genes in a Pakistani RA study sample, we have found 7 T1D-associated loci/genes to be also relevant to RA even after controlling for multiple comparisons.
The most significant association was seen with SKAP2/ rs7804356 on chromosome 7p15.2. Src kinase-associated phosphoprotein 2 (SKAP2) is involved in src signaling pathway. Similar to the previously reported association of the T allele of rs7804356 with T1D, we also found the association of T allele with RA (Barrett et al. 2009). This suggests that T allele acts as a genetic link between T1D and RA. Further investigations for the association of rs7804356 can illustrate the role of risk allele T in key regulatory pathways of autoimmunity.
The second most significant association was observed with GLIS3/ rs7020673. GLIS family zinc finger 3 is a member of the GLI-similar zinc finger protein family. This protein functions as both a repressor and activator of transcription and is specifically involved in the development of pancreatic beta cells, thyroid, eye, liver, and kidney (Lichti-Kaiser et al. 2014). The GLIS3 gene region has been identified as a susceptibility risk locus for both type 1 and type 2 diabetes (Nogueira et al. 2013).
The third most significant association was observed with GSDMB/rs2290400 that is located on chromosome 17q12. Gasdermin B (GSDMB) encodes a member of the gasdermin-domain containing protein family and it plays a role as secretory or metabolic product involved in secretory pathway. GSDMB has previously been shown to be strongly associated with smoking and asthma as well as T1D (Barrett et al. 2009; Marinho et al. 2012).
The fourth most significant association was observed with BACH2/rs11755527 on chromosome 6q15. BTB and CNC homology 1, basic leucine zipper transcription factor 2 (BACH2), is a transcriptional regulator that acts as repressor or activator. It plays an important role in coordinating transcription activation and repression by MAFK (Igarashi et al. 2014). Eyre et al. (Eyre et al. 2010) also tested this SNP (rs11755527) in British Caucasian RA patients but they did not find significant association in their study. This may be due to population differences between the two study samples.
The fifth most significant association was observed with C6orf173/rs9388489 located on chromosome 6q22.32. Chromosome 6 open reading frame 173 (C6orf173) plays a central role in assembly of kinetochore proteins, mitotic progression, and chromosome segregation and has been shown to act as a susceptibility locus for T1D and cardiovascular disease (Barrett et al. 2009; Ding and Kullo 2011).
Two more genetic loci associated with T1D also exhibited significant association with RA, PRKCQ/rs947474, and DLK1/rs941576. Protein kinase C, theta (PRKCQ) gene on chromosome 10p15.1, is involved in activation of the transcription factors NF-kappa B and AP-1, and may link the T cell receptor (TCR) signaling complex to the activation of the transcription factors. The delta-like 1 homolog (DLK1) is considered as a tumor suppressor and is also involved in cell differentiation.
The association of these T1D-associated SNPs with RA indicates that these SNPs not only provide a genetic link between T1D and RA, but also they probably play a role in common biochemical pathways for autoimmune disease pathogenesis. In addition to the potential common genetic background observed for these two autoimmune diseases in this study, it would be also important to identify common environmental factors and/or shared gene-environment interactions that contribute to this picture of common elements in disease pathogenesis.
Conclusion
We have identified at least 7 T1D loci/genes that are also associated with RA risk in Pakistani population. Our study further supports the presence of common biochemical pathways for autoimmune disease pathogenesis. Although our sample size was relatively small, our associations survived after controlling for FDR. This suggests that these associations may be genuine. Additional large studies are required to replicate our findings, which may help to delineate common pathways between different autoimmune diseases.
References
Agrawal S, Desai MP (2003) Simultaneous occurrence of type I diabetes mellitus and juvenile rheumatoid arthritis. Indian Pediatr 40:568–571
Akhter E, Bilal S, Kiani A, Haque U (2011) Prevalence of arthritis in India and Pakistan: a review. Rheumatol Int 31:849–855
Arnett FC, Edworthy SM, Bloch DA, McShane DJ, Fries JF, Cooper NS, Healey LA, Kaplan SR, Liang MH, Luthra HS et al (1988) The American Rheumatism Association 1987 revised criteria for the classification of rheumatoid arthritis. Arthritis Rheum 31:315–324
Barrett JC, Clayton DG, Concannon P, Akolkar B, Cooper JD, Erlich HA, Julier C, Morahan G, Nerup J, Nierras C, Plagnol V, Pociot F, Schuilenburg H, Smyth DJ, Stevens H, Todd JA, Walker NM, Rich SS (2009) Genome-wide association study and meta-analysis find that over 40 loci affect risk of type 1 diabetes. Nat Genet 41:703–707
Becker C, Barbulescu K, Hildner K, Meyer zum Buschenfelde KH, Neurath MF (1998) Activation and methotrexate-mediated suppression of the TNF alpha promoter in T cells and macrophages. Ann N Y Acad Sci 859:311–314
Boscolo P, Youinou P, Theoharides TC, Cerulli G, Conti P (2008) Environmental and occupational stress and autoimmunity. Autoimmun Rev 7:340–343
Burn GL, Svensson L, Sanchez-Blanco C, Saini M, Cope AP (2011) Why is PTPN22 a good candidate susceptibility gene for autoimmune disease? FEBS Lett 585:3689–3698
de Vries R (2011) Genetics of rheumatoid arthritis: time for a change! Curr Opin Rheumatol 23:227–232
Ding K, Kullo IJ (2011) Geographic differences in allele frequencies of susceptibility SNPs for cardiovascular disease. BMC Med Genet 12:55
du Montcel ST, Michou L, Petit-Teixeira E, Osorio J, Lemaire I, Lasbleiz S, Pierlot C, Quillet P, Bardin T, Prum B, Cornelis F, Clerget-Darpoux F (2005) New classification of HLA-DRB1 alleles supports the shared epitope hypothesis of rheumatoid arthritis susceptibility. Arthritis Rheum 52:1063–1068
Eyre S, Hinks A, Bowes J, Flynn E, Martin P, Wilson AG, Morgan AW, Emery P, Steer S, Hocking LJ, Reid DM, Harrison P, Wordsworth P, Thomson W, Worthington J, Barton A (2010) Overlapping genetic susceptibility variants between three autoimmune disorders: rheumatoid arthritis, type 1 diabetes and coeliac disease. Arthritis Res Ther 12:R175
Gregersen PK, Silver J, Winchester RJ (1987) The shared epitope hypothesis. An approach to understanding the molecular genetics of susceptibility to rheumatoid arthritis. Arthritis Rheum 30:1205–1213
Igarashi K, Ochiai K, Itoh-Nakadai A, Muto A (2014) Orchestration of plasma cell differentiation by Bach2 and its gene regulatory network. Immunol Rev 261:116–125
Invernizzi P, Gershwin ME (2009) The genetics of human autoimmune disease. J Autoimmun 33:290–299
Joshi VR (2003) Arthritis in the elderly. J Indian Med Assoc 101:408, 410, 412 passim
Liao KP, Gunnarsson M, Källberg H, Ding B, Plenge RM, Padyukov L, Karlson EW, Klareskog L, Askling J, Alfredsson L (2009) A specific association exists between type 1 diabetes and anti-CCP positive rheumatoid arthritis. Arthritis Rheum 3:653–660
Lichti-Kaiser K, ZeRuth G, Jetten AM (2014) Transcription factor Gli-similar 3 (Glis3): implications for the development of congenital hypothyroidism. J Endocrinol Diabetes Obes 2:1024
Lleo A, Battezzati PM, Selmi C, Gershwin ME, Podda M (2008) Is autoimmunity a matter of sex? Autoimmun Rev 7:626–630
Marinho S, Custovic A, Marsden P, Smith JA, Simpson A (2012) 17q12-21 variants are associated with asthma and interact with active smoking in an adult population from the United Kingdom. Ann Allergy Asthma Immunol 108(402–411), e9
Munakata Y, Kodera T, Saito T, Sasaki T (2005) Rheumatoid arthritis, type 1 diabetes, and Graves’ disease after acute parvovirus B19 infection. Lancet 366:780
Nejentsev S, Howson JM, Walker NM, Szeszko J, Field SF, Stevens HE, Reynolds P, Hardy M, King E, Masters J, Hulme J, Maier LM, Smyth D, Bailey R, Cooper JD, Ribas G, Campbell RD, Clayton DG, Todd JA (2007) Localization of type 1 diabetes susceptibility to the MHC class I genes HLA-B and HLA-A. Nature 450:887–892
Nogueira TC, Paula FM, Villate O, Colli ML, Moura RF, Cunha DA, Marselli L, Marchetti P, Cnop M, Julier C, Eizirik DL (2013) GLIS3, a susceptibility gene for type 1 and type 2 diabetes, modulates pancreatic beta cell apoptosis via regulation of a splice variant of the BH3-only protein Bim. PLoS Genet 9, e1003532
O’Connell JR, Weeks DE (1998) PedCheck: a program for identification of genotype incompatibilities in linkage analysis. Am J Hum Genet 63:259–266
Parkes M, Cortes A, van Heel DA, Brown MA (2013) Genetic insights into common pathways and complex relationships among immune-mediated diseases. Nat Rev Genet 14:661–673
Shera AS, Miyan Z, Basit A, Maqsood A, Ahmadani MY, Fawwad A, Riaz M (2008) Trends of type 1 diabetes in Karachi, Pakistan. Pediatr Diabetes 9:401–406
Smyth DJ, Plagnol V, Walker NM, Cooper JD, Downes K, Yang JH, Howson JM, Stevens H, McManus R, Wijmenga C, Heap GA, Dubois PC, Clayton DG, Hunt KA, van Heel DA, Todd JA (2008) Shared and distinct genetic variants in type 1 diabetes and celiac disease. N Engl J Med 359:2767–2777
Somers EC, Thomas SL, Smeeth L, Hall AJ (2006) Autoimmune diseases co-occurring within individuals and within families: a systematic review. Epidemiology 17:202–217
Symmons DP (2002) Epidemiology of rheumatoid arthritis: determinants of onset, persistence and outcome. Best Pract Res Clin Rheumatol 16:707–722
Torfs CP, King MC, Huey B, Malmgren J, Grumet FC (1986) Genetic interrelationship between insulin-dependent diabetes mellitus, the autoimmune thyroid diseases, and rheumatoid arthritis. Am J Hum Genet 38:170–187
Zonana MF, Reyes E, Weisman AK (2002) Coexistence of four autoimmune diseases in one patient: the kaleidoscope of autoimmunity. J Clin Rheumatol 8:322–325
Acknowledgments
We acknowledge Higher Education Commission for supporting this study within the Pakistan as well as for International Research Support Initiative Program (IRSIP) awarded to Miss Aysha Kiani for University of Pittsburgh, USA. We are thankful to all the physicians for referring the patients and supporting staff for the collection of blood samples. We are also grateful to the patients, control individuals and their family members for their participation and cooperation in this study.
Conflict of interest
All the authors declared that they have no competing interests.
Funding sources
Funds are provided by Higher Education Commission, Islamabad, Pakistan.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kiani, A.K., Jahngir, S., John, P. et al. Genetic link of type 1 diabetes susceptibility loci with rheumatoid arthritis in Pakistani patients. Immunogenetics 67, 277–282 (2015). https://doi.org/10.1007/s00251-015-0839-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-015-0839-0